Cloning and Polymorphisms of Yak Lactate Dehydrogenase b Gene
Abstract
:1. Introduction
2. Results
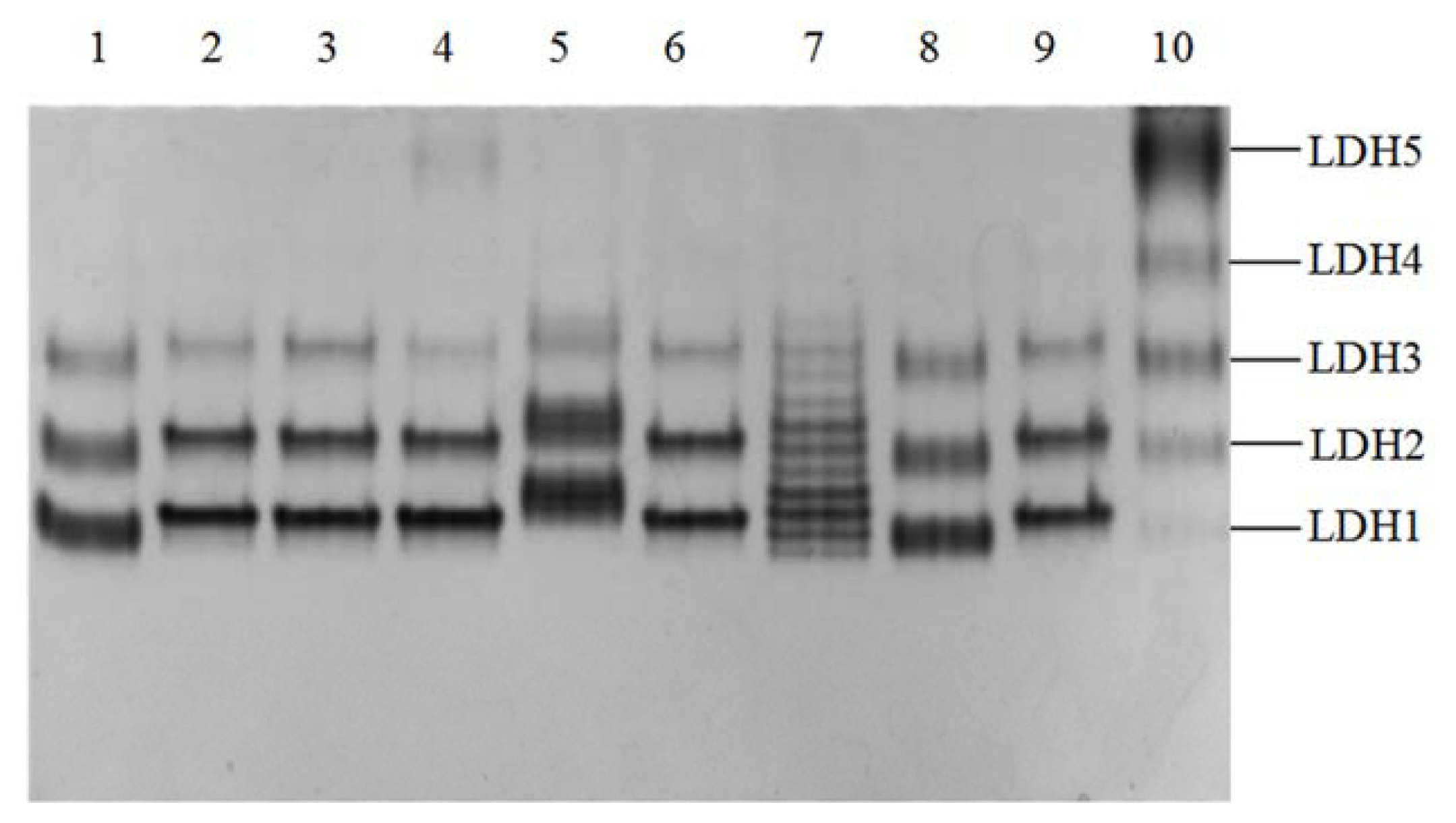
2.1. Phenotypes of Yak LDH1 Isozyme
2.2. Cloning and Sequencing of Yak ldhb cDNA
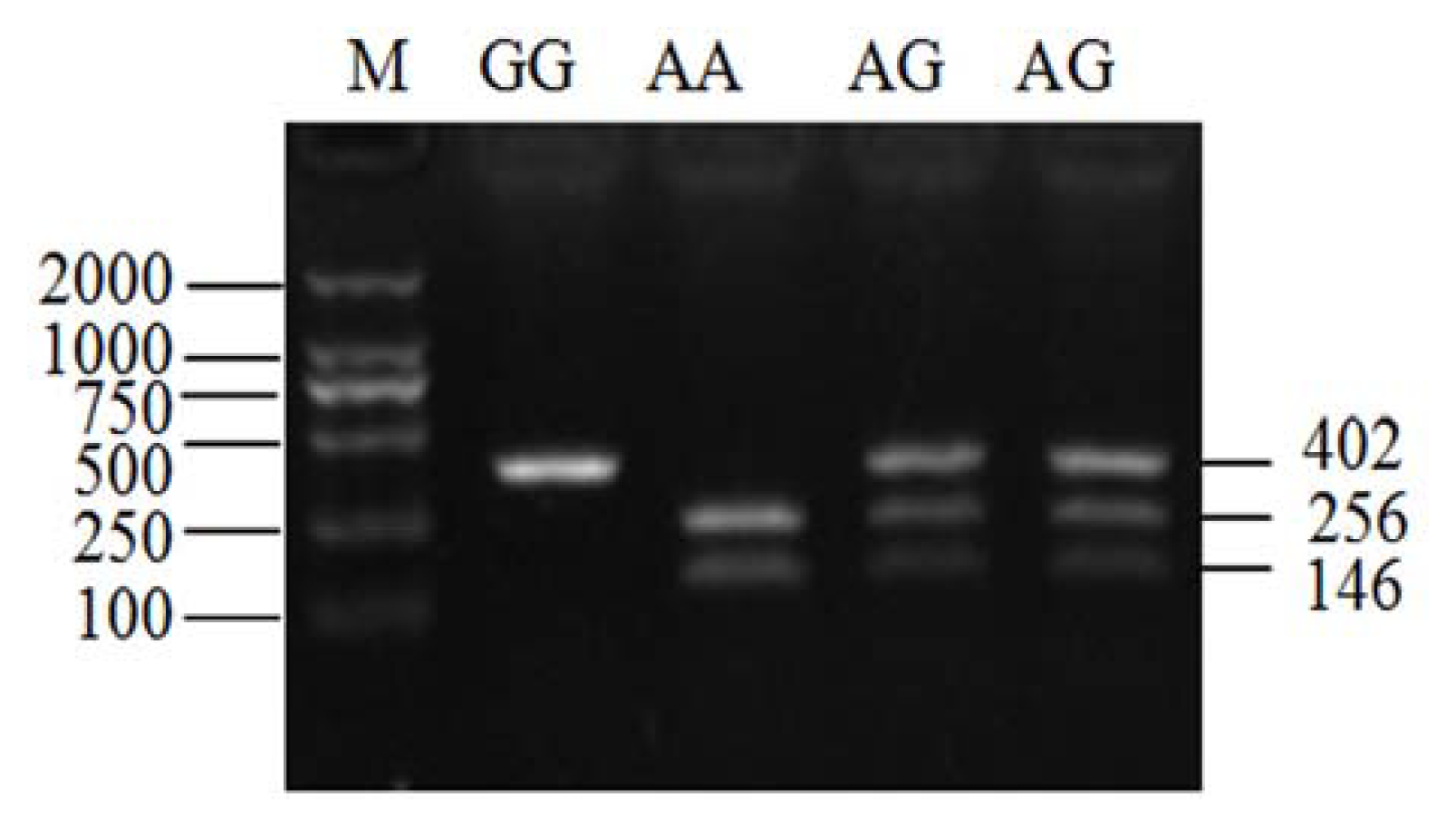
2.3. Genotyping of ldhb G896A in Two Yak Breeds
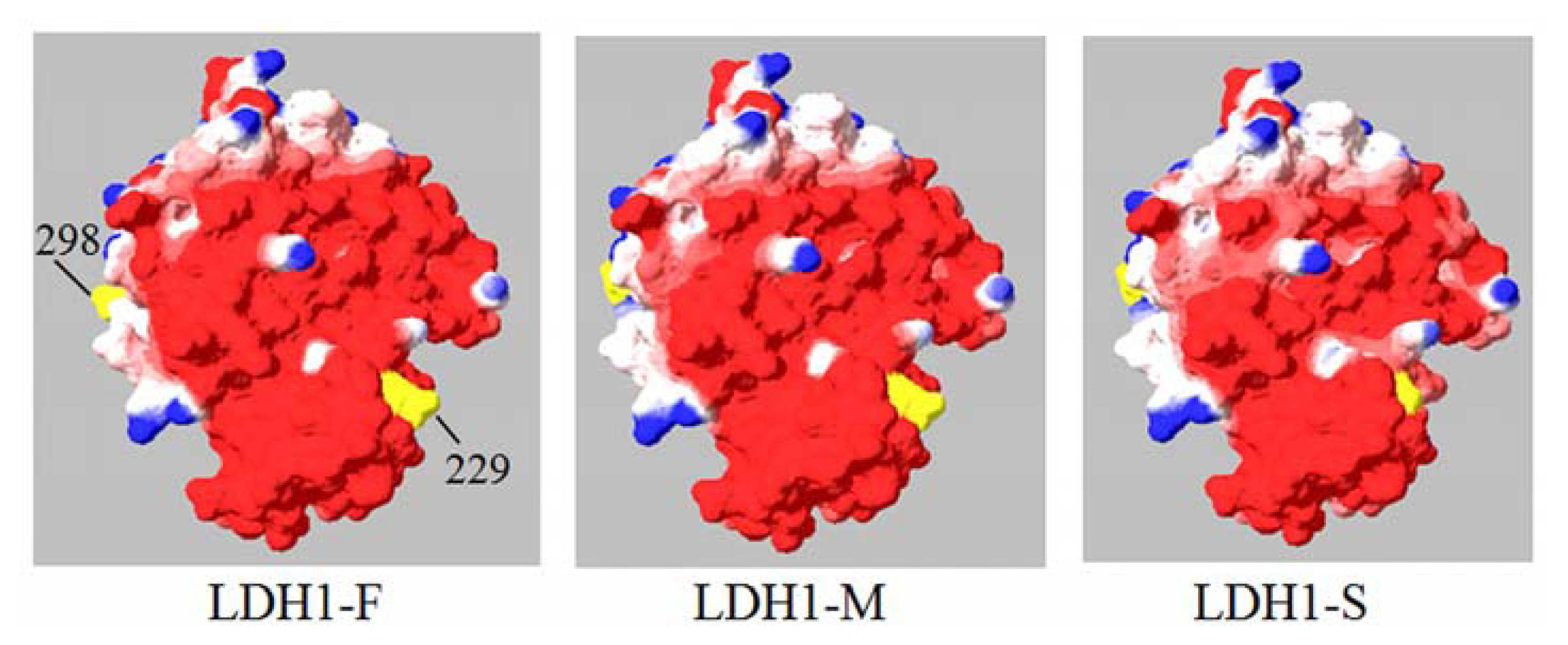
2.4. Molecular Modeling of H Subunit of Yak LDH
3. Discussion
4. Experimental Section
4.1. Animals and Tissue Samples
4.2. Muscle Extracts Preparation
4.3. Electrophoresis of Yak LDH Isozymes
4.4. Cloning and Sequencing of Yak ldhb Alleles
4.5. Assay of ldhb G896A Mutation by PCR-RFLP in Two Yak Breeds
4.6. Homology Modeling of H Subunit of Yak LDH
5. Conclusions
Acknowledgments
Conflict of Interest
References
- Wiener, G.; Han, J.L.; Long, R.J. The Yak, 2nd ed.; The Regional Office for Asia and the Pacific of the Food and Agriculture Organization of the United Nations: Bangkok, Thailand, 2003. [Google Scholar]
- Markert, C.L. Biochemistry and function of lactate dehydrogenase. Cell Biochem. Funct 1984, 2, 131–134. [Google Scholar]
- Li, S.S. Lactate dehydrogenase isoenzymes A (muscle), B (heart) and C (testis) of mammals and the genes coding for these enzymes. Biochem. Soc. Trans 1989, 17, 304–307. [Google Scholar]
- Sheafor, B.A. Metabolic enzyme activities across an altitudinal gradient: an examination of pikas (genus Ochotona). J. Exp. Biol 2003, 206, 1241–1249. [Google Scholar]
- Amano, T.; Yamado, W.; Nabika, T.; Zhang, X.L. Blood protein polymorphisms of Tibetan native cattle, yaks and their hybrid. Rep. Soc. Res. Nativ. Livest 1990, 13, 1–11. [Google Scholar]
- Zheng, Y.C.; Zhao, X.B.; Zhou, J.; Piao, Y.; Jin, S.Y.; He, Q.H.; Hong, J.; Li, N.; Wu, C.X. Identification of yak lactate dehydrogenase B gene variants by gene cloning. Sci. China Life Sci 2008, 51, 431–434. [Google Scholar]
- Hoppeler, H.; Vogt, M.; Weibel, E.R.; Flück, M. Response of skeletal muscle mitochondria to hypoxia. Exp. Physiol 2003, 88, 109–119. [Google Scholar]
- Lin, Y.Q.; Wang, G.S.; Feng, J.; Huang, J.Q.; Xu, Y.O.; Jin, S.Y.; Li, Y.P.; Jiang, Z.R.; Zheng, Y.C. Comparison of enzyme activities and gene expression profiling between yak and bovine skeletal muscles. Livest. Sci 2011, 135, 93–97. [Google Scholar]
- Kuang, L.D.; Zheng, Y.C.; Lin, Y.Q.; Xu, Y.O.; Jin, S.Y.; Li, Y.P.; Dong, F.; Jiang, Z.R. Studies on high altitude adaptation of yak based on genetic variants and activity of lactate dehydrogenase-1. Biochem. Genet 2010, 48, 418–427. [Google Scholar]
- Zhang, L.; Ma, B.; Wu, J.; Fei, C.; Yang, L.; Wan, H. Cloning and characterization of the yak gene coding for calpastatin and in silico analysis of its putative product. Acta Biochim. Pol 2010, 57, 35–41. [Google Scholar]
- Bai, W.L.; Yin, R.H.; Zheng, Y.C.; Ma, Z.J.; Zhong, J.C.; Rin, R.L.; Dou, Q.L.; Zhang, S.C.; Luo, G.B.; Zhao, Z.H. Cloning and molecular characterization of a yak α-lactalbumin cDNA from mammary tissue. Livest. Sci 2010, 129, 122–128. [Google Scholar]
- Scheinfeldt, L.B.; Tishkoff, S.A. Living the high life: High-altitude adaptation. Genome Biol 2010. [Google Scholar] [CrossRef]
- Avivi, A.; Gerlach, F.; Joel, A.; Reuss, S.; Burmester, T.; Nevo, E.; Hankeln, T. Neuroglobin, cytoglobin, and myoglobin contribute to hypoxia adaptation of the subterranean mole rat Spalax. Proc. Natl. Acad. Sci. USA 2010, 107, 21570–21575. [Google Scholar]
- Vogt, M.; Puntschart, A.; Geiser, J.; Zuleger, C.; Billeter, R.; Hoppeler, H. Molecular adaptations in human skeletal muscle to endurance training under simulated hypoxic conditions. J. Appl. Physiol 2001, 91, 173–182. [Google Scholar]
- Natarajan, R.; Fisher, B.J.; Fowler, A.A., III. Regulation of hypoxia inducible factor-1 by nitric oxide in contrast to hypoxia in microvascular endothelium. FEBS Lett 2003, 549, 99–104. [Google Scholar]
- Beall, C.M. Two routes to functional adaptation: Tibetan and Andean high-altitude natives. Proc. Natl. Acad. Sci. USA 2007, 104, S8655–S8660. [Google Scholar]
- Gelfi, C.; de Palma, S.; Ripamonti, M.; Wait, R.; Eberini, I.; Bajracharya, A.; Marconi, C.; Schneider, A.; Hoppeler, H.; Cerretelli, P. New aspects of altitude adaptation in Tibetans: A proteomic approach. FASEB J 2004, 18, 612–614. [Google Scholar]
- Jurie, C.; Ortigues-Marty, I.; Picard, B.; Micol, D.; Hocquette, J.F. The separate effects of the nature of diet and grazing mobility on metabolic potential of muscles from Charolais steers. Livest. Sci 2006, 104, 182–192. [Google Scholar]
- Dietz, A.A.; Lubrano, T. Separation and quantitation of lactic dehydrogenase isoenzymes by disc electrophoresis. Anal. Biochem 1967, 20, 246–257. [Google Scholar]
- ExPASy SIB Bioinformatics Resource Portal. Available online: http://www.expasy.org (accessed on 25 July 2012).
- Amills, M.; Francino, O.; Jansa, M.; Sanchez, A. Isolation of genomic DNA from milk samples by using Chelex resin. J. Dairy Res 1997, 64, 231–238. [Google Scholar]
- Arnold, K.; Bordoli, L.; Kopp, J.; Schwede, T. The SWISS-MODEL Workspace: A web-based environment for protein structure homology modelling. Bioinformatics 2006, 22, 195–201. [Google Scholar]
- Kiefer, F.; Arnold, K.; Künzli, M.; Bordoli, L.; Schwede, T. The SWISS-MODEL repository and associated resources. Nucleic Acids Res 2009, 37, D387–D392. [Google Scholar]
| Species | ldhb | 53 | 204 | 407 | 689 | 700 | 896 |
|---|---|---|---|---|---|---|---|
| Yak | F | G/A | T/C | G/T | A | A/G | A |
| M | G/A | T/C | T | A | A/G | G | |
| S | G | C | T | C | A | G | |
| Cattle | G | T | T | A | A | G |
| Species | LDH1 Phenotype | Amino acid position | ||||
|---|---|---|---|---|---|---|
| 17 | 135 | 229 | 233 | 298 | ||
| Yak | F | Arg/Lys | Gly/Val | Glu | Val/Met | Gln |
| M | Arg/Lys | Val | Glu | Val/Met | Arg | |
| S | Arg | Val | Ala | Met | Arg | |
| Cattle | Arg | Val | Glu | Met | Arg | |
| Species | Phenotype | pI | Molecular weight (Da) | GenBank accession No. |
|---|---|---|---|---|
| Yak | F | 5.87 | 36564.39 | HQ874652 |
| M | 6.03 | 36592.45 | HQ874653 | |
| S | 6.21 | 36534.41 | HQ874654 | |
| Cattle | - | 6.03 | 36592.45 | BC102217 |
| Breed | n | Genotype frequency | Gene frequency | |||
|---|---|---|---|---|---|---|
| AA | AG | GG | A | G | ||
| Jiulong yak | 130 | 0.038 (5) | 0.446 (58) | 0.515 (67) | 0.262 | 0.738 |
| Zhongdian yak | 32 | 0 | 0.375 (12) | 0.625 (20) | 0.188 | 0.812 |
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, G.; Zhao, X.; Zhong, J.; Cao, M.; He, Q.; Liu, Z.; Lin, Y.; Xu, Y.; Zheng, Y. Cloning and Polymorphisms of Yak Lactate Dehydrogenase b Gene. Int. J. Mol. Sci. 2013, 14, 11994-12003. https://doi.org/10.3390/ijms140611994
Wang G, Zhao X, Zhong J, Cao M, He Q, Liu Z, Lin Y, Xu Y, Zheng Y. Cloning and Polymorphisms of Yak Lactate Dehydrogenase b Gene. International Journal of Molecular Sciences. 2013; 14(6):11994-12003. https://doi.org/10.3390/ijms140611994
Chicago/Turabian StyleWang, Guosheng, Xingbo Zhao, Juming Zhong, Meng Cao, Qinghua He, Zhengxin Liu, Yaqiu Lin, Yaou Xu, and Yucai Zheng. 2013. "Cloning and Polymorphisms of Yak Lactate Dehydrogenase b Gene" International Journal of Molecular Sciences 14, no. 6: 11994-12003. https://doi.org/10.3390/ijms140611994





