ScChi, Encoding an Acidic Class III Chitinase of Sugarcane, Confers Positive Responses to Biotic and Abiotic Stresses in Sugarcane
Abstract
:1. Introduction
2. Results and Discussion
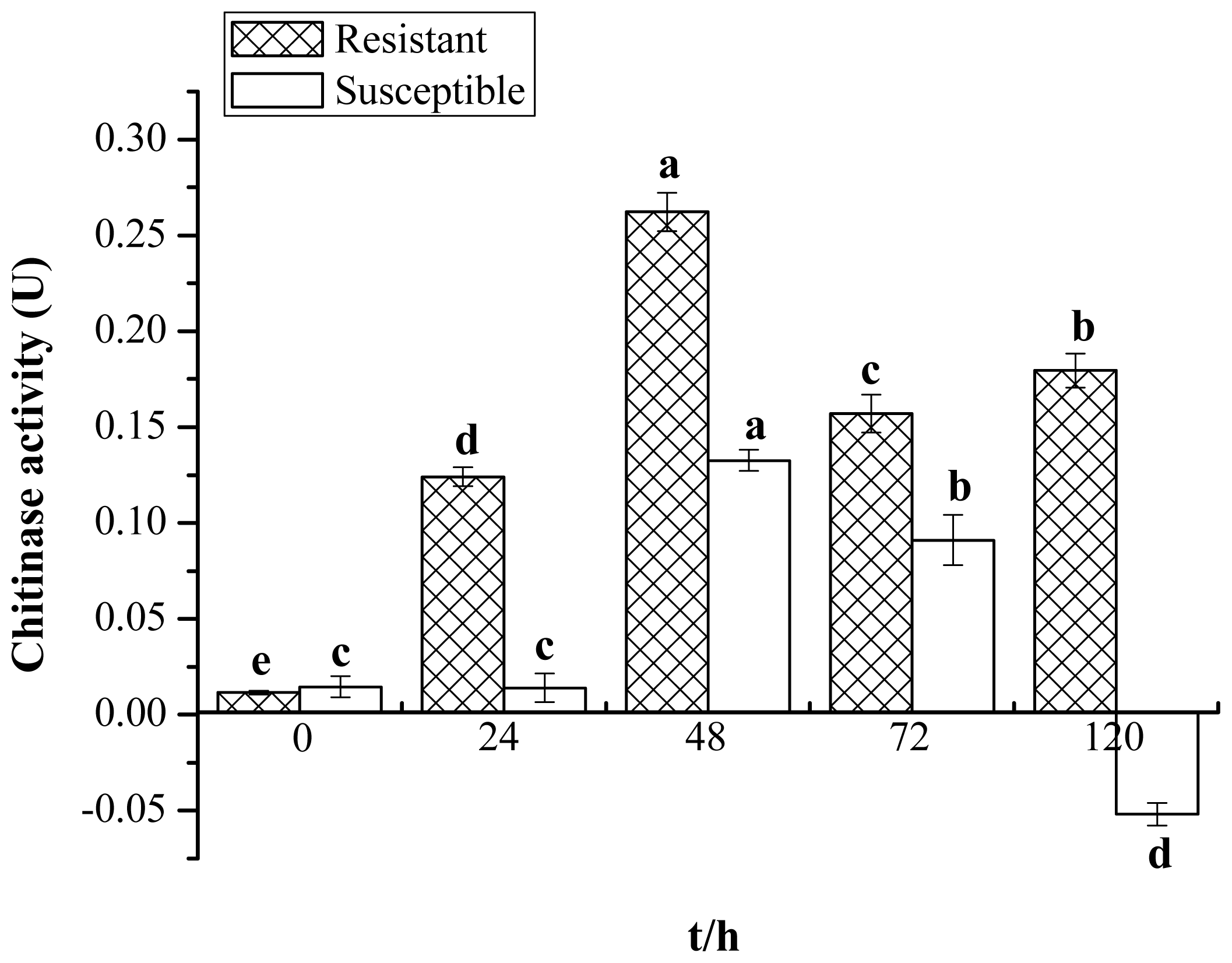
2.1. Enzyme Activity of Chitinase
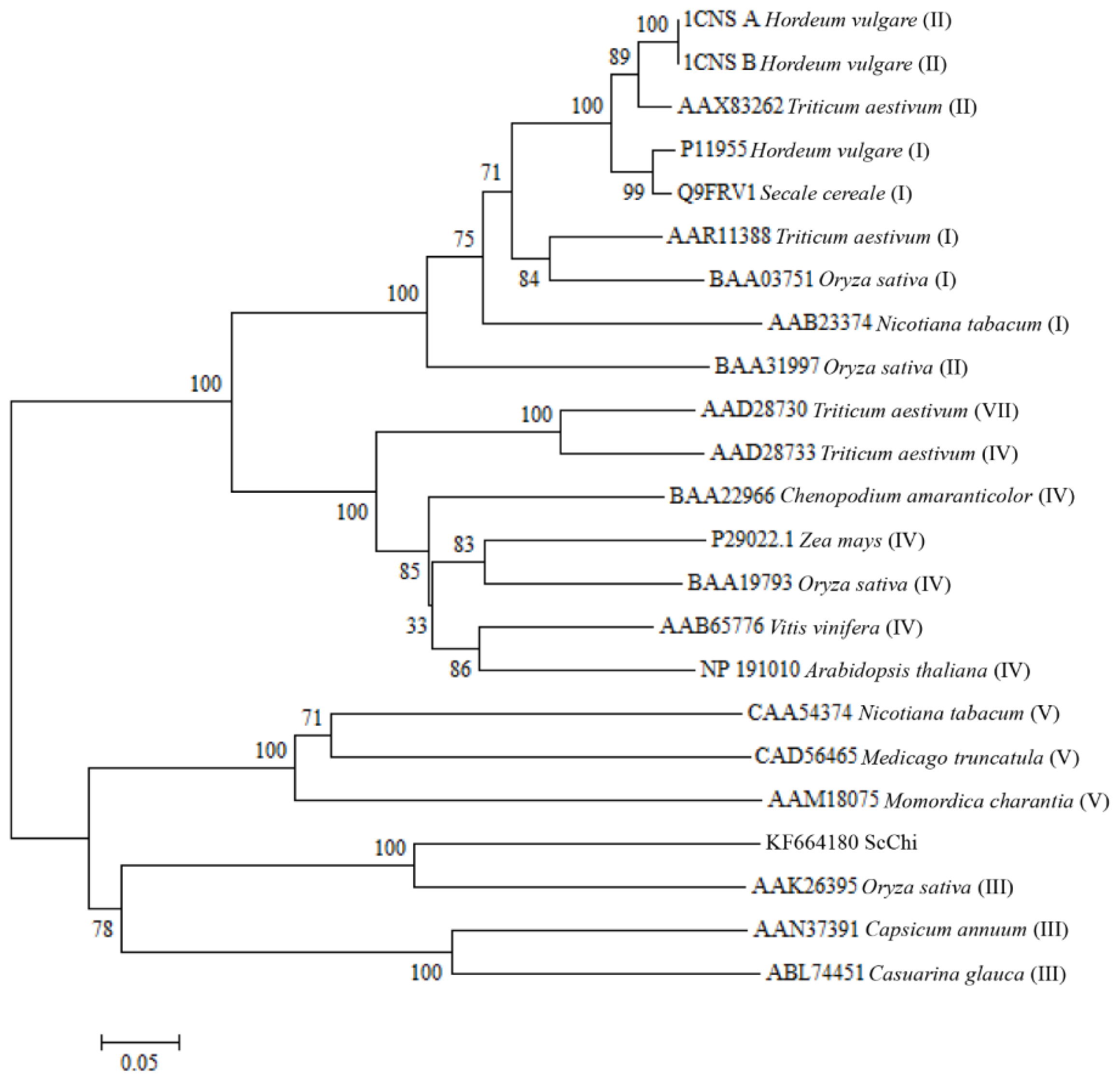
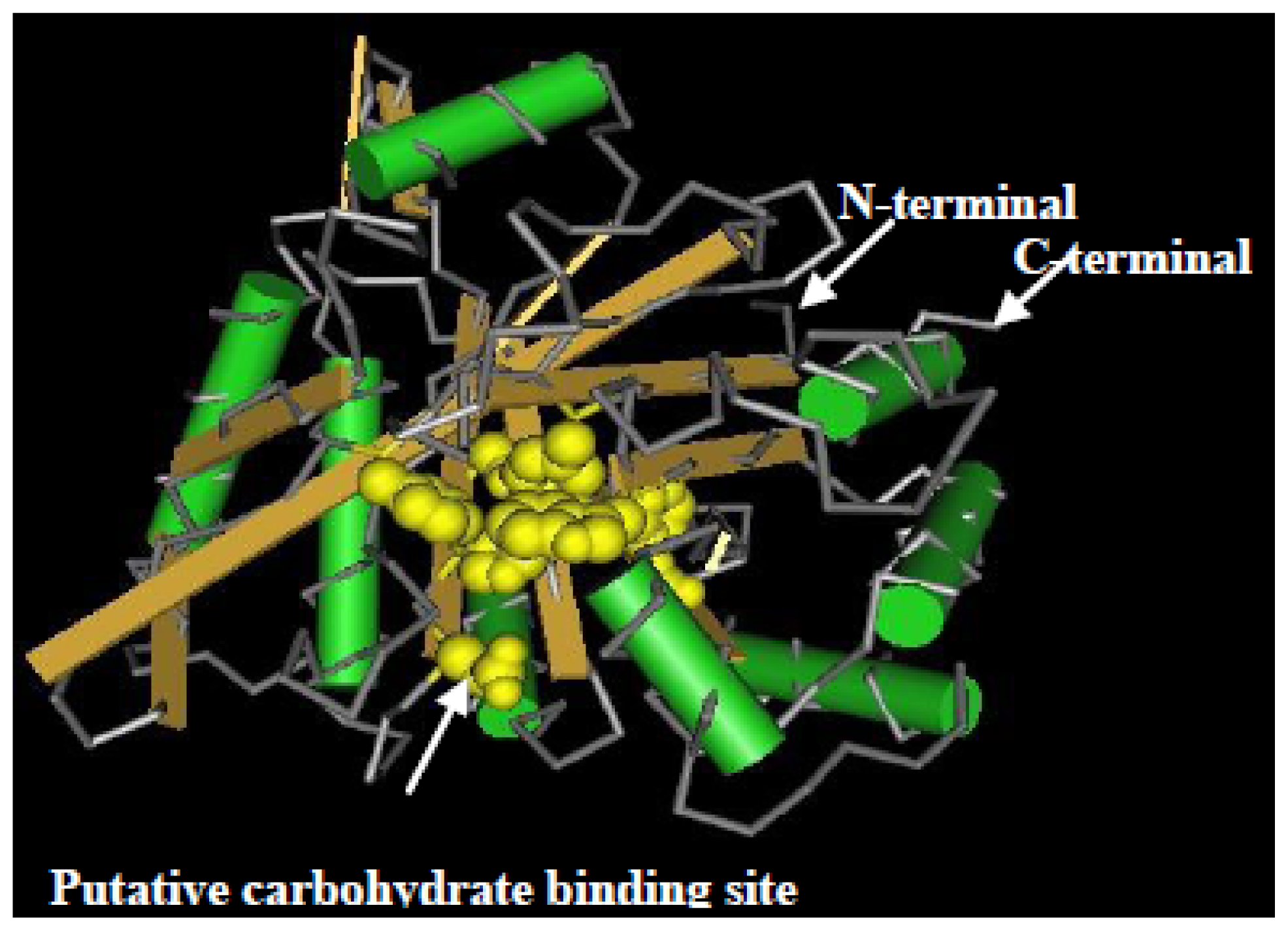
2.2. Cloning and Sequence Analysis of the Chitinase Gene, ScChi
2.3. Subcellular Localization of ScChi
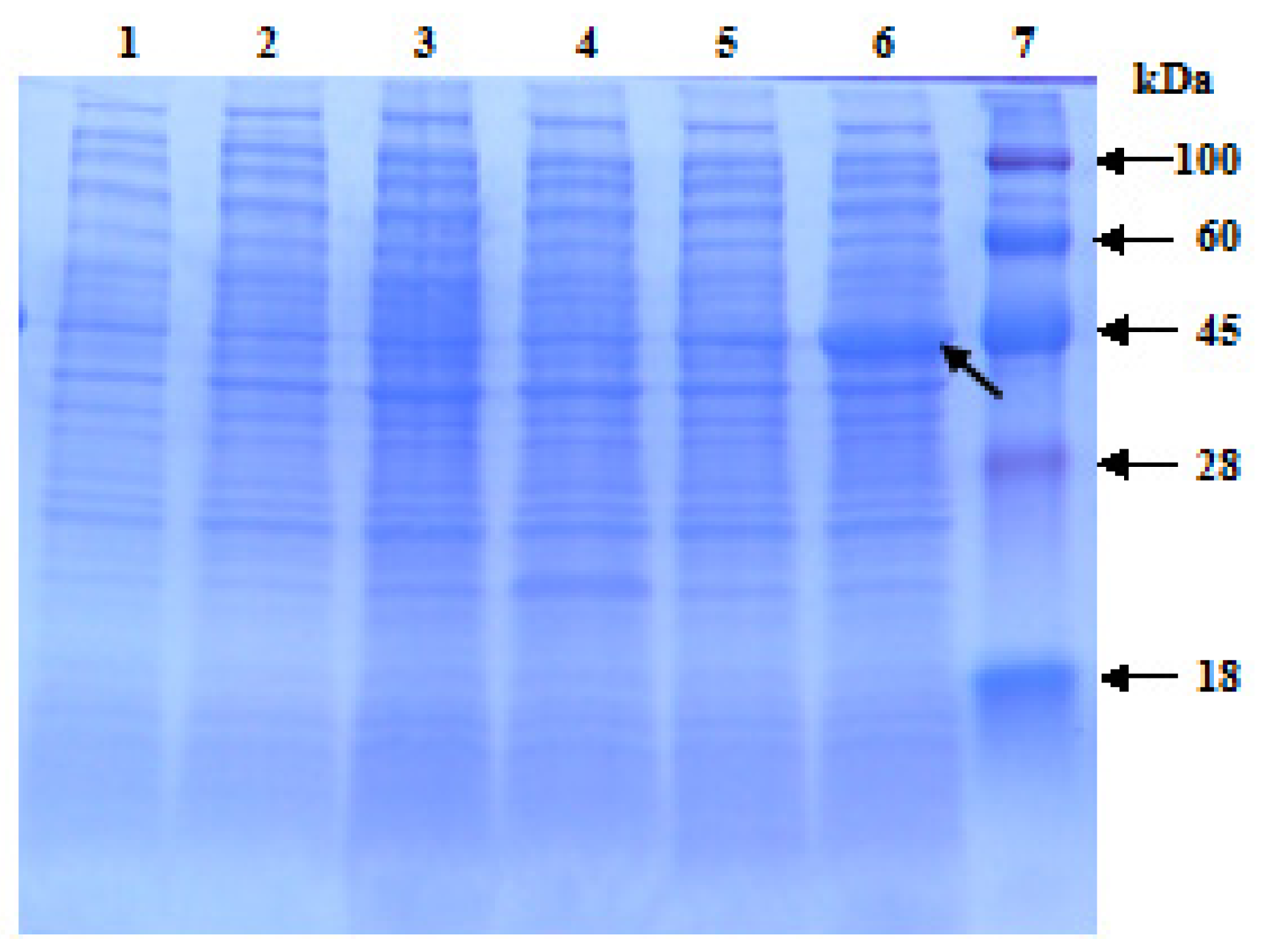
2.4. Prokaryotic Expression of ScChi in E. coli Rosetta Cells
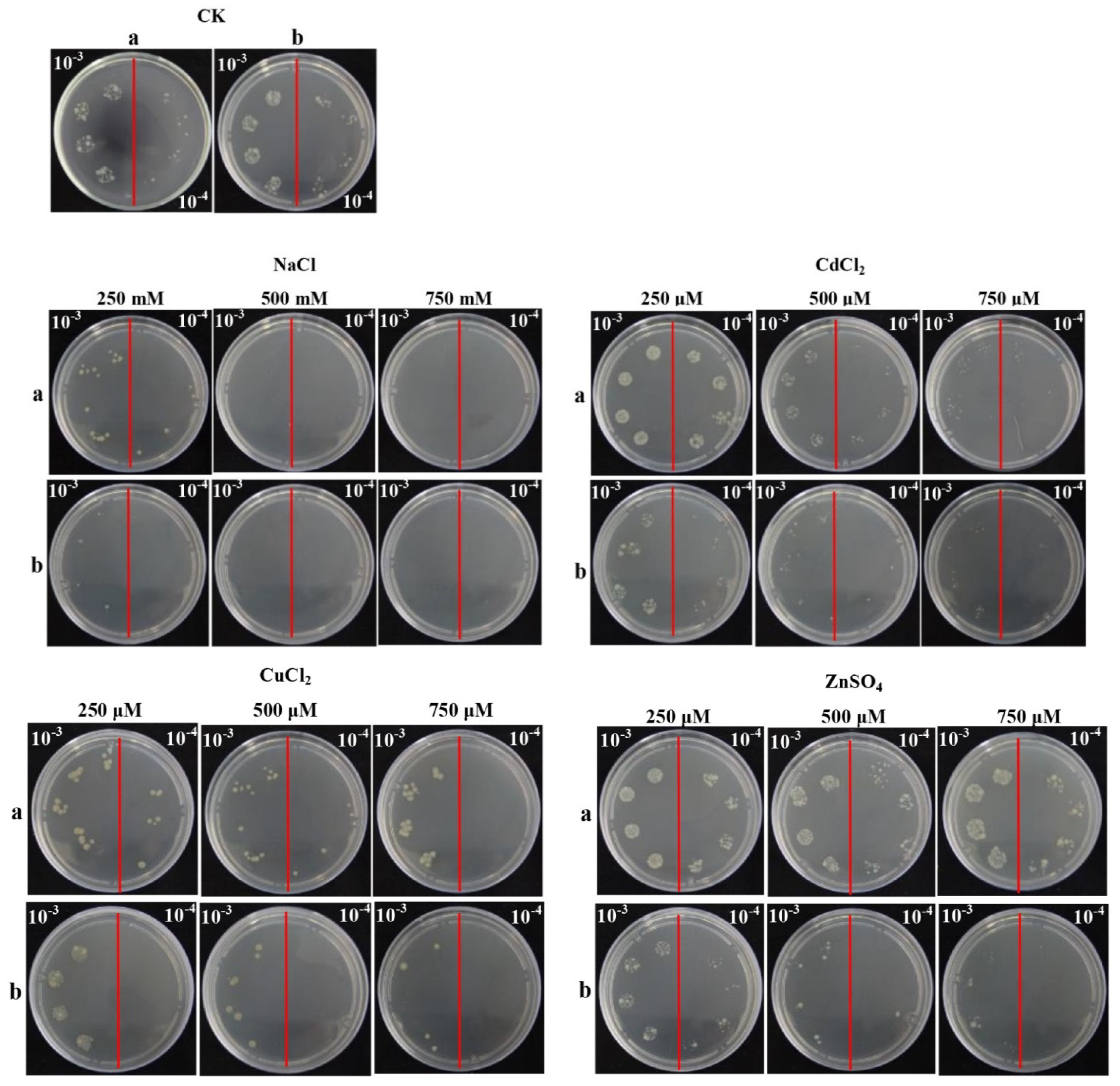
2.5. Overexpression of ScChi in E. coli Enhances Cell Growth under Abiotic Stresses
2.6. Tissue-Specific Expression Analysis of ScChi in Different Sugarcane Tissues
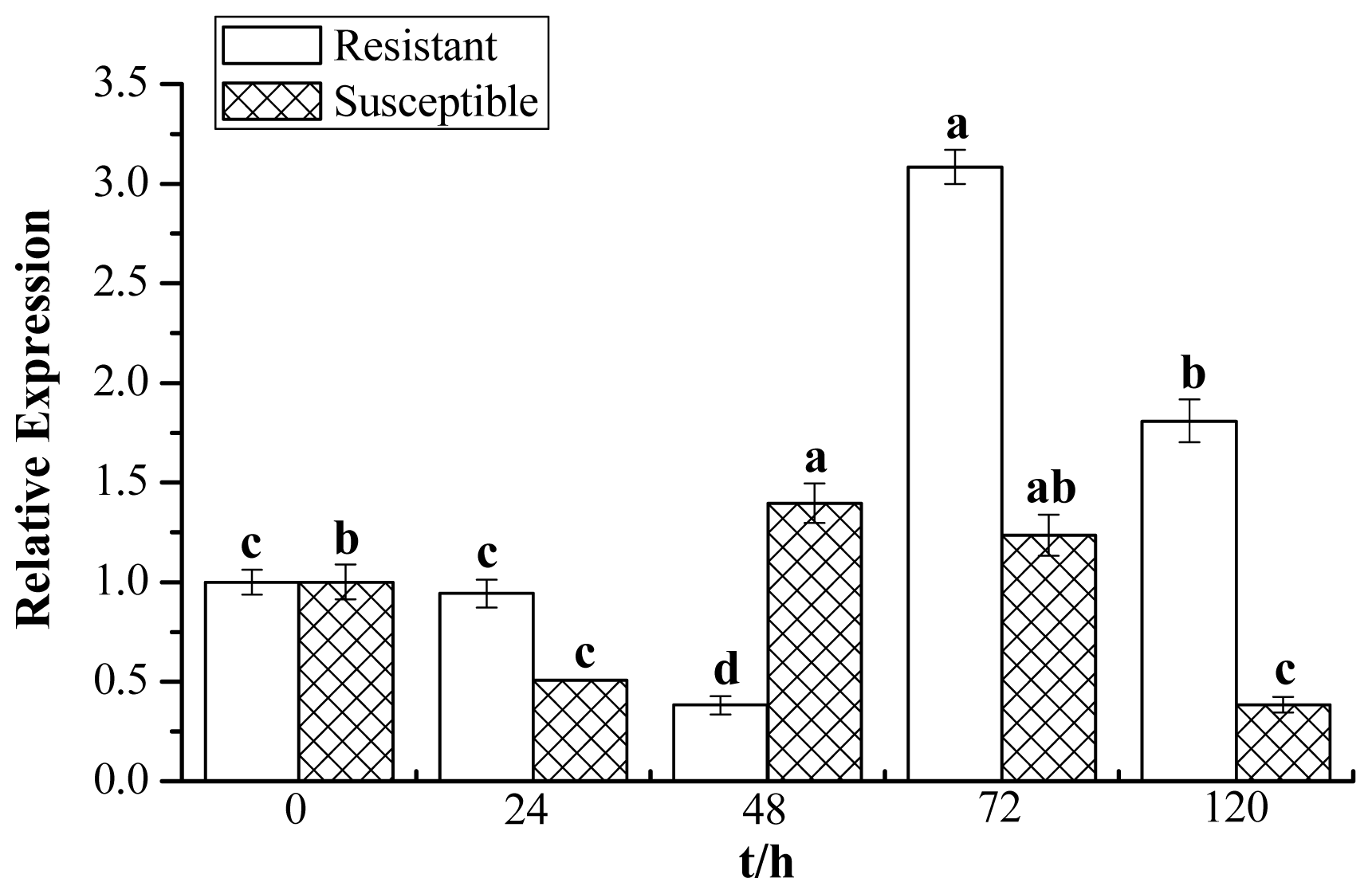
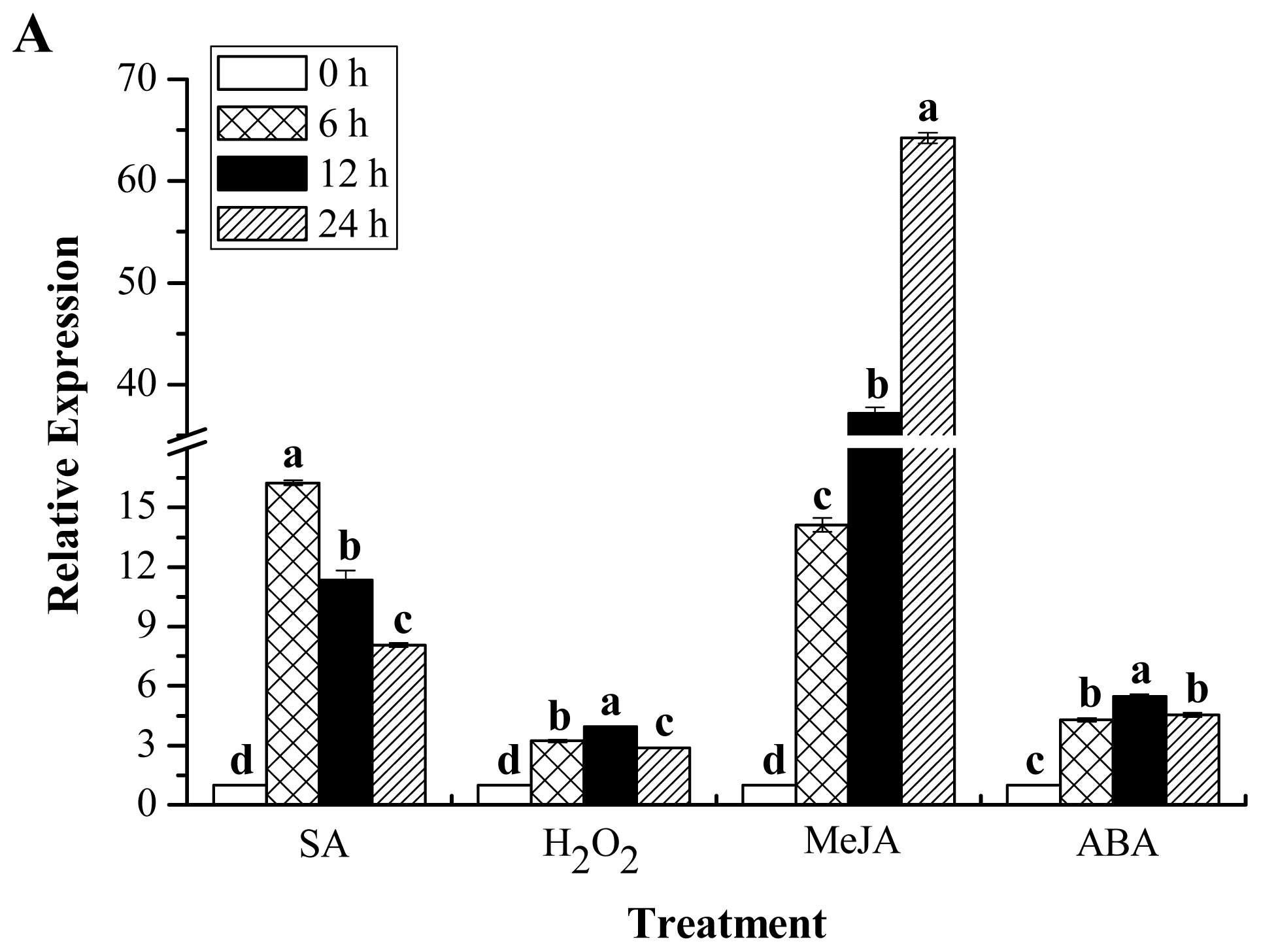
2.7. Expression Profiles of ScChi under Different Stress Treatments
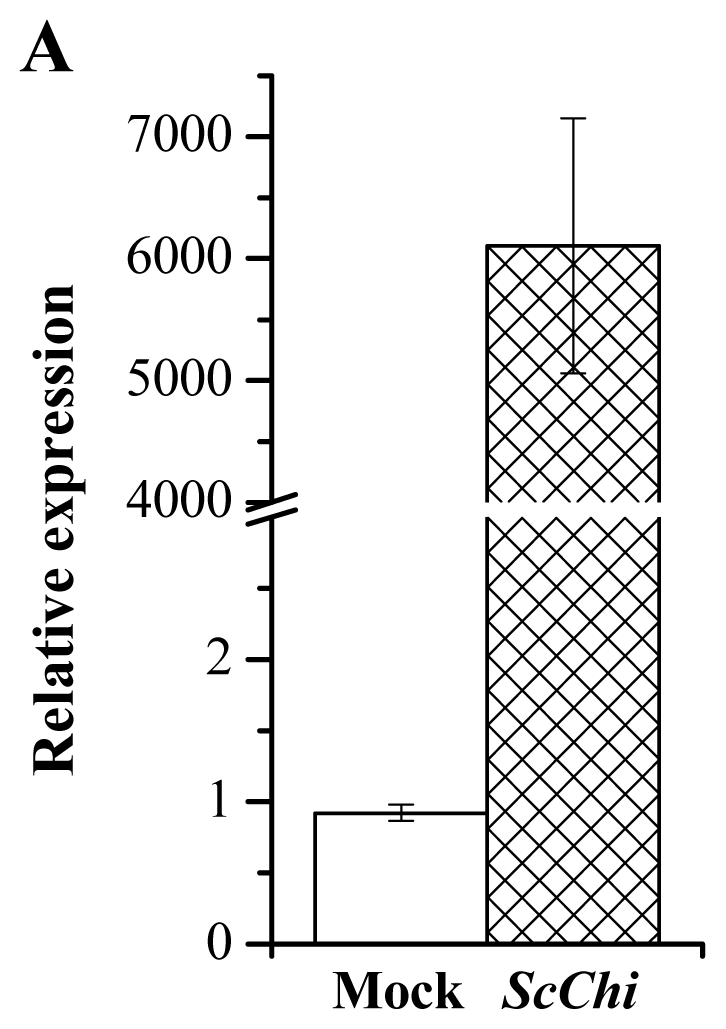
2.8. Transient Expression of ScChi Induces a Defense Response in N. benthamiana
3. Discussion
4. Experimental Section
4.1. Plant Materials and Treatments
4.2. Chitinase Activity Assay
4.3. RNA Extraction and cDNA Synthesis
4.4. Cloning, Sequencing and Bioinformatic Analysis
4.5. Subcellular Localization Assay
4.6. Expression in E. coli Rosetta
4.7. Expression Patterns of ScChi in Different Tissues and Stress Treatments
4.8. Transient Expression of ScChi in N. benthamiana
4.9. Histochemical Assay
4.10. Measurement of Ion Conductivity
5. Conclusions
Acknowledgments
Conflicts of Interest
- Author ContributionsConceived and designed the experiments: Yachun Su, Liping Xu and Youxiong Que. Performed the experiments: Yachun Su, Zhiwei Fu, Yuting Yang, Jinlong Guo, and Shanshan Wang. Analyzed the data: Yachun Su, Liping Xu and Youxiong Que. Contributed reagents/materials/analysis tools: Yachun Su, Liping Xu and Youxiong Que. Wrote the paper: Yachun Su, Liping Xu and Youxiong Que. Revised and approved the final version of the paper: Liping Xu and Youxiong Que.
References
- Walters, D.; Walsh, D.; Newton, A.; Lyon, G. Induced resistance for plant disease control: Maximizing the efficacy of resistance elicitors. Phytopathology 2005, 95, 1368–1373. [Google Scholar]
- Neuhaus, J.M.; Fritig, B.; Linthorst, H.J.M.; Meins, F.; Mikkelsen, J.D.; Ryals, J. A revised nomenclature for chitinase genes. Plant Mol. Biol. Rep 1996, 14, 102–104. [Google Scholar]
- Lawton, K.; Ward, E.; Payne, G.; Moyer, M.; Ryals, J. Acidic and basic class III chitinase mRNA accumulation in response to TMV infection of tobacco. Plant Mol. Biol 1992, 19, 735–743. [Google Scholar]
- Schlumbaum, A.; Mauch, F.; Vogeli, U.; Boller, T. Plant chitinases are potent inhibitors of fungal growth. Nature 1986, 324, 365–367. [Google Scholar]
- Mauch, F.; Mauch-Mani, B.; Boller, T. Antifungal hydrolases in pea tissue II. Inhibition of fungal growth by combinations of chitinase and β-1,3-glucanase. Plant Physiol 1988, 88, 936–942. [Google Scholar]
- Maximova, S.N.; Marelli, J.P.; Young, A.; Pishak, S.; Verica, J.A.; Guiltinan, M.J. Over-expression of a cacao class I chitinase gene in Theobroma cacao L. enhances resistance against the pathogen Colletotrichum gloeosporioides. Planta 2006, 224, 740–749. [Google Scholar]
- Xiao, Y.H.; Li, X.B.; Yang, X.Y.; Luo, M.; Hou, L.; Guo, S.H.; Luo, X.Y.; Pei, Y. Cloning and characterization of a balsam pear class I chitinase gene (Mcchil1) and its ectopic expression enhances fungal resistance in transgenic plants. Biosci. Biotechnol. Biochem 2007, 71, 1211–1219. [Google Scholar]
- Liu, J.J.; Ekramoddoullah, A.K.; Zamani, A. A class IV chitinase is up-regulated by fungal infection and abiotic stresses and associated with slow-canker-growth resistance to Cronartium ribicola in western white pine (Pinus monticola). Phytopathology 2005, 95, 284–291. [Google Scholar]
- Meins, F.; Fritig, B.; Linthorst, H.J.; Mikkelsen, J.D.; Neuhaus, J.M.; Ryals, J. Plant chitinase genes. Plant Mol. Biol. Rep 1994, 12, S22–S28. [Google Scholar]
- Singh, A.; Isaac-Kirubakaran, S.; Sakthivel, N. Heterologous expression of new antifungal chitinase from wheat. Protein Expres. Purif 2007, 56, 100–109. [Google Scholar]
- Hoy, J.W.; Hollier, C.A.; Fontenot, D.B.; Grelen, L.B. Incidence of sugarcane smut in Louisiana and its effects on yield. Plant Dis 1986, 70, 59–60. [Google Scholar]
- Padmanaban, P.; Alexander, K.C.; Shanmugan, N. Effect of smut on growth and yield parameters of sugarcane. Indian Phytopath 1998, 41, 367–369. [Google Scholar]
- Que, Y.X.; Xu, L.P.; Lin, J.W.; Chen, R.K.; Grisham, M.P. Molecular variation of Sporisorium scitamineum in Mainland China revealed by RAPD and SRAP markers. Plant Dis 2012, 96, 1519–1525. [Google Scholar]
- Xu, L.P.; Chen, R.K.; Chen, P.H. Analysis on infection index of smut caused by Ustilago scitaminea in sugarcane segregated population. Chin. J. Trop Crop 2004, 25, 33–36. [Google Scholar]
- Does, M.P.; Houterman, P.M.; Dekker, H.L.; Cornelissen, B.J. Processing, targeting, and antifungal activity of stinging nettle agglutinin in transgenic tobacco. Plant physiol 1999, 120, 421–432. [Google Scholar]
- Kovacs, G.; Sagi, L.; Jacon, G.; Arinaitwe, G.; Busogoro, J.P.; Thiry, E.; Strosse, H.; Swennen, R.; Remy, S. Expression of a rice chitinase gene in transgenic banana (“Gros Michel” AAA genome group) confers resistance to black leaf streak disease. Transgen. Res 2013, 22, 117–130. [Google Scholar]
- Kirubakaran, S.I.; Sakthivel, N. Cloning and overexpression of antifungal barley chitinase gene in Escherichia coli. Protein Expres. Purif 2007, 52, 159–166. [Google Scholar]
- Rahul, P.R.; Kumar, V.G.; Sathyabhama, M.; Viswanathan, R.; Sundar, A.R.; Malathi, P. Characterization and 3D structure prediction of chitinase induced in sugarcane during pathogenesis of Colletotrichum falcatum. J. Plant Biochem. Biot 2013, 1–8. [Google Scholar]
- Lin, S.; Zhou, Y.; Chen, G.; Zhang, Y.; Zhang, Y.; Ning, W.; Pan, D. Molecular responses to the fungal pathogen Gibberella fujikuroi in the leaves of chewing cane (Saccharum officinarum L.). Sugar Tech 2010, 12, 36–46. [Google Scholar]
- Li, D.M.; Staehelin, C.; Wang, W.T.; Peng, S.L. Molecular cloning and characterization of a chitinase-homologous gene from Mikania micrantha infected by Cuscuta campestris. Plant Mol. Biol. Rep 2010, 28, 90–101. [Google Scholar]
- Meuriot, F.; Noquet, C.; Avice, J.C.; Volenec, J.J.; Cunningham, S.M.; Sors, T.G.; Caillot, S.; Ourry, A. Methyl jasmonate alters N partitioning, N reserves accumulation and induces gene expression of a 32-kDa vegetative storage protein that possesses chitinase activity in Medicago sativa taproots. Physiol. Plantarum 2004, 120, 113–123. [Google Scholar]
- Takken, F.L.; Joosten, M.H. Plant resistance genes: Their structure, function and evolution. Eur. J. Plant Pathol 2000, 106, 699–713. [Google Scholar]
- Van-Loon, L.C. Pathogenesis-related proteins. Plant Mol. Biol 1985, 4, 111–116. [Google Scholar]
- Brogue, K.; Chet, I.; Holliday, M.; Cressman, R.; Biddle, P.; Knowlton, S.; Mauvais, C.J.; Broglie, R. Transgenic plants with enhanced resistance to the fungal pathogen Rhizoctonia solani. Science 1991, 254, 1194–1197. [Google Scholar]
- Krishnaveni, S.; Liang, G.H.; Muthukrishnan, S.; Manickam, A. Purification and partial characterization of chitinases from sorghum seeds. Plant Sci 1999, 144, 1–7. [Google Scholar]
- Viswanathan, R.; Malathi, P.; Sundar, A.R.; Aarthi, S.; Premkumari, S.M.; Padmanaban, P. Differential induction of chitinases and thaumatin-like proteins in sugarcane in response to infection by Colletotrichum falcatum causing red rot disease. J. Plant Dis. Protect 2005, 112, 417–425. [Google Scholar]
- Park, S.K.; Kim, C.W.; Kim, H.; Jung, J.S.; Harman, G.E. Cloning and high-level production of a chitinase from Chromobacterium sp. and the role of conserved or nonconserved residues on its catalytic activity. Appl. Microbiol. Biot 2007, 74, 791–804. [Google Scholar]
- Mackenbrock, U.; Vogelsang, R.; Barz, W. Isoflavone and pterocarpan malonylglucosides and β-1,3-glucan-and chitin-hydrolases are vacuolar constituents in chickpea (Cicer arietinum L.). Z. Naturforsch. C 1992, 47, 815–822. [Google Scholar]
- Viswanathan, R.; Samiyappan, R. Antifungal activity of chitinases produced by some fluorescent pseudomonads against Colletotrichum falcatum Went causing red rot disease in sugarcane. Microbiol. Res 2001, 155, 309–314. [Google Scholar]
- Gupta, K.; Agarwal, P.K.; Reddy, M.K.; Jha, B. SbDREB2A, an A-2 type DREB transcription factor from extreme halophyte Salicornia brachiata confers abiotic stress tolerance in Escherichia coli. Plant Cell Rep 2010, 29, 1131–1137. [Google Scholar]
- Guo, J.L.; Xu, L.P.; Fang, J.P.; Su, Y.C.; Fu, H.Y.; Que, Y.X.; Xu, L.P. A novel dirigent protein gene with highly stem-specific expression from sugarcane, response to drought, salt and oxidative stresses. Plant Cell Rep 2012, 31, 1801–1812. [Google Scholar]
- Chaurasia, N.; Mishra, Y.; Ai, L.C. Cloning expression and analysis of phytochelatin synthase (pcs) gene from Anabaena sp. PCC 7120 offering multiple stress tolerance in Escherichia coli. Biochem. Biophys. Res. Commun 2008, 376, 225–230. [Google Scholar]
- Porat, R.; Vinokur, V.; Holland, D.; Gregory-McCollum, T.; Droby, S. Isolation of a citrus chitinase cDNA and characterization of its expression in response to elicitation of fruit pathogen resistance. J. Plant Physiol 2001, 158, 1585–1590. [Google Scholar]
- Levine, A.; Tenhaken, R.; Dixon, R.; Lamb, C. H2O2 from the oxidative burst orchestrates the plant hypersensitive disease resistance response. Cell 1994, 79, 583–593. [Google Scholar]
- Que, Y.X.; Lin, J.W.; Song, X.X.; Xu, L.P.; Chen, R.K. Differential gene expression in sugarcane in response to challenge by fungal pathogen Ustilago scitaminea revealed by cDNA-AFLP. J. Biomed. Biotechnol 2011, 2011. [Google Scholar] [CrossRef]
- Moosawi-Jorf, S.A.; Mahin, B.I. In vitro detection of yeast-like and mycelial colonies of Ustilago scitaminea in tissue-cultured plantlets of sugarcane using polymerase chain reaction. J. Appl. Sci 2007, 7, 3768–3773. [Google Scholar]
- Su, Y.C.; Xu, L.P.; Xue, B.T.; Wu, Q.B.; Guo, J.L.; Wu, L.G.; Que, Y.X. Molecular cloning and characterization of two pathogenesis-related β-1,3-glucanase genes ScGluA1 and ScGluD1 from sugarcane infected by Sporisorium scitamineum. Plant Cell Rep 2013, 32, 1503–1519. [Google Scholar]
- Guo, J.L.; Xu, L.P.; Su, Y.C.; Wang, H.B.; Gao, S.W.; Xu, J.S.; Que, Y.X. ScMT2-1-3, a metallothionein gene of sugarcane, plays an important role in the regulation of heavy metal tolerance/accumulation. BioMed Res 2013, 2013. [Google Scholar] [CrossRef]
- Que, Y.X.; Yang, Z.X.; Xu, L.P.; Chen, R.K. Isolation and identification of differentially expressed genes in sugarcane infected by Ustilago scitaminea. Acta Agron. Sin 2009, 35, 452–458. [Google Scholar]
- Li, S.G.; Gu, J.G.; Jiang, R.B.; Niu, Y.C. The research on biocontrol Trichoderma spp. strains producing chitinase. Chin. Biotech. Bull 2009, 30, 135–138. [Google Scholar]
- CAP3 sequence assembly program. Available online: http://pbil.univ-lyon1.fr/cap3.php (accessed on 1 January 2014).
- ORF Finder online program. Available online: http://www.ncbi.nlm.nih.gov/gorf/gorf.html (accessed on 2 December 2013).
- ProtParam. Available online: http://web.expasy.org/protparam/ (accessed on 1 January 2005).
- SMART. Available online: http://smart.embl-heidelberg.de/ (accessed on 10 October 2011).
- NetNGlyc 1.0 Server. Available online: http://www.cbs.dtu.dk/services/NetNGlyc/ (accessed on 5 June 2013).
- SignalP 4.0 Server. Available online: http://www.cbs.dtu.dk/services/SignalP/ (accessed on 4 June 2013).
- TargetP 1.1 server. Available online: http://www.cbs.dtu.dk/services/TargetP/ (accessed on 4 October 2013).
- TMHMM Server v. 2.0. Available online: http://www.cbs.dtu.dk/services/TMHMM-2.0/ (accessed on 12 June 2013).
- MEGA5.05 software. Available online: http://www.megasoftware.net/ (accessed on 4 May 2011).
- Kumar, S.; Tamura, K.; Nei, M. MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 2004, 5, 150–163. [Google Scholar]
- Hwang, I.S.; Hwang, B.K. Requirement of the cytosolic interaction between pathogenesis-related protein10 and leucine-rich repeat protein1 for cell death and defense signaling in pepper. Plant Cell 2012, 24, 1675–1690. [Google Scholar]
- Su, Y.C.; Guo, J.L.; Ling, H.; Chen, S.S.; Wang, S.S.; Xu, L.P.; Allan, A.C.; Que, Y.X. Isolation of a novel peroxisomal catalase gene from sugarcane, which is responsive to biotic and abiotic stresses. PLos One 2014, 9, e84426. [Google Scholar]
- Que, Y.X.; Xu, L.P.; Xu, J.S.; Zhang, J.S.; Zhang, M.Q.; Chen, R.K. Selection of control genes in real-time qPCR analysis of gene expression in sugarcane. Chin. J. Trop Crop 2009, 30, 274–278. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−Δ ΔCt method. Methods 2001, 25, 402–408. [Google Scholar]
- Huckelhoven, R.; Fodor, J.; Trujillo, M.; Kogel, K.H. Barley Mla and Rar mutants compromised in the hypersensitive cell death response against Blumeria graminis f. sp. hordei are modified in their ability to accumulate reactive oxygen intermediates at sites of fungal invasion. Planta 2000, 212, 16–24. [Google Scholar]






| Primer | Sequence (5′-3′) | Strategy |
|---|---|---|
| ScChi-cDNAF | ATACAGGTCCATTTTGTCAGC | RT-PCR |
| ScChi-cDNAR | AAACCAAGCAAGCCCTAA | RT-PCR |
| ScChi-SublocF | TGCTCTAGAATGACGAACGGCTACC | Subcellular localization vector construction |
| ScChi-SublocR | GGACTAGTGTGGTTGGCAACGATCT | Subcellular localization vector construction |
| ScChi-32aF | CGGGATCCATGACGAACGGCTACCT | prokaryotic expression vector construction |
| ScChi-32aR | CCAAGCTTTCAGTGGTTGGCAACGA | prokaryotic expression vector construction |
| ScChi-QF | ACGGCTACGGCGACAACA | RT-qPCR |
| ScChi-QR | GTCCGCTGACCAGATGAAGAG | RT-qPCR |
| GAPDH-QF | CACGGCCACTGGAAGCA | RT-qPCR |
| GAPDH-QR | TCCTCAGGGTTCCTGATGCC | RT-qPCR |
| ScChi-1301F | TGCTCTAGAATGACGAACGGCTACCTG | Over expression vector construction |
| ScChi-1301R | CGGGATCCTCAGTGGTTGGCAA | Over expression vector construction |
| NtHSR201 F | CAGCAGTCCTTTGGCGTTGTC | RT-qPCR |
| NtHSR201 R | GCTCAGTTTAGCCGCAGTTGTG | RT-qPCR |
| NtHSR203 F | TGGCTCAACGATTACGCA | RT-qPCR |
| NtHSR203 R | GCACGAAACCTGGATGG | RT-qPCR |
| NtHSR51 F | TTGGGCAGAATAGATGGGTA | RT-qPCR |
| NtHSR51 R | TTTGGTGAAAGTCTTGGCTC | RT-qPCR |
| NtNPR1 F | GGCGAGGAGTCCGTTCTTTAA | RT-qPCR |
| NtNPR1 R | TCAACCAGGAATGCCACAGC | RT-qPCR |
| NtPR-1a/c F | AACCTTTGACCTGGGACGAC | RT-qPCR |
| NtPR-1a/c R | GCACATCCAACACGAACCGA | RT-qPCR |
| NtPR2 F | TGATGCCCTTTTGGATTCTATG | RT-qPCR |
| NtPR2 R | AGTTCCTGCCCCGCTTT | RT-qPCR |
| NtPR3 F | CAGGAGGGTATTGCTTTGTTAGG | RT-qPCR |
| NtPR3 R | CGTGGGAAGATGGCTTGTTGTC | RT-qPCR |
| NtEFE26 F | CGGACGCTGGTGGCATAAT | RT-qPCR |
| NtEFE26 R | CAACAAGAGCTGGTGCTGGATA | RT-qPCR |
| NtAccdeaminase F | TCTGAGGTTACTGATTTGGATTGG | RT-qPCR |
| NtAccdeaminase R | TGGACATGGTGGATAGTTGCT | RT-qPCR |
| NtEF1-α F | TGCTGCTGTAACAAGATGGATGC | RT-qPCR |
| NtEF1-α R | GAGATGGGGACAAAGGGGATT | RT-qPCR |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Su, Y.; Xu, L.; Fu, Z.; Yang, Y.; Guo, J.; Wang, S.; Que, Y. ScChi, Encoding an Acidic Class III Chitinase of Sugarcane, Confers Positive Responses to Biotic and Abiotic Stresses in Sugarcane. Int. J. Mol. Sci. 2014, 15, 2738-2760. https://doi.org/10.3390/ijms15022738
Su Y, Xu L, Fu Z, Yang Y, Guo J, Wang S, Que Y. ScChi, Encoding an Acidic Class III Chitinase of Sugarcane, Confers Positive Responses to Biotic and Abiotic Stresses in Sugarcane. International Journal of Molecular Sciences. 2014; 15(2):2738-2760. https://doi.org/10.3390/ijms15022738
Chicago/Turabian StyleSu, Yachun, Liping Xu, Zhiwei Fu, Yuting Yang, Jinlong Guo, Shanshan Wang, and Youxiong Que. 2014. "ScChi, Encoding an Acidic Class III Chitinase of Sugarcane, Confers Positive Responses to Biotic and Abiotic Stresses in Sugarcane" International Journal of Molecular Sciences 15, no. 2: 2738-2760. https://doi.org/10.3390/ijms15022738








