In Silico Identification and Characterization of N-Terminal Acetyltransferase Genes of Poplar (Populus trichocarpa)
Abstract
:1. Introduction
2. Results and Discussion
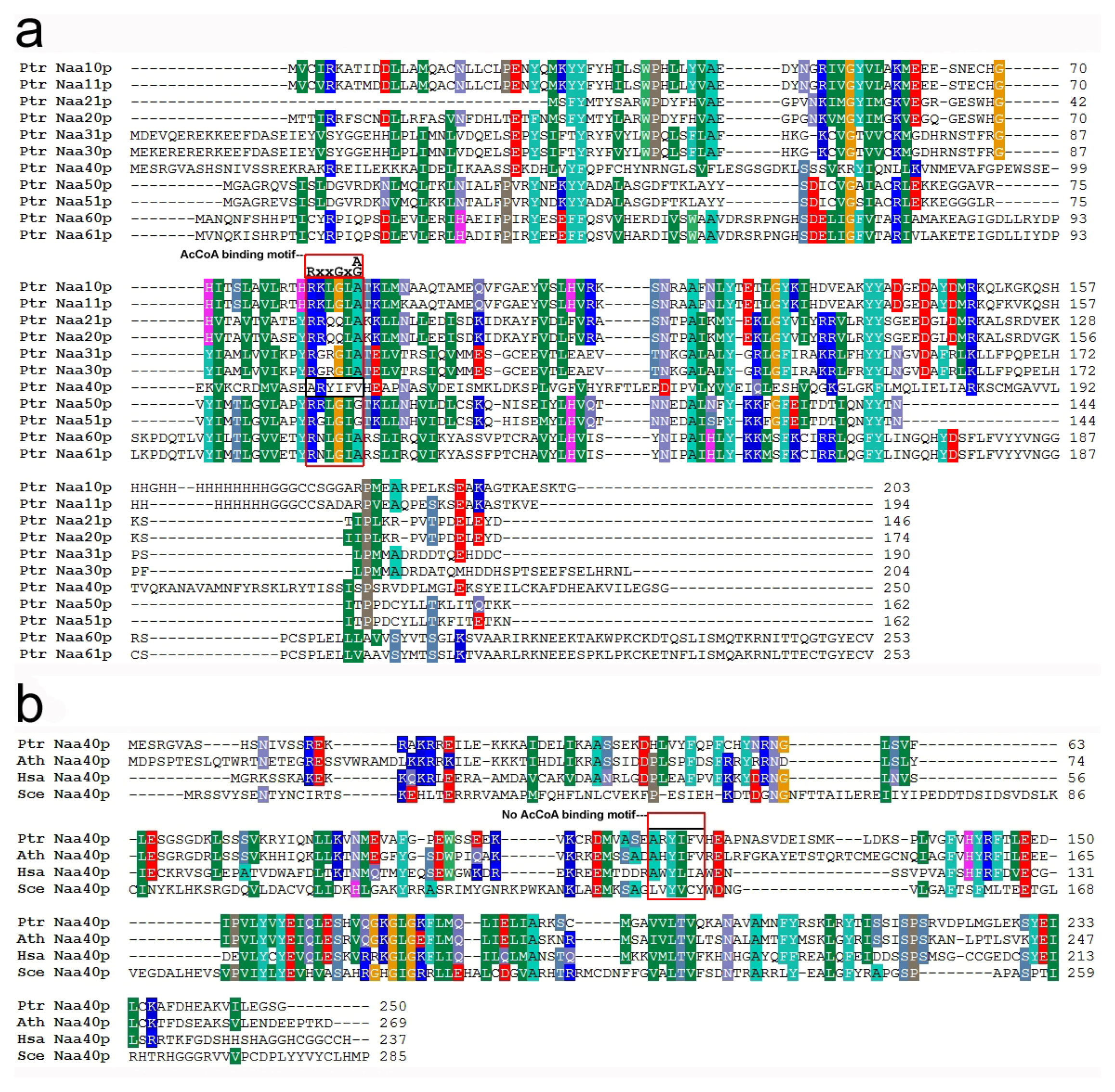
2.1. Identification and Characterization of Genes Encoding Nat Subunits in Populus
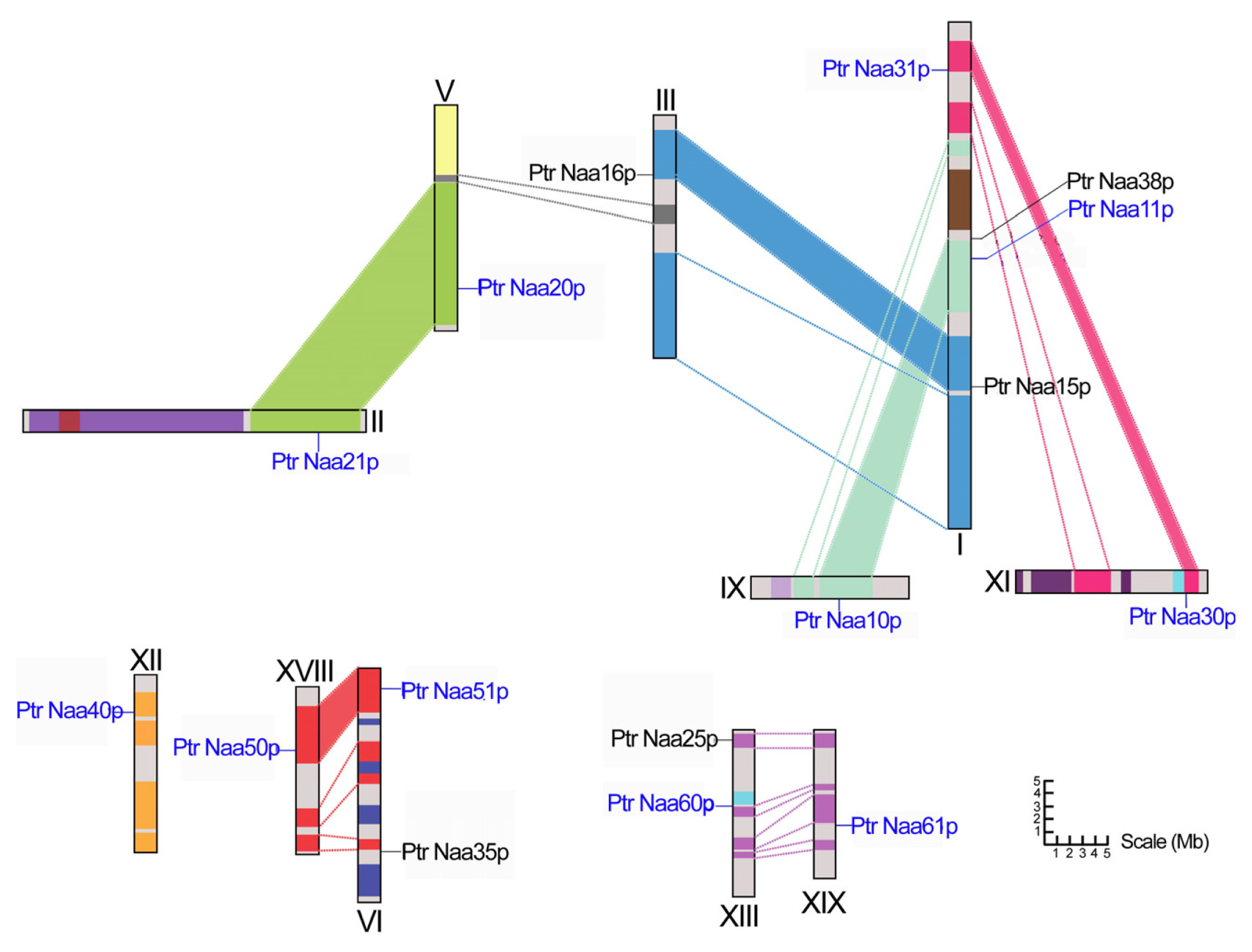
2.2. Chromosomal Location and Duplication of Nat CS Gene in Populus
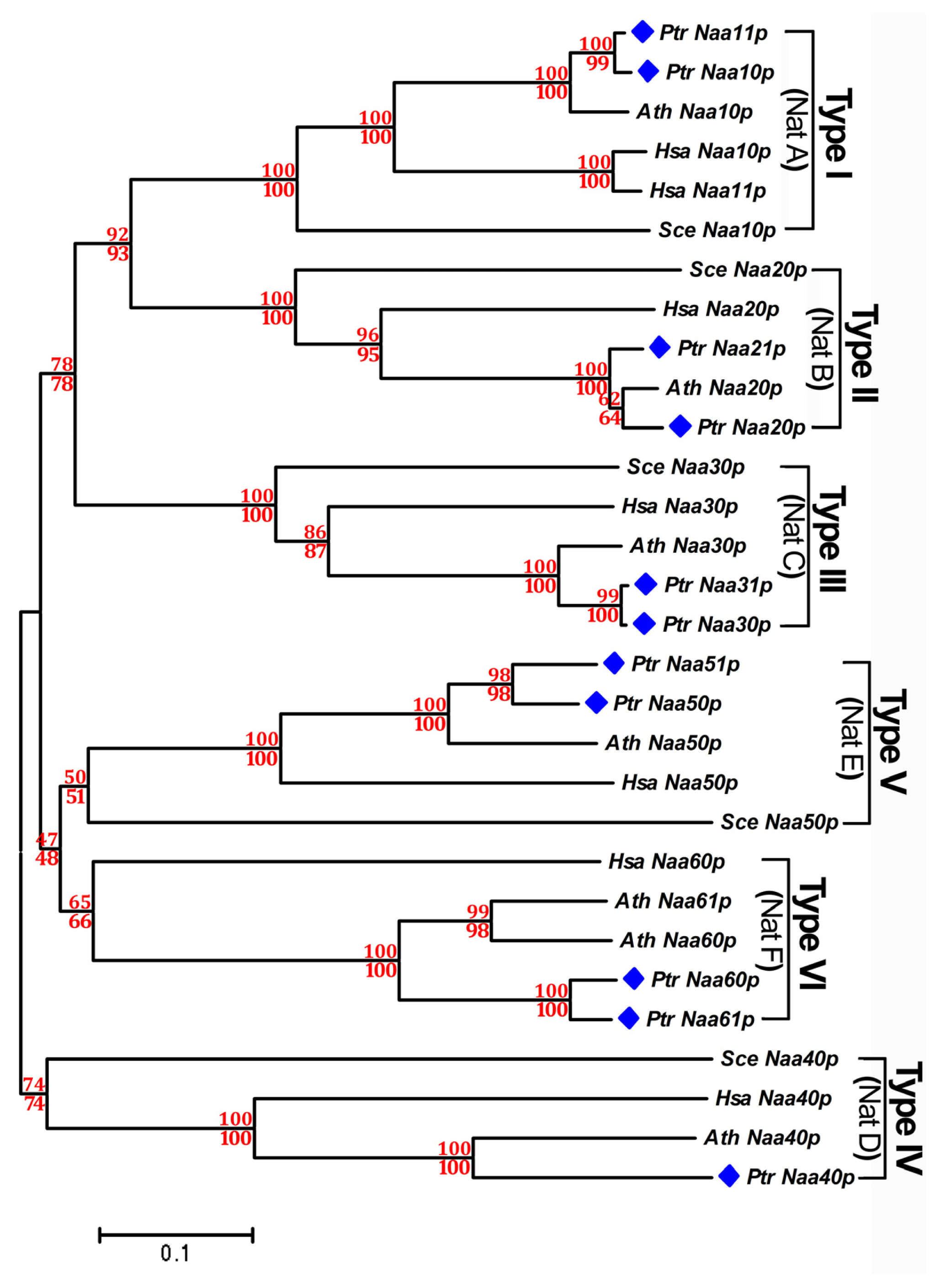
2.3. Phylogenetic Analysis of Nat CS
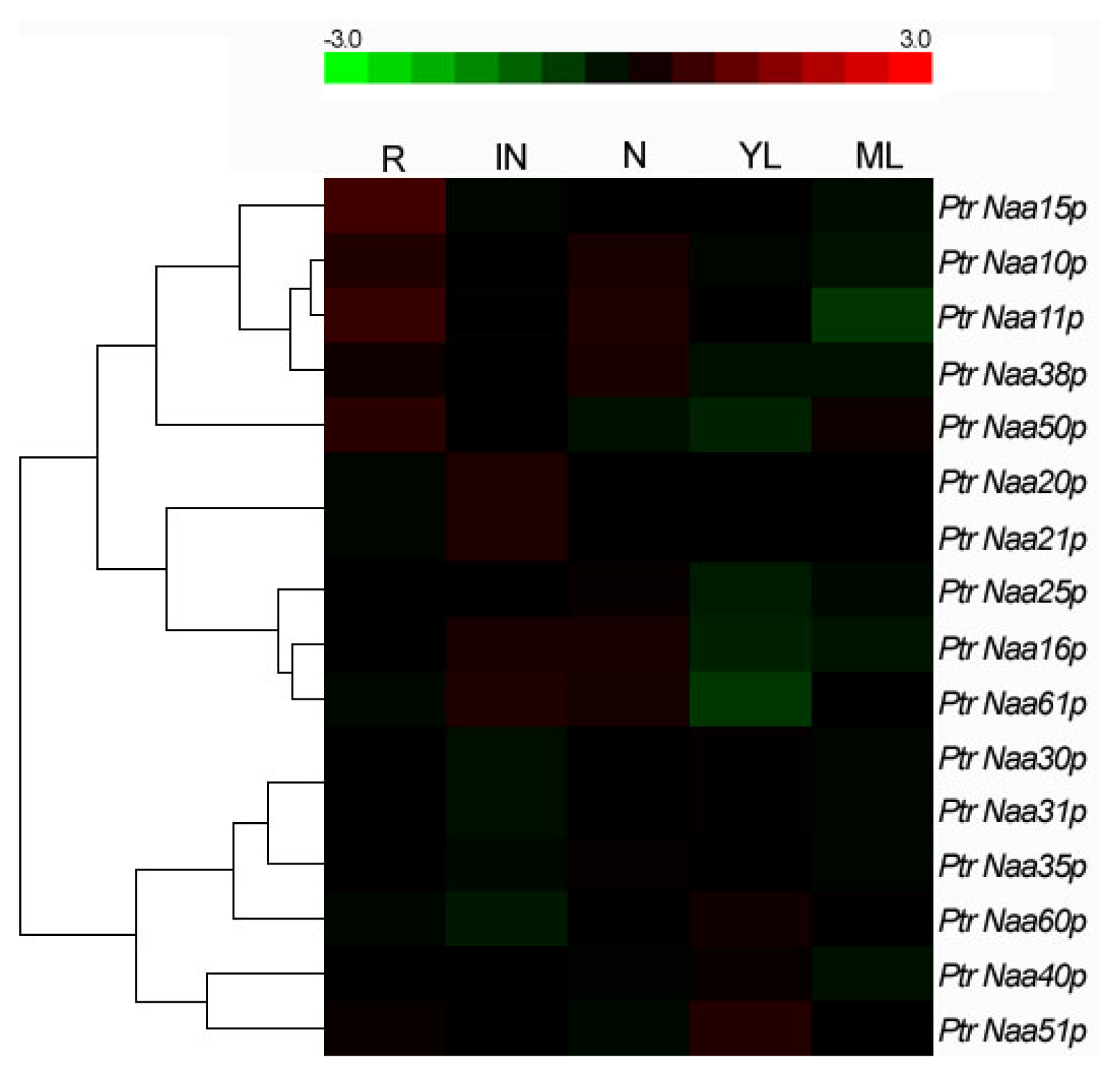
2.4. Tissue Location of Nat CS and AS Gene Expression in Populus
3. Experimental Section
3.1. Acquisition or Establishment of Hidden Markov Model (HMM) Profile Files
3.2. Domain Profile Search
3.3. Chromosomal Location and Phylogenetic Analysis
3.4. In silico Microarray Analysis
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Giglione, C.; Boularot, A.; Meinnel, T. Protein N-terminal methionine excision. Cell Mol. Life Sci 2004, 61, 1455–1474. [Google Scholar]
- Polevoda, B.; Sherman, F. N-terminal acetyltransferases and sequence requirements for N-terminal acetylation of eukaryotic proteins. J. Mol. Biol 2003, 325, 595–622. [Google Scholar]
- Martinez, A.; Traverso, J.A.; Valot, B.; Ferro, M.; Espagne, C.; Ephritikhine, G.; Zivy, M.; Giglione, C.; Meinnel, T. Extent of N-terminal modifications in cytosolic proteins from eukaryotes. Proteomics 2008, 8, 2809–2831. [Google Scholar]
- Ross, S.; Giglione, C.; Pierre, M.; Espagne, C.; Meinnel, T. Functional and developmental impact of cytosolic protein N-terminal methionine excision in Arabidopsis. Plant Physiol 2005, 137, 623–637. [Google Scholar]
- Hwang, C.S.; Shemorry, A.; Varshavsky, A. N-Terminal acetylation of cellular proteins creates specific degradation signals. Science 2010, 327, 973–977. [Google Scholar]
- Shemorry, A.; Hwang, C.-S.; Varshavsky, A. Control of protein quality and stoichiometries by N-terminal acetylation and the N-end rule pathway. Mol. Cell 2013, 50, 540–551. [Google Scholar]
- Geissenhöner, A.; Weise, C.; Ehrenhofer-Murray, A.E. Dependence of ORC silencing function on NatA-mediated Nα acetylation in Saccharomyces cerevisiae. Mol. Cell. Biol 2004, 24, 10300–10312. [Google Scholar]
- Starheim, K.K.; Gevaert, K.; Arnesen, T. Protein N-terminal acetyltransferases: When the start matters. Trends Biochem. Sci 2012, 37, 152–161. [Google Scholar]
- Van Damme, P.; Hole, K.; Pimenta-Marques, A.; Helsens, K.; Vandekerckhove, J.; Martinho, R.G.; Gevaert, K.; Arnesen, T. NatF contributes to an evolutionary shift in protein N-terminal acetylation and is important for normal chromosome segregation. PLoS Genet 2011, 7, e1002169. [Google Scholar]
- Polevoda, B.; Arnesen, T.; Sherman, F. A synopsis of eukaryotic Nα-terminal acetyltransferases: nomenclature, subunits and substrates. BMC Proc 2009, 3, S2. [Google Scholar]
- Eddy, S.R. A new generation of homology search tools based on probabilistic inference. Genome Inform 2009, 23, 205–211. [Google Scholar]
- Tuskan, G.; Difazio, S.; Jansson, S.; Bohlmann, J.; Grigoriev, I.; Hellsten, U.; Putnam, N.; Ralph, S.; Rombauts, S.; Salamov, A. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 2006, 313, 1596–1604. [Google Scholar]
- Polevoda, B.; Sherman, F. Nα-terminal acetylation of eukaryotic proteins. J. Biol. Chem 2000, 275, 36479–36482. [Google Scholar]
- Liu, C.-C.; Zhu, H.-Y.; Dong, X.-M.; Ning, D.-L.; Wang, H.-X.; Li, W.-H.; Yang, C.-P.; Wang, B.-C. Identification and analysis of the acetylated status of poplar proteins reveals analogous N-terminal protein processing mechanisms with other eukaryotes. PLoS One 2013, 8, e58681. [Google Scholar]
- Hollebeke, J.; van Damme, P.; Gevaert, K. N-terminal acetylation and other functions of N-α-acetyltransferases. Biol. Chem 2012, 393, 291–298. [Google Scholar]
- Abdel-Haleem, H. The origins of genome architecture. J. Hered 2007, 98, 633–634. [Google Scholar]
- Hu, R.; Qi, G.; Kong, Y.; Kong, D.; Gao, Q.; Zhou, G. Comprehensive analysis of NAC domain transcription factor gene family in Populus trichocarpa. BMC Plant Biol 2010, 10, 145. [Google Scholar]
- Liu, C.C.; Li, C.M.; Liu, B.G.; Ge, S.J.; Dong, X.M.; Li, W.; Zhu, H.Y.; Wang, B.C.; Yang, C.P. Genome-wide identification and characterization of a dehydrin gene family in poplar (Populus trichocarpa). Plant Mol. Biol. Rep 2011, 30, 848–859. [Google Scholar]
- Barakat, A.; Bagniewska-Zadworna, A.; Choi, A.; Plakkat, U.; DiLoreto, D.S.; Yellanki, P.; Carlson, J.E. The cinnamyl alcohol dehydrogenase gene family in Populus: Phylogeny, organization, and expression. BMC Plant Biol 2009, 9, 26. [Google Scholar]
- Kalluri, U.C.; DiFazio, S.P.; Brunner, A.M.; Tuskan, G.A. Genome-wide analysis of Aux/IAA and ARF gene families in Populus trichocarpa. BMC Plant Biol 2007, 7, 59. [Google Scholar]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol 2011, 28, 2731–2739. [Google Scholar]
- Saitou, N.; Nei, M. The Neighbor-Joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol 1987, 4, 406–425. [Google Scholar]
- Zybailov, B.; Rutschow, H.; Friso, G.; Rudella, A.; Emanuelsson, O.; Sun, Q.; van Wijk, K.J. Sorting signals, N-terminal modifications and abundance of the chloroplast proteome. PLoS One 2008, 3, 1–19. [Google Scholar]
- Polevoda, B.; Sherman, F. Composition and function of the eukaryotic N-terminal acetyltransferase subunits. Biochem. Biophys. Res. Commun 2003, 308, 1–11. [Google Scholar]
- Ohlrogge, J.; Benning, C. Unraveling plant metabolism by EST analysis. Curr. Opin. Plant Biol 2000, 3, 224–228. [Google Scholar]
- Wilkins, O.; Nahal, H.; Foong, J.; Provart, N.J.; Campbell, M.M. Expansion and diversification of the Populus R2R3-MYB family of transcription factors. Plant Physiol 2009, 149, 981–993. [Google Scholar]
- Sjödin, A.; Street, N.R.; Sandberg, G.; Gustafsson, P.; Jansson, S. The Populus Genome Integrative Explorer (PopGenIE): A new resource for exploring the Populus genome. N. Phytol 2009, 182, 1013–1025. [Google Scholar]
- Starheim, K.K.; Gromyko, D.; Velde, R.; Varhaug, J.E.; Arnesen, T. Composition and biological significance of the human Nα-terminal acetyltransferases. BMC Proc 2009, 3, S3. [Google Scholar]
- Larkin, M.; Blackshields, G.; Brown, N.; Chenna, R.; McGettigan, P.; McWilliam, H.; Valentin, F.; Wallace, I.; Wilm, A.; Lopez, R. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar]
- Liu, C.C.; Liu, B.G.; Yang, Z.W.; Li, C.M.; Wang, B.C.; Yang, C.P. Genome-wide identification and in silico analysis of poplar peptide deformylases. Int. J. Mol. Sci 2012, 13, 5112–5124. [Google Scholar]
- Finn, R.; Mistry, J.; Schuster-Bockler, B.; Griffiths-Jones, S.; Hollich, V.; Lassmann, T.; Moxon, S.; Marshall, M.; Khanna, A.; Durbin, R. Pfam: clans, web tools and services. Nucleic Acids Res 2006, 34, D247–D251. [Google Scholar]
- Liu, C.C.; Li, W.; Yang, Z.W.; Liu, B.G.; Ge, S.J.; Zhu, H.Y.; Yang, C.P.; Wei, Z.G. The systematic characterization of poplar CK2α and its theoretical studies on phosphorylation of P-protein C-terminal domain. Afr. J. Microbiol. Res 2011, 5, 4850–4858. [Google Scholar]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser 1999, 41, 95–98. [Google Scholar]
- Liu, C.C.; Lu, T.C.; Li, H.H.; Wang, H.X.; Liu, G.F.; Ma, L.; Yang, C.P.; Wang, B.C. Phosphoproteomic identification and phylogenetic analysis of ribosomal P-proteins in Populus dormant terminal buds. Planta 2010, 231, 571–581. [Google Scholar]
- De Hoon, M.; Imoto, S.; Nolan, J.; Miyano, S. Open source clustering software. Bioinformatics 2004, 20, 1453–1454. [Google Scholar]
- Saldanha, A.J. Java Treeview-extensible visualization of microarray data. Bioinformatics 2004, 20, 3246–3248. [Google Scholar]
| Type | JGI gene and protein ID | Transcript ID | Chromosome location | Protein products | ||
|---|---|---|---|---|---|---|
| NCBI REFseq | Populus genome V2.2 | Protein ID | Novel simplified nomenclature | |||
| NatA CS a | 650021 | XM_002314022.1 | POPTR_0009s06150 | LG_IX:6944007–6945077(−) | XP_002314058.1 | Ptr Naa10p |
| NatA CS | 641307 | XM_002298379.1 | POPTR_0001s26920 | LG_I:18982354–18983685 (+) | XP_002298415.1 | Ptr Naa11p |
| NatA AS b | 548659 | XM_002299594.1 | POPTR_0001s17830 | LG_I:9952442–9966294 (−) | XP_002299630.1 | Ptr Naa15p |
| NatA AS | 553694 | XM_002304144.1 | POPTR_0003s05540 | LG_III:4692360–4705382 (−) | XP_002304180.1 | Ptr Naa16p |
| NatB CS | 818659 | XM_002307550.1 | POPTR_0005s23200 | LG_V:14531524–14534737 (−) | XP_002307586.1 | Ptr Naa20p |
| NatB CS | 643297 | XM_002300805.1 | POPTR_0002s05290 | LG_II:3418242–3421271 (+) | XP_002300841.1 | Ptr Naa21p |
| NatB AS | 571859 | XM_002319920.1 | POPTR_0013s14900 | LG_XIII:12260671–12271953 (−) | XP_002319956.1 | Ptr Naa25p |
| NatC CS | 727122 | XM_002316966.1 | POPTR_0011s14270 | LG_XI:13438711–13441426 (+) | XP_002317002.1 | Ptr Naa30p |
| NatC CS | 642436 | XM_002298895.1 | POPTR_0049s00200 | LG_I:32126776–32129356 (+) | XP_002298931.1 | Ptr Naa31p |
| NatC AS I | 560565 | XM_002308020.1 | POPTR_0006s06370 | LG_VI:3978171–3986294 (+) | XP_002308056.1 | Ptr Naa35p |
| NatC AS II | 641478 | XM_002299954.1 | POPTR_0001s28460 | LG_I:20275848–20278373 (−) | XP_002299990.1 | Ptr Naa38p |
| NatD CS | 729076 | XM_002318277.1 | POPTR_0012s03830 | LG_XII:300904–304114 (−) | XP_002318313.1 | Ptr Naa40p |
| NatE CS | 737117 | XM_002324238.1 | POPTR_0018s01280 | LG_XVIII:5217292–5220254 (+) | XP_002324274.1 | Ptr Naa50p |
| NatE CS | 654093 | XM_002308604.1 | POPTR_0006s26500 | LG_VI:16869902–16872551 (+) | XP_002308640.1 | Ptr Naa51p |
| NatF CS | 834607 | XM_002319219.1 | POPTR_0013s07770 | LG_XIII:7060330–7064827 (+) | XP_002319255.1 | Ptr Naa60p |
| NatF CS | 665408 | XM_002325352.1 | POPTR_0019s07740 | LG_XIX:4276553–4280210 (+) | XP_002325388.1 | Ptr Naa61p |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhu, H.-Y.; Li, C.-M.; Wang, L.-F.; Bai, H.; Li, Y.-P.; Yu, W.-X.; Xia, D.-A.; Liu, C.-C. In Silico Identification and Characterization of N-Terminal Acetyltransferase Genes of Poplar (Populus trichocarpa). Int. J. Mol. Sci. 2014, 15, 1852-1864. https://doi.org/10.3390/ijms15021852
Zhu H-Y, Li C-M, Wang L-F, Bai H, Li Y-P, Yu W-X, Xia D-A, Liu C-C. In Silico Identification and Characterization of N-Terminal Acetyltransferase Genes of Poplar (Populus trichocarpa). International Journal of Molecular Sciences. 2014; 15(2):1852-1864. https://doi.org/10.3390/ijms15021852
Chicago/Turabian StyleZhu, Hang-Yong, Chun-Ming Li, Li-Feng Wang, Hui Bai, Yan-Ping Li, Wen-Xi Yu, De-An Xia, and Chang-Cai Liu. 2014. "In Silico Identification and Characterization of N-Terminal Acetyltransferase Genes of Poplar (Populus trichocarpa)" International Journal of Molecular Sciences 15, no. 2: 1852-1864. https://doi.org/10.3390/ijms15021852




