Identification and Molecular Characterization of a Chitin Deacetylase from Bombyx mori Peritrophic Membrane
Abstract
:1. Introduction
2. Results and Discussion
2.1. Proteomic Analysis of B. mori PM
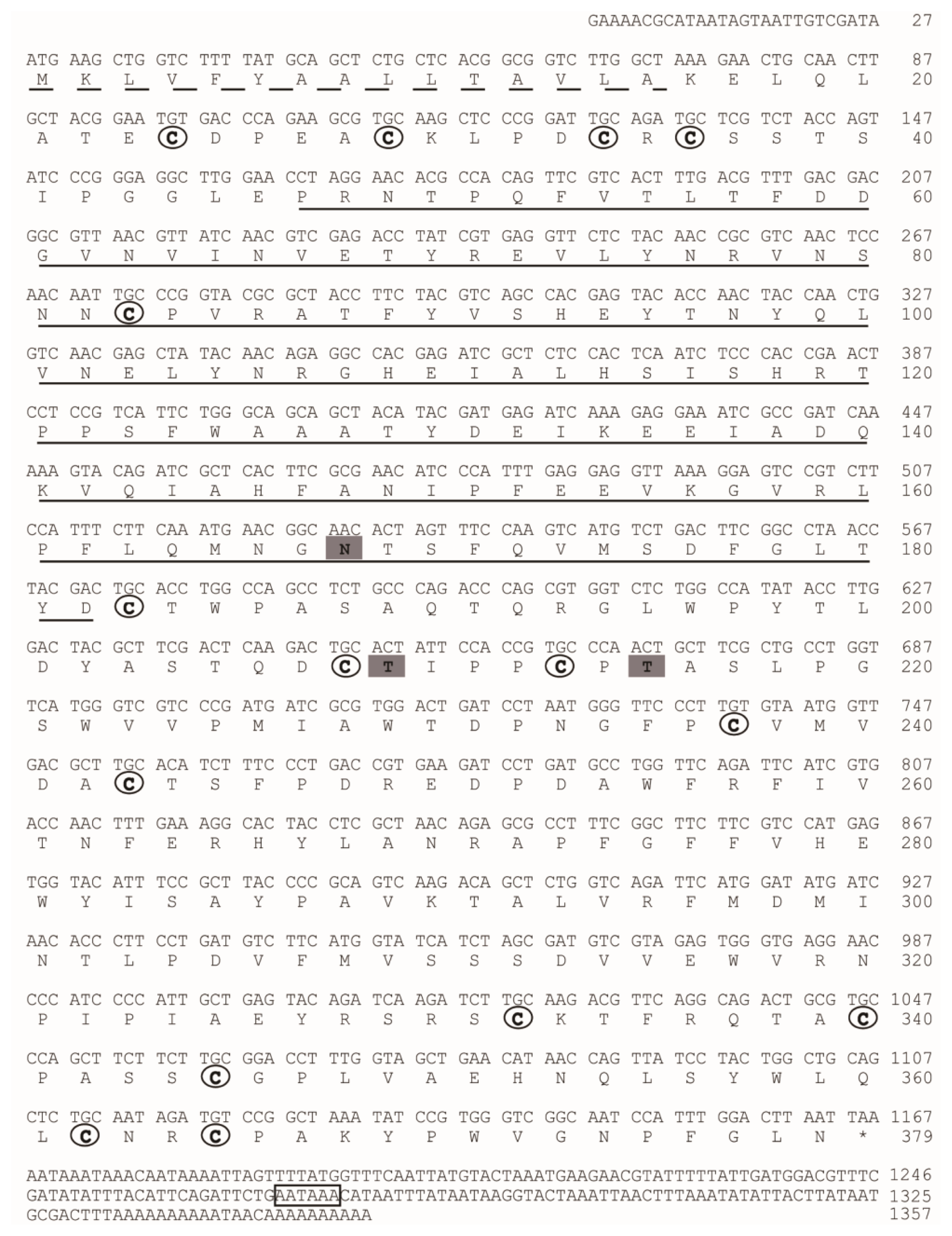
2.2. Sequence Analysis of BmCDA7
2.3. Recombinant Expression and Chitin Deacetylase Activity
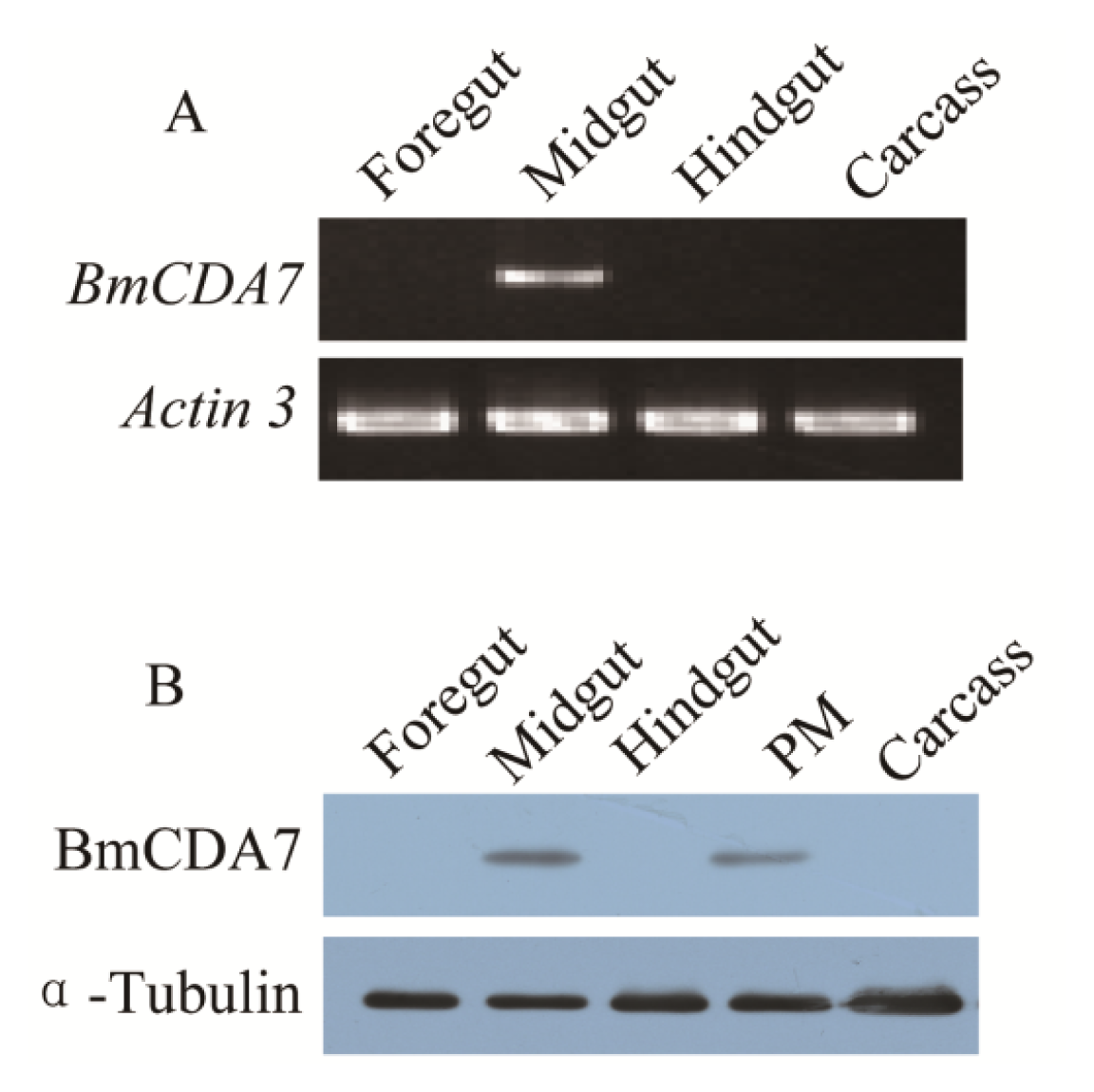
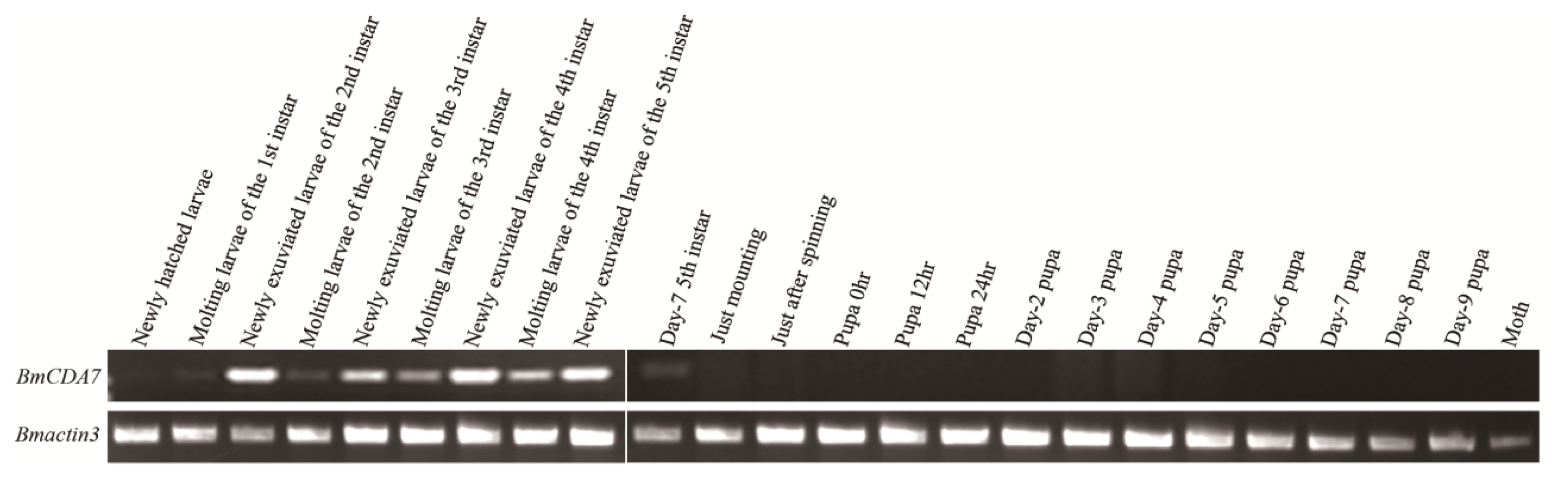
2.4. Expression Profiles and Localization of BmCDA7
3. Experimental Section
3.1. Silkworm Rearing and Sample Preparation
3.2. 2-DE and Image Analysis
3.3. Protein Digestion and Protein Identification by MS
3.4. Rapid Amplification of cDNA 5′ End (5′ RACE) and 3′ RACE
3.5. Phylogenetic Analysis
3.6. Expression Profile Analysis of Putative BmCDA7
3.7. Prokaryotic Expression, Preparation of Antiserum and Immunobloting
3.8. Eukaryotic Expression and Enzyme Assay
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Wang, P.; Granados, R.R. Molecular structure of the peritrophic membrane (PM): Identification of potential PM target sites for insect control. Arch. Insect Biochem. Physiol 2001, 47, 110–118. [Google Scholar]
- Tellam, R.L.; Wijffels, G.; Willadsen, P. Peritrophic matrix proteins. Insect Biochem. Mol. Biol 1999, 29, 87–101. [Google Scholar]
- Hegedus, D.; Erlandson, M.; Gillott, C.; Toprak, U. New insights into peritrophic matrix synthesis, architecture, and function. Annu. Rev. Entomol 2009, 54, 285–302. [Google Scholar]
- Lehane, M.J.; Billingsley, P.F. Biology of the insect midgut. Chapmann Hall Lond 1996, 86–114. [Google Scholar]
- Shen, Z.; Jacobs-Lorena, M. A type I peritrophic matrix protein from the malaria vector Anopheles gambiae binds to chitin. Cloning, expression, and characterization. J. Biol. Chem 1998, 273, 17665–17670. [Google Scholar]
- Wijffels, G.; Eisemann, C.; Riding, G.; Pearson, R.; Jones, A.; Willadsen, P.; Tellam, R. A novel family of chitin-binding proteins from insect type 2 peritrophic matrix. cDNA sequences, chitin binding activity, and cellular localization. J. Biol. Chem 2001, 276, 15527–15536. [Google Scholar]
- Wang, P.; Granados, R.R. An intestinal mucin is the target substrate for a baculovirus enhancin. Proc. Natl. Acad. Sci. USA 1997, 94, 6977–6982. [Google Scholar]
- Cohen, E. Chitin biochemistry: Synthesis, hydrolysis and inhibition. Adv. Insect Physiol 2010, 38, 5–74. [Google Scholar]
- Araki, Y.; Ito, E. A pathway of chitosan formation in Mucor rouxii. Enzymatic deacetylation of chitin. Eur. J. Biochem 1975, 55, 71–78. [Google Scholar]
- Davis, L.L.; Bartnicki-Garcia, S. Chitosan synthesis by the tandem action of chitin synthetase and chitin deacetylase from Mucor rouxii. Biochemistry 1984, 23, 1065–1073. [Google Scholar]
- Alfonso, C.; Nuero, O.M.; Santamaria, F.; Reyes, F. Purification of a heat-stable chitin deacetylase from Aspergillus nidulans and its role in cell wall degradation. Curr. Microbiol 1995, 30, 49–54. [Google Scholar]
- Kauss, H.; Jeblick, W.; Domard, A. The degrees of polymerization and N-acetylation of chitosan determine its ability to elicit callose formation in suspension cells and protoplasts of Catharanthus roseus. Planta 1989, 178, 385–392. [Google Scholar]
- Tsigos, I.; Bouriotis, V. Purification and characterization of chitin deacetylase from Colletotrichum lindemuthianum. J. Biol. Chem 1995, 270, 26286–26291. [Google Scholar]
- Zhao, Y.; Park, R.D.; Muzzarelli, R.A. Chitin deacetylases: Properties and applications. Mar. Drugs 2010, 8, 24–46. [Google Scholar]
- Guo, W.; Li, G.; Pang, Y.; Wang, P. A novel chitin-binding protein identified from the peritrophic membrane of the cabbage looper Trichoplusia ni. Insect Biochem. Mol. Biol 2005, 35, 1224–1234. [Google Scholar]
- Luschnig, S.; Batz, T.; Armbruster, K.; Krasnow, M.A. Serpentine and vermiform encode matrix proteins with chitin binding and deacetylation domains that limit tracheal tube length in Drosophila. Curr. Biol 2006, 16, 186–194. [Google Scholar]
- Wang, S.; Jayaram, S.A.; Hemphala, J.; Senti, K.A.; Tsarouhas, V.; Jin, H.; Samakovlis, C. Septate-junction-dependent luminal deposition of chitin deacetylases restricts tube elongation in the Drosophila trachea. Curr. Biol 2006, 16, 180–185. [Google Scholar]
- Arakane, Y.; Dixit, R.; Begum, K.; Park, Y.; Specht, C.A.; Merzendorfer, H.; Kramer, K.J.; Muthukrishnan, S.; Beeman, R.W. Analysis of functions of the chitin deacetylase gene family in Tribolium castaneum. Insect Biochem. Mol. Biol 2009, 39, 355–365. [Google Scholar]
- Jakubowska, A.K.; Caccia, S.; Gordon, K.H.; Ferre, J.; Herrero, S. Downregulation of a chitin deacetylase-like protein in response to baculovirus infection and its application for improving baculovirus infectivity. J. Virol 2010, 84, 2547–2555. [Google Scholar]
- Toprak, U.; Baldwin, D.; Erlandson, M.; Gillott, C.; Hou, X.; Coutu, C.; Hegedus, D.D. A chitin deacetylase and putative insect intestinal lipases are components of the Mamestra configurata (Lepidoptera: Noctuidae) peritrophic matrix. Insect Mol. Biol 2008, 17, 573–585. [Google Scholar]
- Goldsmith, M.R.; Shimada, T.; Abe, H. The genetics and genomics of the silkworm Bombyx mori. Annu. Rev. Entomol 2005, 50, 71–100. [Google Scholar]
- Ramos, A.; Mahowald, A.; Jacobs-Lorena, M. Peritrophic matrix of the black fly Simulium vittatum: Formation, structure, and analysis of its protein components. J. Exp. Zool 1994, 268, 269–281. [Google Scholar]
- Lehane, M.J.; Allingham, P.G.; Weglicki, P. Composition of the peritrophic matrix of the tsetse fly Glossina morsitans morsitans. Cell Tissue Res 1996, 283, 375–384. [Google Scholar]
- Moskalyk, L.A.; Oo, M.M.; Jacobs-Lorena, M. Peritrophic matrix proteins of Anopheles gambiae and Aedes aegypti. Insect Mol. Biol 1996, 5, 261–268. [Google Scholar]
- Hu, X.; Chen, L.; Xiang, X.; Yang, R.; Yu, S.; Wu, X. Proteomic analysis of peritrophic membrane (PM) from the midgut of fifth-instar larvae Bombyx mori. Mol. Biol. Rep 2012, 39, 3427–3434. [Google Scholar]
- Zhong, X.; Zhang, L.; Zou, Y.; Yi, Q.; Zhao, P.; Xia, Q.; Xiang, Z. Shotgun analysis on the peritrophic membrane of the silkworm Bombyx mori. BMB Rep 2012, 45, 665–670. [Google Scholar]
- Wang, P.; Li, G.; Granados, R.R. Identification of two new peritrophic membrane proteins from larval Trichoplusia ni: Structural characteristics and their functions in the protease rich insect gut. Insect Biochem. Mol. Biol 2004, 34, 215–227. [Google Scholar]
- Dixit, R.; Arakane, Y.; Specht, C.A.; Richard, C.; Kramer, K.J.; Beeman, R.W.; Muthukrishnan, S. Domain organization and phylogenetic analysis of proteins from the chitin deacetylase gene family of Tribolium castaneum and three other species of insects. Insect Biochem. Mol. Biol 2008, 38, 440–451. [Google Scholar]
- Petersen, T.N.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol 1999, 112, 531–552. [Google Scholar]
- Gupta, R.; Jung, E.; Brunak, S. Prediction of N-glycosylation sites in human proteins. In preparation.
- Julenius, K.; Molgaard, A.; Gupta, R.; Brunak, S. Prediction, conservation analysis, and structural characterization of mammalian mucin-type O-glycosylation sites. Glycobiology 2005, 15, 153–164. [Google Scholar]
- Letunic, I.; Doerks, T.; Bork, P. SMART 7: Recent updates to the protein domain annotation resource. Nucleic Acids Res 2012, 40, D302–D305. [Google Scholar]
- Wang, P.; Granados, R.R. Molecular cloning and sequencing of a novel invertebrate intestinal mucin cDNA. J. Biol. Chem 1997, 272, 16663–16669. [Google Scholar]
- Liu, L.; ZHAO, X.P.; Liu, J.J. Screening of chitin deacetylase producing strain. China Brew 2011, 30, 61–64. [Google Scholar]
- Wenling, C.; Duohui, J.; Jiamou, L.; Yandao, G.; Nanming, Z.; Xiufang, Z. Effects of the degree of deacetylation on the physicochemical properties and Schwann cell affinity of chitosan films. J. Biomater. Appl 2005, 20, 157–177. [Google Scholar]
- Ghormade, V.; Kulkarni, S.; Doiphode, N.; Rajamohanan, P.R.; Deshpande, M.V. Chitin deacetylase: A comprehensive account on its role in nature and its biotechnological applications. Curr. Res. Tecnol. Educ. Top. Appl. Micobiol. Microbial. Botechnol 2010, 2, 1054–1066. [Google Scholar]
- Nahar, P.; Ghormade, V.; Deshpande, M.V. The extracellular constitutive production of chitin deacetylase in Metarhizium anisopliae: Possible edge to entomopathogenic fungi in the biological control of insect pests. J. Invertebr. Pathol 2004, 85, 80–88. [Google Scholar]
- Yan, J.X.; Wait, R.; Berkelman, T.; Harry, R.A.; Westbrook, J.A.; Wheeler, C.H.; Dunn, M.J. A modified silver staining protocol for visualization of proteins compatible with matrix-assisted laser desorption/ionization and electrospray ionization-mass spectrometry. Electrophoresis 2000, 21, 3666–3672. [Google Scholar]
- Hou, Y.; Zou, Y.; Wang, F.; Gong, J.; Zhong, X.; Xia, Q.; Zhao, P. Comparative analysis of proteome maps of silkworm hemolymph during different developmental stages. Proteome Sci 2010, 8, 45. [Google Scholar]
- Zhong, X.W.; Zhao, P.; Zou, Y.; Nie, H.Y.; Yi, Q.Y.; Xia, Q.Y.; Xiang, Z.H. Proteomic analysis of the immune response of the silkworm infected by Escherichia coli and Bacillus bombyseptieus. Insect Sci 2012, 19, 559–569. [Google Scholar]
- Xia, Q.; Zhou, Z.; Lu, C.; Cheng, D.; Dai, F.; Li, B.; Zhao, P.; Zha, X.; Cheng, T.; Chai, C.; et al. A draft sequence for the genome of the domesticated silkworm (Bombyx mori). Science 2004, 306, 1937–1940. [Google Scholar]
- Duan, J.; Li, R.; Cheng, D.; Fan, W.; Zha, X.; Cheng, T.; Wu, Y.; Wang, J.; Mita, K.; Xiang, Z.; et al. SilkDB v2.0: A platform for silkworm (Bombyx mori) genome biology. Nucleic Acids Res 2010, 38, D453–D456. [Google Scholar]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol 2011, 28, 2731–2739. [Google Scholar]
- Fan, Y.; Zhang, Y.; Yang, X.; Pei, X.; Guo, S.; Pei, Y. Expression of a Beauveria bassiana chitinase (Bbchit1) in Escherichia coli and Pichia pastoris. Protein Expr. Purif 2007, 56, 93–99. [Google Scholar]
- Li, L.P. Screening of the Strains Producing Chitin Deacetylase. Master Thesis, Shandong Agricultural University, Taian, China, 2007; pp. 19–31. [Google Scholar]
- Srinivasan, V.R. Biotransformation of Chitin to Chitosan. U.S. Patent 5739015, 14 April 1998. [Google Scholar]
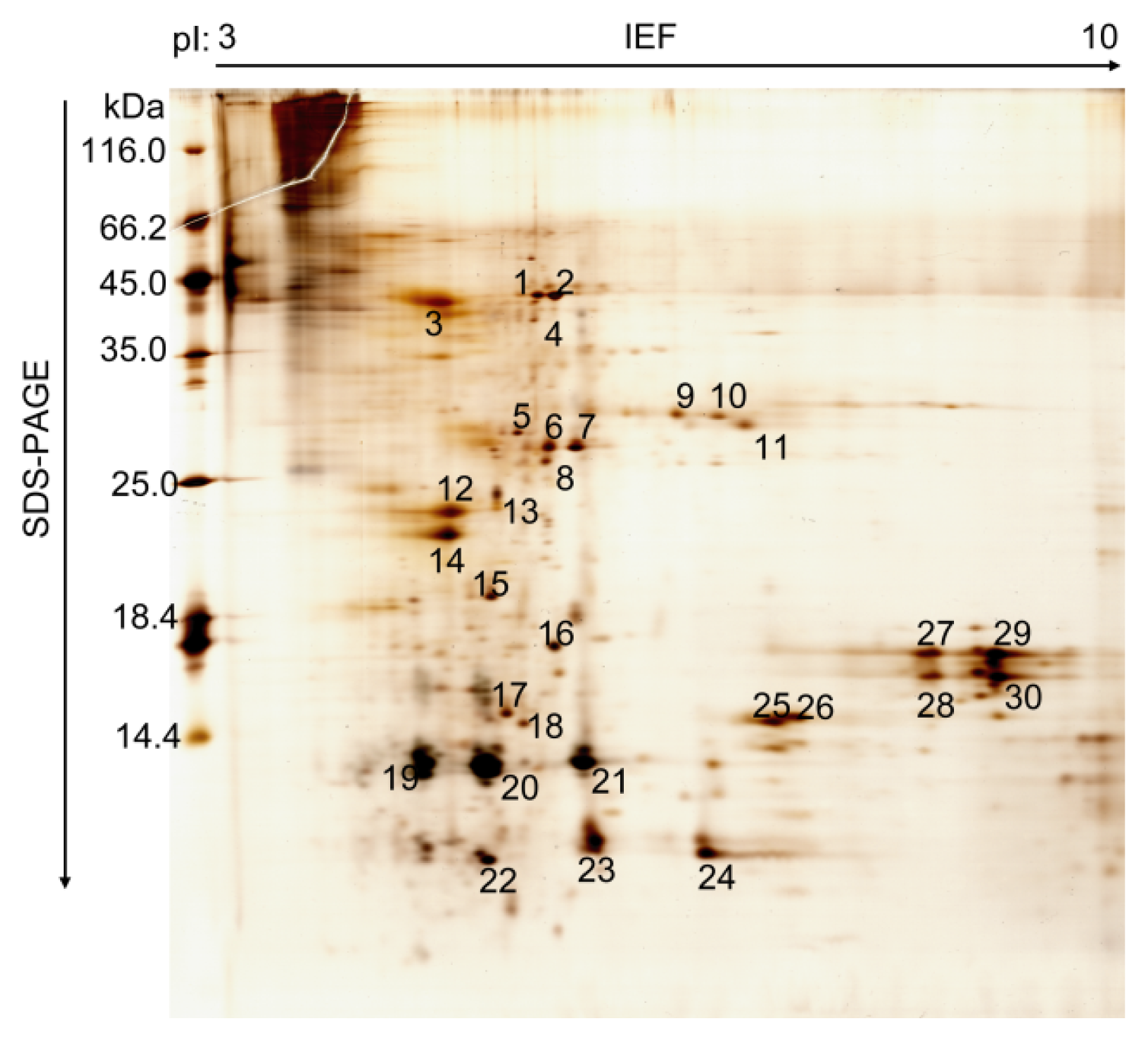


| Spot | Protein Name | Cell Function | NCBI Entry | SilkDB Entry | Mr (kDa) | pI | Seq. (%) |
|---|---|---|---|---|---|---|---|
| 1 | H+ transporting ATP synthase beta subunit isoform 1 | Proton pump, Proton transport | NP_001040450.1 | BGIBMGA012549 | 54.8 | 5.1 | 41.3 |
| 2 | H+ transporting ATP synthase beta subunit isoform 2 | Proton pump, Proton transport | NP_001041705.1 | BGIBMGA012555 | 55.0 | 5.2 | 23.8 |
| 4 | Chitin deacetylase | Enzymatic deacetylation of chitin | - | BGIBMGA013757 | 41.4 | 5.2 | |
| 5 | Unknown | ||||||
| 6 | Actin3 | Cytoskeleton | P04829 | BGIBMGA013945 | 41.9 | 5.3 | 50.0 |
| 7 | Chaperonin | Protein folding | NP_001073348 | BGIBMGA011508 | 59.2 | 5.3 | 24.2 |
| 8–10 | Unknown | ||||||
| 11 | Signal sequence receptor beta subunit | Translocon-associated protein | NP_001040332 | BGIBMGA005769 | 16.4 | 6.9 | |
| 12 | Translationally controlled tumor protein | Translation factor activity | NP_001037572.1 | BGIBMGA003073 | 19.6 | 4.5 | 30.2 |
| 13 | Juvenile hormone binding protein | Carring hormone | - | BGIBMGA010979 | 23.5 | 5.2 | |
| 14 | Unknown | ||||||
| 15 | Chlorophyllide A binding protein precursor | Candidate receptor proteins of Bt | NP_001037071 | BGIBMGA004806 | 315.5 | 5.1 | |
| 16 | ADP-ribosylation factor-like protein | Unknown | NP_001040168 | BGIBMGA010943 | 21.3 | 6.5 | 39.0 |
| 17 | Peritrophic membrane chitin binding protein 2 | Chitin binding protein | - | BGIBMGA001491 | 15.2 | 7.5 | 20.4 |
| 18–24 | Unknown | ||||||
| 25–30 | Triacylglycerol lipase | Lipid metabolism | NP_001040159 | BGIBMGA010400 | 31.8 | 8.8 | 30.8 |
| Absorbance | Activity | |
|---|---|---|
| A400 | A0 | EA |
| 0.36 | 0.21 | 1.85 |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhong, X.-W.; Wang, X.-H.; Tan, X.; Xia, Q.-Y.; Xiang, Z.-H.; Zhao, P. Identification and Molecular Characterization of a Chitin Deacetylase from Bombyx mori Peritrophic Membrane. Int. J. Mol. Sci. 2014, 15, 1946-1961. https://doi.org/10.3390/ijms15021946
Zhong X-W, Wang X-H, Tan X, Xia Q-Y, Xiang Z-H, Zhao P. Identification and Molecular Characterization of a Chitin Deacetylase from Bombyx mori Peritrophic Membrane. International Journal of Molecular Sciences. 2014; 15(2):1946-1961. https://doi.org/10.3390/ijms15021946
Chicago/Turabian StyleZhong, Xiao-Wu, Xiao-Huan Wang, Xiang Tan, Qing-You Xia, Zhong-Huai Xiang, and Ping Zhao. 2014. "Identification and Molecular Characterization of a Chitin Deacetylase from Bombyx mori Peritrophic Membrane" International Journal of Molecular Sciences 15, no. 2: 1946-1961. https://doi.org/10.3390/ijms15021946





