Prediction of Radical Scavenging Activities of Anthocyanins Applying Adaptive Neuro-Fuzzy Inference System (ANFIS) with Quantum Chemical Descriptors
Abstract
:1. Introduction
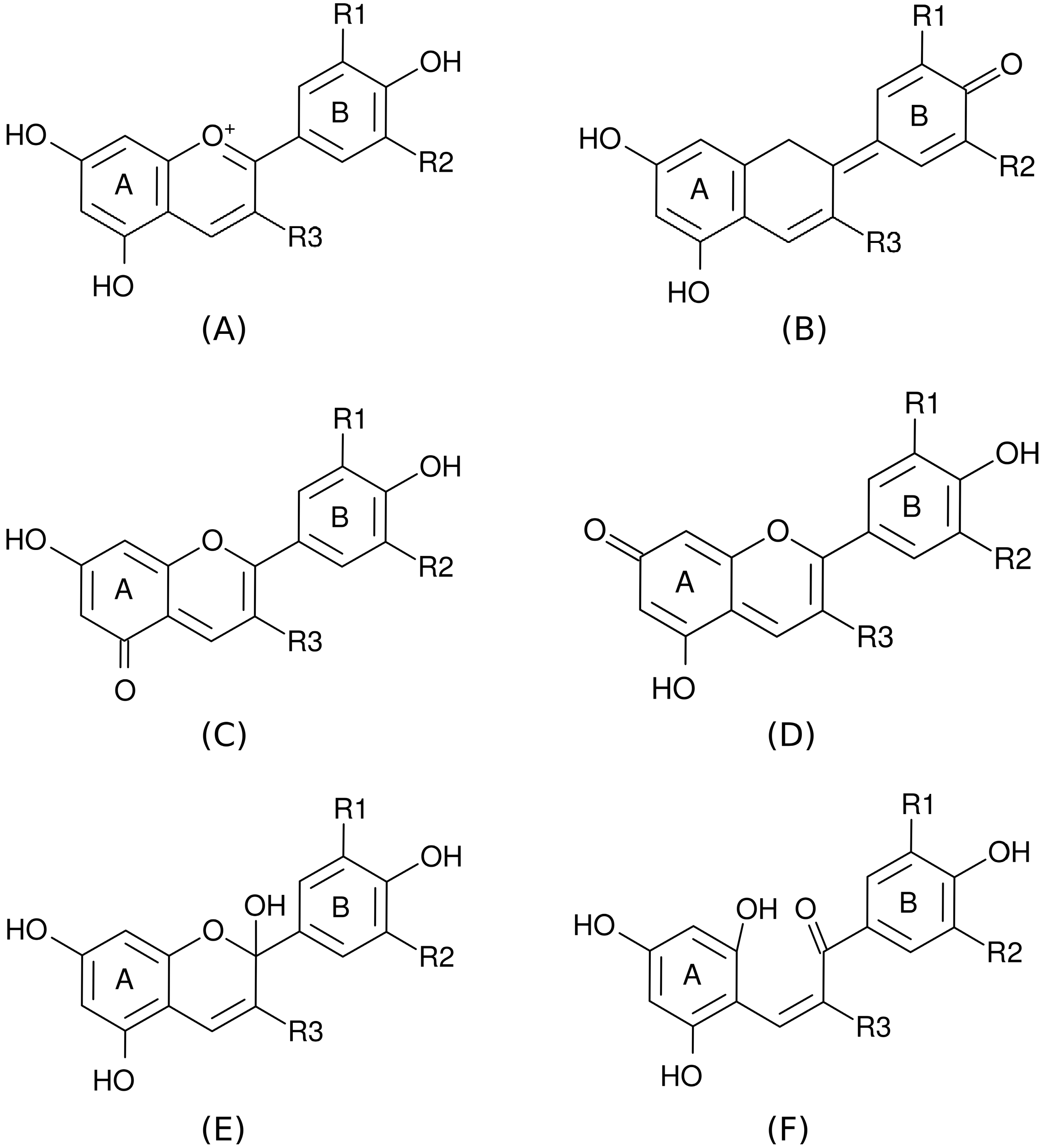
2. Results and Discussion
2.1. Correlation Analysis
| Descriptors | PM6 | PM7 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| FC | QB4' | QB7 | CP | Ch | FC | QB4' | QB7 | CP | Ch | |
| I | 0.554 ** | 0.343 | 0.674 ** | 0.233 | 0.121 | 0.504 * | 0.355 | 0.708 ** | 0.269 | 0.251 |
| A | 0.673 ** | −0.166 | −0.256 | −0.473 * | −0.052 | 0.538 * | −0.206 | −0.037 | −0.289 | −0.199 |
| η | 0.124 | 0.288 | 0.523 * | 0.530 * | 0.121 | 0.263 | 0.310 | 0.418 | 0.378 | 0.298 |
| S | −0.153 | −0.171 | −0.503 * | −0.535 * | −0.138 | −0.238 | −0.132 | −0.398 | −0.411 | −0.284 |
| ω | 0.328 | −0.127 | −0.305 | −0.470 * | −0.054 | 0.202 | −0.112 | −0.085 | −0.326 | −0.212 |
| χ | 0.701 ** | 0.194 | 0.282 | −0.344 | 0.009 | 0.567 ** | 0.087 | 0.551 ** | −0.149 | 0.020 |
| μ | −0.701 ** | −0.194 | −0.282 | 0.344 | −0.009 | −0.567 ** | −0.087 | −0.551 ** | 0.149 | −0.020 |
| Compounds | Method | (1) BDE of FL | BDE of QB7 | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 3 | 5 | 7 | 3' | 4' | 5' | 3 | 5 | 3' | 4' | 5' | |||||||||||||
| cyanidin | PM6 | 80.98 | 84.8 | 90.2 | 82.06 | 83.34 | - | 67.08 | 75.5 | 76.04 | 72.44 | - | |||||||||||
| PM7 | 81.77 | 81.77 | 91.41 | 83.43 | 84.78 | - | 68.98 | 75.72 | 75.54 | 75.16 | - | ||||||||||||
| delphinidin | PM6 | 78.15 | 84.24 | 90.53 | 83.16 | 75.61 | 78.55 | 65.65 | 74.8 | 79.17 | 68.74 | 74.47 | |||||||||||
| PM7 | 82.55 | 85.75 | 92.2 | 85.58 | 77.6 | 82.08 | 68.2 | 75.42 | 78.74 | 71.29 | 75.13 | ||||||||||||
| malvidin | PM6 | 77.33 | 82.05 | 87.19 | - | 73.9 | - | 65.49 | 74.83 | - | 68.04 | - | |||||||||||
| PM7 | 81.19 | 85.22 | 90.98 | - | 75.21 | - | 71.31 | 78.69 | - | 73.36 | - | ||||||||||||
| pelargonidin | PM6 | 81.01 | 84.79 | 90.39 | - | 89.53 | - | 68.45 | 75.5 | - | 74.43 | - | |||||||||||
| PM7 | 81.86 | 85.71 | 91.51 | - | 90.6 | - | 69.68 | 75.89 | - | 76.87 | - | ||||||||||||
| peonidin | PM6 | 80.86 | 84.58 | 90.45 | - | 77.37 | - | 68.38 | 76.99 | - | 69.2 | - | |||||||||||
| PM7 | 81.86 | 85.71 | 91.51 | - | 90.6 | - | 71.53 | 78.3 | - | 73.51 | - | ||||||||||||
| cyanidin-3-coumaroyl-sambubioside-5-galactoside | PM6 | - | - | 88.15 | 80.05 | 82.17 | - | - | - | 74.12 | 75.16 | - | |||||||||||
| PM7 | - | - | 86.57 | 77.29 | 78.88 | - | - | - | 76.74 | 77.52 | - | ||||||||||||
| cyanidin-3-sambubioside-5-galactoside | PM6 | - | - | 89.15 | 80.07 | 82.15 | - | - | - | 74.12 | 75.23 | - | |||||||||||
| PM7 | - | - | 98.43 | 88.66 | 90.97 | - | - | - | 76.74 | 78.07 | - | ||||||||||||
| cyanidin-3,5-diglucoside | PM6 | - | - | 91.23 | 80.23 | 83.08 | - | - | - | 74.85 | 75.92 | - | |||||||||||
| PM7 | - | 85.22 | - | 80.39 | 82.39 | - | - | - | 77.19 | 78.28 | - | ||||||||||||
| cyanidin-3-arabinoside | PM6 | - | 86.49 | 91.25 | 80.53 | 82.61 | - | - | 77.32 | 76.27 | 73.99 | - | |||||||||||
| PM7 | - | 88.14 | 92.69 | 82.63 | 85.54 | - | - | 77.87 | 77.09 | 77.48 | - | ||||||||||||
| cyanidin-3-galactoside | PM6 | - | 93.17 | 93.58 | 81.03 | 83.71 | - | - | 76.74 | 74.76 | 75.55 | - | |||||||||||
| PM7 | - | 96.62 | 96.76 | 84.86 | 85.8 | - | - | 81.26 | 80.34 | 79.3 | - | ||||||||||||
| cyanidin-3-glucoside | PM6 | - | 86.87 | 92.58 | 81.78 | 84.4 | - | - | 76.07 | 74.91 | 75.63 | - | |||||||||||
| PM7 | - | 87.79 | 82.92 | 82.92 | 85.51 | - | - | 77.07 | 77.52 | 77.99 | - | ||||||||||||
| cyanidin-3-rutinoside | PM6 | - | 88.51 | 94.22 | 80.41 | 83.25 | - | - | 76.6 | 74.72 | 76.12 | - | |||||||||||
| PM7 | - | 90.49 | 94.22 | 81.73 | 84.65 | - | - | 80.23 | 77.36 | 78.9 | - | ||||||||||||
| delphinidin-3-glucoside | PM6 | - | 86.92 | 91.58 | 82.36 | 75.31 | 77.35 | - | 76.9 | 78.53 | 69.28 | 74.42 | |||||||||||
| PM7 | - | 89.94 | 92.99 | 84.51 | 77.89 | 80.94 | - | 80.64 | 80.53 | 77.06 | 80.23 | ||||||||||||
| delphinidin-3-rutinoside | PM6 | - | 88.23 | 91.05 | 82.52 | 75.93 | 75.74 | - | 75.22 | 77.2 | 70.95 | 72.55 | |||||||||||
| PM7 | - | 84.54 | 92.68 | 84.59 | 77.79 | 76.34 | - | 72.62 | 80.15 | 73.79 | 73.62 | ||||||||||||
| malvidin-3,5-diglucoside | PM6 | - | - | 86.32 | - | 73.67 | - | - | - | - | 71.07 | - | |||||||||||
| PM7 | - | - | 91.84 | - | 76.05 | - | - | - | - | 72.45 | - | ||||||||||||
| malvidin-3-galactoside | PM6 | - | 87.49 | 89.43 | - | 73.81 | - | - | 74.52 | - | 69.57 | - | |||||||||||
| PM7 | - | 88.43 | 91.56 | - | 75.73 | - | - | 78.65 | - | 72.45 | - | ||||||||||||
| malvidin-3-glucoside | PM6 | - | 85.75 | 89.51 | - | 73.89 | - | - | 76.84 | - | 68.59 | - | |||||||||||
| PM7 | - | 88.63 | 94.26 | - | 78.17 | - | - | 73.52 | - | 71.67 | - | ||||||||||||
| pelargonidin-3-glucoside | PM6 | - | 87.13 | 91.13 | - | 86.79 | - | - | 71.66 | - | 75.48 | - | |||||||||||
| PM7 | - | 86.20 | 93.35 | - | 89.08 | - | - | 72.61 | - | 78.55 | - | ||||||||||||
| peonidin-3-galactoside | PM6 | - | 88.79 | 98.46 | - | 76.39 | - | - | 79.56 | - | 68.31 | - | |||||||||||
| PM7 | - | 88.94 | 92.01 | 78.78 | - | - | 77.10 | - | 72.78 | - | |||||||||||||
| peonidin-3-glucoside | PM6 | - | 88.80 | 90.97 | - | 76.38 | - | - | 74.66 | - | 67.51 | - | |||||||||||
| PM7 | - | 88.57 | 91.51 | - | 78.74 | - | - | 78.95 | - | 72.74 | - | ||||||||||||
| petunidin-3-glucoside | PM6 | - | 86.34 | 90.85 | 82.33 | 74.17 | - | - | 76.79 | 79.31 | 68.40 | - | |||||||||||
| PM7 | - | 88.62 | 94.78 | 87.69 | 78.51 | - | - | 73.73 | 80.33 | 71.51 | - | ||||||||||||
2.2. Prediction of Radical Scavenging Activities of Anthocyanins

| Compounds | (1) Experimental Radical Scavenging Activity | Predicted Radical Scavenging Activity | |
|---|---|---|---|
| PM6 | PM7 | ||
| cyanidin | 33 | 35.3 | 33.2 |
| delphinidin | 42 | 42.6 | 41.0 |
| malvidin | 24 | 26.6 | 26.1 |
| pelargonidin | 31 | 28.3 | 27.6 |
| peonidin | 33 | 31.4 | 29.3 |
| cyanidin-3-coumaroylsambubiose-5-galactoside | 26 | 22.4 | 25.2 |
| cyanidin-3,5-diglucoside | 21 | 22.6 | 20.9 |
| cyanidin-3-arabinoside | 26 | 28.5 | 27.3 |
| cyanidin-3-sambubiose-5-galactoside | 22 | 22.1 | 21.5 |
| cyanidin-3-galactoside | 25 | 30.3 | 30.2 |
| cyanidin-3-glucoside | 32 | 31.1 | 28.8 |
| cyanidin-3-rutinoside | 25 | 27.0 | 27.5 |
| delphinidin-3-glucoside | 42 | 35.4 | 38.1 |
| delphinidin-3-rutinoside | 32 | 33.4 | 30.4 |
| malvidin-3,5-diglucoside | 14 | 16.6 | 14.9 |
| malvidin-3-galactoside | 22 | 22.9 | 21.3 |
| malvidin-3-glucoside | 26 | 22.6 | 21.2 |
| pelargonidin-3-glucoside | 20 | 21.0 | 21.2 |
| peonidin-3-galactoside | 20 | 22.2 | 21.1 |
| peonidin-3-glucoside | 26 | 23.6 | 23.5 |
| petunidin-3-glucoside | 23 | 28.0 | 25.7 |
| (2) Mean absolute error | 2.43 ± 0.35 | 2.06 ± 0.32 | |
| (2) Q-square | 0.82 ± 0.08 | 0.86 ± 0.08 | |
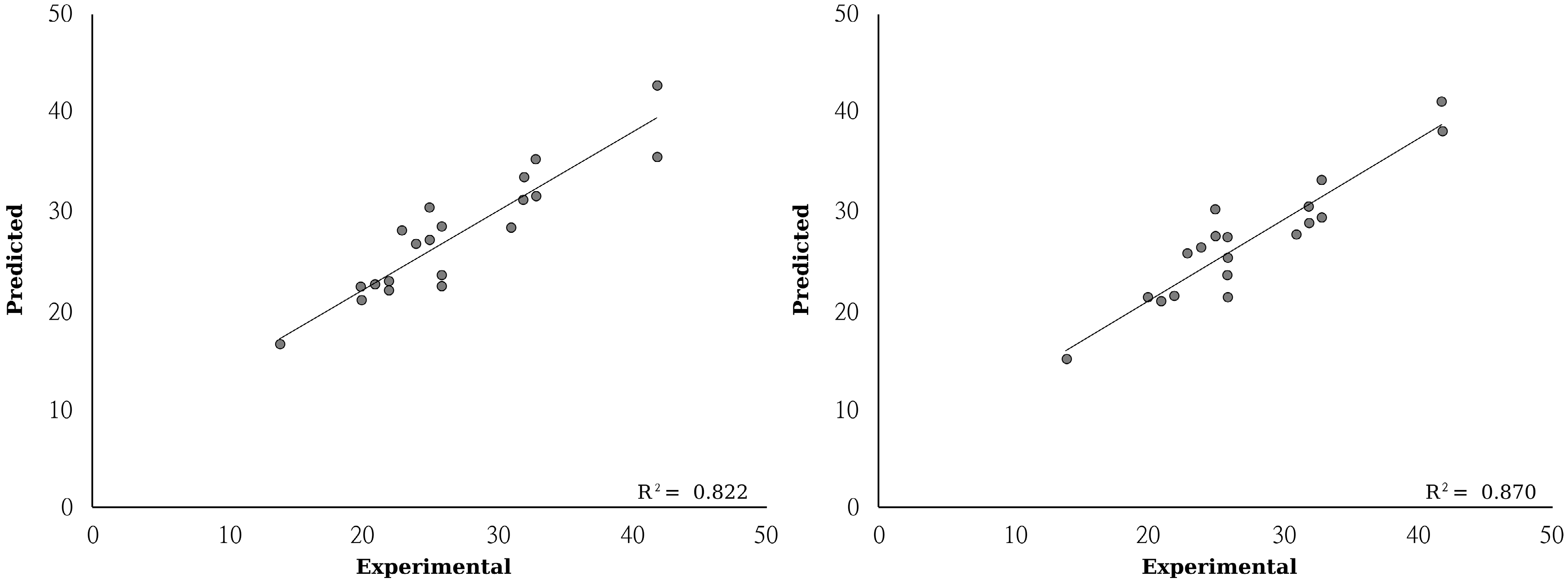
3. Experimental Section
3.1. DPPH Radical Scavenging Activity
| Family | Compounds | (1) nOH | R1 | R2 | (2) R3 | (2) R4 |
|---|---|---|---|---|---|---|
| anthocyanidin | cyanidin | 5 | OH | H | OH | OH |
| delphinidin | 6 | OH | OH | OH | OH | |
| malvidin | 4 | OCH3 | OCH3 | OH | OH | |
| pelargonidin | 4 | H | H | OH | OH | |
| peonidin | 4 | OCH3 | H | OH | OH | |
| anthocyanin | cyanidin-3-coumaroyl-sambubioside-5-galactoside | 3 | OH | H | coumaroyl -sam | gal |
| cyanidin-3-sambubioside-5-galactoside | 3 | OH | H | sam | gal | |
| cyanidin-3-arabinoside | 4 | OH | H | ara | OH | |
| cyanidin-3-galactoside | 4 | OH | H | gal | OH | |
| cyanidin-3-glucoside | 4 | OH | H | glc | OH | |
| cyanidin-3-rutinoside | 4 | OH | H | rut | OH | |
| cyanidin-3,5-diglucoside | 3 | OH | H | glc | glc | |
| delphinidin-3-glucoside | 5 | OH | OH | glc | OH | |
| delphinidin-3-rutinoside | 5 | OH | OH | rut | OH | |
| malvidin-3-galactoside | 3 | OCH3 | OCH3 | gal | OH | |
| malvidin-3-glucoside | 3 | OCH3 | OCH3 | glc | OH | |
| malvidin-3,5-diglucoside | 2 | OCH3 | OCH3 | glc | glc | |
| pelargonidin-3-glucoside | 3 | H | H | glc | OH | |
| peonidin-3-galactoside | 3 | OCH3 | H | gal | OH | |
| peonidin-3-glucoside | 3 | OCH3 | H | glc | OH | |
| petunidin-3-glucoside | 4 | OH | OCH3 | glc | OH |
3.2. Quantum Chemical Descriptors
3.2.1. Molecular Structure Preparation
3.2.2. Calculation of Quantum Chemical Descriptors
3.3. QSAR Model Development
3.3.1. Correlation Analysis
3.3.2. ANFIS

4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Tsuda, T.; Shiga, K.; Ohshima, K.; Kawakishi, S.; Osawa, T. Inhibition of lipid peroxidation and the active oxygen radical scavenging effect of anthocyanin pigments isolated from Phaseolus vulgaris L. Biochem. Pharmacol. 1996, 52, 1033–1039. [Google Scholar] [CrossRef]
- Kong, J.-M.; Chia, L.-S.; Goh, N.-K.; Chia, T.-F.; Brouillard, R. Analysis and biological activities of anthocyanins. Phytochemistry 2003, 64, 923–933. [Google Scholar] [CrossRef]
- Ronald, L.; Wu, X.; Schaich, K. Standardized methods for the determination of antioxidant capacity and phenolics in foods and dietary supplements. J. Agric. Food Chem. 2005, 53, 4290–4302. [Google Scholar] [CrossRef]
- Scott, S.L.; Chen, W.J.; Bakac, A.; Espenson, J.H. Spectroscopic parameters, electrode potentials, acid ionization constants, and electron exchange rates of the 2,2'-azinobis (3-ethylbenzothiazoline-6-sulfonate) radicals and ions. J. Phys. Chem. 1993, 97, 6710–6714. [Google Scholar] [CrossRef]
- Bondet, V.; Brand-Williams, W.; Berset, C. Kinetics and mechanisms of antioxidant activity using the DPPH free radical method. LWT-Food Sci. Technol. 1997, 30, 609–615. [Google Scholar] [CrossRef]
- Amić, D.; Davidovic-Amic, D.; Beslo, D.; Rastija, V.; Lucic, B.; Trinajstic, N. SAR and QSAR of the antioxidant activity of flavonoids. Curr. Med. Chem. 2007, 14, 827–845. [Google Scholar] [CrossRef]
- Mazza, G.; Brouillard, R. Recent developments in the stabilization of anthocyanins in food products. Food Chem. 1987, 25, 207–225. [Google Scholar] [CrossRef]
- Amić, D.; Lučić, B. Reliability of bond dissociation enthalpy calculated by the PM6 method and experimental teac values in antiradical QSAR of flavonoids. Bioorg. Med. Chem. 2010, 18, 28–35. [Google Scholar] [CrossRef]
- Buyukbingol, E.; Sisman, A.; Akyildiz, M.; Alparslan, F.N.; Adejare, A. Adaptive neuro-fuzzy inference system (ANFIS): A new approach to predictive modeling in QSAR applications: A study of neuro-fuzzy modeling of pcp-based NMDA receptor antagonists. Bioorg. Med. Chem. 2007, 15, 4265–4282. [Google Scholar] [CrossRef]
- Kähkönen, M.P.; Heinonen, M. Antioxidant activity of anthocyanins and their aglycons. J. Agric. Food Chem. 2003, 51, 628–633. [Google Scholar] [CrossRef]
- Karelson, M.; Lobanov, V.S.; Katritzky, A.R. Quantum-chemical descriptors in QSAR/QSPR studies. Chem. Rev. 1996, 96, 1027–1044. [Google Scholar] [CrossRef]
- Borkowski, T.; Szymusiak, H.; Gliszczynska-Swiglo, A.; Rietjens, I.M.C.M.; Tyrakowska, B. Radical scavenging capacity of wine anthocyanins is strongly pH-dependent. J. Agric. Food Chem. 2005, 53, 5526–5534. [Google Scholar] [CrossRef]
- Trouillas, P.; Marsal, P.; Siri, D.; Lazzaroni, R.; Duroux, J.-L. A DFT study of the reactivity of OH groups in quercetin and taxifolin antioxidants: The specificity of the 3-OH site. Food Chem. 2006, 97, 679–688. [Google Scholar] [CrossRef]
- Kozlowski, D.; Marsal, P.; Steel, M.; Mokrini, R.; Duroux, J.-L.; Lazzaroni, R.; Trouillas, P. Theoretical investigation of the formation of a new series of antioxidant depsides from the radiolysis of flavonoid compounds. Radiat. Res. 2007, 168, 243–252. [Google Scholar] [CrossRef]
- Mendoza-Wilson, A.M.; Lardizabal-Gutiérrez, D.; Torres-Moye, E.; Fuentes-Cobas, L.; Balandrán-Quintana, R.R.; Camacho-Dávila, A.; Quintero-Ramos, A.; Glossman-Mitnik, D. Optimized structure and thermochemical properties of flavonoids determined by the chih (medium)–DFT model chemistry versus experimental techniques. J. Mol. Struct. 2007, 871, 114–130. [Google Scholar] [CrossRef]
- Osman, A. Multiple pathways of the reaction of 2,2-diphenyl-1-picrylhydrazyl radical (DPPH') with (+)-catechin: Evidence for the formation of a covalent adduct between DPPH' and the oxidized form of the polyphenol. Biochem. Biophys. Res. Commun. 2011, 412, 473–478. [Google Scholar] [CrossRef]
- Matsufuji, H.; Kido, H.; Misawa, H.; Yaguchi, J.; Otsuki, T.; Chino, M.; Takeda, M.; Yamagata, K. Stability to light, heat, and hydrogen peroxide at different pH values and DPPH radical scavenging activity of acylated anthocyanins from red radish extract. J. Agric. Food Chem. 2007, 55, 3692–3701. [Google Scholar] [CrossRef]
- Pasha, F.; Srivastava, H.; Singh, P. Comparative QSAR study of phenol derivatives with the help of density functional theory. Bioorg. Med. Chem. 2005, 13, 6823–6829. [Google Scholar] [CrossRef]
- Chang, H.-J.; Choi, E.H.; Chun, H.S. Quantitative structure-activity relationship (QSAR) of antioxidative anthocyanidins and their glycosides. Food Sci. Biotechnol. 2008, 17, 501–507. [Google Scholar]
- Ghoush, M.A.; Samhouri, M.; Al-Holy, M.; Herald, T. Formulation and fuzzy modeling of emulsion stability and viscosity of a gum–protein emulsifier in a model mayonnaise system. J. Food Eng. 2008, 84, 348–357. [Google Scholar] [CrossRef]
- Amiryousefi, M.R.; Mohebbi, M.; Khodaiyan, F.; Asadi, S. An empowered adaptive neuro-fuzzy inference system using self-organizing map clustering to predict mass transfer kinetics in deep-fat frying of ostrich meat plates. Comput. Electron. Agric. 2011, 76, 89–95. [Google Scholar] [CrossRef]
- Lin, T.-C.; Liu, H.-L.; Kuo, M.-J. Direct adaptive interval type-2 fuzzy control of multivariable nonlinear systems. Eng. Appl. Artif. Intell. 2009, 22, 420–430. [Google Scholar] [CrossRef]
- Jang, J.; Sun, C.; Mizutani, E. Fuzzy inference systems. In Neuro-Fuzzy and Soft Computing: A Computational Approach to Learning and Machine Intelligence; Prentice-Hall: Upper Saddle River, NJ, USA, 1997; pp. 73–91. [Google Scholar]
- Allouche, A.R. Gabedit—A graphical user interface for computational chemistry softwares. J. Comput. Chem. 2010, 32, 174–182. [Google Scholar] [CrossRef]
- Stewart, J.J.P. Optimization of parameters for semiempirical methods V: Modification of NDDO approximations and application to 70 elements. J. Mol. Model. 2007, 13, 1173–1213. [Google Scholar] [CrossRef]
- Stewart, J.J. Optimization of parameters for semiempirical methods VI: More modifications to the NDDO approximations and re-optimization of parameters. J. Mol. Model. 2013, 19, 1–32. [Google Scholar] [CrossRef]
- Stewart, J.J.P. Mopac2012. Available online: http://OpenMOPAC.net (accessed on 12 April 2014).
- The R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2008. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Jhin, C.; Hwang, K.T. Prediction of Radical Scavenging Activities of Anthocyanins Applying Adaptive Neuro-Fuzzy Inference System (ANFIS) with Quantum Chemical Descriptors. Int. J. Mol. Sci. 2014, 15, 14715-14727. https://doi.org/10.3390/ijms150814715
Jhin C, Hwang KT. Prediction of Radical Scavenging Activities of Anthocyanins Applying Adaptive Neuro-Fuzzy Inference System (ANFIS) with Quantum Chemical Descriptors. International Journal of Molecular Sciences. 2014; 15(8):14715-14727. https://doi.org/10.3390/ijms150814715
Chicago/Turabian StyleJhin, Changho, and Keum Taek Hwang. 2014. "Prediction of Radical Scavenging Activities of Anthocyanins Applying Adaptive Neuro-Fuzzy Inference System (ANFIS) with Quantum Chemical Descriptors" International Journal of Molecular Sciences 15, no. 8: 14715-14727. https://doi.org/10.3390/ijms150814715





