The Mitochondrion-Located Protein OsB12D1 Enhances Flooding Tolerance during Seed Germination and Early Seedling Growth in Rice
Abstract
:1. Introduction
2. Results
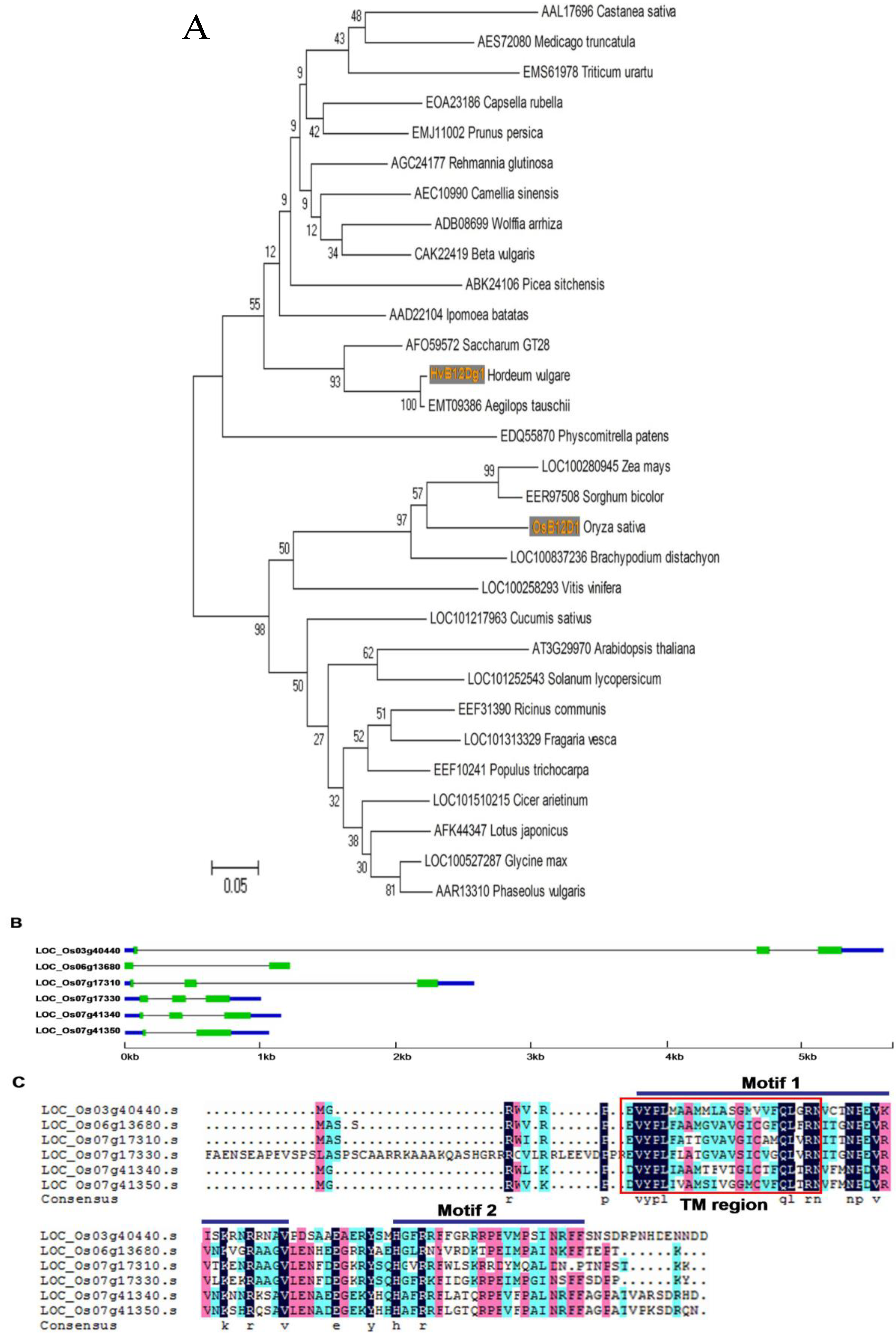
2.1. OsB12D1 Pertains to an Ancient Protein Family
2.2. Gene Structure and Expressional Patterns of OsB12Ds
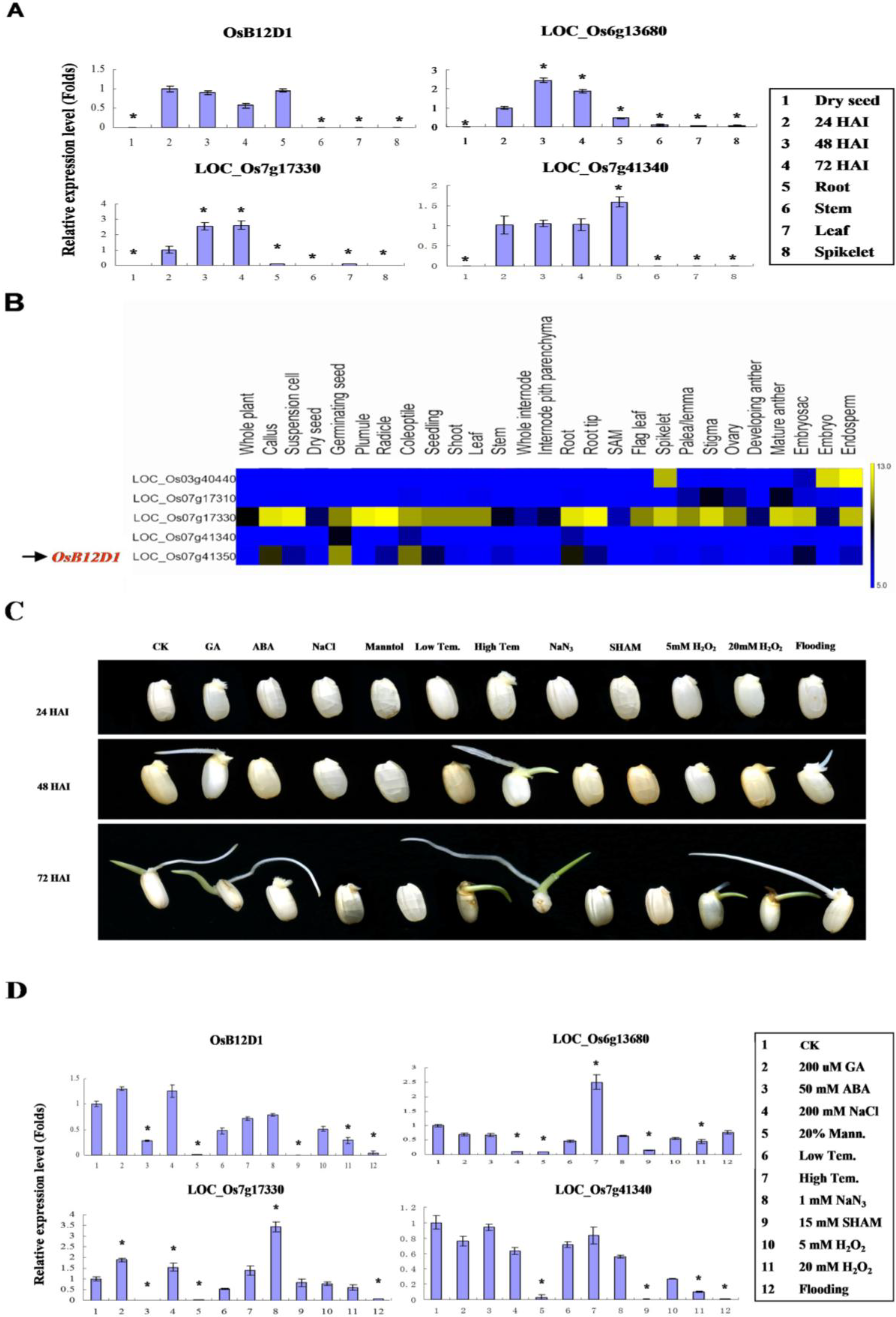
2.3. OsB12D1 Positively Responds to the Anoxia, Submergence and Flooding

2.4. Over Expressing of OsB12D1 Enhances Rice Flooding Tolerance during Seed Germination and Early Seedling Growth

2.5. OsB12D1 Is Mainly Located in the Mitochondrion and Might Enhance Electron Transfer
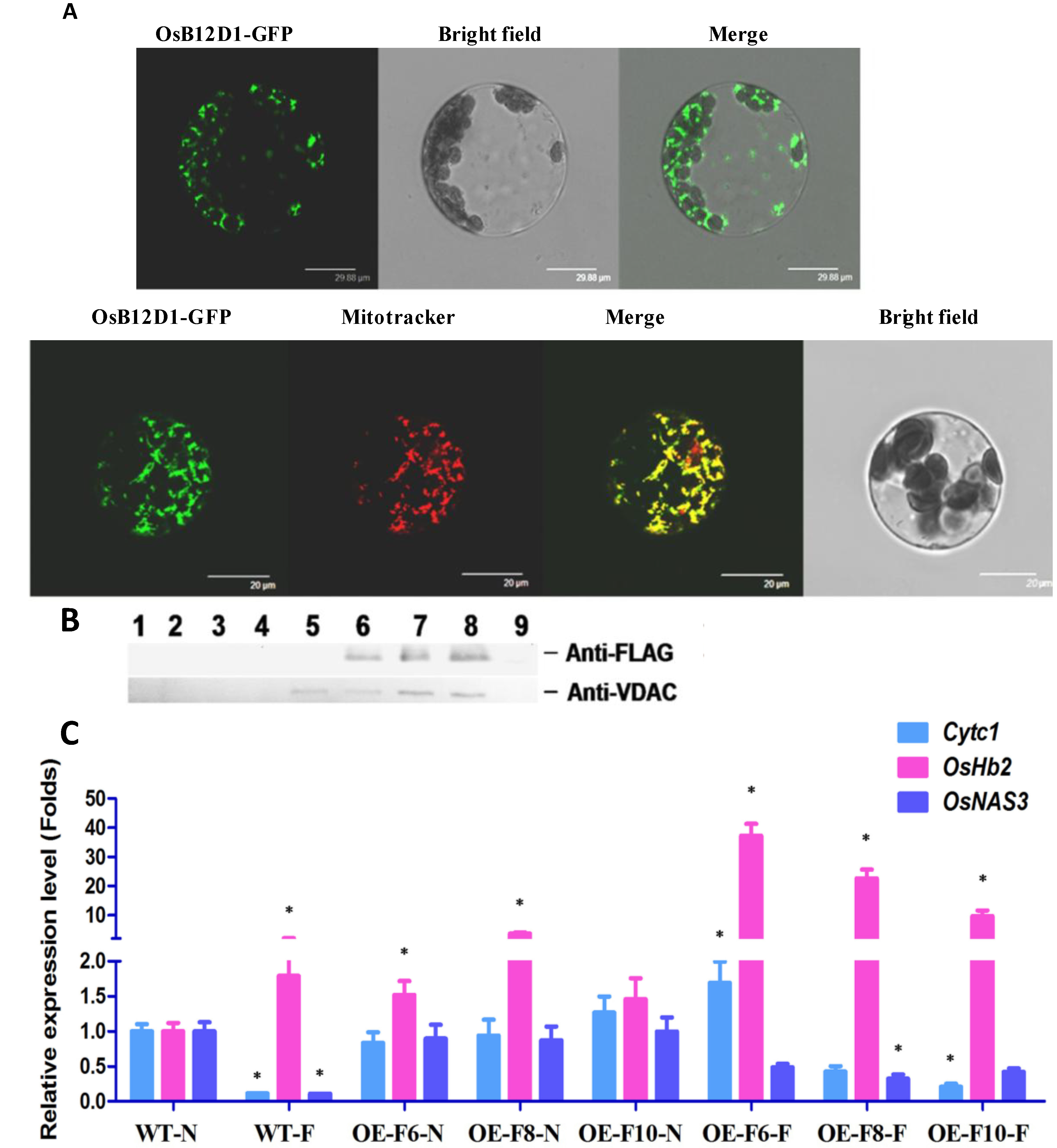
3. Discussion
4. Materials and Methods
4.1. Rice Seed Imbibition and Germinating Conditions
4.2. RNA Extraction and Quantitative Real-Time PCR
4.3. Generation of OsB12D1 Transgenic Rice
4.4. Protoplast Transient Expression Assay
4.5. Mitochondrial Isolation
4.6. Western Blot
4.7. Database Screening and Bioinformatics Analyses
5. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Farrant, J.M.; Moore, J.P. Programming desiccation-tolerance: From plants to seeds to resurrection plants. Curr. Opin. Plant Biol. 2011, 14, 340–345. [Google Scholar] [CrossRef]
- Rajjou, L.; Duval, M.; Gallardo, K.; Catusse, J.; Bally, J.; Job, C.; Job, D. Seed germination and vigor. Annu. Rev. Plant Biol. 2012, 63, 507–533. [Google Scholar] [CrossRef]
- Holdsworth, M.J.; Finch-Savage, W.E.; Grappin, P.; Job, D. Post-genomics dissection of seed dormancy and germination. Trends Plant Sci. 2008, 13, 7–13. [Google Scholar]
- Penfield, S.; King, J. Towards a systems biology approach to understanding seed dormancy and germination. Proc. R. Soc. B 2009, 276, 3561–3569. [Google Scholar] [CrossRef]
- Agrawal, G.K.; Yamazaki, M.; Kobayashi, M.; Hirochika, R.; Miyao, A.; Hirochika, H. Screening of the rice viviparous mutants generated by endogenous retrotransposon Tos17 insertion. Tagging of a zeaxanthin epoxidase gene and a novel ostatc gene. Plant Physiol. 2001, 125, 1248–1257. [Google Scholar] [CrossRef]
- Bentsink, L.; Jowett, J.; Hanhart, C.J.; Koornneef, M. Cloning of DOG1, a quantitative trait locus controlling seed dormancy in Arabidopsis. Proc. Natl. Acad. Sci. USA 2006, 103, 17042–17047. [Google Scholar]
- Guo, X.; Hou, X.; Fang, J.; Wei, P.; Xu, B.; Chen, M.; Feng, Y.; Chu, C. The rice GERMINATION DEFECTIVE 1, encoding a B3 domain transcriptional repressor, regulates seed germination and seedling development by integrating GA and carbohydrate metabolism. Plant J. Cell Mol. Biol. 2013. [Google Scholar] [CrossRef]
- Narsai, R.; Law, S.R.; Carrie, C.; Xu, L.; Whelan, J. In-depth temporal transcriptome profiling reveals a crucial developmental switch with roles for RNA processing and organelle metabolism that are essential for germination in Arabidopsis. Plant Physiol. 2011, 157, 1342–1362. [Google Scholar] [CrossRef]
- Narsai, R.; Howell, K.A.; Carroll, A.; Ivanova, A.; Millar, A.H.; Whelan, J. Defining core metabolic and transcriptomic responses to oxygen availability in rice embryos and young seedlings. Plant Physiol. 2009, 151, 306–322. [Google Scholar] [CrossRef]
- Howell, K.A.; Narsai, R.; Carroll, A.; Ivanova, A.; Lohse, M.; Usadel, B.; Millar, A.H.; Whelan, J. Mapping metabolic and transcript temporal switches during germination in rice highlights specific transcription factors and the role of RNA instability in the germination process. Plant Physiol. 2009, 149, 961–980. [Google Scholar]
- He, D.; Han, C.; Yao, J.; Shen, S.; Yang, P. Constructing the metabolic and regulatory pathways in germinating rice seeds through proteomic approach. Proteomics 2011, 11, 2693–2713. [Google Scholar] [CrossRef]
- Kim, S.T.; Kang, S.Y.; Wang, Y.; Kim, S.G.; Hwang du, H.; Kang, K.Y. Analysis of embryonic proteome modulation by GA and ABA from germinating rice seeds. Proteomics 2008, 8, 3577–3587. [Google Scholar] [CrossRef]
- Zhu, Q.; Zhang, J.; Gao, X.; Tong, J.; Xiao, L.; Li, W.; Zhang, H. The Arabidopsis AP2/ERF transcription factor RAP2.6 participates in ABA, salt and osmotic stress responses. Gene 2010, 457, 1–12. [Google Scholar] [CrossRef]
- Ram, P.C.; Singh, A.K.; Singh, B.B.; Singh, V.K.; Singh, H.P.; Setter, T.L.; Singh, V.P.; Singh, R.K. Environmental characterization of floodwater in Eastern India: Relevance to submergence tolerance of lowland rice. Exp. Agric. 1999, 35, 141–152. [Google Scholar] [CrossRef]
- Ismail, A.M.; Ella, E.S.; Vergara, G.V.; Mackill, D.J. Mechanisms associated with tolerance to flooding during germination and early seedling growth in rice (Oryza sativa). Ann. Bot. 2009, 103, 197–209. [Google Scholar]
- Satler, S.O.; Kende, H. Ethylene and the growth of rice seedlings. Plant Physiol. 1985, 79, 194–198. [Google Scholar] [CrossRef]
- Azuma, T.; Honda, T.; Sadai, A.; Sasayama, D.; Itoh, K. Suppression and promotion of growth by ethylene in rice seedlings depends on ambient humidity. J. Plant Physiol. 2007, 164, 1683–1687. [Google Scholar] [CrossRef]
- Fukao, T.; Xu, K.N.; Ronald, P.C.; Bailey-Serres, J. A variable cluster of ethylene response factor-like genes regulates metabolic and developmental acclimation responses to submergence in rice(W). Plant Cell 2006, 18, 2021–2034. [Google Scholar] [CrossRef]
- Aalen, R.B.; Opsahlferstad, H.G.; Linnestad, C.; Olsen, O.A. Transcripts encoding an oleosin and a dormancy-related protein are present in both the aleurone layer and the embryo of developing barley (Hordeum-vulgare L.) seeds. Plant J. 1994, 5, 385–396. [Google Scholar]
- Jakobsen, K.; Klemsdal, S.S.; Aalen, R.B.; Bosnes, M.; Alexander, D.; Olsen, O.A. Barley aleurone cell development: Molecular cloning of aleurone-specific cDNAs from immature grains. Plant Mol. Biol. 1989, 12, 285–293. [Google Scholar] [CrossRef]
- Aalen, R.B. The transcripts encoding two oleosin isoforms are both present in the aleurone and in the embryo of barley (Hordeum vulgare L.) seeds. Plant Mol. Biol. 1995, 28, 583–588. [Google Scholar] [CrossRef]
- Olsen, O.A.; Jakobsen, K.S.; Schmelzer, E. Development of barley aleurone cells—Temporal and spatial patterns of accumulation of cell-specific messenger-rnas. Planta 1990, 181, 462–466. [Google Scholar]
- Steinum, T.M.; Berner, H.S.; Stacy, R.A.P.; Salehian, Z.; Aalen, R.B. Differential regulation of the barley (Hordeum vulgare) transcripts B22E and B12D in mature aleurone layers. Physiol. Plant. 1998, 102, 337–345. [Google Scholar] [CrossRef]
- Kall, L.; Krogh, A.; Sonnhammer, E.L.L. A combined transmembrane topology and signal peptide prediction method. J. Mol. Biol. 2004, 338, 1027–1036. [Google Scholar] [CrossRef]
- Cao, P.; Jung, K.H.; Choi, D.; Hwang, D.; Zhu, J.; Ronald, P.C. The Rice Oligonucleotide Array Database: An atlas of rice gene expression. Rice 2012, 5, 1939–8433. [Google Scholar]
- Mustroph, A.; Lee, S.C.; Oosumi, T.; Zanetti, M.E.; Yang, H.J.; Ma, K.; Yaghoubi-Masihi, A.; Fukao, T.; Bailey-Serres, J. Cross-kingdom comparison of transcriptomic adjustments to low-oxygen stress highlights conserved and plant-specific responses. Plant Physiol. 2010, 152, 1484–1500. [Google Scholar] [CrossRef]
- Bailey-Serres, J.; Voesenek, L.A.C.J. Flooding stress: Acclimations and genetic diversity. Annu. Rev. Plant Biol. 2008, 59, 313–339. [Google Scholar] [CrossRef]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic. Acids. Res. 2014, 42. [Google Scholar] [CrossRef]
- Zheng, L.Q.; Huang, F.L.; Narsai, R.; Wu, J.J.; Giraud, E.; He, F.; Cheng, L.J.; Wang, F.; Wu, P.; Whelan, J.; et al. Physiological and transcriptome analysis of iron and phosphorus interaction in rice seedlings. Plant Physiol. 2009, 151, 262–274. [Google Scholar] [CrossRef]
- Roy, S.W.; Gilbert, W. The evolution of spliceosomal introns: Patterns, puzzles and progress. Nat. Rev. Genet. 2006, 7, 211–221. [Google Scholar]
- Wu, Y.F.; Zhu, Z.G.; Ma, L.G.; Chen, M.S. The preferential retention of starch synthesis genes reveals the impact of whole-genome duplication on grass evolution. Mol. Biol. Evol. 2008, 25, 1003–1006. [Google Scholar] [CrossRef]
- Aalen, R.B.; Salehian, Z.; Steinum, T.M. Stability of barley aleurone transcripts: Dependence on protein synthesis, influence of the starchy endosperm and destabilization by GA3. Physiol. Plant. 2001, 112, 403–413. [Google Scholar] [CrossRef]
- Lasanthi-Kudahettige, R.; Magneschi, L.; Loreti, E.; Gonzali, S.; Licausi, F.; Novi, G.; Beretta, O.; Vitulli, F.; Alpi, A.; Perata, P. Transcript profiling of the anoxic rice coleoptile. Plant Physiol. 2007, 144, 218–231. [Google Scholar] [CrossRef]
- Jackson, M.B.; Colmer, T.D. Response and adaptation by plants to flooding stress —Preface. Ann. Bot. 2005, 96, 501–505. [Google Scholar] [CrossRef]
- Bewley, J.D. Seed germination and dormancy. Plant Cell 1997, 9, 1055–1066. [Google Scholar] [CrossRef]
- Kumar, V.; Ladha, J.K. Direct seeding of rice: Recent developments and future research needs. Adv. Agron. 2011, 111, 297–413. [Google Scholar] [CrossRef]
- Walker, J.E.; Arizmendi, J.M.; Dupuis, A.; Fearnley, I.M.; Finel, M.; Medd, S.M.; Pilkington, S.J.; Runswick, M.J.; Skehel, J.M. Sequences of 20 subunits of NADH:ubiquinone oxidoreductase from bovine heart mitochondria. Application of a novel strategy for sequencing proteins using the polymerase chain reaction. J. Mol. Biol. 1992, 226, 1051–1072. [Google Scholar] [CrossRef]
- Han, C.; Yang, P.; Sakata, K.; Komatsu, S. Quantitative proteomics reveals the role of protein phosphorylation in rice embryos during early stages of germination. J. Proteome Res. 2014, 13, 1766–1782. [Google Scholar] [CrossRef]
- Kanagarajah, D. The Molecular and Biochernical Characterization of the MLRQ Subunit of NADH: Ubiquinone Oxidoreductase in the Human Mitochondrial Respiratory Chain. Master Thesis, University of Toronto, Toronto, ON, Canada, 2001. [Google Scholar]
- Sowa, A.W.; Duff, S.M.; Guy, P.A.; Hill, R.D. Altering hemoglobin levels changes energy status in maize cells under hypoxia. Proc. Natl. Acad. Sci. USA 1998, 95, 10317–10321. [Google Scholar] [CrossRef]
- Vigeolas, H.; Huhn, D.; Geigenberger, P. Nonsymbiotic hemoglobin-2 leads to an elevated energy state and to a combined increase in polyunsaturated fatty acids and total oil content when overexpressed in developing seeds of transgenic arabidopsis plants. Plant Physiol. 2011, 155, 1435–1444. [Google Scholar] [CrossRef]
- Sharp, R.E.; LeNoble, M.E. ABA, ethylene and the control of shoot and root growth under water stress. J. Exp. Bot. 2002, 53, 33–37. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. Cell Mol. Biol. 1998, 16, 735–743. [Google Scholar] [CrossRef]
- Yoo, S.D.; Cho, Y.H.; Sheen, J. Arabidopsis mesophyll protoplasts: A versatile cell system for transient gene expression analysis. Nat. Protoc. 2007, 2, 1565–1572. [Google Scholar] [CrossRef]
- Howell, K.A.; Millar, A.H.; Whelan, J. Ordered assembly of mitochondria during rice germination begins with pro-mitochondrial structures rich in components of the protein import apparatus. Plant Mol. Biol. 2006, 60, 201–223. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Guo, A.Y.; Zhu, Q.H.; Chen, X.; Luo, J.C. GSDS: A gene structure display server. Yi Chuan Hereditas 2007, 29, 1023–1026. [Google Scholar] [CrossRef]
- Ma, H.; Zhao, J. Genome-wide identification, classification, and expression analysis of the arabinogalactan protein gene family in rice (Oryza sativa L.). J. Exp. Bot. 2010, 61, 2647–2668. [Google Scholar] [CrossRef]
- Bailey, T.L.; Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
He, D.; Zhang, H.; Yang, P. The Mitochondrion-Located Protein OsB12D1 Enhances Flooding Tolerance during Seed Germination and Early Seedling Growth in Rice. Int. J. Mol. Sci. 2014, 15, 13461-13481. https://doi.org/10.3390/ijms150813461
He D, Zhang H, Yang P. The Mitochondrion-Located Protein OsB12D1 Enhances Flooding Tolerance during Seed Germination and Early Seedling Growth in Rice. International Journal of Molecular Sciences. 2014; 15(8):13461-13481. https://doi.org/10.3390/ijms150813461
Chicago/Turabian StyleHe, Dongli, Hui Zhang, and Pingfang Yang. 2014. "The Mitochondrion-Located Protein OsB12D1 Enhances Flooding Tolerance during Seed Germination and Early Seedling Growth in Rice" International Journal of Molecular Sciences 15, no. 8: 13461-13481. https://doi.org/10.3390/ijms150813461




