Phenological, Nutritional and Molecular Diversity Assessment among 35 Introduced Lentil (Lens culinaris Medik.) Genotypes Grown in Saudi Arabia
Abstract
:1. Introduction
2. Results and Discussion
2.1. Phenological Performance
2.2. Proximate Composition
2.3. Anti-Nutritional Factors in Lentil Genotypes
2.4. Antioxidant in Lentil Genotypes
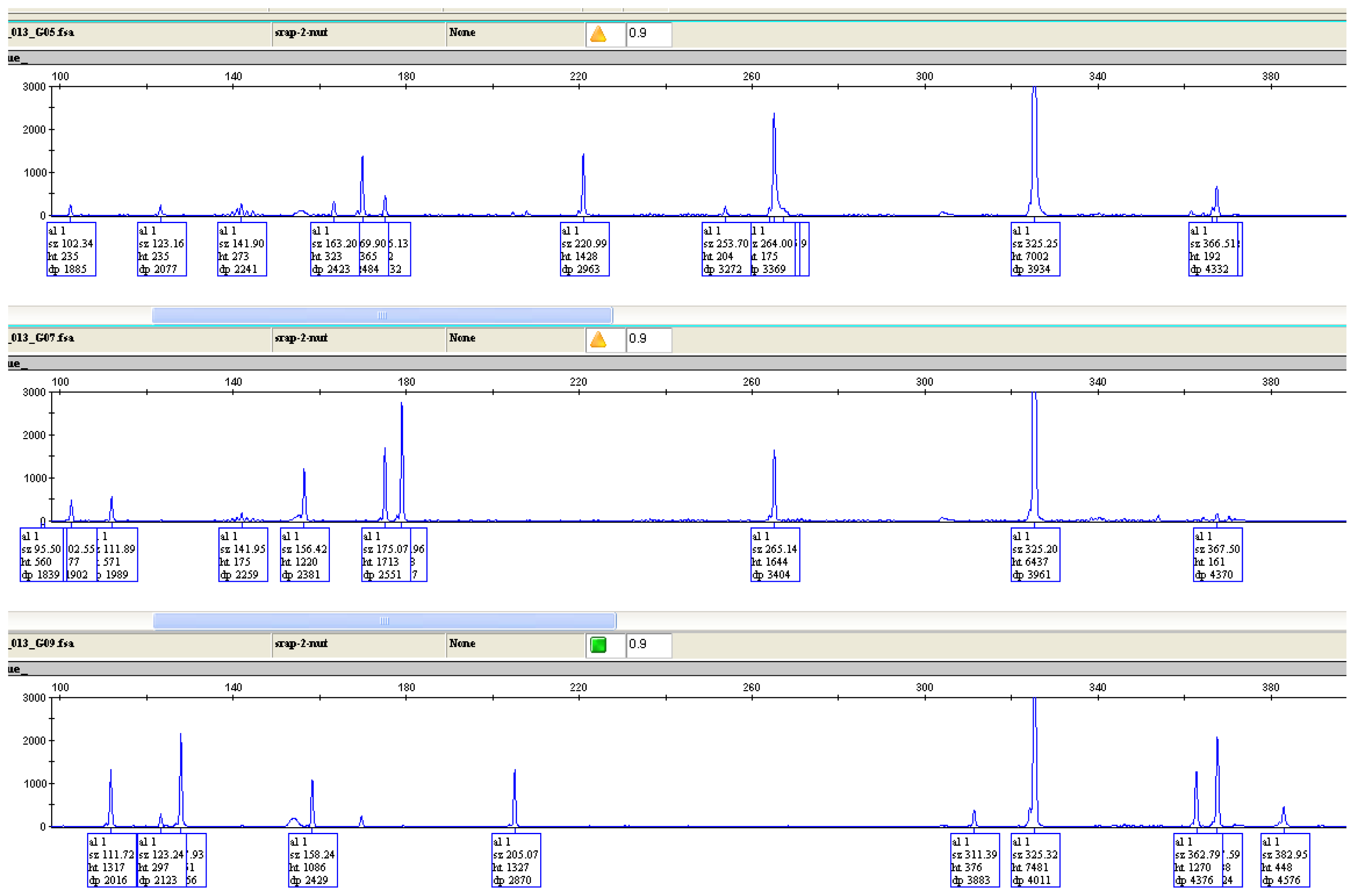
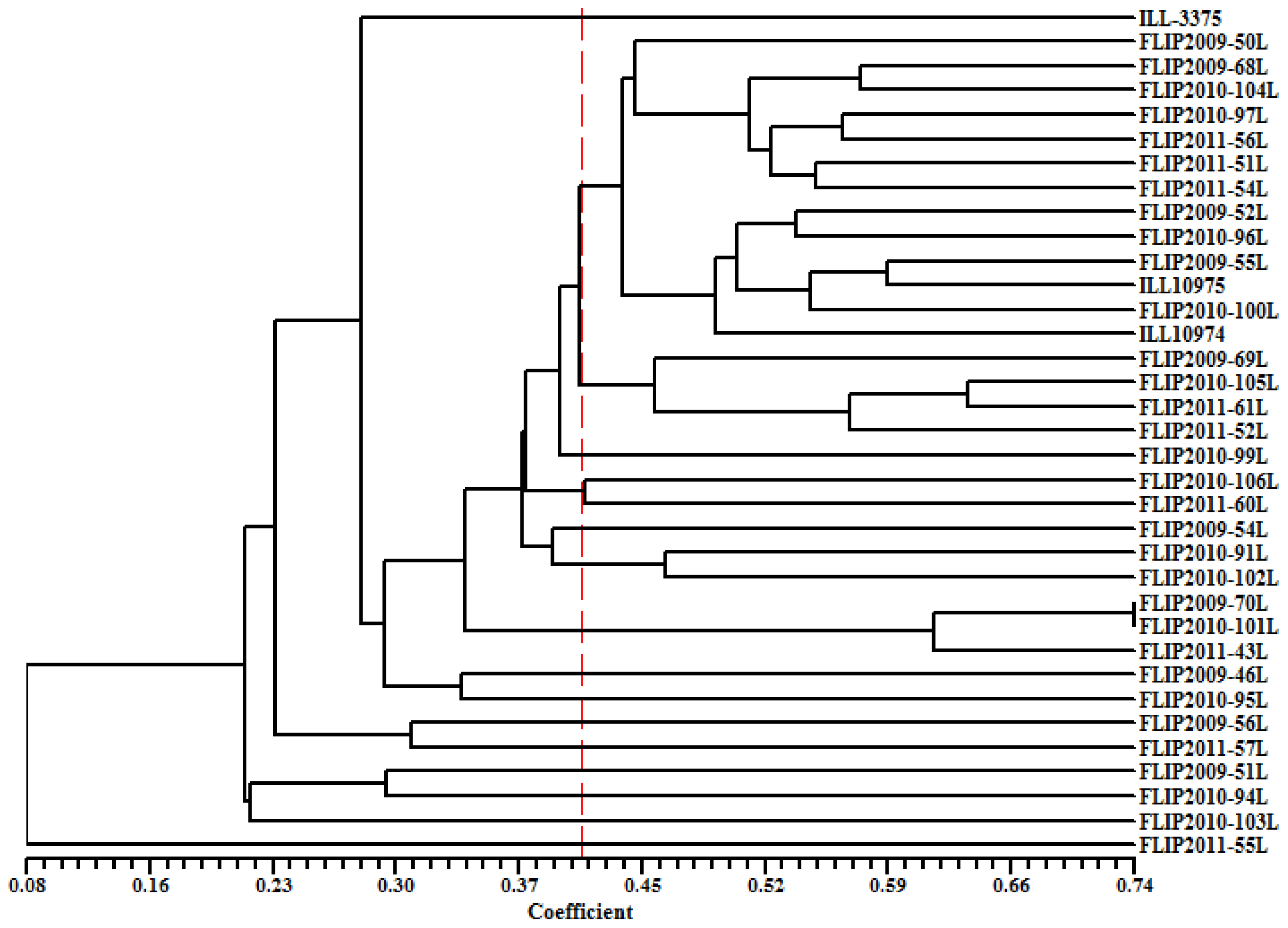
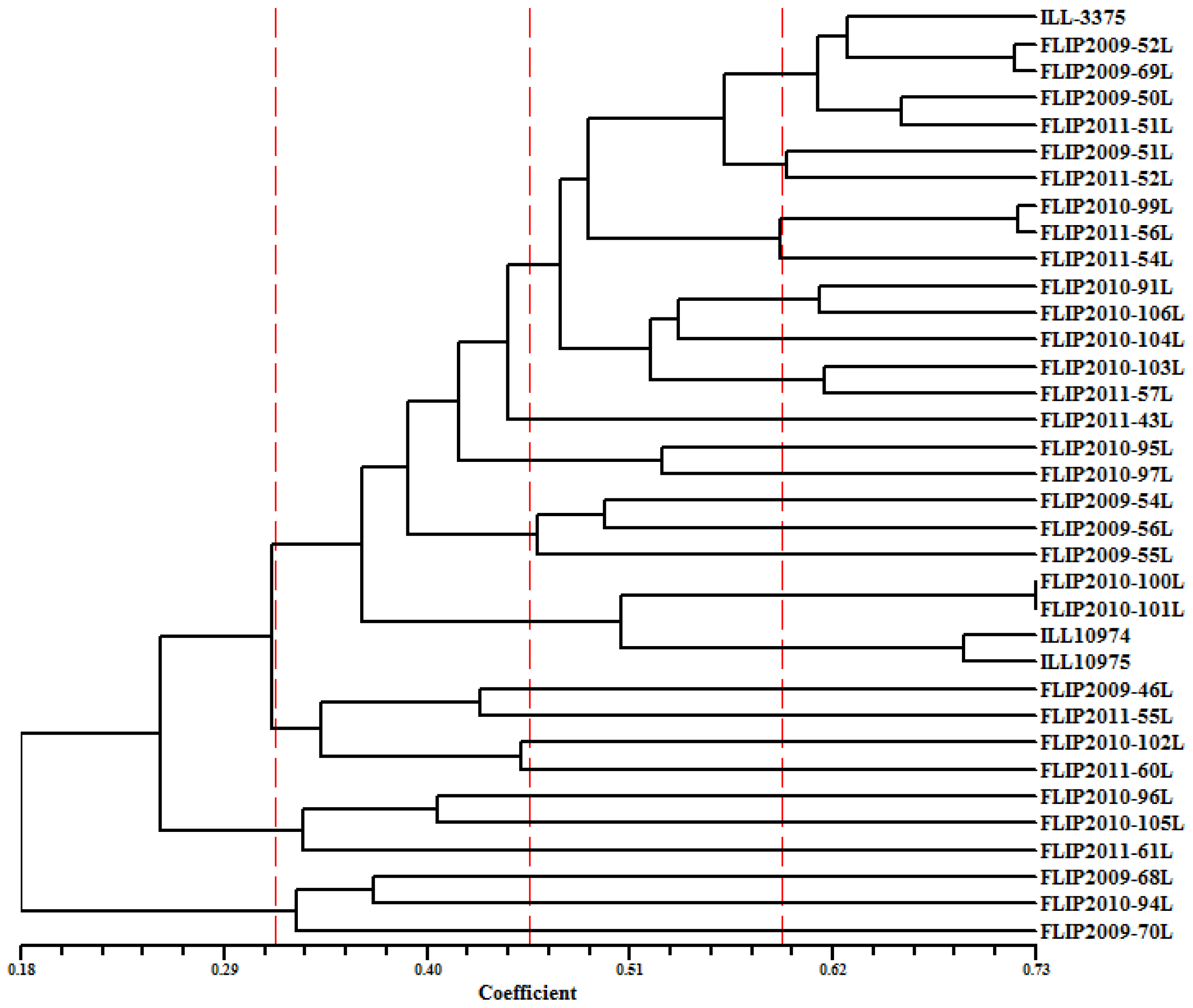
2.5. Genetic Diversity Based on SRAP and AFLP Markers
2.6. Discussion
3. Experimental Section
4. Conclusions
Supplementary Information
ijms-15-00277-s001.pdfAcknowledgments
Conflicts of Interest
References
- Arumuganathan, K.; Earle, E.D. Nuclear DNA content of some important plant species. Plant Mol. Biol 1991, 9, 208–218. [Google Scholar]
- FAOSTAT Database. Available online: http://faostat.fao.org/site/339/default.aspx (accessed on 15 January 2013).
- Callaway, J.C. Hempseed as a nutritional resource: An overview. Euphytica 2004, 140, 65–72. [Google Scholar]
- Handbook of World Food Legumes; Salunkhe, D.; Kadam, S. (Eds.) CRC: Boca Raton, FL, USA, 1989.
- Zheng, G.; Han, H.; Bhatty, R. Physicochemical properties of zero amylose hull-less barley starch. Cereal Chem 1998, 75, 520–524. [Google Scholar]
- Frederick, M.; Cho, S.; Sarker, A.; McPhee, K.; Coyne, C.; Rajesh, P.; Ford, P. Application of biotechnology in breeding lentil for resistance to biotic and abiotic stress. Euphytica 2006, 147, 149–165. [Google Scholar]
- USDA National Nutrient Database for Standard Reference. Available online: http://www.ars.usda.gov/Services/docs.htm?docid=8964 (accessed on 20 December 2013).
- Thavarajah, D.; Thavarajah, P.; Sarker, A.; Materne, M.; Vandemark, G.; Shrestha, R.; Idrissi, O.; Hacikamiloglu, O.; Bucak, B.; Vandenberg, A. A global survey of effects of genotype and environment on selenium concentration in lentils (Lens culinaris L.): Implications for nutritional fortification strategies. Food Chem 2011, 125, 72–76. [Google Scholar]
- Vandenberg, A. Lentil Expansion in Canada. In Milestones in Food Legume Research; Ali, M., Kumar, S., Eds.; Indian Institute of Pulses Research: Kanpur, India, 2009; pp. 58–72. [Google Scholar]
- Hayward, M.D.; Breese, E.L. Population Structure and Variability. In Plant Breeding: Principles and Prospects; Hayward, M.D., Bosemark, N.O., Eds.; Chapman and Hall: London, UK, 1993; pp. 7–29. [Google Scholar]
- Sindhu, J.S.; Mishra, H.O. Genetic variability in Indian microsperma types of lentil. Lentil Exp. News. Serv 1982, 9, 10–11. [Google Scholar]
- Ramgiry, S.R.; Paliwal, K.K.; Tomar, S.K. Variability and correlation of grain yield and other qualitative characters in lentil. LENS Newsl 1989, 16, 19–21. [Google Scholar]
- Sarker, A.; Erskine, W. Utilization of Genetic Resources in Lentil Improvement. Proceedings of the Genetic Resources of Field Crops: Genetic Resources Symposium, EUCARPIA, Poznam, Poland, 16–20 May 2001.
- Tullu, A.; Kusmenoglu, I.; McPhee, K.E.; Muehlbauer, F.J. Characterization of core collection of lentil germplasm for phenology, morphology, seed and straw yields. Genet. Resour. Crop Evol 2001, 48, 143–152. [Google Scholar]
- Fratini, R.; Durán, Y.; García, P.; Perez de la Vega, M. Identification of quantitative trait loci (QTL) for plant structure, growth habit and yield in lentil. Span. J. Agric. Res 2007, 5, 348–356. [Google Scholar]
- Tullu, A.; Taran, B.; Warkentin, T.; Vandenberg, A. Construction of an intraspecific linkage map and QTL analysis for earliness and plant Height in lentil. Crop Sci 2008, 48, 2254–2264. [Google Scholar]
- Zaccardelli, M.; Lupo, F.; Piergiovanni, A.R.; Aghetti, G.; Sonnante, G.; Daminati, M.G.; Sparvoli, F.; Lioi, L. Characterization of Italian lentil (Lens culinaris Medik.) germplasm by agronomic traits, biochemical and molecular markers. Genet. Resour. Crop Evol 2012, 59, 727–738. [Google Scholar]
- Gautam, N.K.; Singh, M.; Khan, Z.; Roy, A.; Akhtar, J.; Ram, B. Assessment of lentil germplasm with respect to agromonic performance and major biotic stress. Legume Res 2013, 36, 214–219. [Google Scholar]
- Roy, S.; Islam, M.A.; Sarker, A.; Malek, M.A.; Rafii, M.Y.; Ismail, M.R. Determination of genetic diversity in lentil germplasm based on quantitative traits. Aust. J. Crop Sci 2013, 7, 14–21. [Google Scholar]
- Erskine, W.; Witcombe, J.R. Lentil Germplasm Catalog; ICARDA: Aleppo, Syria, 1984. [Google Scholar]
- Samec, P.; Nasinec, V. The use of RAPD technique for identification and classification of Pisum sativum L. genotypes. Euphytica 1996, 89, 229–234. [Google Scholar]
- Sonnante, G.; Pignone, D. Assessment of genetic variation in a collection of lentil using molecular tools. Euphytica 2001, 120, 301–307. [Google Scholar]
- Abo-Elwafa, A.; Muraik, K.; Shimada, T. Intra- and inter-specific variation in Lens species revealed by RAPD markers. Theor. Appl. Genet 1995, 90, 335–340. [Google Scholar]
- Sharma, S.K.; Dawson, I.K.; Waugh, R. Relationship among cultivated and wild lentils revealed by RAPD analysis. Theor. Appl. Genet 1995, 91, 647–654. [Google Scholar]
- Sharma, S.K.; Knox, M.R.; Ellis, T.H.N. AFLP analysis of the diversity and phylogeny of Lens and its comparison with RAPD analysis. Theor. Appl. Genet 1996, 93, 751–758. [Google Scholar]
- Ford, R.; Pang, E.C.K.; Taylor, P.W.J. Diversity analysis and species identification in lens using PCR generated markers. Euphytica 1997, 96, 247–255. [Google Scholar]
- Alvarez, M.; Garcia, P.; Perez de la Vega, M. RAPD polymorphism in Spanish lentil landraces and cultivars. J. Genet. Breed 1997, 51, 91–96. [Google Scholar]
- Fikiru, E.; Tesfaye, K.; Bekele, E. Genetic diversity and population structure of Ethiopian lentil (Lens culinaris Medikus) landraces as revealed by ISSR marker. Afr. J. Biotechnol 2007, 6, 1460–1468. [Google Scholar]
- Sonnante, G.; Pignone, D. The major Italian landraces of lentil (Lens culinaris Medik.): Their molecular diversity and origin. Genet. Resour. Crop Evol 2007, 54, 1023–1031. [Google Scholar]
- Toklu, F.; Karaköy, T.; Hakle, I.; Bicer, T.; Brandolini, A.; Kilian, B.; Ozkan, H. Genetic variation among lentil (Lens culinaris Medik.) landraces from Southeast Turkey. Plant Breed 2009, 128, 178–186. [Google Scholar]
- El-Nahas, A.; El-Shazly, H.; Ahmed, S.; Omran, A. Molecular and biochemical markers in some lentil (Lens culinaris Medik.) genotypes. Ann. Agric. Sci 2011, 56, 105–112. [Google Scholar]
- Liu, J.; Guan, J.P.; Xu, D.X.; Zhang, X.Y.; Gu, J.; Zong, X.X. Genetic diversity and population structure in lentil (Lens culinaris Medik.) germplasm detected by SSR Markers. Acta Agron. Sin 2008, 34, 1901–1909. [Google Scholar]
- Babayeva, S.; Akparov, Z.; Abbasov, M.; Mammadov, A.; Zaifizadeh, M.; Street, K. Diversity analysis of Central Asia and Caucasian lentil (Lens culinaris Medik.) germplasm using SSR fingerprinting. Genet. Resour. Crop Evol 2009, 56, 293–298. [Google Scholar]
- Hamwieh, A.; Udupa, S.M.; Sarker, A.; Jung, C.; Baum, M. Development of new microsatellite markers and their application in the analysis of genetic diversity in lentils. Breed. Sci 2009, 59, 77–86. [Google Scholar]
- Kaur, S.; Cogan, N.; Pembleton, L.; Shinozuka, M.; Savin, K.; Materne, M.; Forster, J. Transcriptome sequencing of lentil based on second-generation technology permits large-scale unigene assembly and SSR marker discovery. BMC Genomics 2011. [Google Scholar] [CrossRef]
- Havey, M.J.; Muehlbauer, F.J. Variability for restriction fragment lengths and phylogenies in lentil. Theor. Appl. Genet 1989, 77, 839–843. [Google Scholar]
- Muench, D.G.; Slinkard, A.E.; Scoles, G.J. Determination of genetic variation and taxonomy in lentil (Lens Miller) species by chloroplast DNA polymorphism. Euphytica 1991, 56, 213–218. [Google Scholar]
- Li, G.Y.; Quiros, C.F. Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction, its application to mapping and gene tagging in Brassica. Theor. Appl. Genet 2001, 103, 455–461. [Google Scholar]
- Vos, P.; Hogers, R.; Bleeker, M.; Reijans, M.; Lee, T.V.; Hornes, M.; Frijters, A.; Pot, J.; Peleman, J.; Kuiper, M.; et al. AFLP: A new technique for DNA fingerprinting. Nucleic Acids Res 1995, 23, 4407–4414. [Google Scholar]
- Singh, K.B.; Singh, S. Evaluation of exotic germplasm in lentil. J. Agric. Res 1991, 6, 304–306. [Google Scholar]
- Jain, S.K.; Madaria, S.K.; Rao, S.K.; Nigam, P.K. Analysis of yield factors in lentil. Indian J. Agric. Res 1995, 29, 173–180. [Google Scholar]
- Chakrabotry, M.; Haque, M.F. Genetic variability and component analysis in lentil (Lens culinaris Medik.). J. Res 2000, 25, 199–204. [Google Scholar]
- Singh, I.; Sardana, V.; Sekhon, H. Influence of row spacing and seed rate on seed yield of lentil (Lens culinaris) under different sowing dates. Indian J. Agron 2005, 50, 308–310. [Google Scholar]
- Toklu, F.; Biçer, B.T.; Karaköy, T. Agro-morphological characterization of the Turkish lentil landraces. Afr. J. Biotechol 2009, 8, 4121–4127. [Google Scholar]
- Tuba, B.; Sakar, D. Heritability of yield and its components in lentil (Lens culinaris Medik.). Bulg. J. Agric. Sci 2010, 16, 30–35. [Google Scholar]
- Aghili, P.; Imani, A.; Shahbazi, H.; Alaei, Y. Study of correlation and relationships between seed yield and yield components in Lentil (Lens culinaris Medik). Ann. Biol. Res 2012, 3, 5042–5045. [Google Scholar]
- Sabaghnia, N.; Karimizadeh, R.; Mohammadi, M. Grouping genotypes and test environments by some cluster methods regarding genotype × environment interaction in multi-environment trials. Genetika 2012, 44, 457–473. [Google Scholar] [Green Version]
- Gupta, D.; Sharma, S.K. Evaluation of wild Lens taxa for agro-morphological traits, fungal diseases andmoisture stress in North Western Indian Hills. Genet. Resour. Crop Evol 2006, 53, 1233–1241. [Google Scholar]
- Mondal, M.A.; Puteh, A.; Malek, M.; Roy, S.; Yusop, M. Contribution of morpho-physiological traits on yield of lentil (Lens culinaris Medik). Aust. J. Crop Sci 2013, 7, 167–1172. [Google Scholar]
- Tuba, B.D.; Sakar, D. Studies on variability of lentil genotypes in southern Anatolia of Turkey. Not. Bot. Horti Agrobot. Cluj 2008, 36, 20–24. [Google Scholar]
- Dewan, M.A. Characterization of Lentil Mutants in Respect of Their Morphological and Yield Attributes. Master’s Thesis, Bangladesh Agricultural University, Mymensingh, Bangladesh, 2005. [Google Scholar]
- Wang, N. Quality of Western Canadian Lentils. (accessed on 20 December 2013).
- Mondal, M.M.A.; Howlader, M.H.K.; Akter, M.B.; Dutta, R.K. Evaluation of five advanced lentil mutants in relation to morphophysiological characters and yield. Bangladesh J. Crop Sci 2007, 18, 367–372. [Google Scholar]
- Yadav, S.S.; Phogat, D.S.; Solanki, I.S.; Tomer, T.S. Character association and path analysis in lentil. Indian J. Pulse Res 2003, 16, 22–24. [Google Scholar]
- Anzam, M.S.; Ali, A.; Iqbal, S.M.; Haqqani, A.M. Evaluation and correlation of economically important traits in exotic germplasm of lentil. Int. J. Agric. Biol 2005, 7, 959–961. [Google Scholar]
- Younis, N.; Hanif, M.; Sadiq, S.; Abbas, G.; Asghar, M.J.; Haq, M.A. Estimation of genetic parameters and path analysis in lentil. Pak. J. Agric. Sci 2008, 45, 44–48. [Google Scholar]
- Karadavut, U. Path analysis for yield and yield components in lentil. Turk. J. Field Crop 2009, 14, 97–104. [Google Scholar]
- Jood, S.; Bishnoi, S.; Sharma, A. Chemical analysis and physico-chemical properties of chickpea and lentil cultivars. Nahrung 1998, 42, 71–74. [Google Scholar]
- Boyle, J.L.; Aksay, S.; Roufik, S.; Ribereau, S.; Mondor, M.; Mondor, M.; Farnworth, E.; Rajamohamed, S.H. Comparison of the functional peoperties of pea, chickpea and lentil protein concentrates processed using ultrafiltration and isoelectric precipitation techniques. Food Res. Int 2010, 43, 537–546. [Google Scholar]
- Taleb, M.H.; Khodambashi, M.; Karimi, M. Study of physical and nutritional quality properties in segregating generations of lentil cross. Int. J. Agric. Crop Sci 2013, 5, 2740–2742. [Google Scholar]
- Wang, N.; Daun, J.K. The Chemical Composition and Nutritive Value of Canadian Pulses: Lentil (Lens culinaris); Canadian Grain Commission Grain Research Laboratory: Winnipeg, MB, Canada, 2004; pp. 8–18. [Google Scholar]
- Barulina, H. Lentils of the U.S.S.R. and Other Countries (in Russian); Vsesoiuznaia akademiia sel’sko khoziaistvennyk im. Lenina: Leningrad, Russia, 1930; Volume 40, pp. 1–319. [Google Scholar]
- Solanki, I.S.; Kapoor, A.C.; Singh, U. Nutritional parameters and yield evaluation of newly developed genotypes of lentil (Lens culinaris Medik.). Plant Foods Hum. Nutr 1999, 54, 79–87. [Google Scholar]
- El-Adawy, T.A.; Rahma, E.H.; Eel-Bedawey, A.A.; El-Beltagy, A.E. Nutritional potential and functional properties of germinated mung bean, pea and lentil seeds. Plant Foods Hum. Nutr 2003, 58, 1–13. [Google Scholar]
- Sulieman, M. Physiochemical and Structural Characterization of Germinated and Cooked Lentils (Lens culinaris). Master’s Thesis, University of Khartoum, Khartom, Sudan, 2007. [Google Scholar]
- Wang, N.; Hatcher, D.W.; Toews, R.; Gawalko, E.J. Influence of cooking and dehuling on nutritional composition of several varieties of lentils (Lens culinaris). LWT Food Sci. Technol 2009, 42, 842–848. [Google Scholar]
- Devos, P. Nutritional Value of Lentils and Chickpeas and Changes during Processing. In “Herkes Için Mercimek” Sempozyumu (in Turkish); Toprak Mahsulleri Ofisi: Ankara, Turkey, 1988; pp. 186–196. [Google Scholar]
- Lim, T.K. Edible Medicinal and Non-Medicinal Plants; Springer: Berlin, German, 2012; Volume 2. [Google Scholar]
- Urbano, G.; Porres, J.M.; Frias, J.; Vidal-Valverde, C. Nutritional Value. In Lentil: An Ancient Crop for Modern Times; Yadav, S.S., McNeil, D., Stevenson, P.C., Eds.; Springer: Heidelberg, Germany, 2007; pp. 47–93. [Google Scholar]
- Wang, N.; Daun, J.K. Effects of variety and crude protein content on nutrients and anti-nutrients in lentils (Lens culinaris). Food Chem 2006, 95, 493–502. [Google Scholar]
- Grusak, M. Nutritional and Health-Beneficial Quality. In The Lentil: Botany, Production and Uses; Erskine, W., Muehlbauer, F., Sarker, A., Sharma, B., Eds.; CABI: Oxford, UK, 2009; pp. 368–390. [Google Scholar]
- FAO. Energy and Protein Requirements. Available online: http://www.fao.org/docrep/003/aa040e/aa040e00.htm (accessed on 19 December 2013).
- Lobo, V.; Patil, A.; Phatak, A.; Chandra, N. Free radicals, antioxidants and functional foods: Impact on human health. Pharmacogn. Rev 2010, 4, 118–126. [Google Scholar]
- Prior, R.; Cao, G. Analysis of botanicals and dietary supplements for antioxidant capacity: A review. J. AOAC Int 2000, 83, 950–956. [Google Scholar]
- Amarowicz, R.; Piskuła, M.; Honke, J.; Rudnicka, B.; Troszyńska, A.; Kozłowska, H. Extraction of phenolic compounds from lentil seeds (Lens culinaris) with various solvents. Pol. J. Food Nutr. Sci 1995, 4, 53–62. [Google Scholar]
- Amarowicz, R.; Estrella, I.; Hernández, T.; Dueñas, M.; Troszyńska, A.; Kosińska, A.; Ronald, B. Antioxidant activity of a red lentil extract and its fractions. Int. J. Mol. Sci 2009, 10, 5513–5527. [Google Scholar]
- Amarowicz, R.; Estrella, I.; Hernández, T.; Robredo, S.; Troszyńska, A.; Kosińska, A. Free radical-scavenging capacity, antioxidant activity, and phenolic composition of green lentil (Lens culinaris). Food Chem 2010, 121, 705–711. [Google Scholar]
- Scariot, V.; de Keyser, E.; Handa, T.; de Riek, J. Comparative study of the discriminating capacity and effectiveness of AFLP, STMS and EST markers in assessing genetic relationships among evergreen azales. Plant Breed 2007, 126, 207–212. [Google Scholar]
- Association of Official Analytical Chemists (AOAC), Official Methods of Analysis of AOAC International, 15th ed; AOAC: Washington, DC, USA, 1990.
- Association of Official Analytical Chemists (AOAC), Official Methods of Analysis of AOAC International, 16th ed; AOAC: Washington, DC, USA, 1995.
- Association of Official Analytical Chemists (AOAC), Official Methods of Analysis of AOAC International, 18th ed; AOAC: Washington, DC, USA, 2005.
- Xu, B.J.; Chang, S.K. A comparative study on phenolic profiles and antioxidant activities of legumes as affected by extraction solvents. J. Food Sci 2007, 72, S159–S166. [Google Scholar]
- Llorach, R.; Martínez-Sánchez, A.; Tomás-Barberán, F.A.; Gil, M.I.; Ferreres, F. Characterisation of polyphenols and antioxidant properties of five lettuce varieties and escarole. Food Chem 2008, 108, 1028–1038. [Google Scholar]
- Kakade, M.L.; Simons, N.; Liener, I.E. An evaluation of natural vs. synthetic substrate for measuring the antitryptic activity of soy bean samples. Cereal Chem 1969, 49, 518–526. [Google Scholar]
- Mohamed, A.; Perera, P.; Hafez, Y. New chromophore for phytic acid determination. Cereal Chem 1986, 63, 475–478. [Google Scholar]
- Price, M.L.; Socoyoc, S.V.; Butler, L.G. A critical evaluation of the vanillin reaction as an assay for tannin in sorghum grain. J. Agric. Food Chem 1978, 26, 1214–1218. [Google Scholar]
- Molecular Genetic Analysis of Populations, A Practical Approach, 2nd ed; Hoelzel, A.R. (Ed.) Oxford University Press: New York, NY, USA, 1998.
- Alghamdi, S.S.; Al-Faifi, S.A.; Migdadi, H.M.; Khan, M.A.; El-Harthy, E.H.; Ammar, M.H. Molecular diversity assessment using sequence related amplified polymorphism (SRAP) markers in Vicia Faba L. Int. J. Mol. Sci 2012, 13, 16457–16471. [Google Scholar]
- Wooten, J.A.; Tolley-Jordan, L.R. Validation of phylogenetic signals in amplified fragment length data: Testing the utility and reliability in closely related taxa. BMC Res. Notes 2009. [Google Scholar] [CrossRef]
- Jaccard, P. Nouvelles recherches sur la distribution florale. Bull. Soc. Vaud. Sci. Nat 1908, 44, 223–270. [Google Scholar]
- Rohlf, F.J. Ntsys-pc: Numerical Taxonomy and Multivariate Analysis System, Version 2.2; Exeter Publishing Ltd: New York, NY, USA, 2005. [Google Scholar]
| Character | Mean | SD | Min | Max | CV |
|---|---|---|---|---|---|
| Number of days to 50% flowering | 102.8 | 5.26 | 86 | 112 | 5.1 |
| Number of days to 95% maturing | 135.8 | 4.66 | 121.8 | 142.6 | 3.4 |
| Plant height (cm) | 31 | 2.34 | 27 | 35.7 | 7.5 |
| Number of branches/plant | 3.25 | 0.497 | 2.4 | 4.3 | 15.2 |
| Number of seeds/plant | 28.4 | 5.64 | 18.8 | 42.6 | 19.8 |
| Seed yield/plant | 1.09 | 0.222 | 0.8 | 1.5 | 20.3 |
| S.O.V | DF | No. of Days to 50% Flowering | No. of Days to 95% Maturing | Plant Height (cm) | No. of Branches | No. of Seeds/Plant | Seed Yield/Plant (g) |
|---|---|---|---|---|---|---|---|
| Year (Y) | 1 | 2597.1 ** | 5260.0 ** | 6956.9 ** | 3.55 ns | 6296.5 ** | 7.13 ** |
| Error | 4 | 15.7 | 10.3 | 14.1 | 1.2 | 1.2 | 0.04 |
| Genotype (G) | 34 | 166.2 ** | 130.5 ** | 33.0 ** | 1.50 ** | 191.6 ** | 0.31 ** |
| G × Y | 34 | 14.9 ns | 20.8 ** | 32.3 ** | 0.47 ns | 46.9 ns | 0.06 ns |
| Error | 136 | 17.5 | 11.4 | 14.9 | 0.6 | 34.6 | 0.07 |
| Components | P1 | P2 | P3 | P4 |
|---|---|---|---|---|
| Eigen value | 2.034 | 1.446 | 0.997 | 0.881 |
| Variability (%) | 33.894 | 24.092 | 16.616 | 14.679 |
| Cumulative (%) | 33.894 | 57.986 | 74.602 | 89.281 |
| Eigen Vectors | ||||
| Days to 50% Flowering | 0.502 | −0.467 | −0.049 | 0.24 |
| Days to 95% Maturity | 0.571 | −0.313 | 0.139 | 0.235 |
| Plant Height | −0.028 | −0.333 | 0.718 | −0.604 |
| Number of Branches/Plant | −0.064 | 0.361 | 0.674 | 0.625 |
| Number of Seeds/Plant | 0.432 | 0.503 | 0.073 | −0.271 |
| Seed Yield/Plant | 0.481 | 0.436 | −0.048 | −0.239 |
| Variable | Mean | SD | CV | Min | Max |
|---|---|---|---|---|---|
| Crude protein (%) | 27.346 | 1.273 | 4.66 | 25.3 | 29.3 |
| Moisture (%) | 12.631 | 0.44 | 3.49 | 11.6 | 13.6 |
| Ash (%) | 2.60 | 0.115 | 4.42 | 2.39 | 2.89 |
| Fats (%) | 0.9991 | 0.121 | 12.11 | 0.72 | 1.19 |
| Carbohydrate (%) | 43.997 | 2.215 | 5.03 | 39.5 | 47.1 |
| Essentials Amino Acids (g/kg) | |||||
| Leu | 8.643 | 0.797 | 9.22 | 6.8 | 9.8 |
| Ile | 4.569 | 0.636 | 13.92 | 3.4 | 5.6 |
| Phe | 6.389 | 0.875 | 13.7 | 4.9 | 8.2 |
| Try | 0.7909 | 0.091 | 11.5 | 0.61 | 0.92 |
| His | 4.84 | 0.63 | 13.02 | 3.6 | 6.1 |
| Val | 7.389 | 0.991 | 13.42 | 5.3 | 9 |
| Thr | 6.011 | 1.087 | 18.08 | 4.1 | 7.9 |
| Met | 1.5031 | 0.3127 | 20.8 | 0.85 | 2.1 |
| Lys | 7.309 | 1.018 | 13.94 | 4.5 | 8.6 |
| Arg | 8.526 | 1.001 | 11.74 | 6.6 | 10 |
| Minerals (mg/100 g) | |||||
| Ca | 74.92 | 4.433 | 5.92 | 64.9 | 84.8 |
| P | 412.4 | 66.5 | 16.13 | 286.9 | 546.7 |
| K | 886.1 | 99.6 | 11.24 | 674.4 | 1061.2 |
| Mg | 138.58 | 6.98 | 5.04 | 126.1 | 157.3 |
| Fe | 7.4483 | 0.4455 | 5.98 | 6.57 | 8.57 |
| Mn | 1.5317 | 0.3818 | 24.92 | 1.26 | 2.85 |
| Cu | 1.1337 | 0.1617 | 14.26 | 0.86 | 1.37 |
| Zn | 3.7777 | 0.4809 | 12.73 | 2.63 | 4.51 |
| Antioxidants | |||||
| TPC a (mg GAE/g) | 13.467 | 4.546 | 33.76 | 6.45 | 20.82 |
| TFC b (mg QE/g) | 6.03 | 1.38 | 22.89 | 4.12 | 8.92 |
| DPPH c (μg/g) | 15.092 | 3.036 | 20.12 | 10.61 | 23.26 |
| Anti-nutritional Factors | |||||
| Protease inhibitors (mg/g) | 2.5234 | 0.1847 | 7.32 | 2.08 | 2.78 |
| Tannins (mg/g) | 0.6846 | 0.068 | 9.93 | 0.52 | 0.79 |
| Phytic acids (mg/g) | 0.928 | 0.0908 | 9.79 | 0.72 | 1.09 |
| SRAP | AFLP | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Primer Combination | Total Fragments | Polymorphic Fragments | Polymorphism Rate (%) | PIC | Primer Combination | Total Fragments | Polymorphic Fragments | Polymorphism Rate (%) | PIC |
| ME1/EM1 | 132 | 132 | 100 | 0.924 | ECT/MCTG | 569 | 569 | 100 | 0.968 |
| ME1/EM2 | 393 | 393 | 100 | 0.964 | ETA/MCTC | 375 | 375 | 100 | 0.957 |
| ME2/EM3 | 399 | 399 | 100 | 0.963 | ETC/MCTA | 583 | 583 | 100 | 0.963 |
| ME2/EM4 | 124 | 124 | 100 | 0.908 | ECC/MCCT | 98 | 98 | 100 | 0.894 |
| ME2/EM5 | 708 | 708 | 100 | 0.970 | Total | 1625 | - | - | - |
| ME2/EM6 | 1138 | 1138 | 100 | 0.984 | Mean | 406.25 | - | - | - |
| Total (Mean) | 2894 (482) | - | - | - | - | - | - | - | - |
| Genotype | Pedigree | Origin |
|---|---|---|
| ILL 3375 | ILL 3375 | India |
| FLIP2009-50L | ILL 8090 × ILL 7685 | ICARDA |
| FLIP 2009-51L | ILL 7617 × ILL 4404 | ICARDA |
| FLIP 2009-52L | ILL 5883 × ILL 8113 | ICARDA |
| FLIP 2009-54L | ILL 7012 × ILL 4404 | ICARDA |
| FLIP 2009-55L | ILL 6783 × ILL 98 | ICARDA |
| FLIP 2009-56L | ILL 8077 × ILL 6994 | ICARDA |
| FLIP 2009-64L | - | ICARDA |
| FLIP 2009-68L | ILL 7713 × ILL 7201 | ICARDA |
| FLIP 2009-69L | ILL 790 × ILL 7706 | ICARDA |
| FLIP 2009-70L | ILL 7537 × ILL 4404 | ICARDA |
| FLIP 2010-91L | ILL 8114 × ILL 7555 | ICARDA |
| FLIP 2010-94L | ILL 7620 × ILL 8113 | ICARDA |
| FLIP 2010-95L | ILL 7620 × ILL 8113 | ICARDA |
| FLIP 2010-96L | ILL 7620 × ILL 8113 | ICARDA |
| FLIP 2010-97L | ILL 7620 × ILL 8113 | ICARDA |
| FLIP 2010-99L | ILL 7620 × ILL 8113 | ICARDA |
| FLIP2010-100L | ILL 2501 × ILL 7537 | ICARDA |
| FLIP 2010-101L | ILL 2501 × ILL 7537 | ICARDA |
| FLIP 2010-102L | ILL 4402 × ILL 2501 | ICARDA |
| FLIP 2010-103L | ILL 358 × ILL 87062 | ICARDA |
| FLIP 2010-104L | ILL 6037 × ILL 87062 | ICARDA |
| FLIP 2010-105L | ILL 7723 × ILL 87062 | ICARDA |
| FLIP 2010-106L | ILL 7723 × ILL 87062 | ICARDA |
| FLIP 2011-43L | ILL 7537 × ILL 590 | ICARDA |
| FLIP 2011-51L | ILL 590 × ILL 7979 | ICARDA |
| FLIP 2011-52L | ILL 7010 × ILL 6971 | ICARDA |
| FLIP 2011-54L | ILL 8090 × ILL 7980 | ICARDA |
| FLIP 2011-55L | ILL 8090 × ILL 7980 | ICARDA |
| FLIP 2011-56L | ILL 8090 × ILL 7980 | ICARDA |
| FLIP 2011-57L | ILL 8090 × ILL 7980 | ICARDA |
| ILL 10974 | 96-024L × 99H046 | Australia |
| ILL 10975 | 97-011 × 98H006-99HS001 | Australia |
| FLIP 2011-60L | ILL7723 × ILL8090 | ICARDA |
| FLIP 2011-61L | ILL7537 × ILL590 | ICARDA |
| - | - | - |
| Primer name | Forward 5′–3′ | Primer name | Reverse 5′–3′ |
|---|---|---|---|
| ME1 | TGAGTCCAAACCGGATA | EM1 | GACTGCGTACGAATTAAT |
| ME2 | TGAGTCCAAACCGGAGC | EM2 | GACTGCGTACGAATTTGC |
| ME3 | TGAGTCCAAACCGGAAT | EM3 | GACTGCGTACGAATTGAC |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Alghamdi, S.S.; Khan, A.M.; Ammar, M.H.; El-Harty, E.H.; Migdadi, H.M.; El-Khalik, S.M.A.; Al-Shameri, A.M.; Javed, M.M.; Al-Faifi, S.A. Phenological, Nutritional and Molecular Diversity Assessment among 35 Introduced Lentil (Lens culinaris Medik.) Genotypes Grown in Saudi Arabia. Int. J. Mol. Sci. 2014, 15, 277-295. https://doi.org/10.3390/ijms15010277
Alghamdi SS, Khan AM, Ammar MH, El-Harty EH, Migdadi HM, El-Khalik SMA, Al-Shameri AM, Javed MM, Al-Faifi SA. Phenological, Nutritional and Molecular Diversity Assessment among 35 Introduced Lentil (Lens culinaris Medik.) Genotypes Grown in Saudi Arabia. International Journal of Molecular Sciences. 2014; 15(1):277-295. https://doi.org/10.3390/ijms15010277
Chicago/Turabian StyleAlghamdi, Salem S., Altaf M. Khan, Megahed H. Ammar, Ehab H. El-Harty, Hussein M. Migdadi, Samah M. Abd El-Khalik, Aref M. Al-Shameri, Muhammad M. Javed, and Sulieman A. Al-Faifi. 2014. "Phenological, Nutritional and Molecular Diversity Assessment among 35 Introduced Lentil (Lens culinaris Medik.) Genotypes Grown in Saudi Arabia" International Journal of Molecular Sciences 15, no. 1: 277-295. https://doi.org/10.3390/ijms15010277






