The eTOX Data-Sharing Project to Advance in Silico Drug-Induced Toxicity Prediction
Abstract
:1. Introduction
2. Results and Discussion
2.1. Legacy Data Gathering
| Category | Access | Sharing | Usage |
|---|---|---|---|
| Public | Public upon request | Structure and all available data | Read-across analysis, Models building and validation |
| Non-Confidential | eTOX consortium | Structure and toxicological data | Read-across analysis, Models building and validation |
| Confidential | Honest broker and data owner | Toxicological data | Read-across analysis (without structure query), Models building and validation (without structure query) |
| Private | Data owner | None | Models validation |
- -
- Substance ID: The database substance identifier, created by the EFPIA partner;
- -
- Report ID: The internal report name/identifier;
- -
- Quality assessment: The result/status of the internal quality check;
- -
- Clearance date: The timepoint when the report entered the eTOX extraction process;
- -
- Status: Confidential/non-confidential;
- -
- Progress: Sent to CRO date/Sent to Lhasa Limited date;
- -
- Available at Vitic Nexus eTOX database: The date of the database release that contains the report;
- -
- Progress comments: Comments when applicable;
2.2. Data Integration, Harmonization and Curation
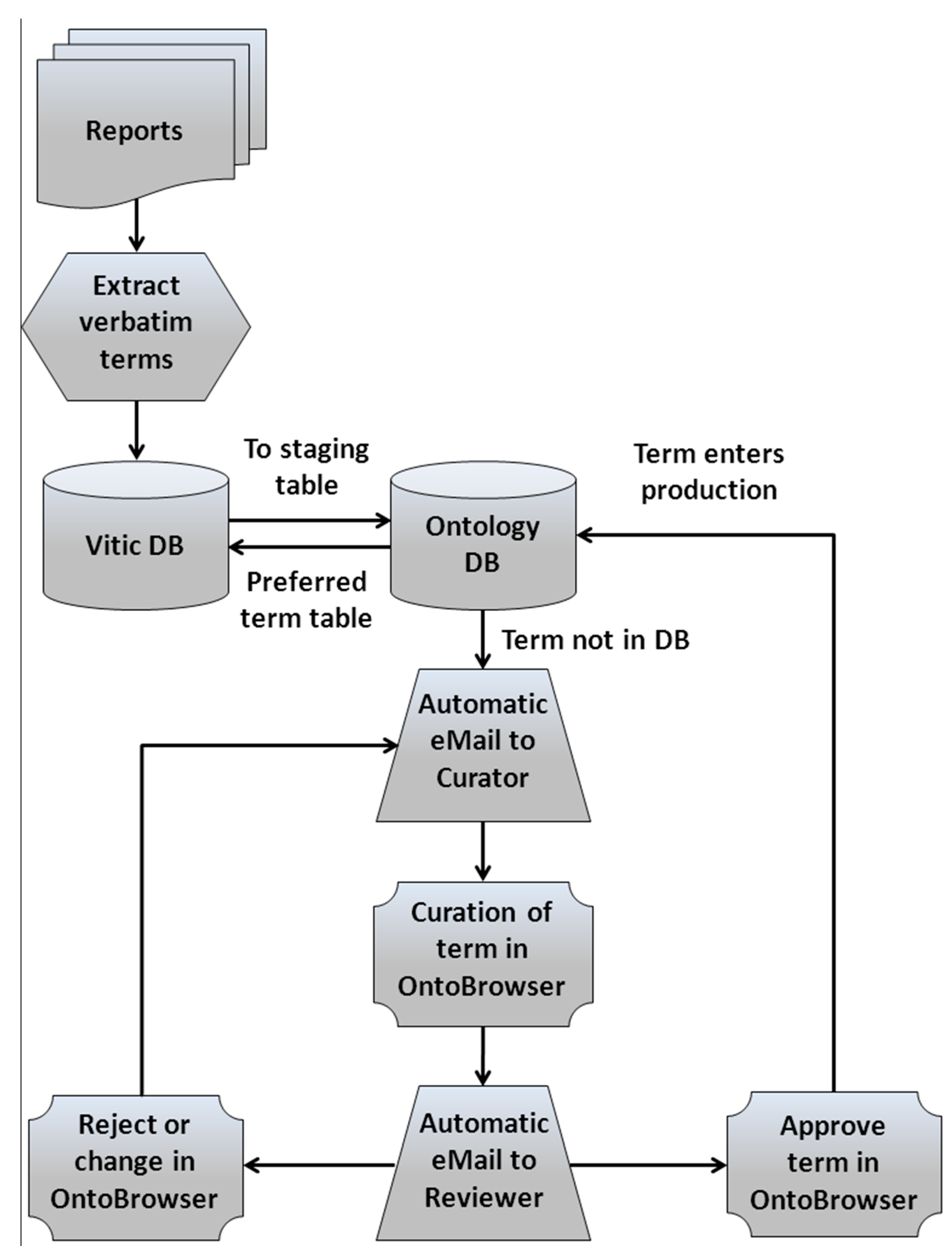
2.3. Data Analysis for Modeling Purposes
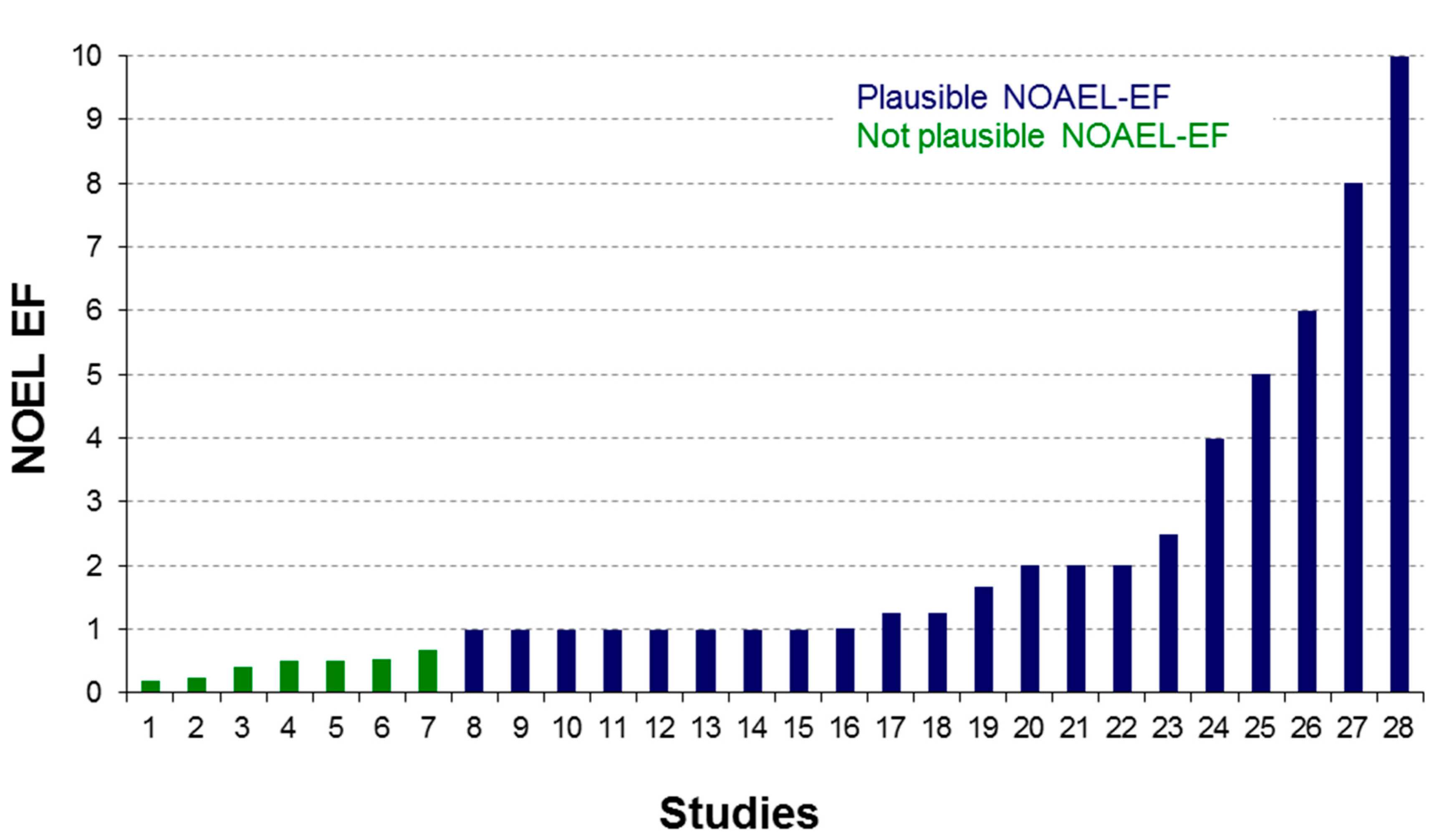
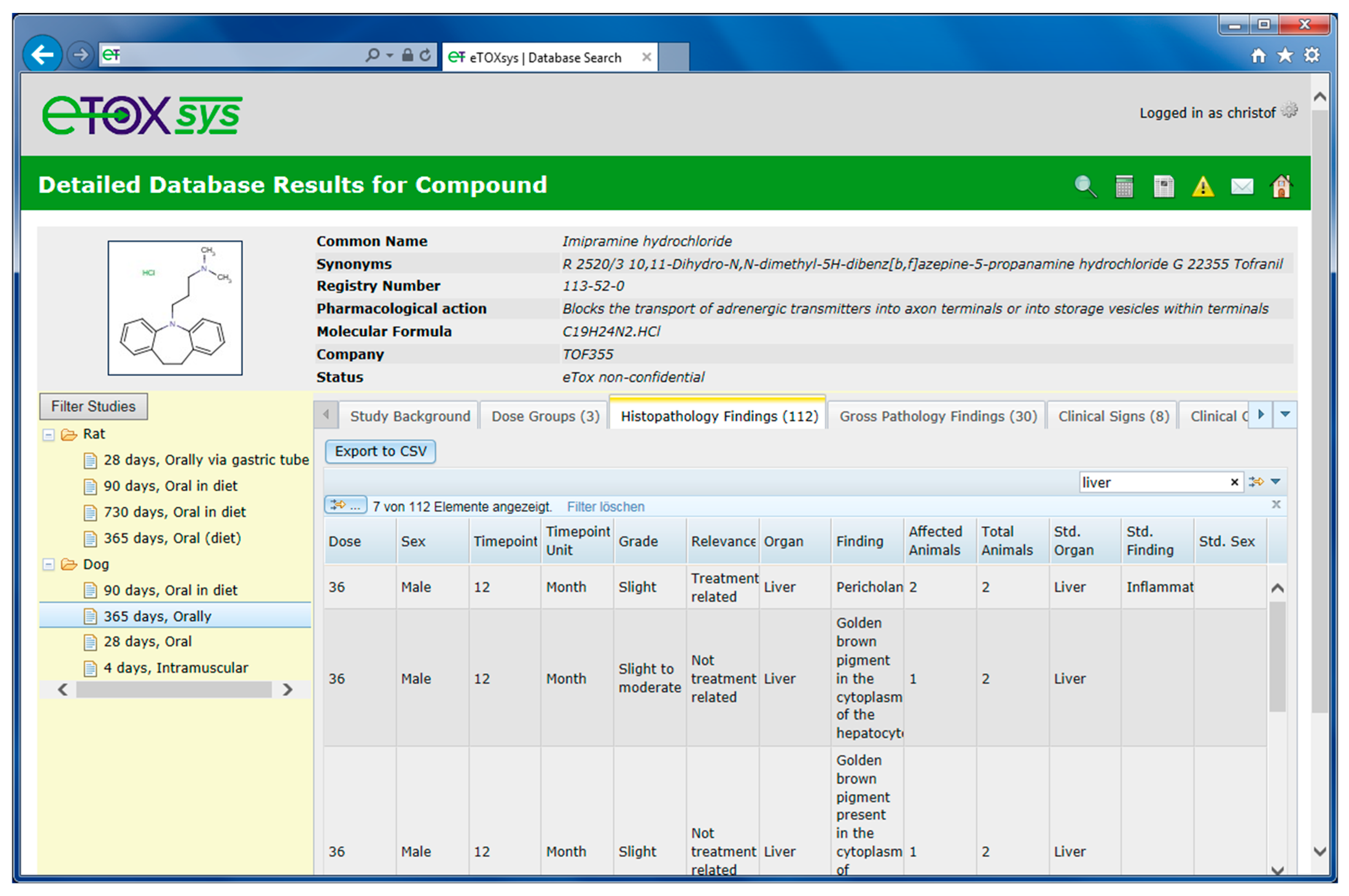
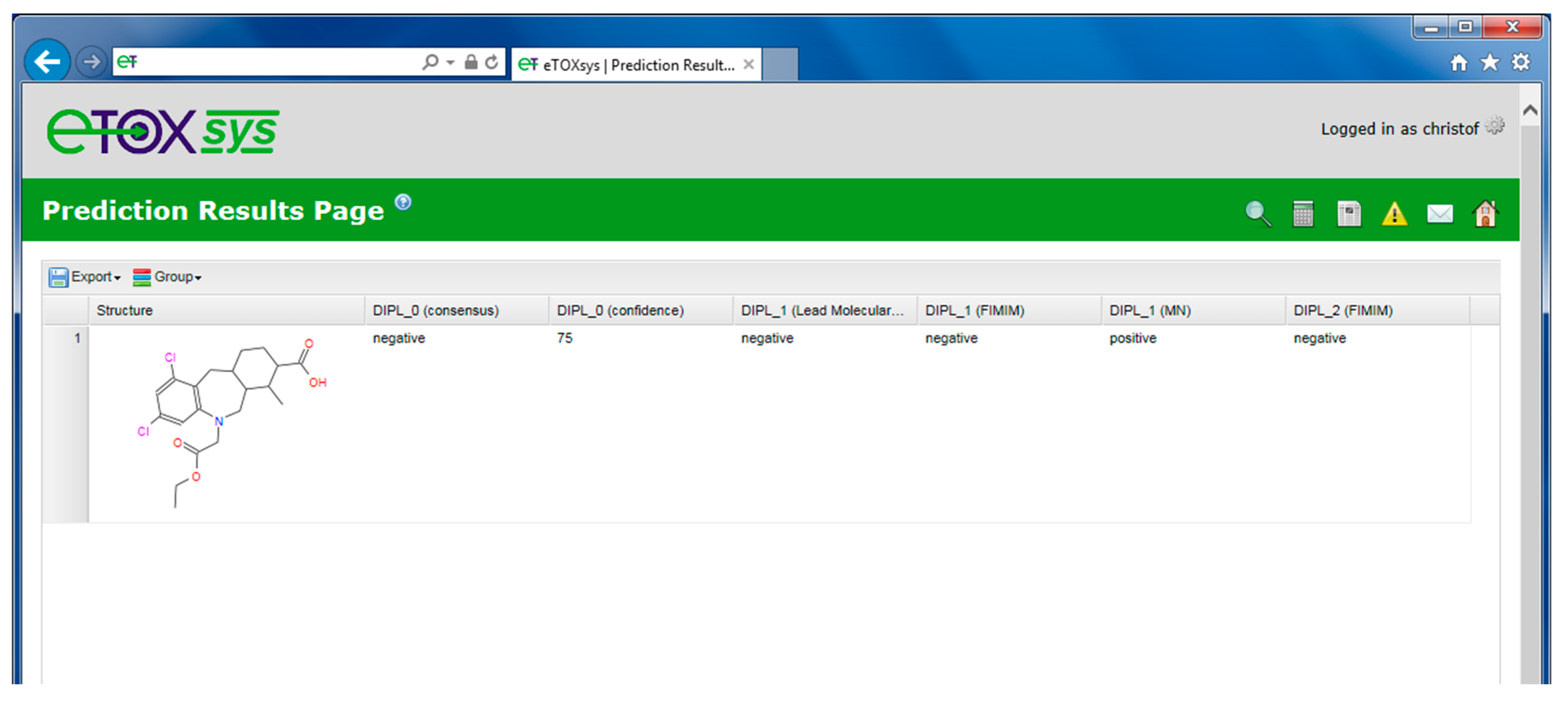
2.4. eTOXsys: The Data Browser and the Predictive System

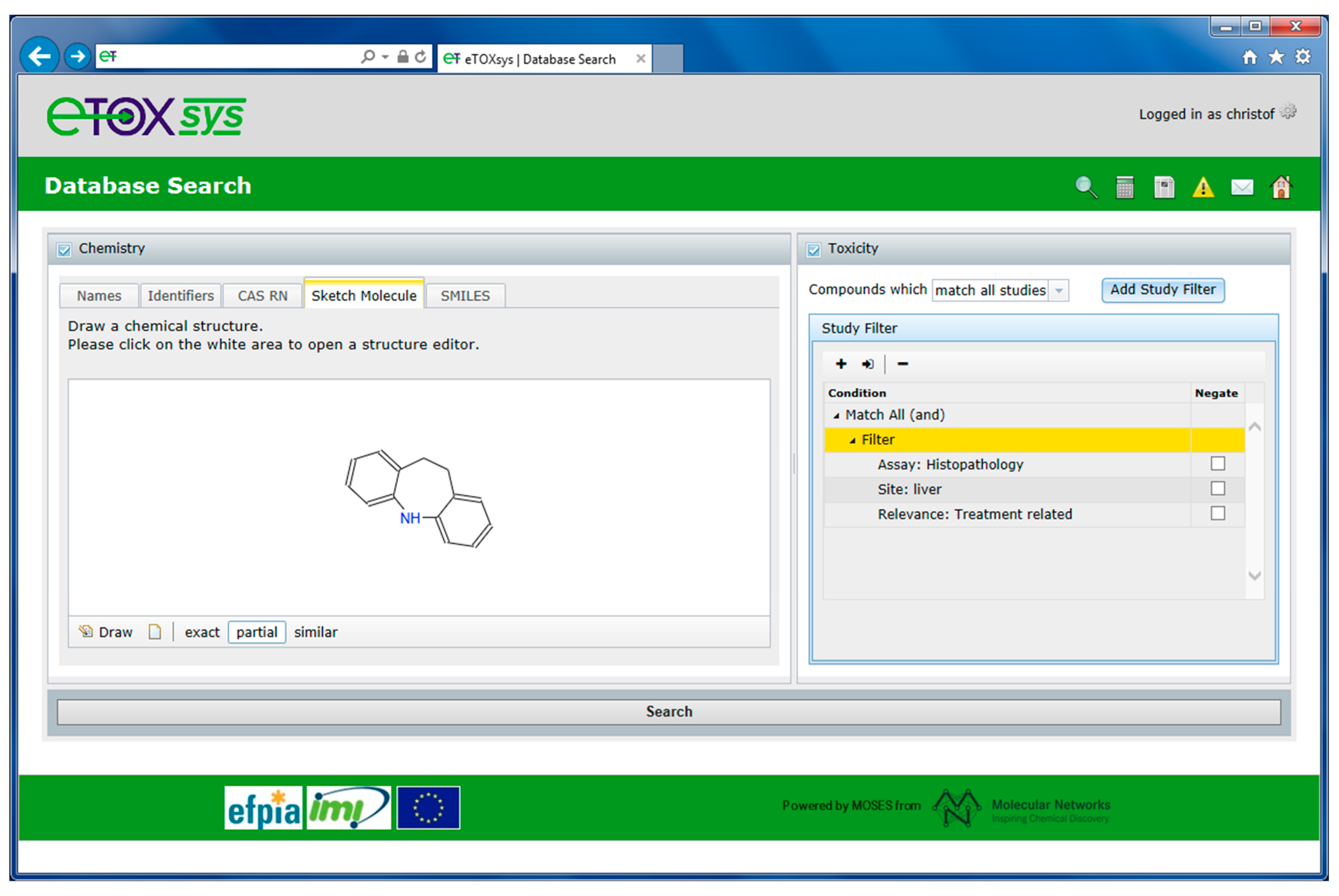
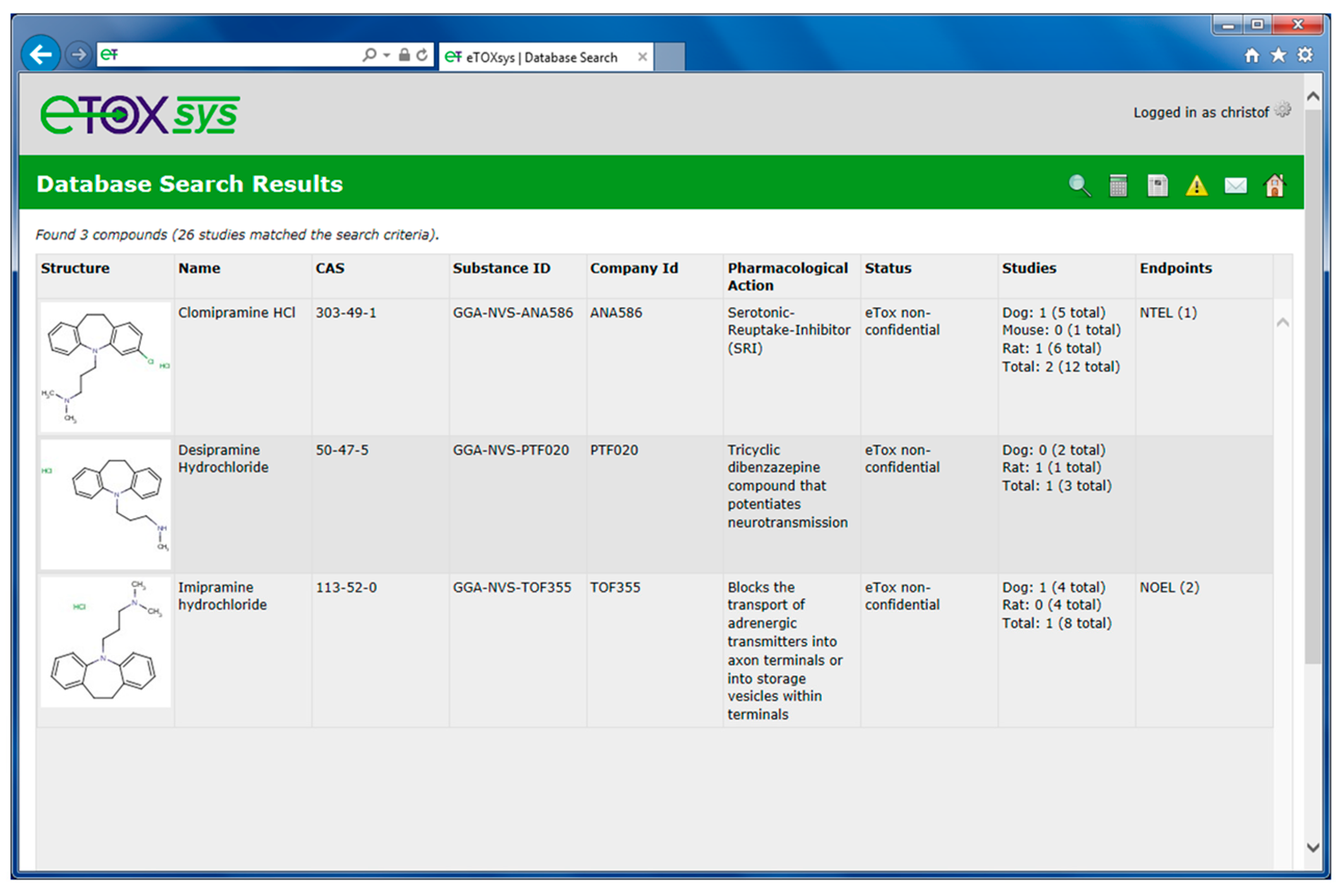


2.5. Ongoing and Future Actions
3. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Sacks, L.V.; Shamsuddin, H.H.; Yasinskaya, Y.I.; Bouri, K.; Lanthier, M.L.; Sherman, R.E. Scientific and regulatory reasons for delay and denial of FDA approval of initial applications for new drugs, 2000–2012. JAMA 2014, 311, 378–384. [Google Scholar] [CrossRef]
- Kramer, J.A.; Sagartz, J.E.; Morris, D.L. The application of discovery toxicology and pathology towards the design of safer pharmaceutical lead candidates. Nat. Rev. Drug Discov. 2007, 6, 636–649. [Google Scholar] [CrossRef] [PubMed]
- FitzGerald, G.A. Testing cardiovascular drug safety and efficacy in randomized trials. Circ. Res. 2014, 114, 1156–1161. [Google Scholar] [CrossRef] [PubMed]
- ToxML. Available online: http://toxml.org (accessed on 31 August 2014).
- DSSTox. Available online: http://www.epa.gov/ncct/dsstox/ (accessed on 31 August 2014).
- ACToR. Available online: http://actor.epa.gov/actor/faces/ACToRHome.jsp (accessed on 31 August 2014).
- Kopp-Schneider, A.; Prieto, P.; Kinsner-Ovaskainen, A.; Stanzel, S. Design of a testing strategy using non-animal based test methods: Lessons learnt from the ACuteTox project. Toxicol. In Vitro 2013, 27, 1395–1401. [Google Scholar] [CrossRef] [PubMed]
- Bushnell, P.J.; Kavlock, R.J.; Crofton, K.M.; Weiss, B.; Rice, D.C. Behavioral toxicology in the 21st century: Challenges and opportunities for 2 behavioral scientists. Neurotoxicol. Teratol. 2010, 32, 313–328. [Google Scholar] [CrossRef] [PubMed]
- Knudsen, T.B.; Houck, K.A.; Sipes, N.S.; Singh, A.V.; Judson, R.S.; Martin, M.T.; Weissman, A.; Kleinstreuer, N.C.; Mortensen, H.M.; Reif, D.M.; et al. Activity profiles of 309 ToxCast™ chemicals evaluated across 292 biochemical targets. Toxicology 2011, 282, 1–15. [Google Scholar]
- Kavlock, R.; Chandler, K.; Houck, K.; Hunter, S.; Judson, R.; Kleinstreuer, N.; Knudsen, T.; Martin, M.; Padilla, S.; Reif, D.; et al. Update on EPA’s ToxCast program: Providing high throughput decision support tools for chemical risk management. Chem. Res. Toxicol. 2012, 25, 1287–1302. [Google Scholar]
- Shah, F.; Greene, N. Analysis of Pfizer compounds in EPA’s ToxCast chemicals-assay space. Chem. Res. Toxicol. 2014, 27, 86–98. [Google Scholar] [CrossRef] [PubMed]
- Attene-Ramos, M.S.; Miller, N.; Huang, R.; Michael, S.; Itkin, M.; Kavlock, R.J.; Austin, C.P.; Shinn, P.; Simeonov, A.; Tice, R.R.; et al. The Tox21 robotic platform for the assessment of environmental chemicals—From vision to reality. Drug Discov. Today 2013, 18, 716–723. [Google Scholar]
- Hardy, B.; Douglas, N.; Helma, C.; Rautenberg, M.; Jeliazkova, N.; Jeliazkov, V.; Nikolova, I.; Benigni, R.; Tcheremenskaia, O.; Kramer, S.; et al. Collaborative development of predictive toxicology applications. J. Cheminform. 2010, 2, 7. [Google Scholar]
- Kohonen, P.; Benfenati, E.; Bower, D.; Ceder, R.; Crump, M.; Cross, K.; Grafström, R.C.; Healy, L.; Helma, C.; Jeliazkova, N.; et al. The ToxBank data warehouse: Supporting the replacement of in vivo repeated dose systemic toxicity testing. Mol. Inf. 2013, 32, 47–63. [Google Scholar]
- Benigni, R.; Battistelli, C.L.; Bossa, C.; Tcheremenskaia, O.; Crettaz, P. New perspectives in toxicological information management, and the role of ISSTOX databases in assessing chemical mutagenicity and carcinogenicity. Mutagenesis 2013, 28, 401–409. [Google Scholar] [CrossRef] [PubMed]
- Tluczkiewicz, I.; Batke, M.; Kroese, D.; Buist, H.; Aldenberg, T.; Pauné, E.; Grimm, H.; Kühne, R.; Schüürmann, G.; Mangelsdorf, I.; et al. The OSIRIS Weight of Evidence approach: ITS for the endpoints repeated-dose toxicity (RepDose ITS). Regul. Toxicol. Pharmacol. 2013, 67, 157–169. [Google Scholar] [PubMed]
- Williams, A.J.; Harland, L.; Groth, P.; Pettifer, S.; Chichester, C.; Willighagen, E.L.; Evelo, C.T.; Blomberg, N.; Ecker, G.; Goble, C.; et al. Open PHACTS: Semantic interoperability for drug discovery. Drug Discov. Today 2012, 17, 1188–1198. [Google Scholar]
- Safe-T. Available online: http://www.imi-safe-t.eu/ (accessed on 31 August 2014).
- COSMOS. Available online: http://www.cosmostox.eu/ (accessed on 31 August 2014).
- Merlot, C. Computational toxicology―A tool for early safety evaluation. Drug Discov. Today 2010, 15, 16–22. [Google Scholar] [CrossRef] [PubMed]
- Richard, A.M.; Yang, C.; Judson, R.S. Toxicity data informatics: Supporting a new paradigm for toxicity prediction. Toxicol. Mech. Methods. 2008, 18, 103–118. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Valerio, L.G., Jr.; Arvidson, K.B. Computational toxicology approaches at the US Food and Drug Administration. Altern. Lab. Anim. 2009, 37, 523–531. [Google Scholar] [PubMed]
- IMI. Available online: http://imi.europa.eu/index_en.html (accessed on 31 August 2014).
- Bril, A.; Canet, E. The innovative medicine initiative (IMI). Med. Sci. 2008, 24, 885–890. [Google Scholar]
- eTOX official website. Available online: http://www.e-TOX.net (accessed on 31 August 2014).
- Briggs, K.; Cases, M.; Heard, D.J.; Pastor, M.; Pognan, F.; Sanz, F.; Schwab, C.H.; Steger-Hartmann, T.; Sutter, A.; Watson, D.K.; et al. Inroads to predict in vivo toxicology—An introduction to the eTOX project. Int. J. Mol. Sci. 2012, 13, 3820–3846. [Google Scholar]
- InnoMed PredTox. Available online: https://www.genedata.com/lp/innomed-predtox.html (accessed on 31 August 2014).
- TG-GATEs. Available online: http://toxico.nibio.go.jp/ (accessed on 31 August 2014).
- DrugMatrix. Available online: https://ntp.niehs.nih.gov/drugmatrix/index.html (accessed on 31 August 2014).
- Li, Q.; Cheng, T.; Wang, Y.; Bryant, S.H. PubChem as a public resource for drug discovery. Drug Discov. Today 2010, 15, 1052–1057. [Google Scholar] [CrossRef] [PubMed]
- Bento, A.P.; Gaulton, A.; Hersey, A.; Bellis, L.J.; Chambers, J.; Davies, M.; Krüger, F.A.; Light, Y.; Mak, L.; McGlinchey, S.; et al. The ChEMBL bioactivity database: An update. Nucleic Acids Res. 2014, 42, D1083–D1090. [Google Scholar] [CrossRef]
- ChEMBL. Available online: https://www.ebi.ac.uk/chembl/ (accessed on 31 August 2014).
- EPARs. Available online: http://www.ema.europa.eu/ema/index.jsp?curl=pages/medicines/landing/epar_search.jsp&mid=WC0b01ac058001d125 (accessed on 31 August 2014).
- DailyMed. Available online: http://dailymed.nlm.nih.gov/ (accessed on 31 August 2014).
- Drugbank. Available online: http://www.drugbank.ca/ (accessed on 31 August 2014).
- Cases, M.; Pastor, M.; Sanz, F. The eTOX library of public resources for in silico toxicity prediction. Mol. Inform. 2013, 32, 24–35. [Google Scholar] [CrossRef]
- eTOXlibrary. Available online: http://cadd.imim.es/etox-library/ (accessed on 31 August 2014).
- RepDose. Available online: http://fraunhofer-repdose.de/ (accessed on 31 August 2014).
- Marchant, C.; Briggs, K.; Long, A. In silico tools for sharing data and knowledge on toxicity and metabolism: Derek for windows, meteor, and vitic. Toxicol. Mech. Methods 2008, 18, 177–187. [Google Scholar] [CrossRef] [PubMed]
- InChi-IUPAC. Available online: http://www.iupac.org/home/publications/e-resources/inchi.html (accessed on 31 August 2014).
- Harland, L.; Larminie, C.; Sansone, S.A.; Popa, S.; Marshall, M.S.; Braxenthaler, M.; Cantor, M.; Filsell, W.; Forster, M.J.; Huang, E.; et al. Empowering industrial research with shared biomedical vocabularies. Drug Discov. Today 2011, 16, 940–947. [Google Scholar]
- SEND. Available online: http://www.cdisc.org/SEND (accessed on 31 August 2014).
- CDISC. Available online: http://www.cdisc.org/mission-and-principles (accessed on 31 August 2014).
- goRENI. Available online: http://www.goreni.org/ (accessed on 24 October 2014).
- Briggs, K.; Barber, C.; Cases, M.; Steger-Hartmann, T. Value of shared preclinical safety studies—The eTOX database. Toxicol. Rep. Submitted. 2014. [Google Scholar]
- Hewitt, M.; Enoch, S.J.; Madden, J.C.; Przybylak, K.R.; Cronin, M.T.D. Hepatotoxicity: A scheme for generating chemical categories for read-across, structural alerts and insights into mechanism(s) of action. Crit. Rev. Toxicol. 2013, 43, 537–558. [Google Scholar] [CrossRef] [PubMed]
- Obiol-Pardo, C.; Gomis-Tena, J.; Sanz, F.; Saiz, J.; Pastor, M. A multiscale simulation system for the prediction of drug-induced cardiotoxicity. J. Chem. Inform. Model. 2011, 51, 483–492. [Google Scholar] [CrossRef]
- Taboureau, O.; Jørgensen, FS. In silico predictions of hERG channel blockers in drug discovery: From ligand-based and target-based approaches to systems chemical biology. Comb. Chem. High Throughput Screen 2011, 14, 375–387. [Google Scholar] [CrossRef] [PubMed]
- Poongavanam, V.; Haider, N.; Ecker, G.F. Fingerprint-based in silico models for the prediction of P-glycoprotein substrates and inhibitors. Bioorg. Med. Chem. 2012, 20, 5388–5395. [Google Scholar] [CrossRef] [PubMed]
- Pinto, M.; Trauner, M.; Ecker, G.F. An in silico classification model for putative ABCC2 substrates. Mol. Inform. 2012, 31, 547–553. [Google Scholar] [CrossRef] [PubMed]
- Perić-Hassler, L.; Stjernschantz, E.; Oostenbrink, C.; Geerke, D.P. CYP 2D6 binding affinity predictions using multiple ligand and protein conformations. Int. J. Mol. Sci. 2013, 14, 24514–24530. [Google Scholar] [CrossRef] [PubMed]
- Vosmeer, C.R.; Pool, R.; van Stee, M.F.; Peric-Hassler, L.; Vermeulen, N.P.E.; Geerke, D.P. Towards automated binding affinity prediction using an iterative linear interaction energy approach. Int. J. Mol. Sci. 2014, 15, 798–816. [Google Scholar] [CrossRef] [PubMed]
- Klepsch, F.; Vasanthanathan, P.; Ecker, G.F. Ligand and structure-based classification models for prediction of P-glycoprotein inhibitors. J. Chem. Inform. Model. 2014, 54, 218–229. [Google Scholar] [CrossRef]
- Carrió, P.; Pinto, M.; Ecker, G.; Sanz, F.; Pastor, M. Applicability domain aNalysis (ADAN): A robust method for assessing the reliability of drug property predictions. J. Chem. Inform. Model. 2014, 54, 1500–1511. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cases, M.; Briggs, K.; Steger-Hartmann, T.; Pognan, F.; Marc, P.; Kleinöder, T.; Schwab, C.H.; Pastor, M.; Wichard, J.; Sanz, F. The eTOX Data-Sharing Project to Advance in Silico Drug-Induced Toxicity Prediction. Int. J. Mol. Sci. 2014, 15, 21136-21154. https://doi.org/10.3390/ijms151121136
Cases M, Briggs K, Steger-Hartmann T, Pognan F, Marc P, Kleinöder T, Schwab CH, Pastor M, Wichard J, Sanz F. The eTOX Data-Sharing Project to Advance in Silico Drug-Induced Toxicity Prediction. International Journal of Molecular Sciences. 2014; 15(11):21136-21154. https://doi.org/10.3390/ijms151121136
Chicago/Turabian StyleCases, Montserrat, Katharine Briggs, Thomas Steger-Hartmann, François Pognan, Philippe Marc, Thomas Kleinöder, Christof H. Schwab, Manuel Pastor, Jörg Wichard, and Ferran Sanz. 2014. "The eTOX Data-Sharing Project to Advance in Silico Drug-Induced Toxicity Prediction" International Journal of Molecular Sciences 15, no. 11: 21136-21154. https://doi.org/10.3390/ijms151121136





