Coactivator Recruitment of AhR/ARNT1
Abstract
:1. Basic Signal Transmission Mechanisms of Nuclear Receptors
2. AhR/ARNT-Related Transcription
3. AhR-Coactivator Interactions
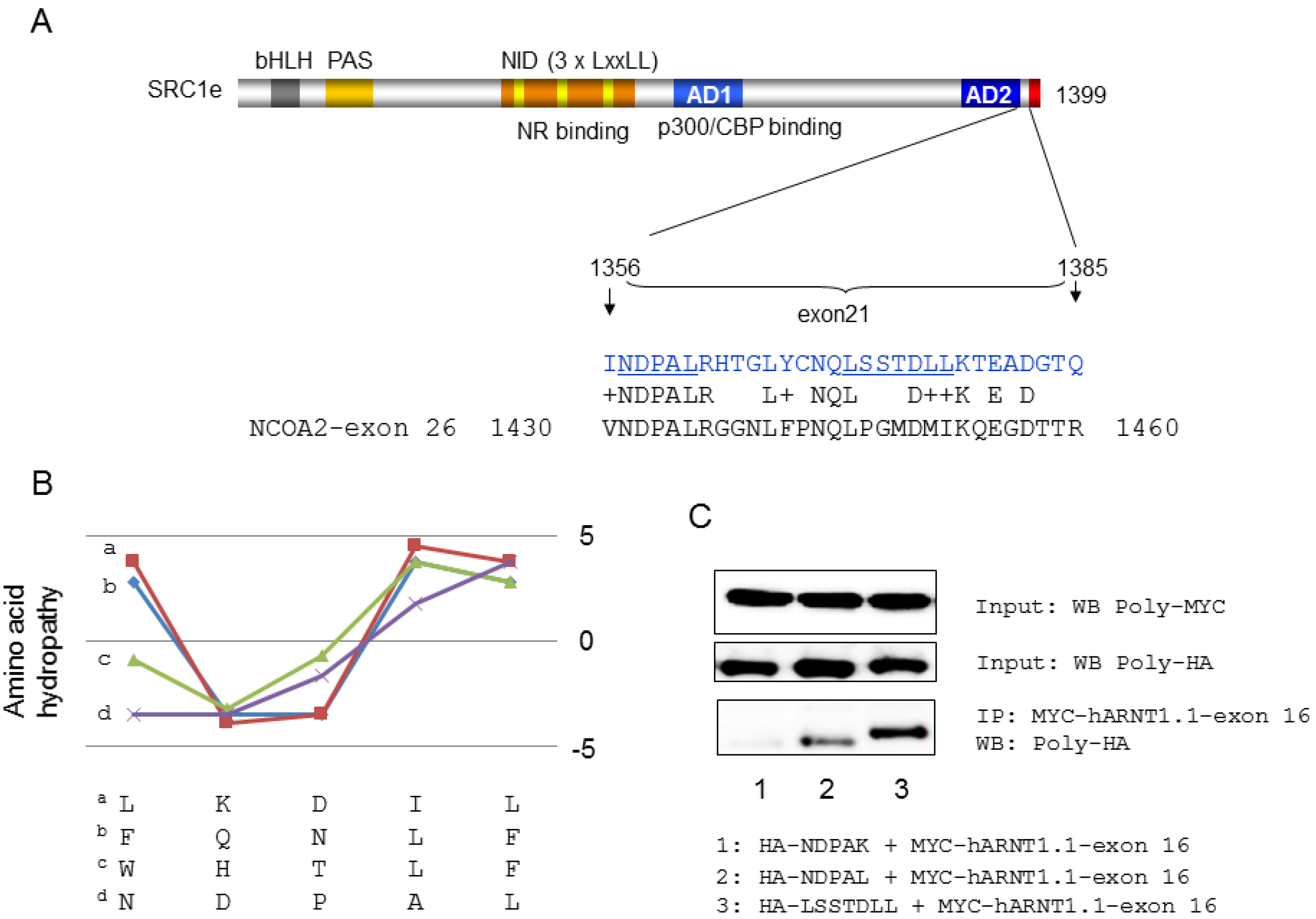
4. Coactivator Recruitment to the AhR/ARNT Complex


5. The Role of TCDD as Endocrine Disrupter
6. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Mangelsdorf, D.J.; Thummel, C.; Beato, M.; Herrlich, P.; Schutz, G.; Umesono, K.; Blumberg, B.; Kastner, P.; Mark, M.; Chambon, P.; et al. The nuclear receptor superfamily: The second decade. Cell 1995, 83, 835–839. [Google Scholar] [CrossRef]
- Germain, P.; Kammerer, S.; Perez, E.; Peluso-Iltis, C.; Tortolani, D.; Zusi, F.C.; Starrett, J.; Lapointe, P.; Daris, J.P.; Marinier, A.; et al. Rational design of RAR-selective ligands revealed by RARbeta crystal stucture. EMBO Rep. 2004, 5, 877–882. [Google Scholar]
- Pratt, W.B.; Toft, D.O. Regulation of signaling protein function and trafficking by the hsp90/hsp70-based chaperone machinery. Exp. Biol. Med. 2003, 228, 111–133. [Google Scholar]
- McInerney, E.M.; Rose, D.W.; Flynn, S.E.; Westin, S.; Mullen, T.M.; Krones, A.; Inostroza, J.; Torchia, J.; Nolte, R.T.; Assa-Munt, N.; et al. Determinants of coactivator LXXLL motif specificity in nuclear receptor transcriptional activation. Genes Dev. 1998, 12, 3357–3368. [Google Scholar] [CrossRef]
- Spencer, T.E.; Jenster, G.; Burcin, M.M.; Allis, C.D.; Zhou, J.; Mizzen, C.A.; McKenna, N.J.; Onate, S.A.; Tsai, S.Y.; Tsai, M.J.; et al. Steroid receptor coactivator-1 is a histone acetyltransferase. Nature 1997, 389, 194–198. [Google Scholar] [CrossRef]
- Sterner, D.E.; Berger, S.L. Acetylation of histones and transcription-related factors. Microbiol. Mol. Biol. Rev. 2000, 64, 435–459. [Google Scholar] [CrossRef]
- Sheppard, H.M.; Harries, J.C.; Hussain, S.; Bevan, C.; Heery, D.M. Analysis of the steroid receptor coactivator 1 (SRC1)-CREB binding protein interaction interface and its importance for the function of SRC1. Mol. Cell. Biol. 2001, 21, 39–50. [Google Scholar] [CrossRef]
- Vo, N.; Goodman, R.H. CREB-binding protein and p300 in transcriptional regulation. J. Biol. Chem. 2001, 276, 13505–13508. [Google Scholar]
- Kazlauskas, A.; Sundstrom, S.; Poellinger, L.; Pongratz, I. The hsp90 chaperone complex regulates intracellular localization of the dioxin receptor. Mol. Cell. Biol. 2001, 21, 2594–2607. [Google Scholar] [CrossRef]
- Carver, L.A.; LaPres, J.J.; Jain, S.; Dunham, E.E.; Bradfield, C.A. Characterization of the Ah receptor-associated protein, ARA9. J. Biol. Chem. 1998, 273, 33580–33587. [Google Scholar]
- Meyer, B.K.; Pray-Grant, M.G.; Vanden Heuvel, J.P.; Perdew, G.H. Hepatitis B virus X-associated protein 2 is a subunit of the unliganded aryl hydrocarbon receptor core complex and exhibits transcriptional enhancer activity. Mol. Cell. Biol. 1998, 18, 978–988. [Google Scholar]
- Whitlock, J.P., Jr. Induction of cytochrome P4501A1. Annu. Rev. Pharmacol. Toxicol. 1999, 39, 103–125. [Google Scholar] [CrossRef]
- Opitz, C.A.; Litzenburger, U.M.; Sahm, F.; Ott, M.; Tritschler, I.; Trump, S.; Schumacher, T.; Jestaedt, L.; Schrenk, D.; Weller, M.; et al. An endogenous tumour-promoting ligand of the human aryl hydrocarbon receptor. Nature 2011, 478, 197–203. [Google Scholar] [CrossRef] [Green Version]
- Adachi, J.; Mori, Y.; Matsui, S.; Takigami, H.; Fujino, J.; Kitagawa, H.; Miller, C.A., 3rd; Kato, T.; Saeki, K.; Matsuda, T. Indirubin and indigo are potent aryl hydrocarbon receptor ligands present in human urine. J. Biol. Chem. 2001, 276, 31475–31478. [Google Scholar] [CrossRef]
- Sinal, C.J.; Bend, J.R. Aryl hydrocarbon receptor-dependent induction of cyp1a1 by bilirubin in mouse hepatoma hepa 1c1c7 cells. Mol. Pharmacol. 1997, 52, 590–599. [Google Scholar]
- Seidel, S.D.; Winters, G.M.; Rogers, W.J.; Ziccardi, M.H.; Li, V.; Keser, B.; Denison, M.S. Activation of the Ah receptor signaling pathway by prostaglandins. J. Biochem. Mol. Toxicol. 2001, 15, 187–196. [Google Scholar] [CrossRef]
- Gonzalez, F.J.; Nebert, D.W. Evolution of the P450 gene superfamily: Animal-plant “warfare”, molecular drive and human genetic differences in drug oxidation. Trends Genet. 1990, 6, 182–186. [Google Scholar]
- Denison, M.S.; Whitlock, J.P., Jr. Xenobiotic-inducible transcription of cytochrome P450 genes. J. Biol. Chem. 1995, 270, 18175–18178. [Google Scholar] [CrossRef]
- Nebert, D.W. Proposed role of drug-metabolizing enzymes: Regulation of steady state levels of the ligands that effect growth, homeostasis, differentiation, and neuroendocrine functions. Mol. Endocrinol. 1991, 5, 1203–1214. [Google Scholar] [CrossRef]
- Kim, J.H.; Stallcup, M.R. Role of the coiled-coil coactivator (CoCoA) in aryl hydrocarbon receptor-mediated transcription. J. Biol. Chem. 2004, 279, 49842–49848. [Google Scholar] [CrossRef]
- Kumar, M.B.; Perdew, G.H. Nuclear receptor coactivator SRC-1 interacts with the Q-rich subdomain of the AhR and modulates its transactivation potential. Gene Exp. 1999, 8, 273–286. [Google Scholar]
- Hestermann, E.V.; Brown, M. Agonist and chemopreventative ligands induce differential transcriptional cofactor recruitment by aryl hydrocarbon receptor. Mol. Cell. Biol. 2003, 23, 7920–7925. [Google Scholar] [CrossRef]
- Endler, A.; Chen, L.; Zhang, J.; Xu, G.T.; Shibasaki, F. Binding of the ERalpha and ARNT1 AF2 domains to exon 21 of the SRC1 isoform SRC1e is essential for estrogen- and dioxin-related transcription. J. Cell Sci. 2012, 125, 2004–2016. [Google Scholar]
- Lipford, J.R.; Deshaies, R.J. Diverse roles for ubiquitin-dependent proteolysis in transcriptional activation. Nat. Cell Biol. 2003, 5, 845–850. [Google Scholar] [CrossRef]
- Hur, E.; Pfaff, S.J.; Payne, E.S.; Gron, H.; Buehrer, B.M.; Fletterick, R.J. Recognition and accommodation at the androgen receptor coactivator binding interface. PLoS Biol. 2004, 2, E274. [Google Scholar] [CrossRef] [Green Version]
- Burd, C.J.; Petre, C.E.; Moghadam, H.; Wilson, E.M.; Knudsen, K.E. Cyclin D1 binding to the androgen receptor (AR) NH2-terminal domain inhibits activation function 2 association and reveals dual roles for AR corepression. Mol. Endocrinol. 2005, 19, 607–620. [Google Scholar] [CrossRef]
- Tetel, M.J.; Giangrande, P.H.; Leonhardt, S.A.; McDonnell, D.P.; Edwards, D.P. Hormone-dependent interaction between the amino- and carboxyl-terminal domains of progesterone receptor in vitro and in vivo. Mol. Endocrinol. 1999, 13, 910–924. [Google Scholar] [CrossRef]
- Alen, P.; Claessens, F.; Verhoeven, G.; Rombauts, W.; Peeters, B. The androgen receptor amino-terminal domain plays a key role in p160 coactivator-stimulated gene transcription. Mol. Cell. Biol. 1999, 19, 6085–6097. [Google Scholar]
- Beischlag, T.V.; Wang, S.; Rose, D.W.; Torchia, J.; Reisz-Porszasz, S.; Muhammad, K.; Nelson, W.E.; Probst, M.R.; Rosenfeld, M.G.; Hankinson, O. Recruitment of the NCoA/SRC-1/p160 family of transcriptional coactivators by the aryl hydrocarbon receptor/aryl hydrocarbon receptor nuclear translocator complex. Mol. Cell. Biol. 2002, 22, 4319–4333. [Google Scholar] [CrossRef]
- Hong, H.; Kohli, K.; Trivedi, A.; Johnson, D.L.; Stallcup, M.R. GRIP1, a novel mouse protein that serves as a transcriptional coactivator in yeast for the hormone binding domains of steroid receptors. Proc. Natl. Acad. Sci. USA 1996, 93, 4948–4952. [Google Scholar]
- Robinson-Rechavi, M.; Carpentier, A.S.; Duffraisse, M.; Laudet, V. How many nuclear hormone receptors are there in the human genome? Trends Genet. 2001, 17, 554–556. [Google Scholar] [CrossRef]
- Shimizu, Y.; Nakatsuru, Y.; Ichinose, M.; Takahashi, Y.; Kume, H.; Mimura, J.; Fujii-Kuriyama, Y.; Ishikawa, T. Benzo[a]pyrene carcinogenicity is lost in mice lacking the aryl hydrocarbon receptor. Proc. Natl. Acad. Sci. USA 2000, 97, 779–782. [Google Scholar] [CrossRef]
- Sorg, O.; Zennegg, M.; Schmid, P.; Fedosyuk, R.; Valikhnovskyi, R.; Gaide, O.; Kniazevych, V.; Saurat, J.H. 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) poisoning in Victor Yushchenko: Identification and measurement of TCDD metabolites. Lancet 2009, 374, 1179–1185. [Google Scholar] [CrossRef]
- Wormke, M.; Stoner, M.; Saville, B.; Walker, K.; Abdelrahim, M.; Burghardt, R.; Safe, S. The aryl hydrocarbon receptor mediates degradation of estrogen receptor alpha through activation of proteasomes. Mol. Cell. Biol. 2003, 23, 1843–1855. [Google Scholar]
- Matthews, J.; Wihlen, B.; Thomsen, J.; Gustafsson, J.A. Aryl hydrocarbon receptor-mediated transcription: Ligand-dependent recruitment of estrogen receptor alpha to 2,3,7,8-tetrachlorodibenzo-p-dioxin-responsive promoters. Mol. Cell. Biol. 2005, 25, 5317–5328. [Google Scholar]
- Beischlag, T.V.; Perdew, G.H. ER alpha-AHR-ARNT protein-protein interactions mediate estradiol-dependent transrepression of dioxin-inducible gene transcription. J. Biol. Chem. 2005, 280, 21607–21611. [Google Scholar] [CrossRef]
- Ohtake, F.; Takeyama, K.; Matsumoto, T.; Kitagawa, H.; Yamamoto, Y.; Nohara, K.; Tohyama, C.; Krust, A.; Mimura, J.; Chambon, P.; et al. Modulation of oestrogen receptor signalling by association with the activated dioxin receptor. Nature 2003, 423, 545–550. [Google Scholar]
- Abdelrahim, M.; Ariazi, E.; Kim, K.; Khan, S.; Barhoumi, R.; Burghardt, R.; Liu, S.; Hill, D.; Finnell, R.; Wlodarczyk, B.; et al. 3-Methylcholanthrene and other aryl hydrocarbon receptor agonists directly activate estrogen receptor alpha. Cancer Res. 2006, 66, 2459–2467. [Google Scholar]
- Min, G.; Kim, H.; Bae, Y.; Petz, L.; Kemper, J.K. Inhibitory cross-talk between estrogen receptor (ER) and constitutively activated androstane receptor (CAR). CAR inhibits ER-mediated signaling pathway by squelching p160 coactivators. J. Biol. Chem. 2002, 277, 34626–34633. [Google Scholar] [CrossRef]
- Reen, R.K.; Cadwallader, A.; Perdew, G.H. The subdomains of the transactivation domain of the aryl hydrocarbon receptor (AhR) inhibit AhR and estrogen receptor transcriptional activity. Arch. Biochem. Biophys. 2002, 408, 93–102. [Google Scholar] [CrossRef]
- Spiegelman, B.M.; Heinrich, R. Biological control through regulated transcriptional coactivators. Cell 2004, 119, 157–167. [Google Scholar] [CrossRef]
- Wenger, R.H. Cellular adaptation to hypoxia: O2-sensing protein hydroxylases, hypoxia-inducible transcription factors, and O2-regulated gene expression. FASEB J. 2002, 16, 1151–1162. [Google Scholar] [CrossRef]
- Brunnberg, S.; Pettersson, K.; Rydin, E.; Matthews, J.; Hanberg, A.; Pongratz, I. The basic helix-loop-helix-PAS protein ARNT functions as a potent coactivator of estrogen receptor-dependent transcription. Proc. Natl. Acad. Sci. USA 2003, 100, 6517–6522. [Google Scholar] [CrossRef]
- Labrecque, M.P.; Takhar, M.K.; Hollingshead, B.D.; Prefontaine, G.G.; Perdew, G.H.; Beischlag, T.V. Distinct roles for aryl hydrocarbon receptor nuclear translocator and ah receptor in estrogen-mediated signaling in human cancer cell lines. PLoS One 2012, 7, e29545. [Google Scholar]
- Ruegg, J.; Swedenborg, E.; Wahlstrom, D.; Escande, A.; Balaguer, P.; Pettersson, K.; Pongratz, I. The transcription factor aryl hydrocarbon receptor nuclear translocator functions as an estrogen receptor beta-selective coactivator, and its recruitment to alternative pathways mediates antiestrogenic effects of dioxin. Mol. Endocrinol. 2008, 22, 304–316. [Google Scholar] [CrossRef]
- Pollenz, R.S.; Davarinos, N.A.; Shearer, T.P. Analysis of aryl hydrocarbon receptor-mediated signaling during physiological hypoxia reveals lack of competition for the aryl hydrocarbon nuclear translocator transcription factor. Mol. Pharmacol. 1999, 56, 1127–1137. [Google Scholar]
- Amazit, L.; Alj, Y.; Tyagi, R.K.; Chauchereau, A.; Loosfelt, H.; Pichon, C.; Pantel, J.; Foulon-Guinchard, E.; Leclerc, P.; Milgrom, E.; et al. Subcellular localization and mechanisms of nucleocytoplasmic trafficking of steroid receptor coactivator-1. J. Biol. Chem. 2003, 278, 32195–32203. [Google Scholar] [CrossRef]
- Maruvada, P.; Baumann, C.T.; Hager, G.L.; Yen, P.M. Dynamic shuttling and intranuclear mobility of nuclear hormone receptors. J. Biol. Chem. 2003, 278, 12425–12432. [Google Scholar]
- Matthews, J.; Gustafsson, J.A. Estrogen receptor and aryl hydrocarbon receptor signaling pathways. Nucl. Recept. Signal 2006, 4, e016. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Endler, A.; Chen, L.; Shibasaki, F. Coactivator Recruitment of AhR/ARNT1. Int. J. Mol. Sci. 2014, 15, 11100-11110. https://doi.org/10.3390/ijms150611100
Endler A, Chen L, Shibasaki F. Coactivator Recruitment of AhR/ARNT1. International Journal of Molecular Sciences. 2014; 15(6):11100-11110. https://doi.org/10.3390/ijms150611100
Chicago/Turabian StyleEndler, Alexander, Li Chen, and Futoshi Shibasaki. 2014. "Coactivator Recruitment of AhR/ARNT1" International Journal of Molecular Sciences 15, no. 6: 11100-11110. https://doi.org/10.3390/ijms150611100



