Effects of RNAi-Mediated Knockdown of Histone Methyltransferases on the Sex-Specific mRNA Expression of Imp in the Silkworm Bombyx mori
Abstract
:1. Introduction
2. Results and Discussion
2.1. Results
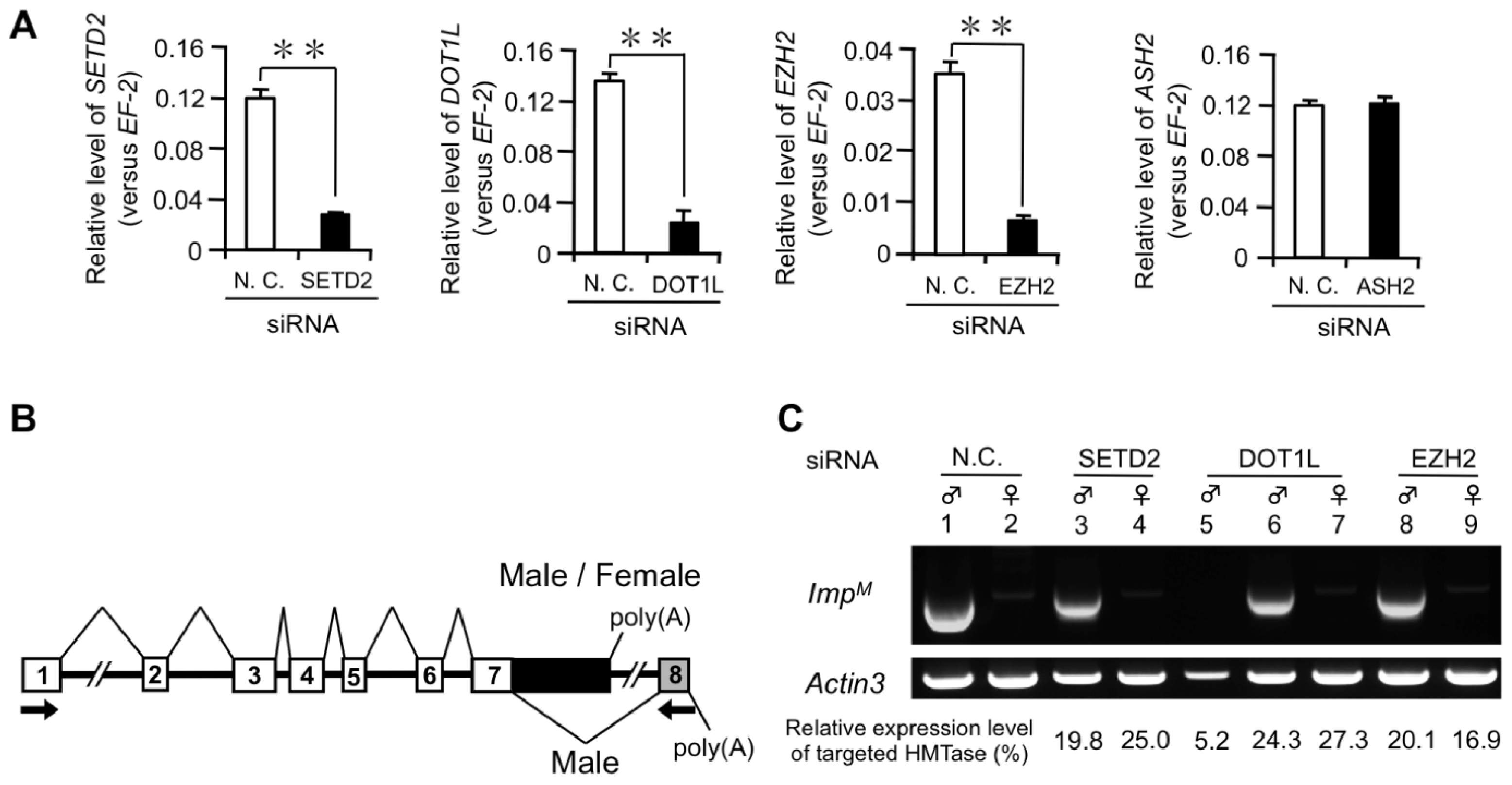
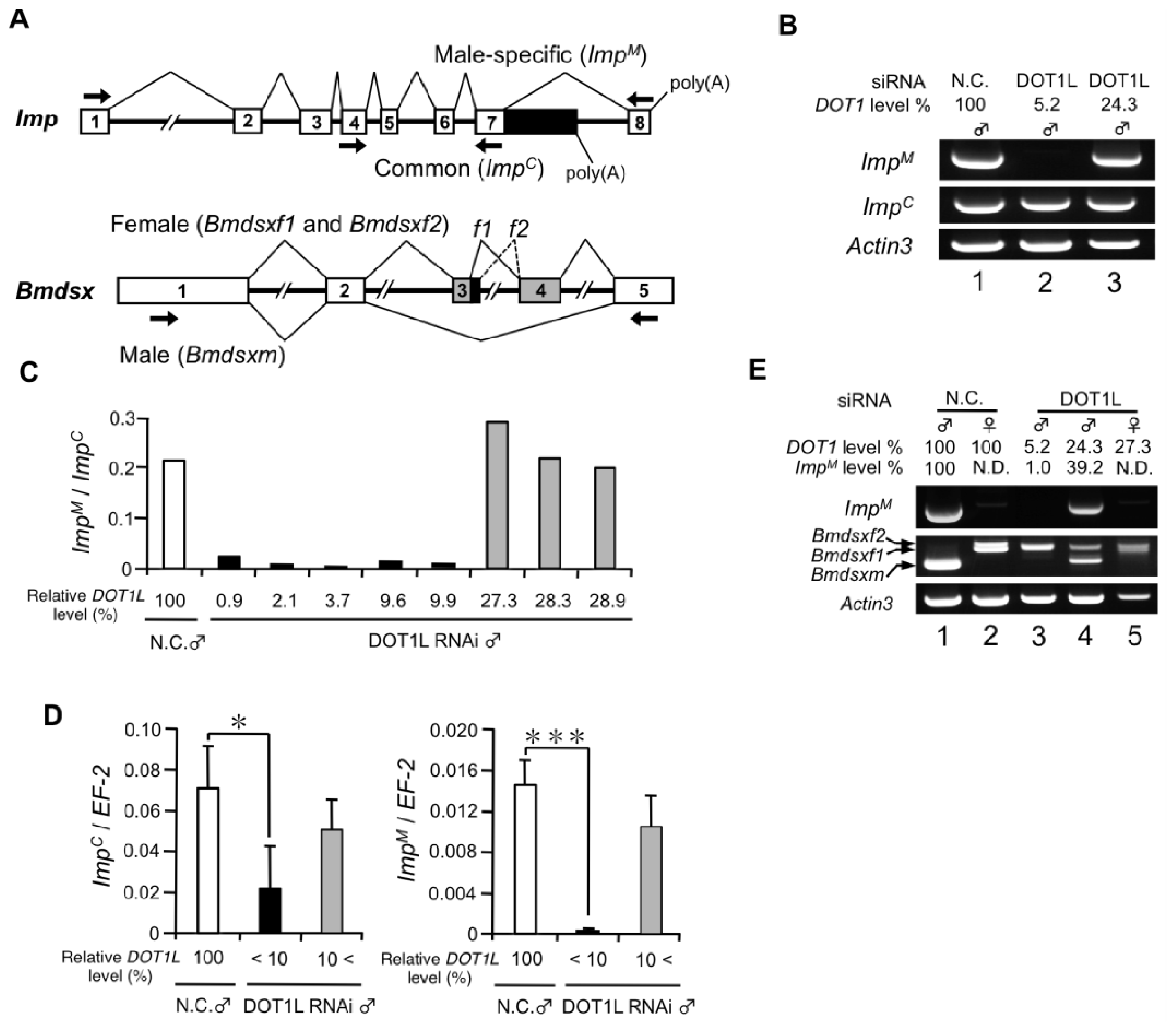
2.1.1. Knockdown of DOT1L Abolished Male-Specific Expression of the Imp mRNA
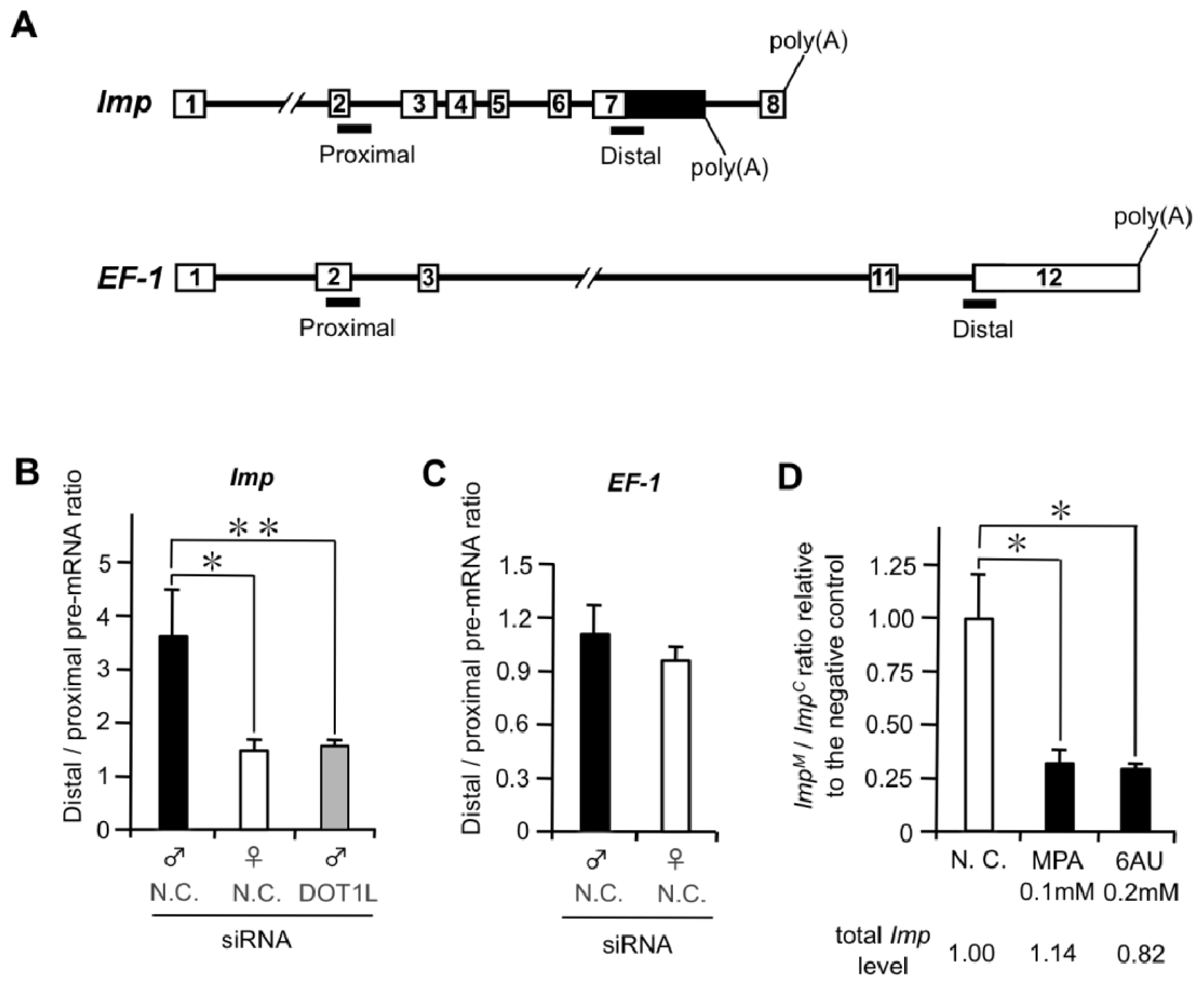
2.1.2. DOT1L Knockdown Affects Male-Specific Splicing of Imp Pre-mRNA
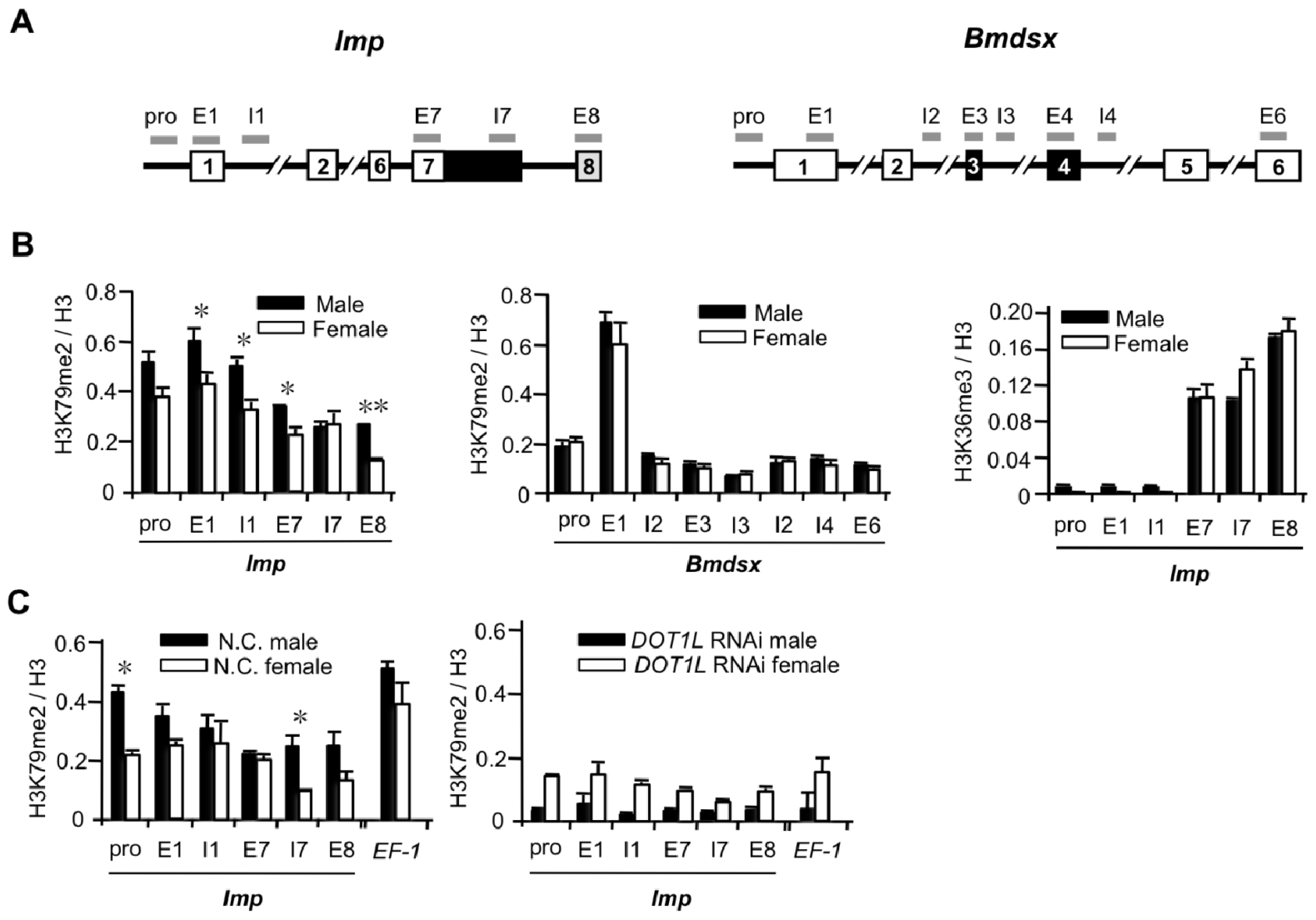
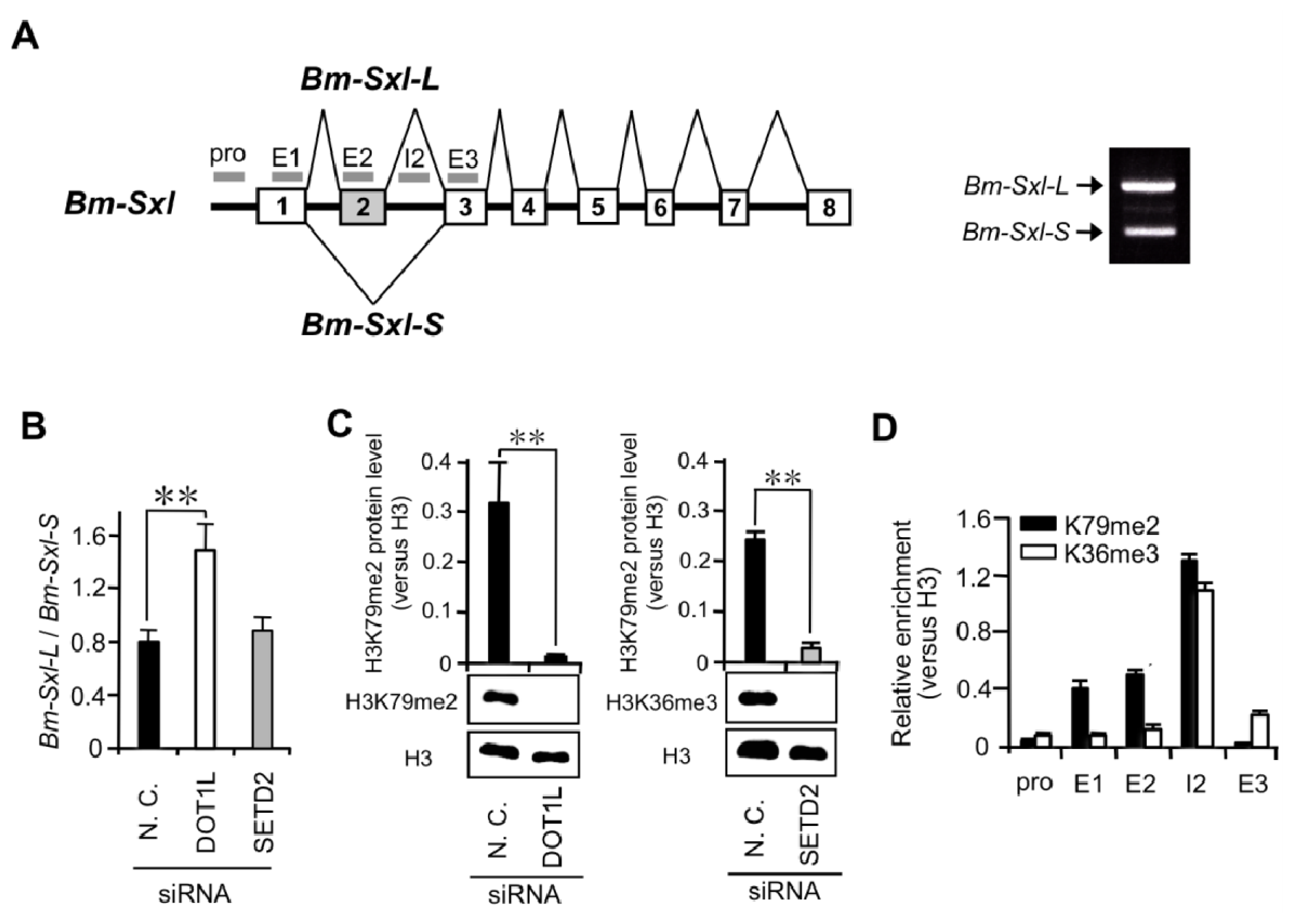
2.1.3. High Levels of H3K79me2 Favor Inclusion of Male-Specific Exon of Imp
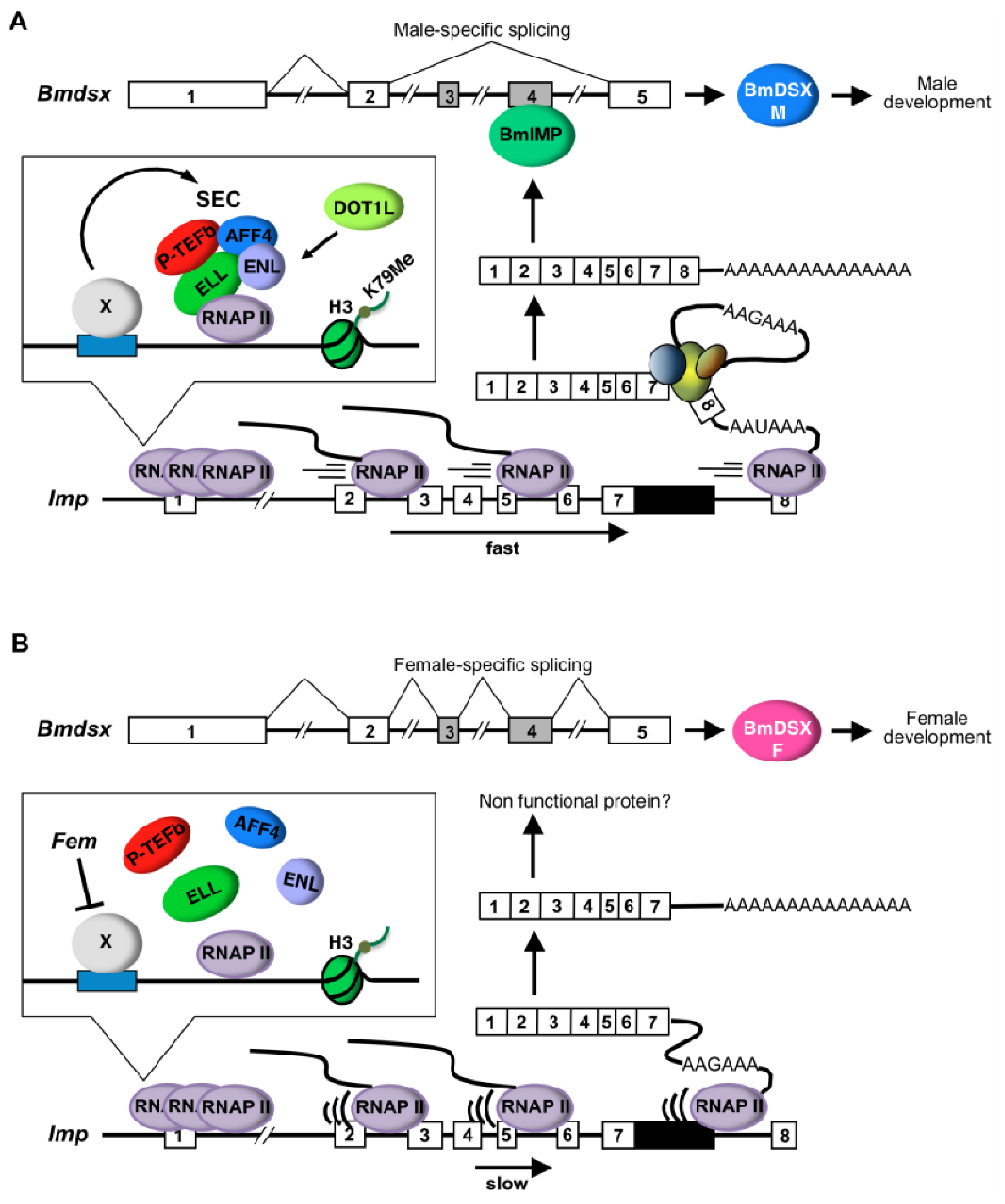
2.1.4. Male and Female Differences in RNAP II Processivity in the Imp Gene
2.1.5. Suppression of Male-Specific Imp Pre-mRNA Splicing by Inhibitors of Nucleotide Biosynthesis
2.1.6. Embryonic Lethality Caused by DOT1L Knockdown
2.2. Discussion
3. Experimental Section
3.1. Silkworm Strains
3.2. Preparation of siRNAs
3.3. Injection of siRNAs into Eggs
3.4. Extraction of Total RNA and Genomic DNA
3.5. Reverse Transcription (RT)-PCR Analyses
3.6. Quantitative Real-Time RT-PCR (qRT-PCR)
3.7. ChIP Experiments
3.8. Mycophenolic Acid (MPA) and 6-Azauracil (6AU) Treatments
4. Conclusions
Acknowledgments
Conflicts of Interest
- Author ContributionsMasataka Suzuki wrote and prepared all parts of this manuscript. He designed and arranged all the experiments presented in this manuscript. He acquired government funds to perform this study.Haruka Ito performed all the experiments described above. Some experiments were originally designed based on her idea.Fugaku Aoki made a great contribution to perform ChIP-qPCR analysis and gave us helpful comments on preparing this manuscript based on his deep knowledge about chromatin modifications.
References
- Bandziulis, R.J.; Swanson, M.S.; Dreyfuss, G. RNA-binding proteins as developmental regulators. Genes Dev 1989, 3, 431–437. [Google Scholar]
- Green, M.R. Biochemical mechanisms of constitutive and regulated pre-mRNA splicing. Annu. Rev. Cell Biol 1991, 7, 559–599. [Google Scholar]
- Wang, E.T.; Sandberg, R.; Luo, S.; Khrebtukova, I.; Zhang, L.; Mayr, C.; Kingsmore, S.F.; Schroth, G.P.; Burge, C.B. Alternative isoform regulation in human tissue transcriptomes. Nature 2008, 456, 470–476. [Google Scholar]
- Black, D.L. Finding splice sites within a wilderness of RNA. RNA 1995, 1, 763–771. [Google Scholar]
- Chou, M.Y.; Rooke, N.; Turck, C.W.; Black, D.L. hnRNP H is a component of a splicing enhancer complex that activates a c-src alternative exon in neuronal cells. Mol. Cell. Biol 1999, 19, 69–77. [Google Scholar]
- Expert-Bezancon, A.; Le Caer, J.P.; Marie, J. Heterogeneous nuclear ribonucleoprotein (hnRNP) K is a component of an intronic splicing enhancer complex that activates the splicing of the alternative exon 6A from chicken beta-tropomyosin pre-mRNA. J. Biol. Chem 2002, 277, 16614–16623. [Google Scholar]
- Li, X.; Shambaugh, M.E.; Rottman, F.M.; Bokar, J.A. SR proteins Asf/SF2 and 9G8 interact to activate enhancer-dependent intron D splicing of bovine growth hormone pre-mRNAin vitro. RNA 2000, 6, 1847–1858. [Google Scholar]
- Modafferi, E.F.; Black, D.L. Combinatorial control of a neuron-specific exon. RNA 1999, 5, 687–706. [Google Scholar]
- Alló, M.; Schor, I.E.; Muñoz, M.J.; de la Mata, M.; Agirre, E.; Valcárcel, J.; Eyras, E.; Kornblihtt, A.R. Chromatin and alternative splicing. Cold Spring Harb. Symp. Quant. Biol 2010, 75, 103–111. [Google Scholar]
- Luco, R.F.; Allo, M.; Schor, I.E.; Kornblihtt, A.R.; Misteli, T. Epigenetics in alternative pre-mRNA splicing. Cell 2011, 144, 16–26. [Google Scholar]
- Luco, R.F.; Misteli, T. More than a splicing code: Integrating the role of RNA, chromatin and non-coding RNA in alternative splicing regulation. Curr. Opin. Genet. Dev 2011, 21, 366–372. [Google Scholar]
- Spies, N.; Nielsen, C.B.; Padgett, R.A.; Burge, C.B. Biased chromatin signatures around polyadenylation sites and exons. Mol. Cell 2009, 36, 245–254. [Google Scholar]
- Schwartz, S.; Meshorer, E.; Ast, G. Chromatin organization marks exon-intron structure. Nat. Struct. Mol. Biol 2009, 16, 990–995. [Google Scholar]
- Tilgner, H.; Nikolaou, C.; Althammer, S.; Sammeth, M.; Beato, M.; Valcárcel, J.; Guigó, R. Nucleosome positioning as a determinant of exon recognition. Nat. Struct. Mol. Biol 2009, 16, 996–1001. [Google Scholar]
- Schor, I.E.; Rascovan, N.; Pelisch, F.; Alló, M.; Kornblihtt, A.R. Neuronal cell depolarization induces intragenic chromatin modifications affecting NCAM alternative splicing. Proc. Natl. Acad. Sci. USA 2009, 106, 4325–4330. [Google Scholar]
- Kolasinska-Zwierz, P.; Down, T.; Latorre, I.; Liu, T.; Liu, X.S.; Ahringer, J. Differential chromatin marking of introns and expressed exons by H3K36me3. Nat. Genet 2009, 41, 376–381. [Google Scholar]
- Luco, R.F.; Pan, Q.; Tominaga, K.; Blencowe, B.J.; Pereira-Smith, O.M.; Misteli, T. Regulation of alternative splicing by histone modifications. Science 2010, 327, 996–1000. [Google Scholar]
- Sims, R.J., 3rd; Millhouse, S.; Chen, C.F.; Lewis, B.A.; Erdjument-Bromage, H.; Tempst, P.; Manley, J.L.; Reinberg, D. Recognition of trimethylated histone H3 lysine 4 facilitates the recruitment of transcription postinitiation factors and pre-mRNA splicing. Mol. Cell 2007, 28, 665–676. [Google Scholar]
- Saint-André, V.; Batsché, E.; Rachez, C.; Muchardt, C. Histone H3 lysine 9 trimethylation and HP1γ favor inclusion of alternative exons. Nat. Struct. Mol. Biol 2011, 18, 337–344. [Google Scholar]
- Zhou, Y.; Lu, Y.; Tian, W. Epigenetic features are significantly associated with alternative splicing. BMC Genomics 2012, 13, 123. [Google Scholar]
- Hashimoto, H. The role of the W chromosome for sex determination in the silkwormBombyx mori. Jpn. J. Genet 1933, 8, 245–258. [Google Scholar]
- Niimi, T.; Sahara, K.; Oshima, H.; Yasukouchi, Y.; Ikeo, K.; Traut, W. Molecular cloning and chromosomal localization of the Bombyx Sex-lethal gene. Genome 2006, 49, 263–268. [Google Scholar]
- Mita, K.; Kasahara, M.; Sasaki, S.; Nagayasu, Y.; Yamada, T.; Kanamori, H.; Namiki, N.; Kitagawa, M.; Yamashita, H.; Yasukochi, Y.; et al. The genome sequence of silkwormBombyx mori. DNA Res 2009, 11, 27–35. [Google Scholar]
- Suzuki, M.G.; Funagume, S.; Kanda, T.; Tamura, T.; Shimada, T. Role of the male BmDSX protein in the sexual differentiation ofBombyx mori. Evol. Dev 2005, 7, 58–68. [Google Scholar]
- Suzuki, M.G.; Ohbayashi, F.; Mita, K.; Shimada, T. The mechanism of sex-specific splicing at the doublesex gene is different between Drosophila melanogaster andBombyx mori. Insect Biochem. Mol. Biol 2001, 31, 1201–1211. [Google Scholar]
- Suzuki, M.G.; Imanishi, S.; Dohmae, N.; Nishimura, T.; Shimada, T.; Matsumoto, S. Establishment of a novel in vivo sex-specific splicing assay system to identify a trans-acting factor that negatively regulates splicing of Bombyx mori dsx female exons. Mol. Cell. Biol 2008, 28, 333–343. [Google Scholar]
- Suzuki, M.G.; Imanishi, S.; Dohmae, N.; Asanuma, M.; Matsumoto, S. Identification of a male-specific RNA binding protein that regulates sex-specific splicing of Bmdsx by increasing RNA binding activity of BmPSI. Mol. Cell. Biol 2010, 30, 5776–5786. [Google Scholar]
- Suzuki, M.G.; Kobayashi, S.; Aoki, F. Male-specific splicing of the silkworm Imp gene is maintained by an autoregulatory mechanism. Mech. Dev 2014, 131, 47–56. [Google Scholar]
- Terenius, O.; Papanicolaou, A.; Garbutt, J.S.; Eleftherianos, I.; Huvenne, H.; Kanginakudru, S.; Albrechtsen, M.; An, C.; Aymeric, J.L.; Barthel, A.; et al. RNA interference in Lepidoptera: An overview of successful and unsuccessful studies and implications for experimental design. J. Insect Physiol 2011, 57, 231–245. [Google Scholar] [Green Version]
- Yamaguchi, J.; Mizoguchi, T.; Fujiwara, H. siRNAs induce efficient RNAi response in Bombyx mori. PLoS One 2011, 6, e25469. [Google Scholar]
- Nguyen, A.T.; Zhang, Y. The diverse functions of Dot1 and H3K79 methylation. Genes Dev 2011, 25, 1345–1358. [Google Scholar]
- Schübeler, D.; MacAlpine, D.M.; Scalzo, D.; Wirbelauer, C.; Kooperberg, C.; van Leeuwen, F.; Gottschling, D.E.; O’Neill, L.P.; Turner, B.M.; Delrow, J.; et al. The histone modification pattern of active genes revealed through genome-wide chromatin analysis of a higher eukaryote. Genes Dev 2004, 18, 1263–1271. [Google Scholar]
- Morillon, A.; Karabetsou, N.; Nair, A.; Mellor, J. Dynamic lysine methylation on histone H3 defines the regulatory phase of gene transcription. Mol. Cell 2005, 18, 723–734. [Google Scholar]
- Pokholok, D.K.; Harbison, C.T.; Levine, S.; Cole, M.; Hannett, N.M.; Lee, T.I.; Bell, G.W.; Walker, K.; Rolfe, P.A.; Herbolsheimer, E.; et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 2005, 122, 517–527. [Google Scholar]
- Kadener, S.; Fededa, J.P.; Rosbash, M.; Kornblihtt, A.R. Regulation of alternative splicing by a transcriptional enhancer through RNA pol II elongation. Proc. Natl. Acad. Sci. USA 2002, 99, 8185–8190. [Google Scholar]
- De la Mata, M.; Alonso, C.R.; Kadener, S.; Fededa, J.P.; Blaustein, M.; Pelisch, F.; Cramer, P.; Bentley, D.; Kornblihtt, A.R. A slow RNA polymerase II affects alternative splicingin vivo. Mol. Cell 2003, 12, 525–532. [Google Scholar]
- Nakanishi, T.; Nakano, A.; Nomura, K.; Sekimizu, K.; Natori, S. Purification, gene cloning, and gene disruption of the transcription elongation factor S-II inSaccharomyces cerevisiae. J. Biol. Chem 1992, 267, 13200–13204. [Google Scholar]
- Nakanishi, T.; Shimoaraiso, M.; Kubo, T.; Natori, S. Structure-function relationship of yeast S-II in terms of stimulation of RNA polymerase II, arrest relief, and suppression of 6-azauracil sensitivity. J. Biol. Chem 1995, 270, 8991–8995. [Google Scholar]
- Powell, W.; Reines, D. Mutations in the second largest subunit of RNA polymerase II cause 6-azauracil sensitivity in yeast and increased transcriptional arrest in vitro. J. Biol. Chem. 1996, 271, 6866–6873. [Google Scholar]
- Franklin, T.J.; Cook, J.M. The inhibition of nucleic acid synthesis by mycophenolic acid. Biochem. J 1969, 113, 515–524. [Google Scholar]
- Exinger, F.; Lacroute, F. 6-Azauracil inhibition of GTP biosynthesis inSaccharomyces cerevisiae. Curr. Genet 1992, 22, 9–11. [Google Scholar]
- Uptain, S.M.; Kane, C.M.; Chamberlin, M.J. Basic mechanisms of transcript elongation and its regulation. Annu. Rev. Biochem 1997, 66, 117–172. [Google Scholar]
- Howe, K.J.; Kane, C.M.; Ares, M., Jr. Perturbation of transcription elongation influences the fidelity of internal exon inclusion inSaccharomyces cerevisiae. RNA 2003, 9, 993–1006. [Google Scholar]
- Masumoto, M.; Yaginuma, T.; Niimi, T. Functional analysis of Ultrabithorax in the silkworm, Bombyx mori, using RNAi. Dev. Genes Evol 2009, 219, 437–444. [Google Scholar]
- Llamazares, S.; Moreira, A.; Tavares, A.; Girdham, C.; Spruce, B.A.; Gonzalez, C.; Karess, R.E.; Glover, D.M.; Sunkel, C.E. polo encodes a protein kinase homolog required for mitosis inDrosophila. Genes Dev 1991, 5, 2153–2165. [Google Scholar]
- Pinto, P.A.; Henriques, T.; Freitas, M.O.; Martins, T.; Domingues, R.G.; Wyrzykowska, P.S.; Coelho, P.A.; Carmo, A.M.; Sunkel, C.E.; Proudfoot, N.J.; et al. RNA polymerase II kinetics in poilo polyadenylation signal selection. EMBO J 2011, 30, 2431–2444. [Google Scholar]
- De la Mata, M.; Lafaille, C.; Kornblihtt, A.R. First come, first served revisited: Factors affecting the same alternative splicing event have different effects on the relative rates of intron removal. RNA 2010, 16, 904–912. [Google Scholar]
- Luo, Z.; Lin, C.; Shilatifard, A. The super elongation complex (SEC) family in transcriptional control. Nat. Rev. Mol. Cell Biol 2012, 13, 543–547. [Google Scholar]
- Shilatifard, A.; Lane, W.S.; Jackson, K.W.; Conaway, R.C.; Conaway, J.W. An RNA polymerase II elongation factor encoded by the human ELL gene. Science 1996, 271, 1873–1876. [Google Scholar]
- Shilatifard, A.; Duan, D.R.; Haque, D.; Florence, C.; Schubach, W.H.; Conaway, J.W.; Conaway, R.C. ELL2, a new member of an ELL family of RNA polymerase II elongation factors. Proc. Natl. Acad. Sci. USA 1997, 94, 3639–3643. [Google Scholar]
- Bitoun, E.; Oliver, P.L.; Davies, K.E. The mixed-lineage leukemia fusion partner AF4 stimulates RNA polymerase II transcriptional elongation and mediates coordinated chromatin remodeling. Hum. Mol. Gen 2007, 16, 92–106. [Google Scholar]
- Mueller, D.; Bach, C.; Zeisig, D.; Garcia-Cuellar, M.P.; Monroe, S.; Sreekumar, A.; Zhou, R.; Nesvizhskii, A.; Chinnaiyan, A.; Hess, J.L.; et al. A role for the MLL fusion partner ENL in transcriptional elongation and chromatin modification. Blood 2007, 110, 4445–4454. [Google Scholar]
- Milcarek, C.; Albring, M.; Langer, C.; Park, K.S. The eleven-nineteen lysine-rich leukemia gene (ELL2) influences the histone H3 protein modifications accompanying the shift to secretory immunoglobulin heavy chain mRNA production. J. Biol. Chem 2011, 286, 33795–33803. [Google Scholar]
- Colgan, D.F.; Manley, J.L. Mechanism and regulation of mRNA polyadenylation. Genes Dev 1997, 11, 2755–2766. [Google Scholar]
- Tian, B.; Hu, J.; Zhang, H.; Lutz, C.S. A large-scale analysis of mRNA polyadenylation of human and mouse genes. Nucleic Acids Res 2005, 33, 201–212. [Google Scholar]
- Kornblihtt, A.R. Chromatin, transcript elongation and alternative splicing. Nat. Struct. Mol. Biol 2006, 13, 5–7. [Google Scholar]
- Jones, B.; Su, H.; Bhat, A.; Lei, H.; Bajko, J.; Hevi, S.; Baltus, G.A.; Kadam, S.; Zhai, H.; Valdez, R.; et al. The histone H3K79 methyltransferase Dot1L is essential for mammalian development and heterochromatin structure. PLoS Genet 2008, 4, e1000190. [Google Scholar]
- Shanower, G.A.; Muller, M.; Blanton, J.L.; Honti, V.; Gyurkovics, H.; Schedl, P. Characterization of the grappa gene, the Drosophila histone H3 lysine 79 methyltransferase. Genetics 2005, 169, 173–184. [Google Scholar]
- Brock, H.W.; van Lohuizen, M. The Polycomb group—No longer an exclusive club. Curr. Opin. Genet. Dev 2001, 11, 175–181. [Google Scholar]
- Suzuki, M.G.; Suzuki, K.; Aoki, F.; Ajimura, M. Effect of RNAi-mediated knockdown of the Bombyx mori transformer-2 gene on the sex-specific splicing of Bmdsx pre-mRNA. Int. J. Dev. Biol 2012, 56, 693–699. [Google Scholar]
- Koike, Y.; Mita, K.; Suzuki, M.G.; Maeda, S.; Abe, H.; Osoegawa, K.; deJong, P.J.; Shimada, T. Genomic sequence of a 320-kb segment of the Z chromosome of Bombyx mori containing a kettin ortholog. Mol. Genet. Genomics 2003, 269, 137–149. [Google Scholar]
| siRNA | Injected eggs | Early or mid-stage embryonic lethal | Late-stage embryonic lethal | Viable |
|---|---|---|---|---|
| N. C. siRNA | 115 | 58 (50.4%) | 32 (27.8%) | 25 (21.7%) |
| DOT1L siRNA | 263 | 188 (71.5%) | 74 (28.1%) | 1 (0.4%) (male) |
| Target gene | siRNA | Sense | Antisense |
|---|---|---|---|
| SETD2 | 393 | UGCCAGCUCUGAGUCUGAUUCAAU | AUUGAAUCAGACUCAGAGCUGGCA |
| 490 | CAGUGUAGCUCAAGAGAUATT | UAUCUCUUGAGCUACACUGTT | |
| DOT1L | 243 | UUCCAAAGCAACUACAGAAUCGAUG | CAUCGAUUCUGUAGUUGCUUUGGAA |
| 353 | AUUUACUCGCUUUACUUUGTT | CAAAGUAAAGCGAGUAAAUTT | |
| EZH2 | 38 | GACAACCCAACAGGUACCAAUAAGA | UCUUAUUGGUACCUGUUGGGUUGUC |
| 224 | CGACGGGAAAGUGCAUGGUGAUAAA | UUUAUCACCAUGCACUUUCCCGUCG | |
| ASH2 | 157 | GACCGGCCUCUAGUCAAAUUCAAGA | UCUUGAAUUUGACUAGAGGCCGUC |
| 167 | UAGUCAAAUUCAAGAGCCACCUGUA | UACAGGUGGCUCUUGAAUUUGACUA |
| Target gene | Primers | Sequence (5′ to 3′) | Denature | Annealing | Elongation | N° cycles |
|---|---|---|---|---|---|---|
| Bmdsx | FDSX-F2 | CGCCTTACCGCAGACAGGCAG | 98 °C | 57 °C | 57 °C | 35 |
| FDSX-R4 | GCGCAGTGTCGTCGCTACAAGG | 10 s | 30 s | 60 s | ||
| ImpM | BmIMPF1 | ATGGACGGTGACATGTCTCAAG | 98 °C | 55 °C | 57 °C | 35 |
| BmIMPR1 | TCATCCCGCCTCAGACGATTG | 10 s | 30 s | 90 s | ||
| ImpC | IMPE4F1 | TCCCATAATAATCTCATTGGAC | 98 °C | 55 °C | 57 °C | 35 |
| IMPE7R1 | AATGTGAACGGTGGTCTCGTG | 10 s | 30 s | 90 s | ||
| Actin3 | BA3F1 | AGATGACCCAGATCATGTTCG | 98 °C | 57 °C | 57 °C | 26 |
| BASR1 | GAGATCCACATCTGTTGGAAG | 10 s | 30 s | 30 s | ||
| m | BmSxlF1 | ATTAATCATCATAAAGCTACG | 98 °C | 57 °C | 57 °C | 35 |
| BmSxlR1 | AATCCGTAACTGTAGCCAGTC | 10 s | 30 s | 30 s |
| Target gene | Primers | Sequence (5′ to 3′) |
|---|---|---|
| SETD2 | SETD2qPCRF1 | CCTACAGGACATCTGGAGTTAC |
| SETD2qPCRR1 | GAATCAGTACCAGCATTTAGATG | |
| DOT1L | dot1qPCRF1 | AGAATCCGAACGACTCGACAG |
| dot1qPCRR1 | CTGTTCTTGGTCTTCGTTCAAC | |
| EZH2 | EZH2F1 | GGTGTAGTGACAACCCAACAG |
| EZH2R1 | TCTTAACTCCTGAGCTGTTCC | |
| ASH2 | ASHF2 | GGGGACCAGGTTCCACGAGTC |
| ASHR2 | TACAGGTGGCTCTTGAATTTG | |
| Bmdsx promoter | BmdsxproF1 | TGCATGTTTCTTATTAATCAGCTAG |
| BmdsxproR1 | GTAAATTTCGTAAAAGCTGACCAG | |
| Bmdsx E1 | ChIPEx1F | CCTGTACCACCAGTGGTGAAG |
| ChIPEx1R | CTGACGGCGGTGGAGCGTATG | |
| Bmdsx I2 | ChIPI2F | CTACATGAACAGTACCAGTCAG |
| ChIPI2R | GTAAGTACAAACTAAATAGCGTTC | |
| Bmdsx E3 | ChIPEx3F | GTCGACGAGTACGCGAGGAAG |
| ChIPEx3R | TGTGATGCATGTATCTGTCGC | |
| Bmdsx I3 | ChIPI3F | GTAACTGACCTTCTTGCTAATC |
| ChIPI3R | CTGTGCCATTTTATTAATATCGTC | |
| Bmdsx E4 | ChIPEx4F | ATATAAGTGGTGTACTGTCTTC |
| ChIPEx4R | CCATAGATCCAATGTTACGAC | |
| Bmdsx I4 | ChIPI4F | GTTCAAACACATCGAAGCTAC |
| ChIPI4R | GTCCGAGATAGACTGGCCTTG | |
| Bmdsx E6 | ChIPEx6F | GGCACAGCGCCGACAAGTAAG |
| ChIPEx6R | ATTGTCTGTAGATATTCGTGATC | |
| BmIMP promoter1S | IMPproF2 | TTAAGCATTTAATTATAAGAAGATC |
| IMPproR2 | CTAGAATCTGCGATTACATAC | |
| BmIMP E1S | IMPE1F1 | TCCGTTCAGTACTCGCTATAC |
| IMPE1R1 | TCTTACCTATCGTCATAGATTC | |
| BmIMP I1S | IMPI1F2 | ATTTGGTAAAATAGTCTCGTATC |
| IMPI1R2 | ACCTTGTGATACGGGGTTAAC | |
| BmIMP pro1L | IMPE1LproF1 | GCTGCCCCACCCTTTAAACCG |
| IMPE1LproR1 | CTCGATCGTGCTGACTCTAGC | |
| BmIMP E1L | IMPE1F1 | TTTCAAGTATACTCCTTCTATAG |
| IMPE1R1 | TTCGCCATTTTGAGCAGATTG | |
| BmIMP I1L | IMPI1F3 | CAAATGGGCACATATTGTTGG |
| IMPI1R3 | GTTTAAGCGCTTTCGTGATGG | |
| BmIMP E7 | IMPE7F1 | ATGCGGGAAGAAGGTTTTATG |
| IMPE7R1 | AATGTGAACGGTGGTCTCGTG | |
| BmIMP I7 | IMPI7F1 | GTGCATAAATCCACAGAACAG |
| IMPI7R1 | TTACTCAGAAACTCAGAAGTAC | |
| BmIMP E8 | IMPE8F1 | CGTCTGAGGCGGGATGAGAAC |
| IMPE8R1 | TAAATTCGCCGCAATCAGCAG | |
| EF1 promoter | EF-1proF1 | TATATCAATTTTGGTGCAAGAATGG |
| EF-1proR1 | GTAATAATATTCTATTCTATCCACCG | |
| EF-1 E2 | EF-1E2F1 | TGGCGATGGAGGCGGAGAAG |
| EF-1E2R1 | CTCAACTTCCCAGCTGTCTGC | |
| EF-1 I3 | EF-1I3F1 | ACTTACTTATTTATGATCATGCGTC |
| EF-1I3R1 | GCTAACCACAATTATATTTGTGGAG | |
| EF-1 E6 | EF-1E6F1 | TACAGGTCATTTCTGCACGTAAG |
| EF-1E6R1 | TCATCCCAGTTAACTGTTGGATC | |
| EF-1 I11 | EF-1I11F1 | TTCATGGACTACATTTTACCTTGG |
| EF-1I11R1 | CTAAGCTCTTCTAAAAGAGATGAGC | |
| EF-1 I12 | EF-1I12F1 | GCATTAATATTAATTCCACCACAAG |
| EF-1I12R1 | CACACCTCACTGCTCTTCCGC | |
| Bm-Sxl promoter | Sxli1F1 | GGCTAAACTATCTTCAACAAG |
| Sxli1R1 | CGGTCACCGTTCTCGTGAAAG | |
| Bm-Sxl E1 | Sxle1F1 | GCCAGTCCAAATGGACGAATC |
| Sxle1R1 | GTTCACTGACTTTCGAGTGAG | |
| Bm-Sxl E2 | Sxle2F1 | GCCTACTCGAACAATAAAAAAG |
| Sxle2R1 | TTCCAAAGAATTGAAACTCCTG | |
| Bm-Sxl I2 | Sxli2F1 | TGAATCAGAACATCTCATTTGG |
| Sxli2R1 | CCAAGCCGCTGCCTACCTAAC | |
| Bm-Sxl E3 | Sxle3F1 | CGAGGCAGAGCGGGTTCGAAC |
| Sxle3R1 | CCTTCATCACTCGACAGCTCTC | |
| BmIMP Proximal | IMPE2F1 | ATCCTCAAAGGTACTCATCAG |
| IMPE2I2R | GCATGCATCACTCAACAATAC | |
| BmIMP Distal | IMPE7F2 | GGATCATCGGCAAAGGCGGAC |
| IMPI7R2 | AGCACTTGGATCATTCATACC | |
| EF-1 Proximal | EF-1E2F1 | TGGCGATGGAGGCGGAGAAGG |
| EF-1E2I2R | GAAAAAGAAGAAAGCATTCATGC | |
| EF-1 Distal | EF-1I11E12F | ATGATACTGTATTAACTGCATTC |
| EF-1E12R2 | TTCAGGATTTTGAGACCCTGG | |
| BmIMP E7-E8 | IMPE7F1 | ATGCGGGAAGAAGGTTTTATG |
| BmIMPR1 | TCATCCCGCCTCAGACGATTG | |
| BmIMP E1-E3 | BmIMPF1 | ATGGACGGTGACATGTCTCAAG |
| IMPE3R1 | CATCCATTCAACCCGTTTATG | |
| BmIMP E1S | BmIMPF1 | ATGGACGGTGACATGTCTCAAG |
| IMPE2R2 | GCCTGCTCTGGACTCTCGAAG | |
| BmIMP E1L | IMPE1LF1 | AATCTGCTCAAAATGGCGAAG |
| IMPE2R2 | GCCTGCTCTGGACTCTCGAAG | |
| Bm-Sxl S | BmSxlF2 | ACTCGCGTTACCTATTTAAC |
| BmSxlSR1 | GTACTGCTGTTGGATTTGGTC | |
| Bm-Sxl L | BmSxlF2 | ACTCGCGTTACCTATTTAAC |
| BmSxlLR1 | CTGCTGTTGGATTTGATTTTC |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Suzuki, M.G.; Ito, H.; Aoki, F. Effects of RNAi-Mediated Knockdown of Histone Methyltransferases on the Sex-Specific mRNA Expression of Imp in the Silkworm Bombyx mori. Int. J. Mol. Sci. 2014, 15, 6772-6796. https://doi.org/10.3390/ijms15046772
Suzuki MG, Ito H, Aoki F. Effects of RNAi-Mediated Knockdown of Histone Methyltransferases on the Sex-Specific mRNA Expression of Imp in the Silkworm Bombyx mori. International Journal of Molecular Sciences. 2014; 15(4):6772-6796. https://doi.org/10.3390/ijms15046772
Chicago/Turabian StyleSuzuki, Masataka G., Haruka Ito, and Fugaku Aoki. 2014. "Effects of RNAi-Mediated Knockdown of Histone Methyltransferases on the Sex-Specific mRNA Expression of Imp in the Silkworm Bombyx mori" International Journal of Molecular Sciences 15, no. 4: 6772-6796. https://doi.org/10.3390/ijms15046772




