Connexin 26 is Down-Regulated by KDM5B in the Progression of Bladder Cancer
Abstract
:1. Introduction
2. Results and Discussion
2.1. The Expression Levels of Cx26 Were Inhibited by KDM5B
2.2. The Expression of KDM5B is Up-regulated and Cx26 is Down-Regulated in the Progression of Bladder Cancer
2.3. The Stringent Converse Relationship between Cx26 and KDM5B
3. Experimental Section
3.1. Cell Lines and Cell Culture
3.2. KDM5B Expression Constructs
3.3. ShRNA Constructs for KDM5B Gene Silencing
3.4. Transfection of T24 Cells, Western Blotting and Immunocytochemistry
3.5. Reverse Transcription Polymerase Chain Reaction (RT-PCR) and Real Time qPCR
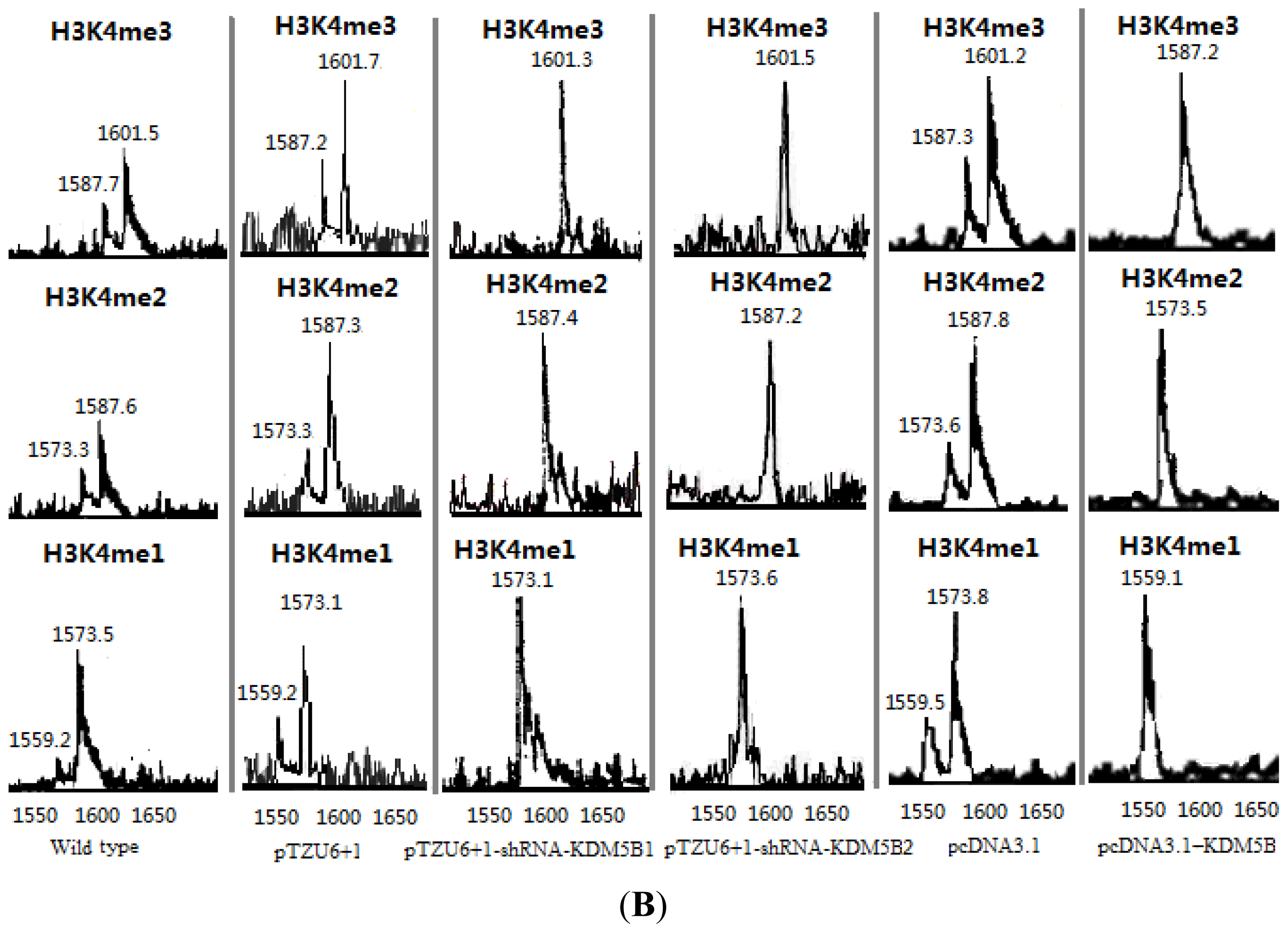
3.6. Demethylation Reactions and MALDI-TOF Mass Spectrometry
3.7. Statistical Analysis
4. Conclusions
Supplementary Information
ijms-14-07866-s001.pdfAcknowledgments
Conflict of Interest
References
- Parkin, D.M.; Pisani, P.; Ferlay, J. Global cancer statistics. CA Cancer J. Clin 2008, 49, 33–64. [Google Scholar]
- Kotake, T. Clinical problems in patients who were exposed to carcinogens. Hinyokika Kiyo 1989, 35, 2063–2069. [Google Scholar]
- Hou, J.; Wu, J.; Dombkowski, A.; Zhang, K.; Holowatyj, A.; Boerner, J.L.; Yang, Z.Q. Genomic amplification and a role in drug-resistance for the KDM5A histone demethylase in breast cancer. Am. J. Transl. Res 2012, 4, 247–256. [Google Scholar]
- Li, W.; Zhao, L.; Zang, W.; Liu, Z.; Chen, L.; Liu, T.; Xu, D.; Jia, J. Histone demethylase JMJD2B is required for tumor cell proliferation and survival and is overexpressed in gastric cancer. Biochem. Biophys. Res. Commun 2011, 416, 372–378. [Google Scholar]
- Pereira, F.; Barbachano, A.; Silva, J.; Bonilla, F.; Campbell, M.J.; Munoz, A.; Larriba, M.J. KDM6B/JMJD3 histone demethylase is induced by vitamin D and modulates its effects in colon cancer cells. Hum. Mol. Genet 2011, 20, 4655–4665. [Google Scholar]
- Xiang, Y.; Zhu, Z.; Han, G.; Ye, X.; Xu, B.; Peng, Z.; Ma, Y.; Yu, Y.; Lin, H.; Chen, A.P.; et al. JARID1B is a histone H3 lysine 4 demethylase up-regulated in prostate cancer. Proc. Natl. Acad. Sci. USA 2007, 104, 19226–19231. [Google Scholar]
- Kristensen, L.H.; Nielsen, A.L.; Helgstrand, C.; Lees, M.; Cloos, P.; Kastrup, J.S.; Helin, K.; Olsen, L.; Gajhede, M. Studies of H3K4me3 demethylation by KDM5B/Jarid1B/PLU1 reveals strong substrate recognition in vitro and identifies 2,4-pyridine-dicarboxylic acid as an in vitro and in cell inhibitor. FEBS J 2012, 279, 1905–1914. [Google Scholar]
- Hayami, S.; Yoshimatsu, M.; Veerakumarasivam, A.; Unoki, M.; Iwai, Y.; Tsunoda, T.; Field, H.I.; Kelly, J.D.; Neal, D.E.; Yamaue, H.; et al. Overexpression of the JmjC histone demethylase KDM5B in human carcinogenesis: Involvement in the proliferation of cancer cells through the E2F/RB pathway. Mol. Cancer 2010, 9, 59–72. [Google Scholar]
- Catchpole, S.; Spencer-Dene, B.; Hall, D.; Santangelo, S.; Rosewell, I.; Guenatri, M.; Beatson, R.; Scibetta, A.G.; Burchell, J.M.; Taylor-Papadimitriou, J. PLU-1/JARID1B/KDM5B is required for embryonic survival and contributes to cell proliferation in the mammary gland and in ER+ breast cancer cells. Int. J. Oncol 2011, 38, 1267–1277. [Google Scholar]
- Goodenough, D.A.; Goliger, J.A.; Paul, D.L. Connexins, connexons, and intercellular communication. Annu. Rev. Biochem 1996, 65, 475–502. [Google Scholar]
- Klein, G. Cancer, apoptosis, and nonimmune surveillance. Cell Death Differ 2004, 11, 13–17. [Google Scholar]
- Trosko, J.E.; Ruch, R.J. Cell-cell communication and carcinogenesis. Front. Biosci 1998, 3, 208–236. [Google Scholar]
- Grossman, H.B.; Liebert, M.; Lee, I.W.; Lee, S.W. Decreased connexin expression and intercellular communication in human bladder cancer cells. Cancer Res 1994, 54, 3062–3065. [Google Scholar]
- Sun, Y.; Zhao, X.; Yao, Y.; Qi, X.; Yuan, Y.; Hu, Y. Connexin 43 interacts with Bax to regulate apoptosis of pancreatic cancer through a gap junction-independent pathway. Int. J. Oncol 2012, 41, 941–948. [Google Scholar]
- Li, Z.; Zhou, Z.; Welch, D.R.; Donahue, H.J. Expressing connexin 43 in breast cancer cells reduces their metastasis to lungs. Clin. Exp. Metastasis 2008, 25, 893–901. [Google Scholar]
- Ezumi, K.; Yamamoto, H.; Murata, K.; Higashiyama, M.; Damdinsuren, B.; Nakamura, Y.; Kyo, N.; Okami, J.; Ngan, C.Y.; Takemasa, I.; et al. Aberrant expression of connexin 26 is associated with lung metastasis of colorectal cancer. Clin. Cancer Res 2008, 14, 677–684. [Google Scholar]
- Ozawa, H.; Matsunaga, T.; Kamiya, K.; Tokumaru, Y.; Fujii, M.; Tomita, T.; Ogawa, K. Decreased expression of connexin-30 and aberrant expression of connexin-26 in human head and neck cancer. Anticancer Res 2007, 27, 2189–2195. [Google Scholar]
- Chen, Y.; Huhn, D.; Knosel, T.; Pacyna-Gengelbach, M.; Deutschmann, N.; Petersen, I. Downregulation of connexin 26 in human lung cancer is related to promoter methylation. Int. J. Cancer 2005, 113, 14–21. [Google Scholar]
- Jamieson, S.; Going, J.J.; D’Arcy, R.; George, W.D. Expression of gap junction proteins connexin 26 and connexin 43 in normal human breast and in breast tumours. J. Pathol 1998, 184, 37–43. [Google Scholar]
- Gee, J.; Tanaka, M.; Grossman, H.B. Connexin 26 is abnormally expressed in bladder cancer. J. Urol 2003, 169, 1135–1137. [Google Scholar]
- Tanaka, M.; Grossman, H.B. Connexin 26 gene therapy of human bladder cancer: Induction of growth suppression, apoptosis, and synergy with Cisplatin. Hum. Gene Ther 2001, 12, 2225–2236. [Google Scholar]
- Tanaka, M.; Fraizer, G.C.; de La Cerda, J.; Cristiano, R.J.; Liebert, M.; Grossman, H.B. Connexin 26 enhances the bystander effect in HSVtk/GCV gene therapy for human bladder cancer by adenovirus/PLL/DNA gene delivery. Gene Ther 2001, 8, 139–148. [Google Scholar]
- Chen, S.; Ma, J.; Wu, F.; Xiong, L.J.; Ma, H.; Xu, W.; Lv, R.; Li, X.; Villen, J.; Gygi, S.P.; et al. The histone H3 Lys 27 demethylase JMJD3 regulates gene expression by impacting transcriptional elongation. Genes Dev 2012, 26, 1364–1375. [Google Scholar]
- Liang, C.Y.; Hsu, P.H.; Chou, D.F.; Pan, C.Y.; Wang, L.C.; Huang, W.C.; Tsai, M.D.; Lo, W.S. The histone H3K36 demethylase Rph1/KDM4 regulates the expression of the photoreactivation gene PHR1. Nucleic Acids Res 2011, 39, 4151–4165. [Google Scholar]
- Liu, Z.; Zhou, S.; Liao, L.; Chen, X.; Meistrich, M.; Xu, J. Jmjd1a demethylase-regulated histone modification is essential for cAMP-response element modulator-regulated gene expression and spermatogenesis. J. Biol. Chem 2010, 285, 2758–2770. [Google Scholar]
- Berger, S.L. The complex language of chromatin regulation during transcription. Nature 2007, 447, 407–412. [Google Scholar]
- Kouzarides, T. Chromatin modifications and their function. Cell 2007, 128, 693–705. [Google Scholar]
- Lan, F.; Collins, R.E.; de Cegli, R.; Alpatov, R.; Horton, J.R.; Shi, X.; Gozani, O.; Cheng, X.; Shi, Y. Recognition of unmethylated histone H3 lysine 4 links BHC80 to LSD1-mediated gene repression. Nature 2007, 448, 718–722. [Google Scholar]
- Burchardt, M.; Burchardt, T.; Shabsigh, A.; de La Taille, A.; Benson, M.C.; Sawczuk, I. Current concepts in biomarker technology for bladder cancers. Clin. Chem 2000, 46, 595–605. [Google Scholar]
- Datta, A.; Adelson, M.E.; Mogilevkin, Y.; Mordechai, E.; Sidi, A.A.; Trama, J.P. Oncoprotein DEK as a tissue and urinary biomarker for bladder cancer. BMC Cancer 2011, 11, 234–240. [Google Scholar]
- Johnen, G.; Gawrych, K.; Bontrup, H.; Pesch, B.; Taeger, D.; Banek, S.; Kluckert, M.; Wellhausser, H.; Eberle, F.; Nasterlack, M.; et al. Performance of survivin mRNA as a biomarker for bladder cancer in the prospective study UroScreen. PLoS ONE 2012, 7, e35363. [Google Scholar]
- Li, F.; Chen, D.N.; He, C.W.; Zhou, Y.; Olkkonen, V.M.; He, N.; Chen, W.; Wan, P.; Chen, S.S.; Zhu, Y.T.; et al. Identification of urinary Gc-globulin as a novel biomarker for bladder cancer by two-dimensional fluorescent differential gel electrophoresis (2D-DIGE). J. Proteomics 2012, 77, 225–236. [Google Scholar]
- Urquidi, V.; Chang, M.; Dai, Y.; Kim, J.; Wolfson, E.D.; Goodison, S.; Rosser, C.J. IL-8 as a urinary biomarker for the detection of bladder cancer. BMC Urol. 2012, 12. [Google Scholar] [CrossRef]
- Klose, R.J.; Zhang, Y. Regulation of histone methylation by demethylimination and demethylation. Nat. Rev. Mol. Cell Biol 2007, 8, 307–318. [Google Scholar]
- Shin, S.; Janknecht, R. Diversity within the JMJD2 histone demethylase family. Biochem. Biophys. Res. Commun 2007, 353, 973–977. [Google Scholar]
- Tan, H.; Wu, S.; Wang, J.; Zhao, Z.K. The JMJD2 members of histone demethylase revisited. Mol. Biol. Rep 2008, 35, 551–556. [Google Scholar]
- Cruciani, V.; Mikalsen, S.O. The vertebrate connexin family. Cell. Mol. Life Sci 2006, 63, 1125–1140. [Google Scholar]
- Free Statistics and Forecasting Software. Spearman Rank Correlation—Free Statistics Software (Calculator). Available online: http://www.wessa.net/rwasp_spearman.wasp/ (accessed on 20 February 2013).




© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, X.; Su, Y.; Pan, J.; Zhou, Z.; Song, B.; Xiong, E.; Chen, Z. Connexin 26 is Down-Regulated by KDM5B in the Progression of Bladder Cancer. Int. J. Mol. Sci. 2013, 14, 7866-7879. https://doi.org/10.3390/ijms14047866
Li X, Su Y, Pan J, Zhou Z, Song B, Xiong E, Chen Z. Connexin 26 is Down-Regulated by KDM5B in the Progression of Bladder Cancer. International Journal of Molecular Sciences. 2013; 14(4):7866-7879. https://doi.org/10.3390/ijms14047866
Chicago/Turabian StyleLi, Xin, Yongping Su, Jinhong Pan, Zhansong Zhou, Bo Song, Enqing Xiong, and Zhiwen Chen. 2013. "Connexin 26 is Down-Regulated by KDM5B in the Progression of Bladder Cancer" International Journal of Molecular Sciences 14, no. 4: 7866-7879. https://doi.org/10.3390/ijms14047866



