Abstract
Amino acid substitutions in influenza A virus are the main reasons for both antigenic shift and virulence change, which result from non-synonymous mutations in the viral genome. Nucleocapsid protein (NP), one of the major structural proteins of influenza virus, is responsible for regulation of viral RNA synthesis and replication. In this report we used LC-MS/MS to analyze tryptic digestion of nucleocapsid protein of influenza virus (A/Puerto Rico/8/1934 H1N1), which was isolated and purified by SDS poly-acrylamide gel electrophoresis. Thus, LC-MS/MS analyses, coupled with manual de novo sequencing, allowed the determination of three substituted amino acid residues R452K, T423A and N430T in two tryptic peptides. The obtained results provided experimental evidence that amino acid substitutions resulted from non-synonymous gene mutations could be directly characterized by mass spectrometry in proteins of RNA viruses such as influenza A virus.
1. Introduction
Influenza virus has long been a global health threat since 1918 [1]. Although the annually circulating strains of influenza virus are not very virulent, there is still concern that the genome of these seasonal virus strains can mutate to acquire the ability to cause mortality in humans [2,3]. Additionally, for the avian influenza virus strains that usually show no adaptation to a human host, the virus genome can mutate to allow the virus to cross the species barrier to infect humans. For example, H5N1, H1N1 and recently reported H7N9 virus strains have shown their ability to cause severe infections in humans [4–6].
Mutations in viral genomes, some of which are non-synonymous mutations and thus result in amino acid substitutions, are often detected by gene sequencing [7–9]. With the introduction of soft ionization techniques such as ESI and MALDI, characterization of large biomolecules such as proteins has been achieved with high sensitivity and accuracy. Mass spectrometry has been used to analyze several mutations in hemoglobin variants [10,11]. Up to seven amino acid substitutions in HA of influenza A virus were revealed by mass spectrometry [12].
As influenza A virus has a relatively high mutation rate, there will always be an urgent need to detect variation in amino acid sequences resulting from non-synonymous SNPs that may have functional consequences. While both DNA and RNA have served as targets for most genotyping screen strategies, the other major functional molecule, protein, has recently been explored as a source for proteotyping, wherein a variety of protein forms from a single gene are characterized through sophisticated mass spectrometric techniques [13]. Similar to DNA/RNA-based genotyping, proteotyping strategy can be applied on either a single protein [14] or on a proteome-wide scale [15]. Because influenza A virus continues to mutate to evolve, the previously established DNA/RNA-based PCR approaches often fail to detect the newly emerging strains due to sequence variation in primer and probe [16]. However, for the protein-based proteotyping strategy, the mutated peptides or modified peptides van be detected without the concerns in PCR approaches. Therefore, once the proteotyping strategy is optimized for any given strain, it should be effective to detect an array of isoforms of viral proteins, including the peptides upon modification and amino acid substitution. In this study, we report the characterization of the nucleocapsid protein (isolated and purified by SDS-PAGE) of influenza A virus by mass spectrometry. By manual interpretation of the MS/MS data, three amino acid substitutions were identified. The results indicated that mass spectrometry coupled with de novo peptide sequencing had the power to characterize the amino acid substitutions in proteins of RNA viruses such as influenza A virus.
2. Results and Discussion
2.1. Identification of NP Protein
Influenza virus was inoculated in chick embryos. Several serial passages were performed to enhance the rates of multigenic mutations. The virus particles were purified from the collected allantoic fluid and then lysed and separated on 12% SDS-PAGE. After staining with Colloidal Coomassie G250, two major bands were found at 15 and 56 kDa, respectively (Figure 1). The band at 56 kDa was cut off and subject to in-gel tryptic digestion. LC-MS/MS analysis of the obtained peptide mixture coupled with protein database searching identified a total of 18 unique peptides of nucleocapsid protein from influenza virus (A/Puerto Rico/8/1934 H1N1) (Table 1).

Figure 1.
Identification of nucleocapsid protein from purified influenza virus (A/Puerto Rico/8/1934 H1N1). The purified virus was lysed and separated on 12% SDS-PAGE. The upper band around 56 kDa was cut off and subjected to in-gel digestion, followed by mass spectrometric analysis. Database searching identified 18 tryptic peptides (labeled with red) from nucleocapsid protein of influenza A virus. Manual interpretation of the obtained MS/MS data identified three amino acid substitutions (R452K, T423A and N430T, highlighted with yellow) within two tryptic peptides (labeled with green).

Table 1.
Summary of tryptic peptides identified in nucleocapsid protein from influenza virus (A/Puerto Rico/8/1934 H1N1) by database searching.
Besides the peptides identified by database searching, two additional mutated peptides were determined by manual interpretation of the available data, in which three amino acid substitutions were identified. Accordingly, both database searching and manual interpretation of the obtained LC-MS/MS data allowed the assignment of a total of 20 unique peptide sequences.
2.2. Identification of AA Substitution of R452K
Interpretation of the MS/MS spectrum of the doubly-charged ion peak MP1 at m/z 856.40 (Figure 2) allowed the identification of a partial sequence of ESA, considering the ion series of m/z 1449.73, 1320.71, 1233.69, 1162.64 at the high mass end of the spectrum were y type fragment ions y13, y12, y11, y10, respectively. The sequence of ESA was readily to be located in one of the tryptic peptides of NP: MMESARPEDVSFQGR (447–461) with theoretical m/z value of 870.40 for its doubly-charged ion. Thus, a nominal mass shift of −28 Da was observed for the detected doubly-charged ion of MP1 in comparison with the molecular weight of the theoretical sequence of MMESARPEDVSFQGR (447–461) in NP, which might result from amino acid substitution of one of five residues in the theoretical sequence: R→Q/K, V→A, M→C, D→S or E→T. The possibility for amino acid substitution of M→C was readily eliminated because the Cys (C) residue would be chemically alkylated during sample preparation if Methionine (M447/448) was mutated into Cys (C). Noticeably, the fragment ion y9 at m/z 1034.53 adjacent to y10 ion (m/z 1162.64) in the high mass range of the MS/MS spectrum indicated that the residue next to the Alanine (A451) should be either K or Q, considering that the calculated difference between 1162.64 and 1034.53 was identical to the nominal mass of 128 of either of these two amino acid residues. In addition, substitution of Arginine (R452) with either Lysine (K) or Glutamine (Q) was also confirmed by the detection of the base peak at m/z 129, which was the immonium ion of either K or Q. Although both K and Q residues had identical nominal mass of 128, the exact masses of them were different (K with 128.095 and Q with 128.058). The precise mass difference between y9 and y10 was calculated as 128.11, suggesting that the R452 was substituted by K, but not Q. This conclusion was well supported by the precise mass data of the immonium ion detected at m/z 129.11, which was much closer to the theoretic mass data of immonium ion of K (129.1022) than that of Q (129.0659). The assignments of most y series ions (from y5 to y13) clearly demonstrated the internal sequence of PEDV, eliminating the possibilities of amino acid substitutions at E454, D455 and V456. It should be noticed that there was a Proline (P) in the sequence, at which internal fragmentation could occur. Some internal sequences such as PE, PED, PEDV and PEDVS were detected and assigned, confirming that amino acid substitution should occur at R452 but not E454, D455 and V456. Additionally, detection of some of the a and b series ions such as a2, a3, b2 and b3 indicated that E449 was not subject to amino acid substitution, confirming the substitution of R452→K.
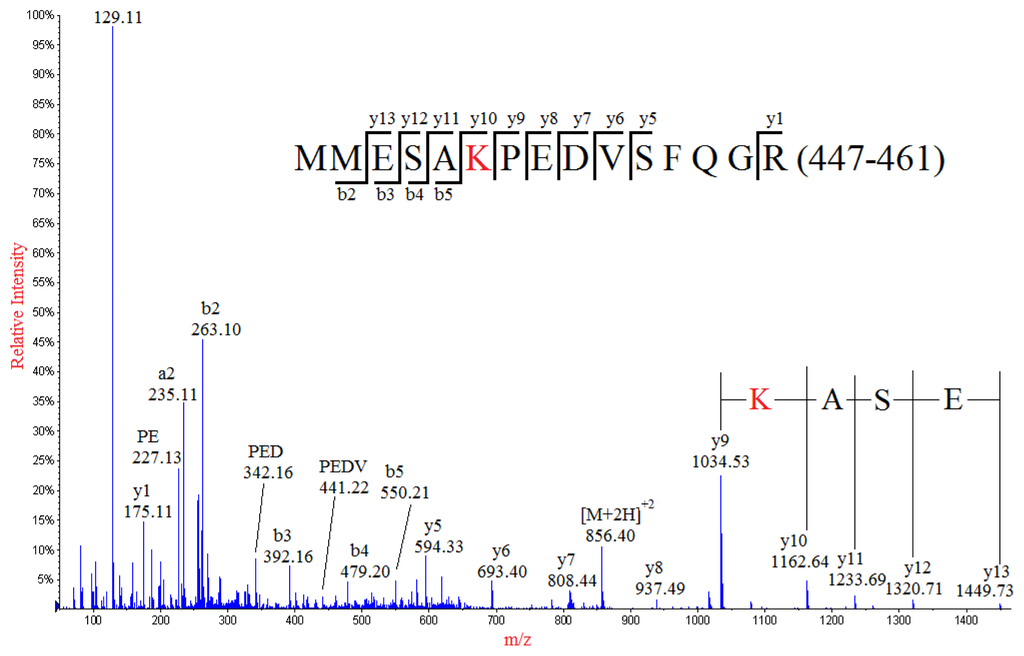
Figure 2.
MS/MS spectrum of the doubly-charged ion at m/z 856.40 from the analysis of peak MP1. The mutated peptide (MMESAKPEDVSFQGR) of a normal sequence (residues 447–461) from tryptic digestion of nucleocapsid protein was identified, in which the R452 was substituted with K.
2.3. Identification of AA Substitution T423A and N430T
De novo sequencing of the MS/MS spectrum of a doubly-charged ion peak MP2 at m/z 720.36 identified a partial sequence of TIMAAFT with y series ions at m/z 1368.68, 1267.71, 1154.61, 1023.56, 952.53, 881.46, 734.42 and 633.35 (Figure 3), which was not found in the theoretical sequence of NP. However, investigation of the theoretical sequence of NP revealed a sequence of TIMAAFN (424–430), which was identical to the deduced sequence except for the N430 residue. Therefore amino acid substitution of N430→T was identified, which resulted in a mass shift of −13.01 Da. The identified sequence was contained in a tryptic peptide of NP: TTIMAAFNGNTEGR (423–436), of which the calculated m/z value of the doubly-charged ion was 741.86. However, a nominal mass shift of −43 but not −13.01 Da (N430→T) was observed for MP2 when compared to the theoretical sequence of TTIMAAFNGNTEGR (423–436), suggesting that there might be at least one additional amino acid substitution in the sequence, which resulted in an additional mass shift of −30.01 Da. Investigation of the rest of the residues of the tryptic peptide sequence (423–436) indicated that there were three amino acid residues that could result in a mass shift of −30.01 Da upon substitution: T423→A, T433→A, E434→T. The detection of the y series ion y13 at m/z 1368.66, as well as b3-H2O, b3, b2-H2O and b2 ions, indicated that the first three residues in the peptide were ATI, thus confirming the identification of substitution of T423→A. Therefore, the peak MP2 was identified as the tryptic peptide in the residues from 423 to 436 with the two substitutions, namely T423→A and N430→T.
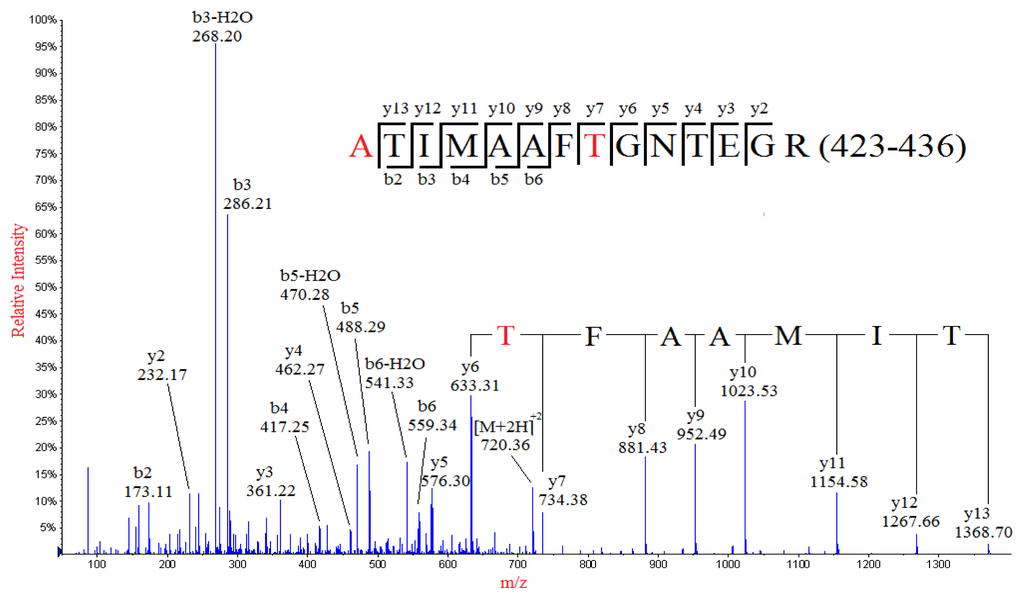
Figure 3.
MS/MS spectrum of the doubly-charged ion at m/z 720.36 from the analysis of peak MP2. The mutated peptide (ATIMAAFTGNTEGR) of a normal sequence (residues 423–436) from tryptic digestion of nucleocapsid protein was identified, in which the T423 and N430 were substituted with A and T, respectively.
2.4. Bioinformatics Analysis
The sequences of nucleocapsid proteins were exclusively retrieved from “The FLU project” at GenBank. A protein sequence database containing the retrieved sequences and the mutated sequence was built and subject to multiple alignment and linkage tree analysis (Figure 4). The output file containing the whole tree data can be found in supplemental materials.
Figure 4.
Part of detailed tree picture generated from linkage analysis of mutated nucleocapsid protein identified by MS/MS, by which the neighbor-joining method of clustering was used. The strain containing the mutated nucleocapsid protein (A/Puerto Rico/8/1934 (H1N1.mu)), as well as its original strain (A/Puerto Rico/8/1934 (H1N1)), is labeled with red rectangular box.
3. Experimental Section
3.1. Chemicals and Materials
Sequencing-grade TPCK-modified trypsin was purchased from Promega (Madison, WI, USA). Bradford protein assay kit, ammonium bicarbonate, dithiothreitol (DTT), iodoacetamide (IAA) were purchased from Bio-Rad (Hercules, CA, USA). All the other chemicals were purchased from Sigma-Aldrich (St. Louis, MO, USA). Influenza virus (A/Puerto Rico/8/1934 H1N1) was propagated in a biosafety level 2 (BL-2) containment facilities. Ultra-pure water was prepared by a MilliQ water purification system (Millpore, Bedford, MA, USA).
3.2. Virus Cultivation and Purification
Embryonated chicken eggs were inoculated with the influenza A virus (A/Puerto Rico/8/1934 H1N1) and incubated for 72 h at 37 °C. The allantoic fluid was harvested, followed by centrifugation at 5000 rpm for 15 min. The virus in the allantoic fluid was pelleted through a 4-step discontinuous gradient cushion consisting of 30%, 40%, 50% and 60% (w/v) sucrose, in a SW40 Ti rotor (Beckman-coulter, Fullerton, CA, USA) at 35,000 rpm at 4 °C for 60 min. The virus band between 40% and 50% sucrose was carefully collected, and suspended in 10 mM Tris-HCl pH 8.0, 150 mM NaCl. Aliquots of the purified virus sample were kept at 4 °C.
3.3. SDS-PAGE
The purified virus particles were lysed with 2× Laemmli sample buffer and kept at 95 °C for 5 min. The protein concentration was assayed with Micro BCA (bicinchoninic acid) protein assay kit (Pierce, Rockford, IL, USA). Electrophoretic separation was performed in a Mini-Cell system (Bio-Rad, Hercules, CA, USA), and run in 12% tris-glycine-SDS polyacrylamide gels with a 5% stacking gel. After electrophoresis, the gels were stained with colloidal Coomassie G250 and scanned with a calibrated densitometer (GS800, Bio-Rad).
3.4. In-Gel Digestion
Protein bands of interest were cut off from gels and washed with Milli-Q water three times. Then the gel pieces were destained with a solution of 50 mM NH4HCO3 in 50% ACN until the Coomassie blue in the gel became invisible. The destained gel pieces were reduced in 10 mM DTT, 50 mM NH4HCO3 aqueous solution at 60 °C for 60 min, followed by alkylation in 50 mM IAA, 50 mM NH4HCO3 aqueous solution at room temperature in dark for 30 min. The gel pieces were dehydrated with ACN, and then incubated in freshly prepared digestion solution containing 50 mM NH4HCO3 and 0.1 g/L TPCK-trypsin overnight at 37 °C. The resulting tryptic peptides were extracted with 5% trifluoroacetic acid (TFA) in 60% ACN and stored at −20 °C until LC-MS/MS analysis.
3.5. Capillary LC-MS/MS Analysis
The tryptic peptides were lyophilized and redissolved in high performance liquid chromatography (HPLC) buffer A (0.1% formic acid) and then separated on a C18 column (100 mm × 180 μm i.d.). The elution gradient was from 5% to 40% buffer B (0.1% formic acid, 99% ACN, flow rate: 0.2 μL/min) for 90 min. The eluted peptides were then analyzed on an ABI QSTAR spectrometer using information dependent acquisition mode (IDA; Analyst QS, Applied Biosystems, Carlsbad, CA, USA) by selecting the three most intense ions for MS/MS analysis. A survey scan of 300–2000 Da was collected for 3 s followed by 5 s MS/MS scans of 40–1500 Da using the standard rolling collision energy settings. The dynamic exclusion time was set as 1.5 min.
MASCOT generic files were generated from the obtained MS data by using a script embedded in the Analyst QS 2.0 software (MDS Sciex, South San Francisco, CA, USA) and used to search against the Swiss-Prot protein database on a local MASCOT server (version 2.1, Matrix Science, London, UK). One missed cleavage was allowed. Carbamidomethylation of cysteines was specified as fixed modification, whereas oxidation of methionine was selected as variable modification. The mass tolerance was set to 0.3 and 0.6 Da for peptide and MS/MS ion masses, respectively. Manual de novo sequencing of peptide tandem mass spectra was performed with the aid of Pepsea (1.1) in Analyst QS 2.0 software (MDS Sciex).
3.6. Bioinformatics Analysis
The mutated nucleocapsid protein containing three amino acid substitutions was analyzed by using a suite of bioinformatics tools at NCBI ( http://www.ncbi.nlm.nih.gov/genomes/FLU/FLU.html) [17].
4. Conclusions
We herein identified by LC-MS/MS analysis three amino acid substitutions in nucleocapsid protein from influenza virus (A/Puerto Rico/8/1934 H1N1). The three amino acid substitutions were located in two tryptic peptides of the nucleocapsid protein. One of identified amino acid substitutions, R452K, was located within the tryptic peptide MP1 (447–461), whereas the other two amino acid substitutions, T423A and N430T, were located within tryptic peptide MP2 (423–436). Both of the peptides were identified through manual interpretation of the relating MS/MS data, which included both calculation of high resolution MS data and assignment of fragment ions in MS/MS data. The outcome of this study indicated that the MS/MS analysis of amino acid substitutions might be useful in investigating the antigens from influenza viruses.
Acknowledgments
This work was supported by Major State Basic Research Development Program of China (973 Program) (2011CB503900), Natural Science Foundation of China (81070078, 21175055, 81270157), Beijing Municipal Natural Science Foundation (7102158), Jilin Province Science and Technology Department (20110739), Jilin University Bethune Project B (2012210).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Tang, J.W.; Shetty, N.; Lam, T.T.; Hon, K.L. Emerging, novel, and known influenza virus infections in humans. Infect. Dis. Clin. N. Am 2010, 24, 603–617. [Google Scholar]
- Mitchell, R.; Taylor, G.; McGeer, A.; Frenette, C.; Suh, K.N.; Wong, A.; Katz, K.; Wilkinson, K.; Amihod, B.; Gravel, D. Canadian Nosocomial Infection Surveillance Program. Understanding the burden of influenza infection among adults in Canadian hospitals: A comparison of the 2009–2010 pandemic season with the prepandemic and postpandemic seasons. Am. J. Infect. Control 2013, 41, 1032–1037. [Google Scholar]
- Sheu, T.G.; Fry, A.M.; Garten, R.J.; Deyde, V.M.; Shwe, T.; Bullion, L.; Peebles, P.J.; Li, Y.; Klimov, A.I.; Gubareva, L.V. Dual resistance to adamantanes and oseltamivir among seasonal influenza A(H1N1) viruses: 2008–2010. J. Infect. Dis 2011, 203, 13–17. [Google Scholar]
- Punpanich, W.; Chotpitayasunondh, T. A review on the clinical spectrum and natural history of human influenza. Int. J. Infect. Dis 2012, 16, e714–e723. [Google Scholar]
- Dortmans, J.C.; Dekkers, J.; Wickramasinghe, I.N.; Verheije, M.H.; Rottier, P.J.; van Kuppeveld, F.J.; de Vries, E.; de Haan, C.A. Adaptation of novel H7N9 influenza A virus to human receptors. Sci. Rep 2013, 3, 3058. [Google Scholar]
- Crusat, M.; Liu, J.; Palma, A.S.; Childs, R.A.; Liu, Y.; Wharton, S.A.; Lin, Y.P.; Coombs, P.J.; Martin, S.R.; Matrosovich, M.; et al. Changes in the hemagglutinin of H5N1 viruses during human infection-influence on receptor binding. Virology 2013, 447, 326–337. [Google Scholar]
- Mahony, J.B. Nucleic acid amplification-based diagnosis of respiratory virus infections. Expert Rev. Anti-Infect. Therapy 2010, 8, 1273–1292. [Google Scholar]
- Takayama, I.; Nakauchi, M.; Fujisaki, S.; Odagiri, T.; Tashiro, M.; Kageyama, T. Rapid detection of the S247N neuraminidase mutation in influenza A(H1N1)pdm09 virus by one-step duplex RT-PCR assay. J. Virol. Methods 2013, 188, 73–75. [Google Scholar]
- Rodríguez-Sánchez, B.; Alonso, M.; Catalán, P.; Sánchez Conde, M.; González-Candelas, F.; Giannella, M.; Bouza, E.; García de Viedma, D. Genotyping of a nosocomial outbreak of pandemic influenza A/H1N1 2009. J. Clin. Virol 2011, 52, 129–132. [Google Scholar]
- Haynes, C.A.; Guerra, S.L.; Fontana, J.C.; DeJesús, V.R. HPLC-ESI-MS/MS analysis of hemoglobin peptides in tryptic digests of dried-blood spot extracts detects HbS, HbC, HbD, HbE, HbO-Arab, and HbG-Philadelphia mutations. Clin. Chim. Acta 2013, 424, 191–200. [Google Scholar]
- Théberge, R.; Infusini, G.; Tong, W.; McComb, M.E.; Costello, C.E. Top-down analysis of small plasma proteins using an LTQ-Orbitrap. Potential for mass spectrometry-based clinical assays for transthyretin and hemoglobin. Int. J. Mass Spectrom 2011, 300, 130–142. [Google Scholar]
- Liu, N.; Wang, G.; Lee, K.C.; Guan, Y.; Chen, H.; Cai, Z. Mutations in influenza virus replication and transcription: Detection of amino acid substitutions in hemagglutinin of an avian influenza virus (H1N1). FASEB J 2009, 23, 3377–3382. [Google Scholar]
- Roth, M.J.; Parks, B.A.; Ferguson, J.T.; Boyne, M.T., II; Kellehe, N.L. Proteotyping: Population proteomics of human leukocytes using top down mass spectrometry. Anal. Chem 2008, 80, 2857–2866. [Google Scholar]
- Liu, N.; Song, W.; Lee, K.C.; Wang, P.; Chen, H.; Cai, Z. Identification of amino acid substitutions in avian influenza virus (H5N1) matrix protein 1 by using nanoelectrospray MS and MS/MS. J. Am. Soc. Mass Spectrom 2009, 20, 312–320. [Google Scholar]
- Sheynkman, G.M.; Shortreed, M.R.; Frey, B.L.; Scalf, M.; Smith, L.M. Large-scale mass spectrometric detection of variant peptides resulting from nonsynonymous nucleotide differences. J. Proteome Res 2014, 13, 228–240. [Google Scholar]
- Lalle, E.; Bordi, L.; Castilletti, C.; Meschi, S.; Selleri, M.; Carletti, F.; Lapa, D.; Travaglini, D.; Ippolito, G.; Capobianchi, M.R.; et al. Design and clinical application of a molecular method for detection and typing of the influenza A/H1N1pdm virus. J. Virol. Methods 2010, 163, 486–488. [Google Scholar]
- Influenza Virus Resource. Available online: http://www.ncbi.nlm.nih.gov/genomes/FLU/FLU.html (accessed on 26 January 2014).
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).