2.1. Discovery of MICALs
During the search for CasL SH3 domain interactors among proteins recombinantly expressed from a thymus cDNA library, Suzuki
et al. [
1] identified a novel cytoplasmic protein, which they named MICAL1 from the Molecule Interacting with CasL. CasL (also known as HEF1 and NEDD9) belongs to the Cas family of proteins. It is a docking protein important for several functions among which are T cell receptor and β1 integrin-induced responses such as interleukin-2 production and migratory response, which all involve cytoskeleton rearrangements. Sequence analysis of the cDNA encoding MICAL revealed that it is a multidomain protein coupling an
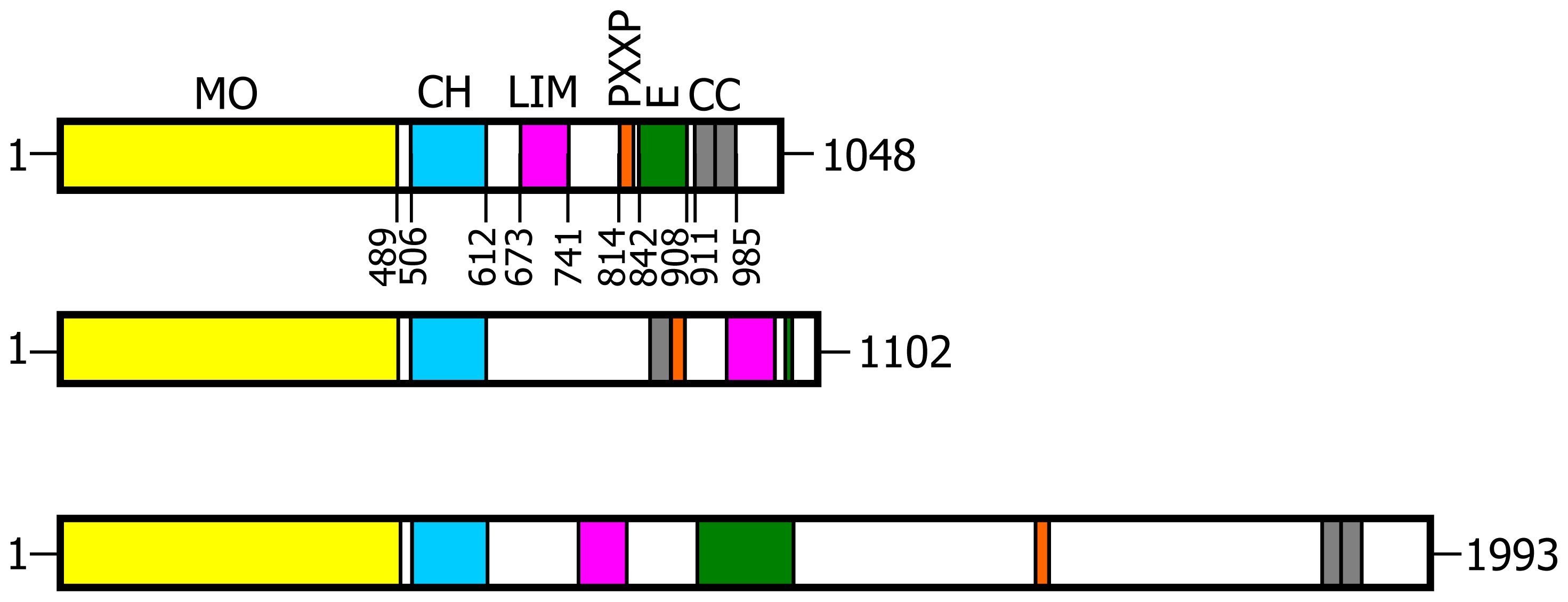
N-terminal flavoprotein-like domain to regions known to mediate protein-protein interactions, namely: a calponin homology (CH) domain typical of actin binding proteins; a LIM domain (from the three gene products Lin-11, Isl-1 and Mec-3, [
17,
18]), which contains two zinc fingers; a Pro-rich region with the PXKP signature for the interaction with SH3 domains; a
C-terminal region with motifs suggesting the formation of coiled-coil structures (
Figure 1). Co-localization and co-immunoprecipitation experiments, following expression of various MICAL1 constructs in the cells, demonstrated that MICAL1 interacts with CasL through its Pro-rich region [
1]. MICAL1 was also found to co-localize with vimentin, the main component of intermediate filaments, and to interact with it with its
C-terminal region [
1]. Although the interaction with vimentin still awaits independent confirmation [
8,
19], these observations led to the proposal that MICAL1 participates in the cascade of events downstream of CasL phosphorylation in response to β1 integrin and/or T cell receptor stimulation, providing a physical link with cytoskeleton proteins [
1]. Independent studies in the Kolodkin group on the intracellular components that transduce the signal initiated by the interaction of semaphorins with their plexin receptors on neurons led to identify MICAL1 as a novel interactor of PlexA cytoplasmic domain in an embryonic Drosophila cDNA library [
2]. While in Drosophila there is only one gene encoding MICAL, vertebrates contain three genes encoding MICAL isoforms indicated as MICAL1, MICAL2 and MICAL3 (
Figure 1) [
1,
2,
20]. Furthermore, MICAL-like forms, also encoded by different genes, have been identified [
1,
2,
21,
22], but they will not be discussed further due to the absence of the flavoprotein region. Genetic studies showed that inactivation of the single MICAL gene of Drosophila leads to phenotypes similar to those obtained by inactivating either the semaphorin 3A (Sema3A) or plexin A (PlexA) [
2]. By introducing the rather drastic triple G-to-W substitution in the GXGXXG motif in the unique putative adenylate-binding site of MICAL flavoprotein-like
N-terminal domain, it was also concluded that the integrity of such domain is required for MICAL function [
2]. This conclusion was independently confirmed by Beuchle
et al. [
23] who correlated point mutations in the Drosophila MICAL gene, all causing a milchstrasse phenotype, with alterations of myofilaments organization and synaptic structure. Evidence in support of the essentiality of the
N-terminal region for MICAL function was later obtained also in mammalian neuronal and non-neuronal cells [
24,
25].
Overall, these studies demonstrated a role of MICAL in transducing the signal initiated by the interaction of semaphorins with their plexin receptors on the target cells, which leads to local (actin) cytoskeleton disassembly, as exemplified by axon growth cone collapse [
2]. They also provided the first evidence that the
N-terminal putative flavoprotein domain may be essential for MICAL function [
2,
23]. The latter is not restricted to neuronal cells as demonstrated by, e.g., the original work of Suzuki
et al. in T cells [
1], that of Beuchle
et al. [
23] in Drosophila somatic muscle cells, of Fisher
et al. [
19] in rat and by the more recent experiments of Giridharan
et al. [
25] in non-neuronal mammalian cells.
The recent review of Hung
et al. [
9] provides an excellent historical overview on the discovery of semaphorins as chemorepellent protein factors capable to induce the collapse of the axon growth cone through local disassembly of actin filaments and, therefore, to control the direction of axon growth towards its target cell. The review also nicely describes the experimental approaches that led to the identification and characterization of several proteins participating in the transduction of semaphorin signaling. With other recent articles [
8,
16], it also summarizes the evidence for the role, distribution and expression patterns of MICAL forms as gathered through genetic and cell biology studies on different cell types. The original discovery of chemorepellent and chemoattractive cues stimulated a variety of studies aiming to understand how cytoskeleton dynamics is controlled, at the molecular level, in different cells. Such studies led to the discovery of a large number of proteins, including GTPases, GTPase-activating proteins and GTP-binding proteins, the microtubule regulators tau and the collapsin response mediator proteins (CRMP), several actin polymerizing/depolymerizing and bundling factors and, as a novel member, MICALs. However, in spite of the fact that a great deal of work has already been done, how the dynamics of the cytoskeleton is controlled in the cell is far from being understood. The growing number of interconnected signaling pathways and of the (macro)molecules participating in the process is actually demonstrating the complexity of the control of cytoskeleton dynamics. For example, although the role and mechanism of action of semaphorins has been initially studied in neurons, it is now clear that they participate in the control of the cytoskeleton in a broad variety of cells, through their receptors and downstream effectors. As a result, understanding the molecular details of the semaphorin-initiated signaling pathway is also of interest in order to design ways to modulate their action in the context of neurodegenerative diseases, of cell metastatization and migration, and of angiogenesis as part of cancer therapy, of pathogen infection or to promote axon regeneration in, e.g., spinal cord injury [
7–
9,
11,
12,
14,
26–
37]. In this respect, understanding MICAL mechanism of action is important in that it may provide key information for the understanding of the semaphorin pathway as well as tools to control it [
11,
14,
26,
31,
37]. In support of biomedically relevant outcomes of studies on MICAL are several observations. The expression of MICAL forms increases at sites of spinal cord injury in model animals, thus supporting the hypothesis that inhibiting its function may be beneficial to promote neuronal regeneration [
38]. MICAL is also involved in myofilaments organization and synaptic structure [
23,
25] suggesting that modulation of MICAL activity may be of interest also in the context of neuromuscular disorders, including amyotrophic lateral sclerosis [
31]. A link between MICAL function and vesicle trafficking between the endoplasmic reticulum and the Golgi complex was also found [
19,
39]. Furthermore, the overexpression of MICAL2 alternative splicing variants associates with particularly aggressive prostate cancers forms [
37]. Lowering their levels through small interfering RNA technology significantly decreases cell growth [
37]. Finally, the work of Xue
et al. [
40] on zebrafish suggests that MICAL forms may also be important for cardiovascular development.
2.2. Hypotheses on MICAL Function
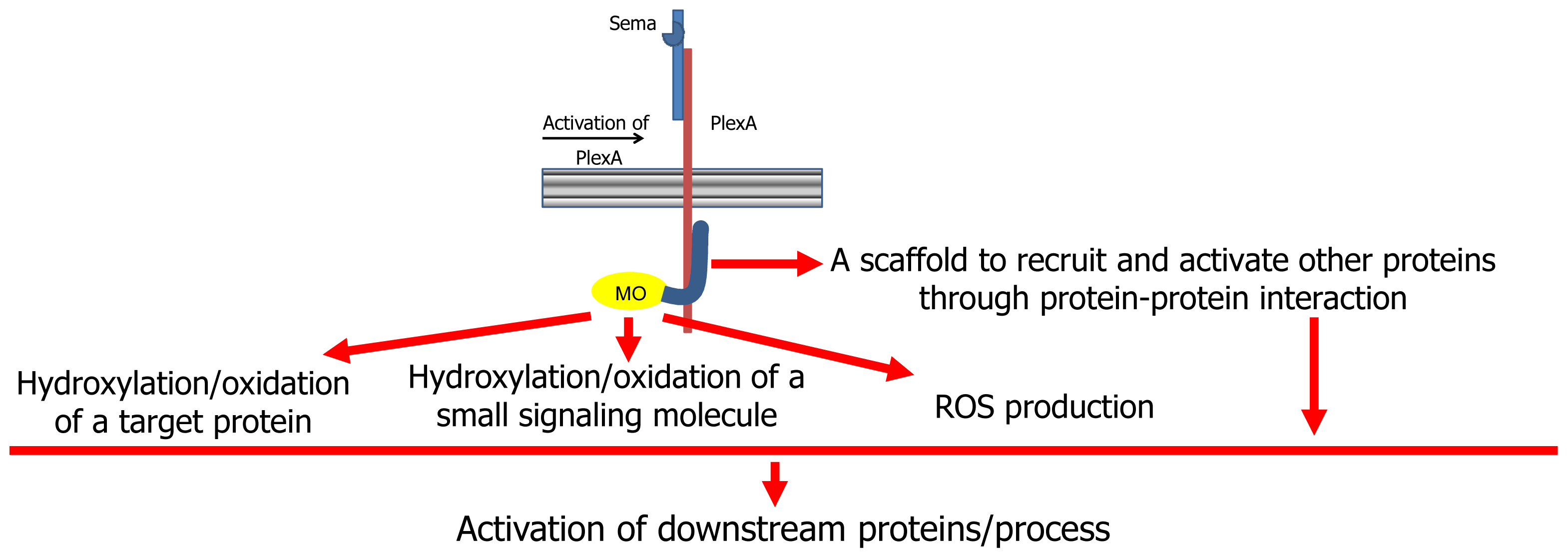
On the basis of the body of current knowledge on MICALs, several hypotheses on its mode of action have been made (
Figure 2, [
7,
9,
13]). According to a first hypothesis, MICAL may simply act as a scaffold protein. When the semaphorin (Sema1A in Drosophila, Sema3A in mammals) interacts with its plexin A receptor directly (as in Drosophila) or through the neuropilin coreceptors (in mammals), several proteins are recruited to the (activated) plexin cytoplasmic domain. Among them is MICAL, which interacts with plexin through its
C-terminal region (
Figure 2, [
2]).
This scaffold hypothesis is supported by the presence in MICAL of protein interaction domains (
Figure 1) and by the observation that MICAL1 binds CasL with its Pro-rich region [
1], with the rab 1 GTPase with its last ~145 residues [
41] and with CRMP forms with its
N-terminal region including the flavoprotein, CH and LIM domains [
24]. Thus, following semaphorin-plexin interaction, MICAL binds to the cytoplasmic region of plexin where it helps recruiting other proteins and promoting the modulation of their activity with respect to the downstream events (
Figure 2).
The scaffold model would explain how MICAL may interfere with CasL [
1,
8,
13] or CRMP [
16,
24] functions by sequestering them. In support of a scaffold role is the finding that MICAL1 modulates apoptosis by binding NDR1/2 (nuclear Dbf2-related) kinases, through its part spanning from the LIM domain to the
C-terminus [
42]. By competing with MST1 for NDR binding, MICAL prevents mammalian Ste-20-like kinase (MST1)-induced NDR activation [
42]. Finally, there are several MICAL-like proteins lacking the
N-terminal flavoprotein-like region, which also participate in functions linked to cytoskeleton dynamics such as endosomal recycling and establishment of cell-cell contacts [
22,
43–
51].
The presence of the
N-terminal region with sequence similarity to several flavoproteins of the oxidase and monooxygenase (hydroxylase) class, which appeared to be essential for MICAL function [
2], led to alternative hypotheses linking MICAL function to the catalytic activity of its
N-terminal flavoprotein domain ([
13],
Figure 2).
This hypothesis is supported by several experiments that showed that the
N-terminal flavoprotein-like domain of MICAL is essential for its function, while the other domains are important for its localization [
2,
9,
24,
25,
52,
53]. The binding of MICAL to plexin cytosolic region (and/or further interaction with other proteins) would activate MICAL’s enzymatic function.
According to one hypothesis, MICAL flavoprotein domain would catalyze an oxidase reaction by mediating the transfer of reducing equivalents from a small molecule to molecular oxygen with the release of hydrogen peroxide (or superoxide anion) as the signaling molecule responsible of the downstream events [
7,
9,
13,
16].
In support of this hypothesis is the finding that CRMP2 may be oxidized in response to semaphorin stimulation leading to formation of a homodimer stabilized by a disulfide bridge [
54]. Oxidized CRMP2 may form a transient disulfide-linked complex with thioredoxin, which would stimulate CRMP2 inactivation through phosphorylation by glycogen synthase kinase-3, leading to growth cone collapse [
54].
As discussed below, the MICAL flavoprotein-like domain actually exhibits a NAD(P)H oxidase activity, which would be consistent with the hypothesis. As an alternative, the small molecule being oxidized by MICAL would be the actual effector, but no evidence in support of this hypothesis is available. Finally, MICAL could covalently modify, through oxidation or hydroxylation, a protein side chain, leading to a modulation of its function. As discussed below, on the basis of the structure of MICAL’s
N-terminal region [
3,
4] and of experimental evidence, actin [
52,
53] and the collapsin responsive mediator protein-1 (CRMP1) [
24] may be good candidates as the protein substrates.
Furthermore, the similarity of the
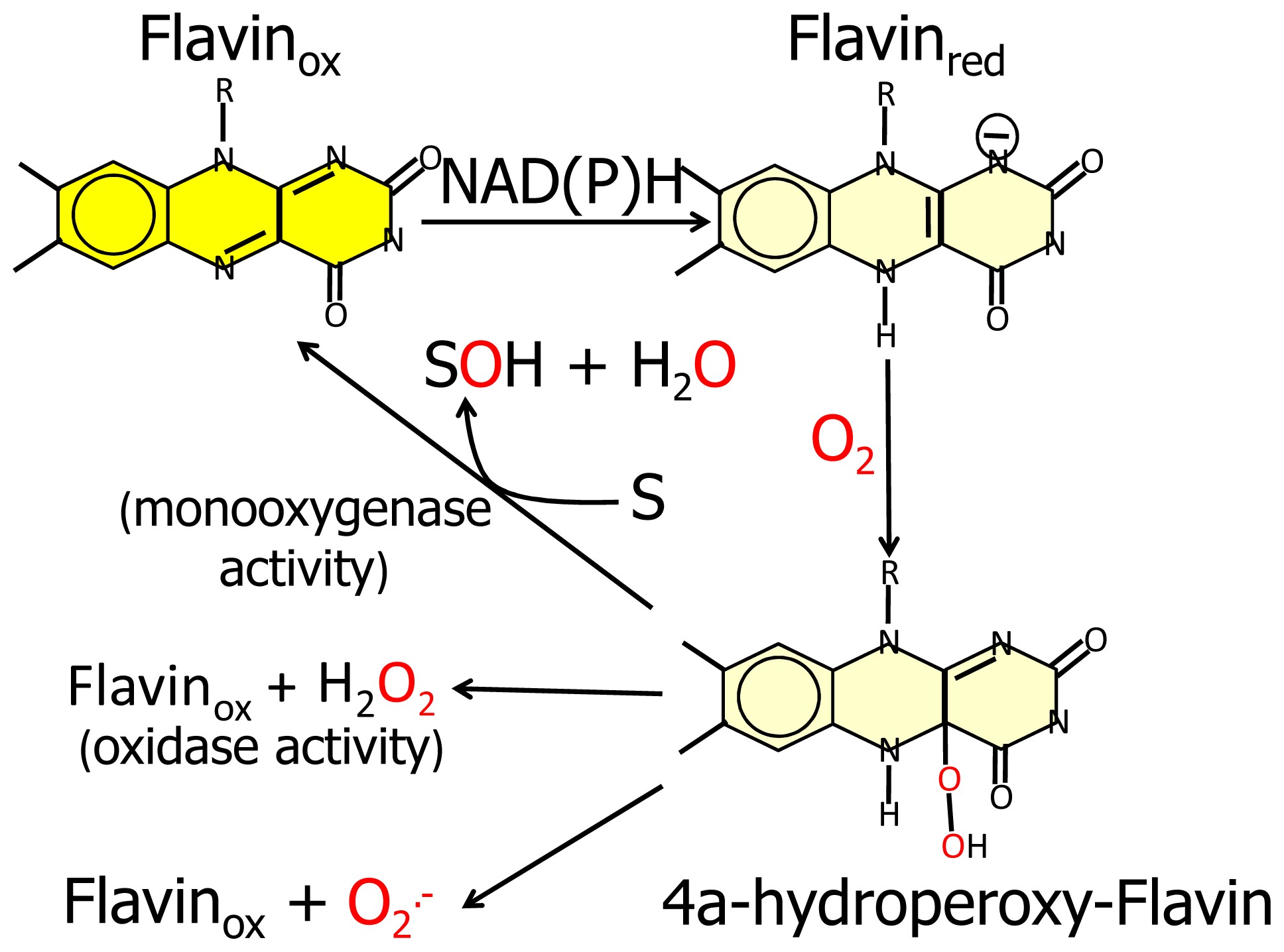
N-terminal protein region with FAD-dependent monooxygenases would suggest that MICAL catalyzes a hydroxylation rather than an oxidase reaction. In this case no reactive oxygen species would be released since it is well known that FAD containing monooxgenases/hydroxylases insert one oxygen atom from molecular oxygen in the hydroxylatable substrate and release the second oxygen atom in a water molecule (
Figure 3, [
5,
6,
55]). In this reaction, reducing equivalents would be provided by reduced pyridine nucleotides, NADH or NADPH, possibly linking the control of cytoskeleton dynamics to the primary cell energy metabolism and/or signaling pathways involving NAD derivatives like, e.g., polyADP ribose [
56].
Should MICAL modify a protein side-chain, one of the relevant biological issues would be the reversibility of the modification. Cysteine oxidation leading to formation of disulfides may be reversed by systems like that formed by the thioredoxin/thioredoxin reductase couple [
57], while methionine oxidation may be reversed by methionine sulfoxide reductases [
58]. Modification of other side chains may require novel enzymatic activities, which would lead to other exciting discoveries.
Since FAD-dependent monooxygenases can catalyze a NAD(P)H oxidase reaction in the absence of the substrate to be hydroxylated (
Figure 3), it cannot be ruled out that MICALs may differentially control the oxidase and the monooxygenase activities depending on the MICAL isoform and/or the presence of interacting (macro)molecules.
Whether MICAL becomes catalytically active only when it is bound to plexin, or other proteins determine its activation still needs to be established. The work of Schmidt
et al. [
24], later confirmed by Giridharan
et al. [
25], supported the hypothesis that the activity of the
N-terminal catalytic domain is autoinhibited by the region
C-terminal to the LIM domain (see below). Binding of plexin A and other interactors to such a
C-terminal region would remove the inhibition activating MICAL reaction. Furthermore, removal of the physical interaction between the
N-terminal and the
C-terminal regions may allow binding of the actual (protein) substrate to the
N-terminal catalytic flavoprotein domain. Substrate binding may be assisted by the adjacent domains, e.g., the CH and LIM domains.
An important contribution to the understanding of MICAL and, therefore, of semaphorin signaling and of the control of cytoskeleton dynamics may be given by in depth in vitro biochemical studies of MICAL forms. These studies are still limited, and will be reviewed in this article.
2.4. Analysis of MICAL Primary Structure
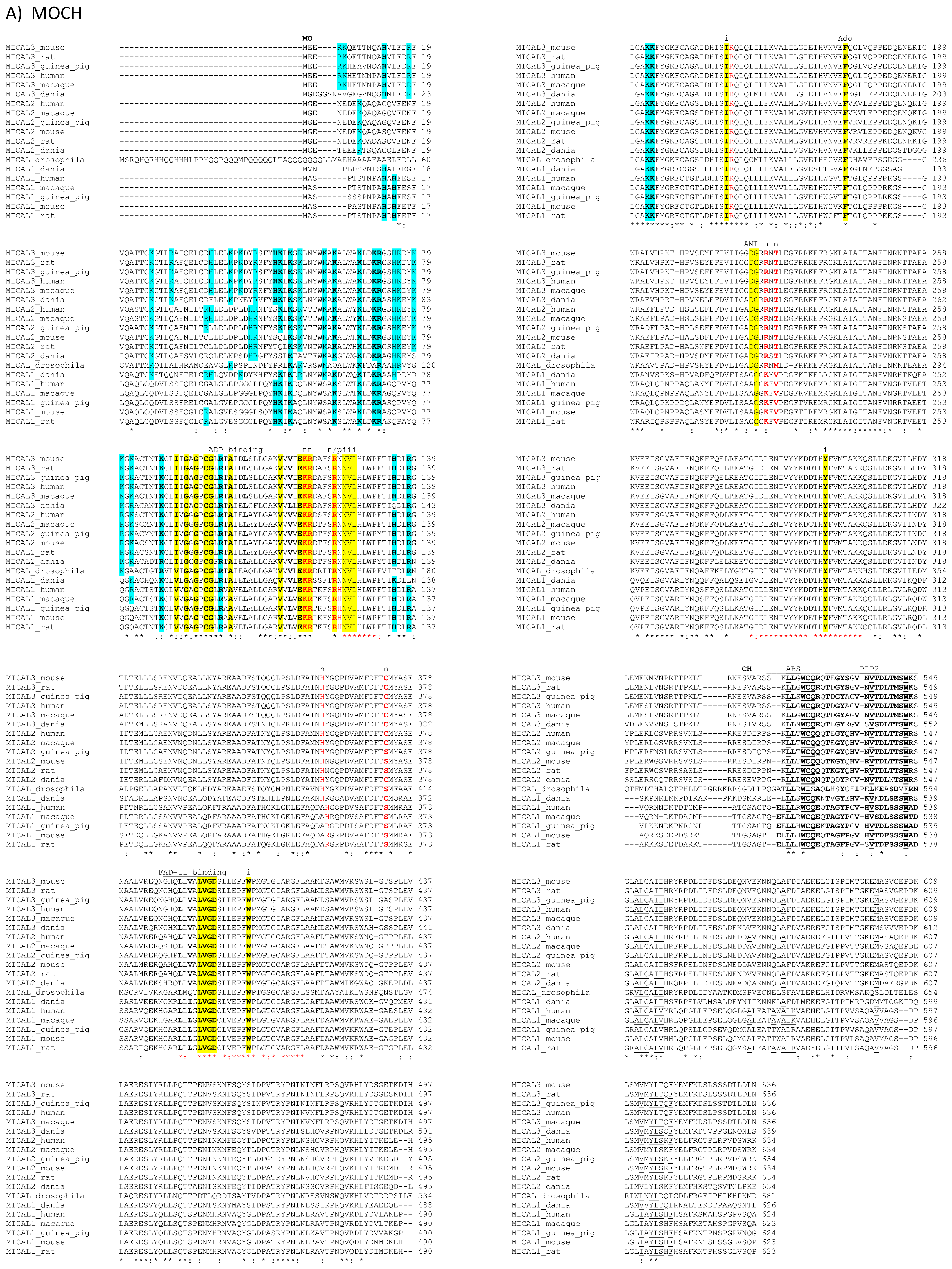
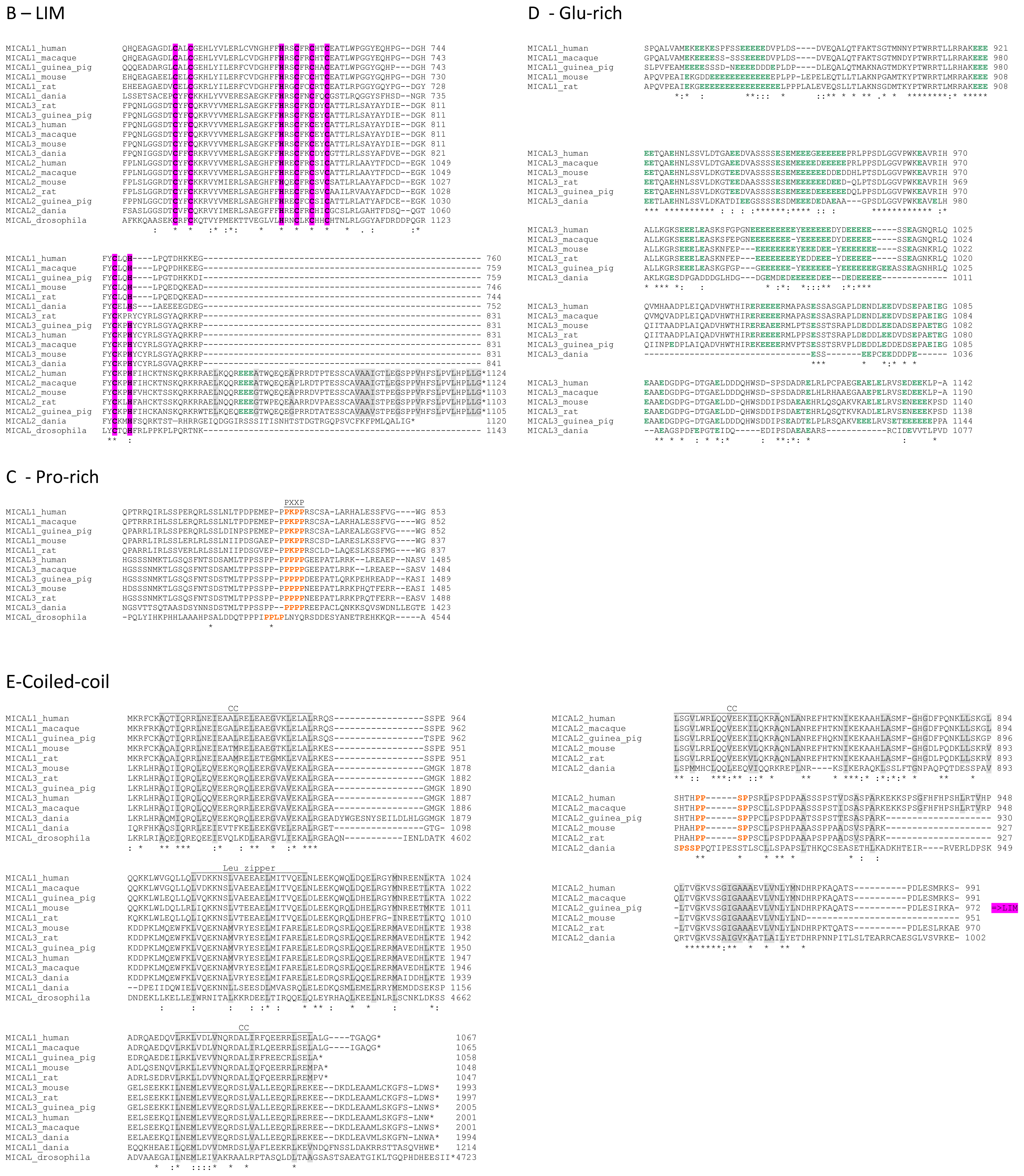
The primary structures of representative MICAL forms are compared in
Figure 4, which has also been annotated on the basis of information deriving from structural studies [
3,
4,
65]. In the following text and in
Figure 4 the residue numbering is given for mouse MICAL1, for which high resolution structural models are available [
3,
4].
In the
N-terminal part of the protein the presence of a flavoprotein domain was initially hypothesized on the basis of the presence of regions matching the consensus sequence for the formation of a Rossman fold for the binding of the ADP moiety of FAD or pyridine nucleotides (residues 87–114, [
62]) and of the so-called second FAD consensus sequence identified by Eggink
et al. (residues 383–393,
Figure 4, [
63,
69]).
An overall sequence similarity with proteins of the
p-hydroxybenzoate hydroxylase (PHBH) structural family ([
5,
6] and references therein) was found beyond the first ≈80 residues of MICALs leading to the hypothesis of the flavoprotein nature of its ≈500 residues
N-terminal domain (
Figures 4 and
5). This hypothesis was later confirmed by the determination of the three dimensional structure of mouse MICAL1
N-terminal region [
3,
4] and by
in vitro studies on the corresponding region of mouse [
4] and human [
70] MICAL1.
In the N-terminal segment (~80 residues), some residues are conserved in all MICALs. Other residues are only conserved within each MICAL 1–3 class and may be diagnostic for the classification of MICAL forms. Drosophila MICAL contains an ≈40 residues N-terminal extension with respect to mammalian MICALs. However, it is not a distinctive feature of insects MICALs, for which most of the sequences in databanks are not complete.
The flavoprotein-like domain is followed by a calponin homology (CH) domain (residues 506–612,
Figures 1 and
4, [
65]). Sequence (
Figure 4) and structural analysis [
65] identified it as a type 2 CH domain (see below). The LIM domain (residues 673–741), which forms zinc finger structures [
17,
18], is present in one copy in all MICAL forms with the conserved Cys and His residues implicated in zinc ions coordination. However, in MICAL2 the region separating it from the CH domain is longer than in MICAL1 and 3 (
Figures 1 and
4). Furthermore, it is between a possible coiled-coil motif and the
C-terminus, which may also form a coiled-coil structure. In contrast, such coiled-coil motifs are only
C-terminal to the LIM domain in MICAL1 and 3.
In the
C-terminal region, a Pro-rich region matching the PXXP motif typical of proteins that interact with SH3 domains is found in MICAL1 [
1] and MICAL3 only (
Figure 4C). However, a Pro-rich region possibly matching the PXXP consensus sequence is actually found also in MICAL2, but in the region containing the putative coiled-coil motifs (
Figure 4E). MICAL1 and 3 exhibit Glu (and Asp)-rich regions of different length (
Figure 4D). They may also contribute to protein-protein interactions, but their precise role in MICALs is not known. In MICAL2, a Glu-rich region is found in the
C-terminal protein segment following the LIM domain, but it is very short and is unlikely to play a specific role (
Figure 4B).
The final part of MICAL1 and 3 may form coiled-coil structures similar to that found in the α domain of proteins of the Ezrin, Radizin and Moesin (ERM) family as pointed out by Terman
et al. [
2]. The number and distribution of heptad repeats vary among MICALs (
Figure 4E). In MICAL1, one of the coiled-coil motifs (residues 964–986) corresponds to the leucine zipper identified by Suzuki
et al. [
1].
The last four residues of MICAL have been suggested to match the consensus sequence (S/TXV) for the formation of a PDZ binding motif [
2,
23], but comparison of multiple sequences does not allow us to confirm the hypothesis (
Figure 4E).
2.5. High Resolution Structure of MICAL Forms
A breakthrough in the understanding of the possible function of MICAL was the determination of the high resolution structure of the
N-terminal flavoprotein-like region of mouse MICAL1 by two groups [
3,
4]. Confirmation of the presence of a CH and of a LIM domain in MICALs was obtained by nuclear magnetic resonance (NMR, see below). The 1–489 region of mouse MICAL1 was produced in
E. coli cells in fusion with an
N-terminal His
6-tag [
3,
4]. Well-diffracting crystals were obtained by Siebold
et al. [
3] using such a protein form, while the Nadella
et al. [
4] crystals were obtained using an engineered protein form, which carried the Lys-to-Ala double substitution of surface residues 141 and 142. As expected, the structures of the as-isolated (oxidized) proteins (2 Å resolution, PDB ID 2BRA for [
4]; 1.45 Å resolution, 2BRY for [
3]) are similar to each other.
A homodimer was found in the crystallographic asymmetric unit, although the protein is monomeric in solution. Only a few disordered segments are present so that the details of the structure can be examined. Each monomer contains one FAD coenzyme and shows an N-terminal four-helix bundle region (residues 1–85) followed by a core region (residues 86–442) and a C-terminal segment (residues 443–489).
The core region exhibits the PHBH fold, which is typical of several FAD-containing oxidases and monooxygenases. In particular the highest structural similarity is with PHBH (1PBE) and phenol hydroxylase (1PN0). Thus, the core of MICAL flavoprotein domain is bilobal in shape. The larger portion contains the FAD binding region and is formed by residues 86–234 and 367–444 (the boundaries of domains are as described in [
3] and are, as expected, similar to those described in [
4]). This region should also contain the NAD(P) binding site of MICAL by analogy with the enzymes of the PHBH class (see below). The region between residues 235 and 366 forms the smaller lobe of the protein, which completes the active site. This 235–366 region was defined as the monooxygenase domain by Siebold
et al. [
3], but we will indicate it as the substrate binding domain in order to adopt the nomenclature used for PHBH [
68,
71].
The flavoprotein domain of mouse MICAL1 shows a simplified topology with respect to PHBH, which may allow for a degree of flexibility higher than in PHBH. In PHBH the polypeptide chain passes several times between the FAD and the substrate binding domains. In MICAL, the FAD binding region and the substrate binding domain are only connected by a long two-stranded β sheet formed by β strands β9 and β15 (
Figures 5 and
6). Such a difference is due to the absence, in MICAL, of regions corresponding to PHBH β5 and β6 strands and α4 helix (
Figure 5), which are replaced by α-helix 11 of MICAL’s substrate binding domain.
Both the N-terminal 1–85 region and the C-terminal 442–489 segment are ordered and make extensive contacts with the core region. Not surprisingly, the C-terminal 443–489 region of MICAL, which connects the flavoprotein domain to the CH domain, has no similarity with PHBH C-terminal segment, which forms the interface domain stabilizing the functional homodimer.
The
N-terminal 1–85 region, which has no counterpart in PHBH, contains several basic residues. Some of them are conserved across all MICALs or are found in most MICAL sequences (H11, H13, H49, K50, K52, K61, K66, K(or R) 69, R(or K)70, K(or R)86,
Figures 4 and
7). These residues, as well as several conserved residues in the initial part of the FAD domain (R98, K142, K141, R136, H133,
Figures 4 and
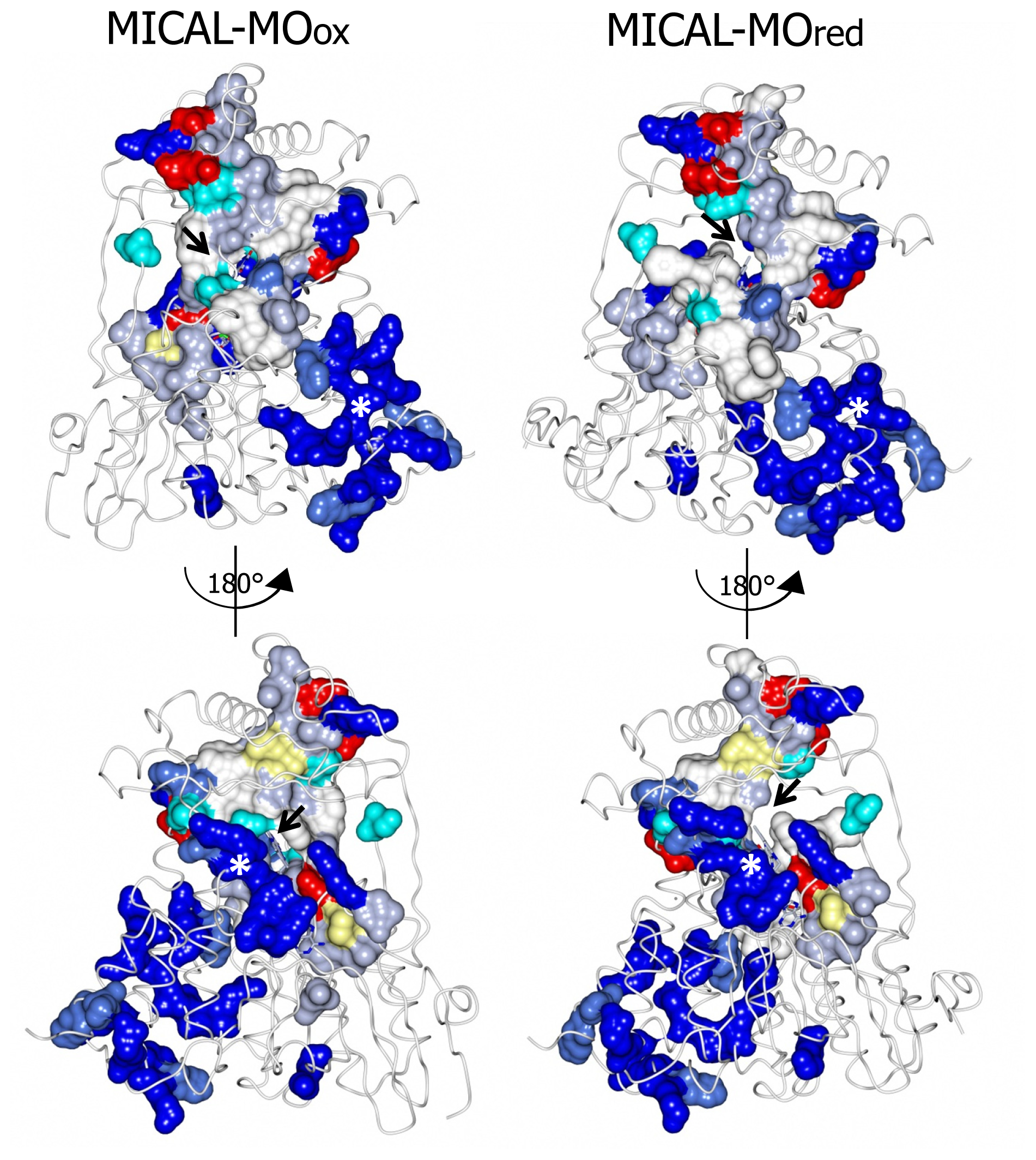
7), contribute to different extents to the formation of one extended patch of basic potential found on one face of the protein (
Figures 7 and
8). Such a positively charged region is on the same side of the protein as several strictly conserved segments of MICALs (residues 121–130, 278–300, 390–411), which surround the FAD isoalloxazine ring and may contribute to the definition of MICAL (protein) substrate binding region and catalytic site. Due to its location and to the acidity of several cytoskeletal proteins, including actin, the
N-terminal 1–85 basic region of MICAL has been proposed to be important to promote the interaction with the putative protein substrate [
3,
4].
A second patch of basic potential is found on the opposite face of the protein (
Figures 7 and
8; [
3,
4]). It is formed by the MICAL’s conserved residues K115, R116, R121, H122, R158, K221, R356 (
Figure 4). Sequence (
Figure 5) and structural comparison between MICAL and PHBH indicates that these residues may contribute to NADPH binding. Determination of the structure of the R220Q variant of PHBH (with the flavin in the reduced form) in complex with NADP(H) ([
64], PDB ID 1K0J) showed that NADPH binds in an extended conformation on the protein surface in a groove that crosses the FAD binding site and is lined by several protein side-chains and by the pyrophosphate and adenosyl moieties of FAD. Residues K115, R116, R121, K221, V223 and S368 of MICAL may correspond to R33, Q34, R42, H162, R269 and I164 of PHBH, which participate in NADP(H) binding ([
4,
64],
Figure 5).
As in enzymes of the PHBH structural family, FAD is bound to MICAL in an extended conformation. Its adenylate moiety is bound to the Rossman fold identified by the GXGXXG motif at positions 91–96 [
62,
68,
71]. A β strand (β17, [
4]) formed by residues 384–390 interacts with the ribityl 3′OH and corresponds to the Eggink’s second FAD consensus sequence [
63] detected in MICAL’s primary structure (
Figure 4A). Both the dimethylbenzene and the pyrimidine rings of FAD are exposed to solvent (
Figures 6,
7,
8) on the two opposite faces of the protein. This is at variance with PHBH in which the dimethylbenzene ring of the flavin points towards the solvent where NADPH binds, while the pyrimidine ring is buried within the protein. Several of the residues interacting with FAD isoalloxazine ring belong to the highly conserved MICAL’s regions (see above).
The side-chain of the conserved Trp400 stacks against the
re face of the flavin isoalloxazine ring and must undergo conformational changes in order to allow the productive binding of the nicotinamide portion of NADPH during the enzyme reductive half reaction (see below). On the isoalloxazine
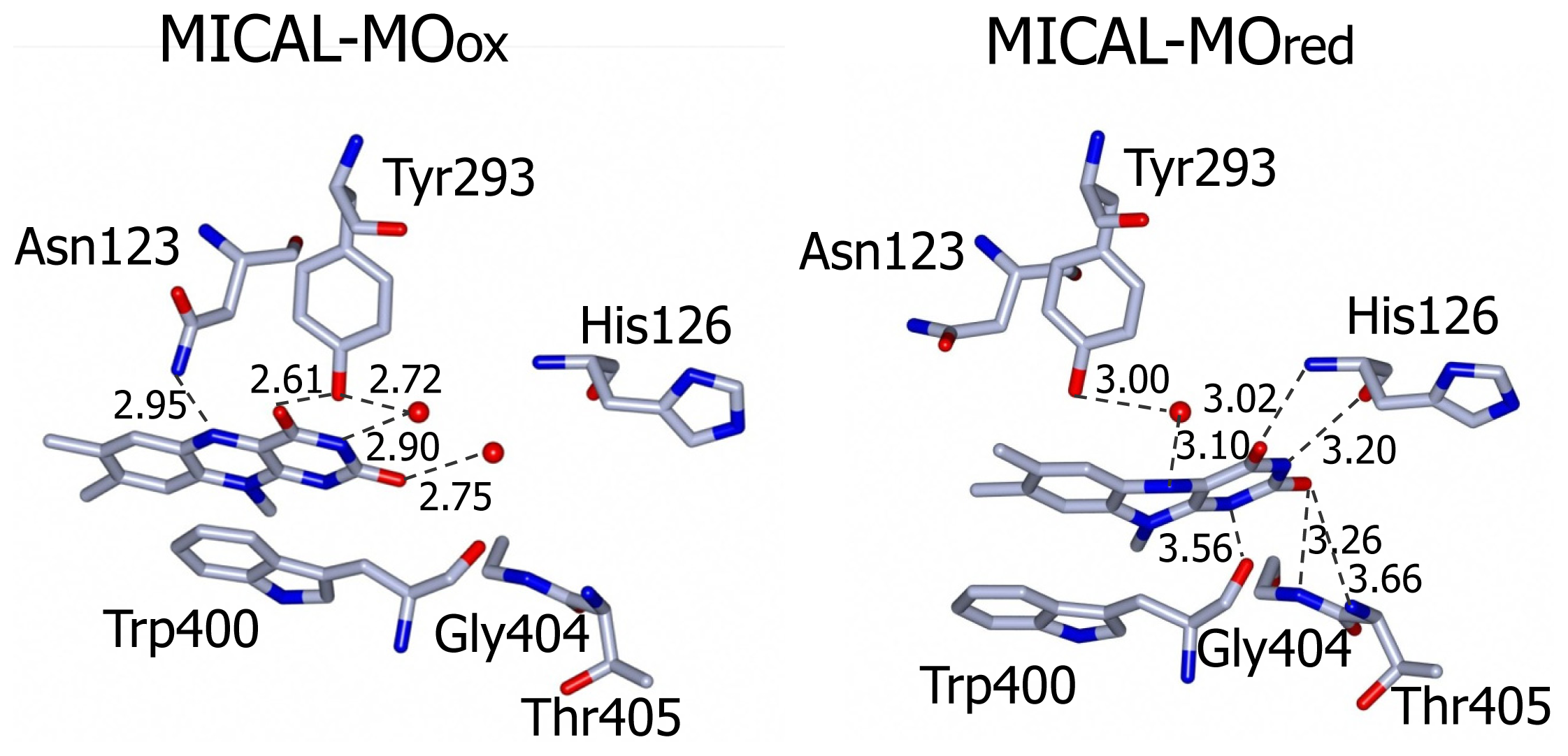
si side, van der Walls interactions are made with Ile157 side chain. Asn123 side-chain amide is at hydrogen bonding distance from the flavin N5 position and is held in place by a complex network of hydrogen bonding interactions between the amide oxygen atom and Asn243, Thr291 and Asp360. The side-chain hydroxyl group of Tyr293 is hydrogen bonded to FAD O(4) contributing to the stabilization of the observed conformation. The FAD N(1), O(2) and N(3) atoms interact with water molecules. Wat71 is held in place by Tyr293 and Val124, while Wat 95 may interact with Asp 393 side-chain carboxylate and with the ribityl 2′OH group (
Figure 9).
In support of the hypothesis that MICAL is a monooxygenase of the PHBH class is the fact that the conformation of the flavin cofactor corresponds to the “flavin out” conformation of PHBH. Such a “flavin out” conformation is observed only with some mutant forms of PHBH or in the presence of the alternative substrate 2,4 dihydroxybenzoate (see [
5,
6] and references therein), while, in most cases, PHBH is in the “flavin in” state.
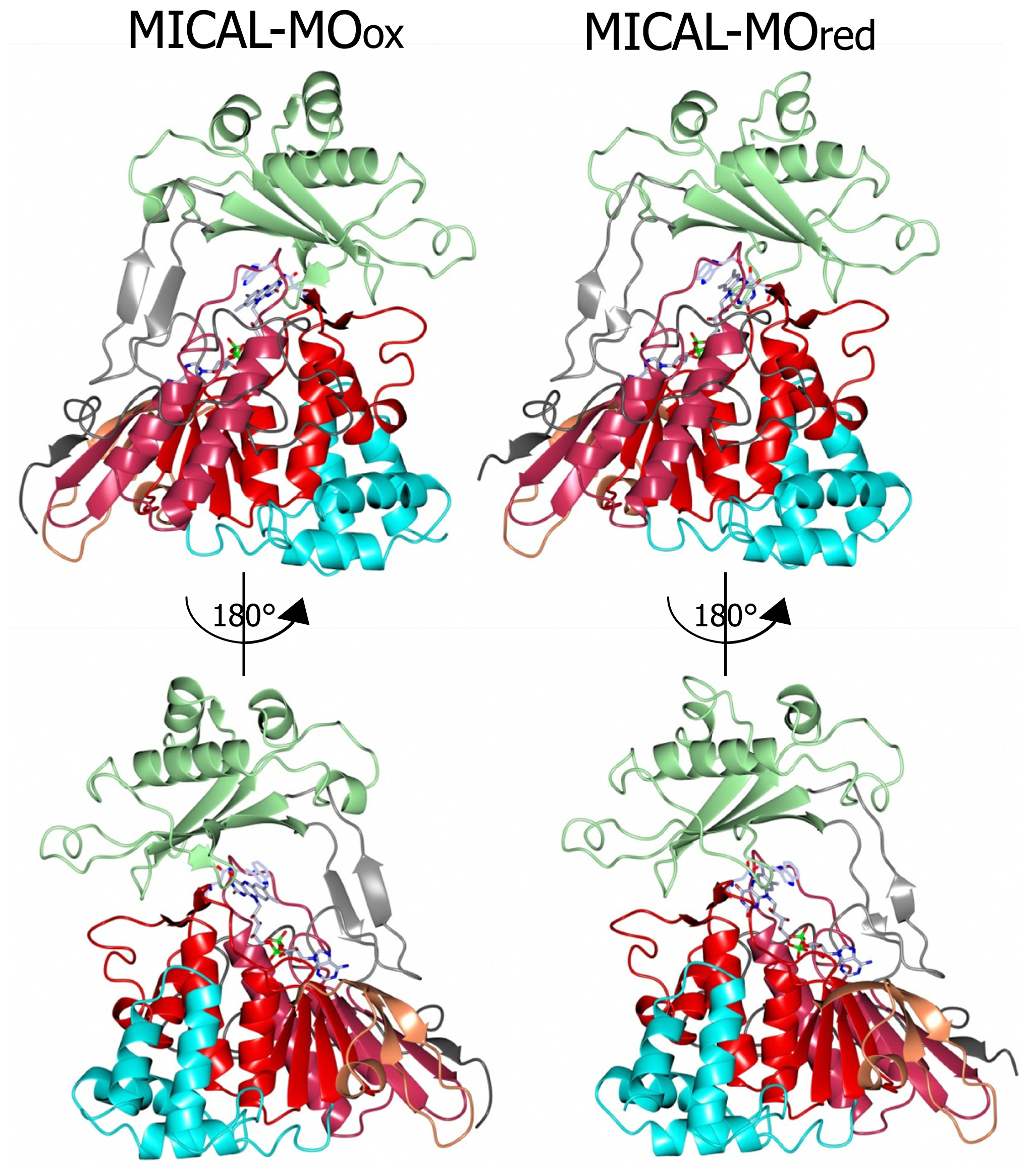
Incubation of MICAL crystals with NADPH led to the determination of the structure of the protein with FAD in the reduced form (as deduced from the bent conformation of the isoalloxazine ring) in a conformation corresponding to the “flavin in” state of PHBH (PDB ID: 2C4C, chain A, [
3],
Figures 6,
7,
8,
9). No NADP(H) was bound to the protein preventing the identification of its binding site. In one of the chains (chain B) both “flavin out” and “flavin in” conformations are observed.
The switch of FAD from the “out” to the “in” conformation occurred through a rotation (~20°) of the ribityl C1–C2 bond, which was accompanied by domain reorientation. In the “in” conformation the isoalloxazine ring is shielded from solvent and buried at the interface between the flavin and the substrate binding domain (compare left and right panels of
Figure 8).
The three water molecules interacting with the flavin N(1), O(2), N(3) and the ribityl 2′OH are displaced with their position occupied by the FAD pyrimidine ring, which now interacts with His126 main chain amide and carbonyl groups, Gly404 and Thr405 (
Figure 9). Tyr293 side chain hydroxyl group now interacts through a water molecule with the FAD N(5) position. The ring stacking interaction between the flavin isoalloxazine ring and Trp400 side-chain is lost, but Trp400 main chain oxygen atom is at hydrogen bonding distance from the reduced FAD N(5) atom (
Figure 9). It has been proposed [
3] that the loss of hydrogen bonding interaction between Tyr293 of the substrate binding domain and FAD O(4) upon flavin reduction may trigger the conformational switch from the “flavin out” to the “flavin in” state. The latter is accompanied by conformational changes of the polypeptide chain with changes in secondary structure elements in the substrate binding domain (compare
Figures 6 and
7 left and right panels). The 6.5° difference in the relative position of the FAD and substrate binding domains of oxidized and reduced MICAL is larger than that observed in PHBH due to the simpler topology of MICAL as compared to PHBH. The β sheet formed by β9 and β15 connecting the FAD and the substrate binding domain has been proposed to act as a hinge during domains reorientation [
3].
The MICAL region corresponding to the substrate binding cavity of PHBH shows no similarity to PHBH, regardless of the protein conformations selected for comparison [
3,
4]. On the contrary, comparison of the structures of MICAL in the oxidized (“flavin out”) and reduced (“flavin in”) forms reveals how the hydroxylatable substrate may access the reduced flavin [
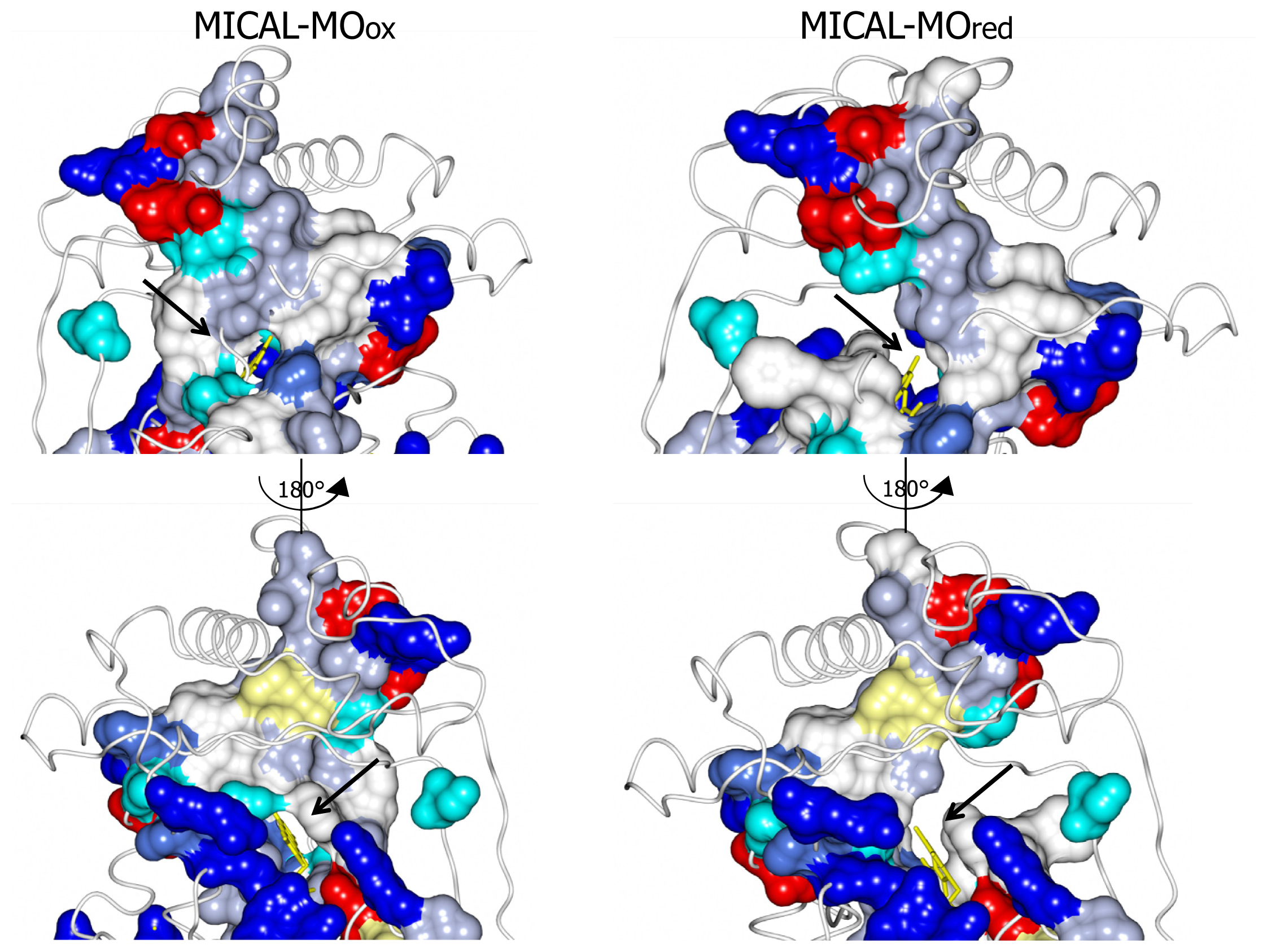
3]. The domain reorientation that accompanies the switch from the “flavin out” to the “flavin in” conformation leads to the formation of a channel connecting the protein surface to the active site above the flavin isoalloxazine ring (highlighted in
Figure 8). In this conformation the central ring of FAD, with the reactive C4a-N5 position, becomes solvent accessible. The residues lining this channel are conserved in MICALs suggesting an important functional role (
Figure 4). The channel, which appears to be rather hydrophobic, is roughly located on the side of the protein where the positively charged N-terminal domain unique to MICAL lies.
These observations, together with the presence of typical protein interaction domains in full-length MICAL, support the hypothesis that the
N-terminal flavoprotein-like domain catalyzes a monooxygenase reaction and that the substrate may be a protein side-chain. The
N-terminal positively charged region of the MICAL monooxygenase domain may contribute to the binding and orientation of the protein substrate with the side-chain to be modified accessing the catalytic site through the tunnel. The MICAL’s additional domains may affect substrates binding as well as the conformational changes that are likely to take place during the catalytic cycle of MICAL, by analogy with PHBH mechanism ([
5,
6] and references therein). Starting from the oxidized (“flavin out”) state, binding of NADPH should trigger a first conformational change involving reorientation of Trp400 side-chain and allowing correct positioning of the NAD(P)H nicotinamide ring for hydride transfer to the flavin N(5). Next the protein should switch to the “flavin in” conformation in which reduced FAD can react with molecular oxygen to generate the 4a-hydroperoxide intermediate in a protected environment that would allow hydroxyl transfer to the (protein) substrate and release of a water molecule rather than of hydrogen peroxide or superoxide (
Figure 3). When the MICAL (protein) substrate would come into play in the catalytic cycle cannot be predicted from structural analyses. PHBH is virtually inactive in the absence of the substrate to be hydroxylated, exhibiting a very low NADPH oxidase activity. Although NADPH can bind to the free enzyme, it can reduce the flavin coenzyme (in the “flavin out” state) only when the hydroxylatable substrate is present [
5,
6]. In PHBH, NADP
+ is released prior to the switch to the “in” conformation so that no contribution to the control of oxygen reactivity is made by the oxidized nicotinamide product. This is at variance with the role of NAD(P)
+ in the catalytic cycle of another important class of monooxygenases exemplified by the flavin monooxygenase (FMO) from
Methylophaga sp. strain SK1 [
73]. In this class of monooxygenases, bound NADP
+ contributes to promote the hydroxylation reaction [
74–
78].
The structures of the CH domain of human MICAL1 (PDB ID: 2DK9, [
65,
79]; PDB ID: 1WYL, no accompanying paper), MICAL2 (PDB ID: 2E9K, no accompanying publication) and MICAL3 (PDB ID:2D88, no accompanying paper) and of the LIM domain of human MICAL1 (residues 687–755, PDB ID 2CO8; no accompanying publication) were determined by NMR confirming their sequence-based identification (
Figure 4A,B). These studies also demonstrated that it is in principle feasible to produce these protein fragments and, e.g., to study their ability to interact with MICAL interactors or the MICAL monooxygenase domain (and other truncated forms) and how they modulate MICAL’s catalytic properties.
Calponin homology (CH) domains are protein interacting domains that are often found in actin and microtubule binding proteins [
66,
67,
80,
81]. In most of the F-actin binding or bundling proteins two tandem CH domains are present and form the actin binding site. MICALs contain a single type 2 CH domain ([
59,
65],
Figure 4), which is predicted not to be sufficient to bind F-actin [
80,
81]. Accordingly, no binding between the isolated MICAL1 CH domain and F-actin could be detected [
65]. Thus, it has been proposed that the CH domain may assist regions of the MICAL’s monooxygenase domain in (protein) substrate binding either by completing the binding site, or by establishing the initial interaction with the protein substrate and presenting it to the catalytic domain. In this respect, the work of Hung
et al. [
52,
53], which will be discussed below, demonstrated for the first time that the MICAL monooxygenase-like domain is sufficient to interact with F-actin causing its depolymerization in a NADPH-dependent reaction.
LIM domains (from the initials of the homeodomain proteins Lin11, Isl-1 and Mec-3, [
17,
18]) are cysteine- and histidine-rich, zinc-coordinating domains, which contain two tandemly repeated zinc fingers. The LIM domain is a protein interaction module found in structural proteins, kinases and transcription factors with roles in gene expression, cell adhesion and signal transduction [
17,
18]. No information on its precise role in MICAL is yet known.
2.6. Catalytic Activities of MICAL
In vitro mechanistic studies of MICAL forms are very much needed to complement genetic and in cell studies as well as structural work. However, in spite of the fact that the protein was first identified in 2002 [
1,
2] and structures of the monooxygenase domain are available since December 2005 [
3,
4], very little mechanistic work has been done on isolated protein forms. However, essential key observations have been reported recently using mouse [
4], human [
70] and Drosophila MICAL forms [
52,
53]. The mouse MICAL1 monooxygenase domain has been produced by Siebold
et al. [
3] and Nadella
et al. [
4] with the aim to solve its three-dimensional structure by X-ray crystallography. As a complement to the structural studies, Nadella
et al. [
4] reported some kinetic properties of the mouse MICAL1 monooxygenase domain, which we shall here indicate as MICAL-MO or simply MO.
A coupled assay with horse radish peroxidase (HRP) and amplex red was used to determine that mouse MICAL-MO catalyzes the oxidation of NAD(P)H with production of hydrogen peroxide. The enzyme was reported to prefer NADPH over NADH as the reductant and to exhibit a
kcat of 77 s
−1 and a
KM for NADPH of 222 μM at pH 7.0, which was indicated as the pH optimum for the activity. (−) epigallocatechin gallate (EGCG) was reported to be a potent inhibitor of mouse MICAL-MO domain (
Ki, 2 μM), non-competitive with respect to NADPH. This result is very interesting in the light of previous work that demonstrated that treatment of dorsal ganglion cells with EGCG mimics MICAL inactivation by making cells insensitive to semaphorins [
2,
38]. Furthermore, xanthofulvin, a natural compound structurally related to EGCG, was also found to inhibit semaphorin signaling and to promote neuronal regeneration following spinal cord injury [
82,
83]. Forms of the Drosophila MICAL comprising the monooxygenase domain (MICAL-MO) or both the monooxygenase and the CH domain (MICAL-MOCH) have been produced as fusions with
N- or
C-terminal Nus and His tags [
20,
52,
53]. The purified proteins have been mainly used to study their interaction and effect on actin [
52,
53], but no information is available on flavin content and spectral properties of the proteins, nor on the kinetics of the catalyzed reactions.
More recently, the monooxygenase domain of human MICAL1 was produced in
E. coli cells without or with
N- or
C-terminal His-tags [
70]. The purified proteins contained stoichiometric amounts of FAD and were indistinguishable from each other with respect to the absorbance and fluorescence spectra and the steady-state kinetic parameters of the NAD(P)H oxidase reaction. Thus, most of the experiments designed to carry out the basic characterization of the protein have been carried out with the most abundant
C-terminally His-tagged MICAL-MO form.
The absorbance spectrum of human MICAL-MO exhibits maxima at 278, 376 and 457 nm with a
A278/
A457 ratio of ~13. The calculated extinction coefficient at 457 nm was 8.1 mM
−1 cm
−1 (
Figure 10). The broad absorbance band extending to 700 nm agrees well with the presence of Trp400 side-chain, which is positioned essentially parallel to the FAD isoalloxazine ring in the structure of the “as isolated” (oxidized) mouse MICAL-MO at a distance suitable for the formation of a charge-transfer complex (
Figure 9, [
3,
4]). Such an absorption band might provide a tool to monitor the conformational changes that are predicted to take place during the catalytic cycle of MICAL-MO. The FAD bound to human MICAL-MO does not interact with sulfite nor stabilizes 1-electron reduced semiquinone forms of the flavin during photoreduction in the presence of EDTA and deazariboflavin, chemical reduction with dithionite or NADPH or exposure to air of the reduced enzyme. From anaerobic NADPH titration at pH 7.0 a midpoint potential of ≈ −150 mV was calculated for the
Eox/
Ered couple. These properties support the functional similarity of MICAL-MO with enzymes of the FAD-dependent monooxygenase family whose prototype is PHBH, rather than typical FAD-dependent oxidases [
55,
84], which also exhibit the PHBH fold (e.g.,
d-amino acid oxidase; [
85]). No spectral changes were observed by titrating the enzyme with NADP
+, a fact that may limit the study of the interaction between MICAL-MO and the pyridine nucleotide substrate/product in future work.
As in the case of the mouse protein, human MICAL-MO catalyzes NAD(P)H oxidation in the presence of atmospheric oxygen with a preference for NADPH over NADH in terms of both
kcat/
KNAD(P)H and
kcat (
Table 2). By comparing the rate and extent of NADPH oxidation (spectrophotometrically) and of oxygen consumption (with an oxygen electrode), in the presence and absence of catalase and/or superoxide dismutase, it was concluded that one NADPH is oxidized to NADP
+ per molecule of oxygen being reduced to hydrogen peroxide.
The 1:1 stoichiometry between oxidized NADPH and hydrogen peroxide being produced was confirmed by quantifying the hydrogen peroxide product with HRP and amplex red or
o-dianisidine. Interestingly, it was found that the NADPH oxidase activity of MICAL-MO could not be measured in a coupled assay with HRP and amplex red or
o-dianisidine due to interference of NADPH with the HRP reaction and a stimulation of NADPH oxidation by HRP and amplex red in the presence of H
2O
2. Thus, H
2O
2 production by MICAL-MO NADPH oxidase activity could only be reliably quantified in a two-step assay in which NADPH was allowed to be oxidized by MICAL-MO, the reaction was stopped by rapidly oxidizing residual NADPH (with glutamate synthase and its substrates as a suitable dehydrogenase, [
87]), and
o-dianisidine and HRP were eventually added to quantify H
2O
2.
Both the
kcat (~4 s
−1) and the K
M for NADPH (20–30 μM) were one order of magnitude lower than the values reported for the mouse species (77 s
−1 and 222 μM, respectively, [
4]). The differences in the
KM values for NADPH is due to the presence of 0.1 M KCl in the assays done with the mouse enzyme and to the sensitivity of MICAL-MO to the ionic strength of the medium and the type of anions (see
Table 2 and below). In the light of the strong sequence similarity between the mouse and human proteins, the different
kcat values are more likely due to a combination of the assay methods adopted (direct measurement of NADPH oxidation for the human protein versus the coupled HRP/amplex red assay for the mouse protein) and other experimental factors, rather than actual species-dependent differences. The NADPH oxidase activity of human MICAL-MO is significantly higher than that of PHBH, taken as the reference enzyme. The latter is virtually inactive in the absence of the
p-hydroxybenzoate substrate [
5,
6]. Whether the MICAL-MO NADPH oxidase activity is physiologically relevant is unclear. MICAL-MO NADPH oxidase activity is very sensitive to ionic strength and type of anions (see below) so that it is expected that the reaction is very slow in the cell.
However, whether such a sensitivity is removed by the other MICAL’s domains and the interacting proteins will need to be established.
In vitro, the detectable, NADPH oxidase activity of MICAL-MO provides a tool to study, perhaps for the first time, the reaction between a monooxygenase of the PHBH class and NADPH in the absence of the substrate undergoing hydroxylation. As a first step in such a direction, the reaction between human MICAL-MO and NADPH was studied in a stopped-flow under anaerobic conditions leading to the observation that the bound FAD is converted to the 2-electron reduced hydroquinone form without the detectable formation of intermediates. The apparent
Kd for NADPH and the
kred were similar to those measured under steady-state conditions in the same buffer, including 10% glycerol (
Table 2) that lowers
kcat and increases
KM for NADPH (
Table 2). Thus, it appears that the enzyme NADPH oxidase activity is limited by the rate of the enzyme reductive half reaction.
Human MICAL-MO was found to be very sensitive to the ionic strength of the solvent and to the type of anions present, which all specifically lowered kcat/KNADPH.
Phosphate, chloride and acetate salts competed directly for NADPH binding regardless of the counter ion and with decreasing potency in the stated order. The lack of “special” effect of cations such as Mg
2+ or Ca
2+ is informative in the light of the role played by these ions
in vivo. Interestingly, by varying the concentration of Tris and Bis-tris buffer,
kcat/
KNADPH seemed to depend on electrostatics so that the dependence of the parameter from ionic strength could be well fitted with the Debye-Huckel equation [
70]. The effects of anions on MICAL-MO further supports its functional similarity with enzymes of the PHBH family. In PHBH, anions compete with NADPH phosphate groups for the binding to positively charged residues on the protein (
Figures 7 and
8). Furthermore, electrostatics plays an important role for both NADPH binding and catalysis [
5,
6,
88].
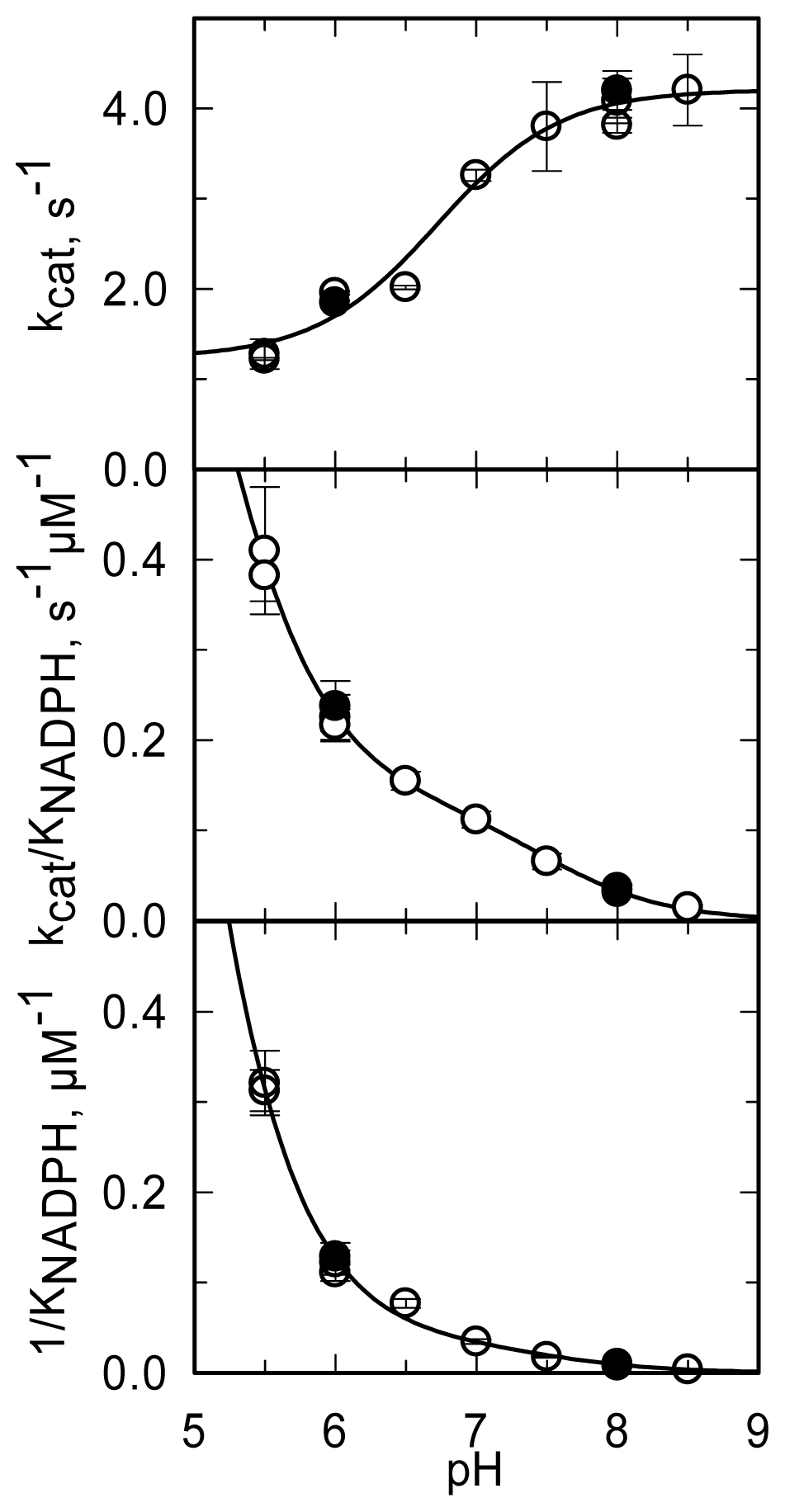
The pH dependence of the steady-state kinetic parameters of the NADPH oxidase reaction was studied in a buffer system designed to maintain essentially constant ionic strength across the pH interval being explored and to minimize interference of specific anions (
Figure 11,
Table 3).
kcat increases from a low constant value (at low pH) to a limiting maximum value (at high pH) as a group with an apparent pK
a of 6.7 dissociates.
kcat/
KNADPH and 1/
KNADPH showed similar pH dependencies. They decreased from high (non-limiting) values (at low pH) to low values (at high pH) as groups with apparent pK
a of 4.6–4.9 and ~7.5 deprotonated. The
kcat,
kcat/
KNADPH and 1/
KNADPH profiles (
Figure 11) are all qualitatively similar to those obtained with PHBH when the interaction with NADPH was studied [
88]. As proposed for PHBH, also in MICAL they may reflect an increased positive charge of the protein at low pH, which favors the interaction with the negatively charged NADPH. With respect to the group that needs to be deprotonated in order to observe maximum
kcat, the group in the mouse MICAL-MO active site which is likely to exhibit a pK
a of 6.7 is His126 (
Figure 9). This residue is conserved in all MICALs and may stabilize the “flavin in” conformation through H-bonds between its main chain atoms and the isoalloxazine N(3) and O(4) atoms. Neutralization of the side-chain of this residue upon deprotonation may favor the “flavin out” conformation, which is the conformation that allows hydride transfer from NADPH to the flavin N5 position.
Interestingly,
kcat and, to a greater extent,
kcat/
KNADPH were found to be sensitive to solvent viscosity, consistently with the hypothesis that one or more of the conformational changes predicted to take place during MICAL-MO catalytic cycle is sufficiently slow to contribute to the determination of
kcat and
kcat/
KNADPH. With respect to the enzyme reductive half reaction, a minimal scheme including essential conformational changes would involve: (i) binding of NADPH to the oxidized species in the “flavin out” conformation with Trp400 stacking against the isoalloxazine ring; (ii) repositioning of Trp400 and of NADPH to reach the correct geometry for hydride transfer from the NADPH nicotinamide ring to the FAD isoalloxazine ring; (iii) hydride transfer with formation of FAD hydroquinone; (iv) switch from the “flavin out” to the “flavin in” conformation, which leads to formation of the tunnel connecting the surface of the protein with the active site (
Figure 8); (v) completion of the monooxygenase/hydroxylation reaction.
This minimal scheme is certainly not sufficient to describe the details of the reaction since it does not take into account several relevant aspects such as, for example: (i) the existence of a third “open” conformation, as in PHBH; (ii) if NADPH binds to both “flavin out” and “flavin in” conformations, or just to the “out” state; (iii) whether NADP
+ dissociates prior to the switch form the “out” to the “in” conformation, as in PHBH; (iv) whether the physiological substrate undergoing hydroxylation binds to both conformations and/or (v) how its binding affects the out/in states and NADPH/NADP
+ binding and reactivity. Whether the NADPH oxidase activity of MICAL-MO requires the switch from the “out” to the “in” conformation is also not clear. In principle, reduced FAD may react with molecular oxygen when in the “out” conformation leading to production of hydrogen peroxide. In this respect a great deal of work has been done recently to elucidate how flavoenzymes react with oxygen (see [
74] for a recent review and a collection of relevant references). Thus, it would be of great interest to apply similar experimental and theoretical approaches to the understanding of the reactivity with oxygen of MICAL in its various conformations.
Among essential steps towards the understanding of MICAL-MO physiological role is the identification of the physiological (hydroxylatable) substrate and of specific inhibitors.
A survey of potential substrates among amino acids, (poly)amines and aromatic compounds was done with the human MICAL-MO [
70]. The molecules either had no effect on the reaction velocity or acted as inhibitors. Benzoate derivatives were mild inhibitors (estimated
Ki values in the 0.1–1 mM range). They yielded competitive or non-competitive inhibition patterns with respect to NADPH, and their potency increased as the number of hydroxyl groups increased. This result correlated with the proposal that EGCG is a potent and selective inhibitor of mouse MICAL-MO ([
4],
Ki, 2 μM). Testing the effect of this compound on human MICAL-MO confirmed that EGCG is a non-competitive inhibitor of MICAL-MO with similar effects on slopes and intercepts of double reciprocal plots (
Table 4). However, EGCG was a less potent inhibitor of human MICAL-MO (
Ki, 17 μM) than reported for the mouse protein (
Ki, 2 μM, [
4]). The difference may be related to the assay format (see above) by taking into account that EGCG, as a catechol, is a potent H
2O
2 scavenger. Interestingly, during spectrophotometric titrations, EGCG caused human MICAL-MO aggregation/denaturation at concentrations just above its K
i (17 μM, [
70]), while no absorption changes were observed at lower concentrations. This result casts some doubt on the conclusion that EGCG is a selective inhibitor of MICAL-MO. It might even be suggested that some of the observed inhibitory effects of EGCG on semaphorin-induced growth cone collapse [
2] may be due to its effect on protein aggregation/denaturation. The similarity of EGCG with xanthofulvin (SM-216289), which was reported to be a selective inhibitor of semaphorin of therapeutic interest [
82,
83], may lead to the proposal that also this compound may in part function as a radical scavenger or by denaturing semaphorin.
2.7. The Actin Depolymerizing Activity of MICAL
A great deal of work has been done to test the hypothesis that actin is a MICAL substrate. Fundamental
in vitro experiments were done using Drosophila MICAL forms comprising the MO (MICAL-MO) and both the MO and CH domains (MICAL-MOCH). The
in vitro work complemented genetic studies in Drosophila in which the actin-rich bristles were used as a model to study the effect of MICAL forms on actin organization [
52,
53].
Several tests were used to study the effect of Drosophila MICAL forms on globular (monomeric) actin (G-actin) and on the filamentous form (F-actin). Similar results were obtained with the Drosophila MICAL-MO and MICAL-MOCH forms indicating that the CH domain is not essential for MICAL-actin interaction. Both MICAL forms were found to co-sediment with F-actin, demonstrating an interaction between the proteins (
Figure 12). When G-actin was allowed to polymerize in the presence of MICAL forms and NADPH, a significant fraction of actin (and MICAL) was found in the supernatant from a high-speed centrifugation indicating that MICAL caused F-actin depolymerization. By exploiting the fact that pyrenyl-actin fluorescence is significantly greater in F-actin than in G-actin (λ
ex, 365 nm; λ
em, 407 nm), it was shown that Drosophila MICAL-MO and MICAL-MOCH interfere with actin polymerization and cause actin depolymerization in a fashion that is dependent on NADPH and MICAL concentration (
Figure 12). In these experiments, initially, fluorescence increases due to the prevailing actin polymerization. Eventually, fluorescence decreases due to actin depolymerization until a steady-state is reached. Addition of MICAL forms and NADPH to pre-polymerized pyrene-labeled F-actin led to observing fluorescence decrease, consistently with actin depolymerization. In these and several other experiments presented by Hung
et al. [
52,
53], the observed depolymerizing activity was associated with the MO domain, as no differences were observed when experiments were carried out with the MICAL-MO or MICAL-MOCH forms. Importantly, MICAL appears to interact along the F-actin filament rather than at one of its ends and several actin bundling and capping proteins have no effect on MICAL depolymerizing action [
52,
53]. Interestingly, it was also shown that direct contact between MICAL forms and F-actin was required for the depolymerization [
52]. This finding led the authors to rule out that MICAL’s effect on F-actin is due to diffusible hydrogen peroxide resulting from the MICAL’s NADPH oxidase activity, thus supporting the hypothesis that MICAL catalyzes a monooxygenase reaction. However, it cannot be ruled out that MICAL may catalyze a NADPH oxidase reaction when bound to F-actin and that it is the high local hydrogen peroxide concentration that causes F-actin depolymerization. Finally, by reisolating actin after incubation with MICAL-MOCH and NADPH under polymerization conditions, it was shown that actin was unable to repolymerize [
52]. Modification of Met44 and Met47 into methionine sulfoxide was detected by mass spectrometry in MICAL-treated actin. Met44 was identified as the site of specific modification by generating Drosophila Met44Leu and Met47Leu actin variants. The proteins were able to polymerize and to bind MICAL in the absence of NADPH, but the Met44Leu and the Met44Leu/Met47Leu variants were no longer sensitive to MICAL and NADPH [
52].
Met44 is in the actin
d-loop at the pointed end of the actin monomer. In G-actin, the loop is very flexible and Met44 is among the residues that are the most sensitive to oxidation [
89–
92]. High resolution models of F-actin have not been obtained yet making it difficult to discuss Met44 role in F-actin and its accessibility to MICAL. In the F-actin model of Oda
et al. [
93], the
d-loop makes extensive (both electrostatic and hydrophobic) longitudinal contacts with the adjacent monomer along the filament, which is a left handed, two-chained long helix [
92]. Thus, the
d-loop and its precise conformation are believed to be important for F-actin stability so that chemical modifications of one or more of its residues may indeed destabilize the filament. Although Met44 is partially protected from oxidizing agents in F-actin [
90], it may lie toward the outside of the F-actin structure (with Met47, [
90]) so that it may be accessible to MICAL-MO. This may also be possible in the light of the conformational flexibility of the
d-loop in actin filaments as tested by cross-linking experiments with forms of yeast actin in which the
d-loop was subjected to Cys-scanning [
94]. In this respect it is of interest that expression of the Met44Cys yeast actin variant in yeast led to no viable colonies supporting a crucial role of this residue [
94]. For MICAL-MO to modify actin Met44, MICAL should be able to extract the residue from the interface between actin monomers in the filament. Conversion of Met44 into methionine sulfoxide would locally destabilize the filament promoting its fragmentation. Modeling studies on actin filaments indicated a conformational heterogeneity of actin monomers along the filament, which may affect its local stability [
95]. Thus, Met44 could be accessible to MICAL at least in some parts of the polymer. In this respect, similar modeling studies may greatly contribute to the understanding of MICAL-actin interactions.
In support of the identification of Met44 as the actin residue specifically targeted by MICAL is the fact that introducing the Met44Leu variant into
Drosophila bristles, a phenotype similar to that obtained with a MICAL loss-of-function mutant is observed. Interestingly, the dominant Met44Thr mutation of skeletal muscle actin is associated with nemaline myotrophy, in which actin accumulation and aggregation are observed [
96].
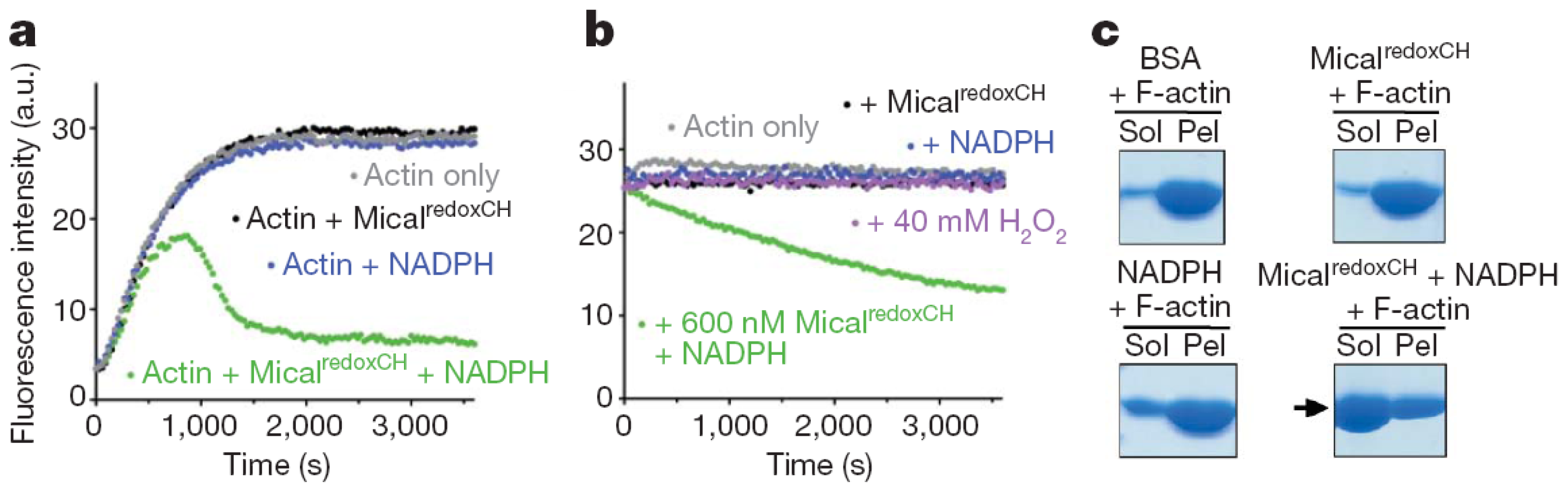
The human MICAL-MO was also found to exhibit an F-actin depolymerizing activity [
70]. The reaction was monitored fluorimetrically by exploiting the different fluorescence intensity of pyrene-labeled G- and F-actin (
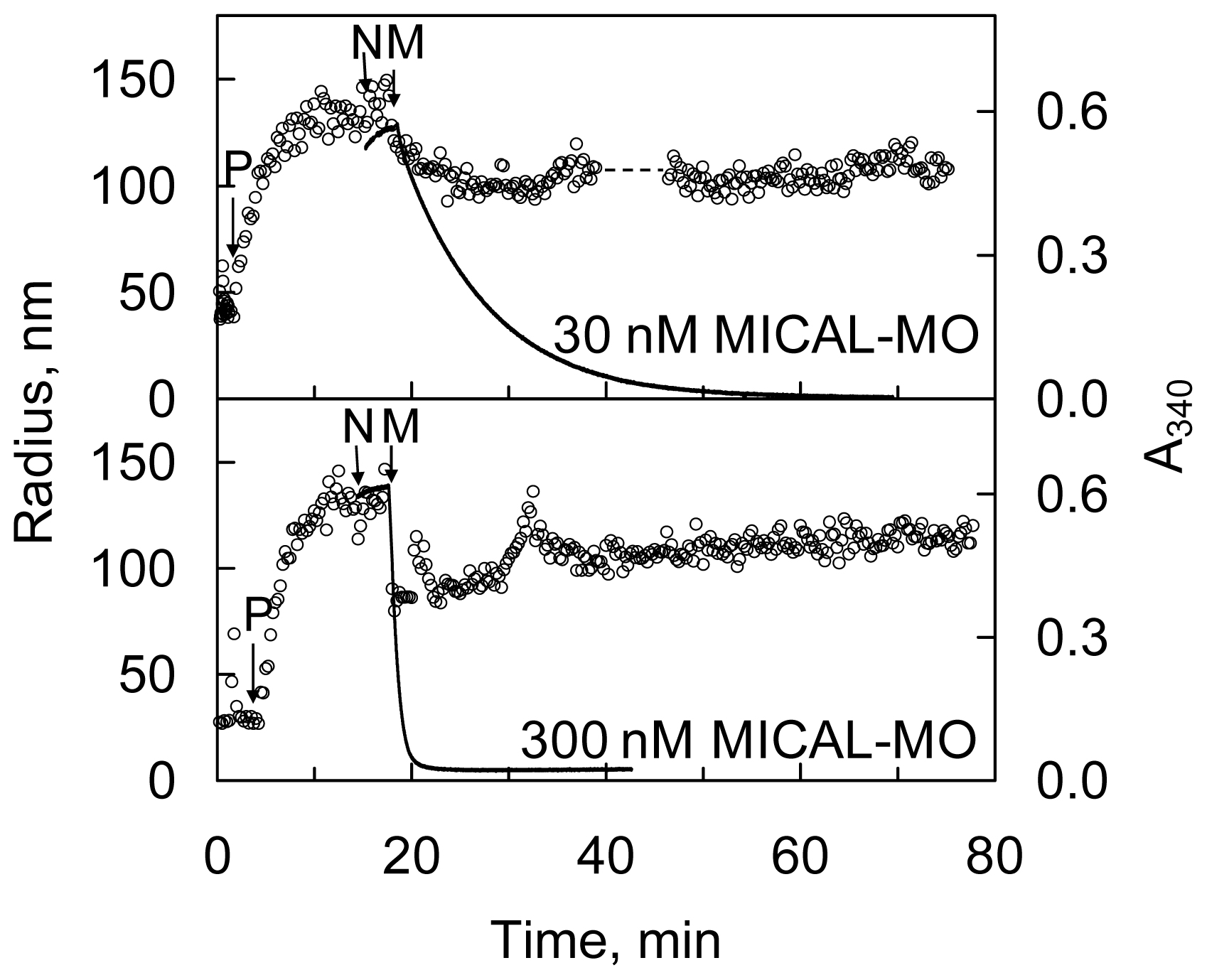
Figure 13) and by sedimentation assays. Dynamic light scattering (DLS) was also used to determine the average radius of the species in solution. The time-course and extent of NAD(P)H oxidation were monitored spectrophotometrically (
Figures 14 and
15).
Addition of human MICAL-MO to pre-polymerized pyrenyl-labeled actin, in a buffer stabilizing F-actin (F-buffer), after NADPH, led to a decrease of fluorescence, which was dependent on MICAL and NADPH concentration (
Figure 13). Also some human MICAL-MO was found to co-sediment with F-actin. When pre-polymerized F-actin was incubated with human MICAL-MO and NADPH in F-buffer, a fraction of actin was found in the supernatant after high-speed centrifugation confirming MICAL- and NADPH-dependent F-actin depolymerization.
The rate and extent of NAD(P)H consumption were monitored spectrophotometrically in the presence of G- and F-actin demonstrating that F-actin, but not G-actin, significantly stimulates NAD(P)H oxidation by lowering the
KM value for NADPH (50-fold) and increasing k
cat (3- or 6-fold at 2.4 μM actin or extrapolated at infinite actin concentration, respectively) when compared to the values measured in the absence of actin, but under the same buffer conditions (
Table 5) resulting in a ~500–800– fold
kcat/
KNADPH increase.
These results are those expected for an authentic monooxygenase in the presence of its substrate. A
KM for F-actin of 4.7 μM was determined at 300 μM NADPH. A decrease of the intensity of the signal was also detected by DLS. It paralleled the decrease of absorbance at 340 nm confirming the F-actin depolymerizing effect of MICAL-MO in the presence of NAD(P)H (
Figure 15). Some technically critical points were highlighted by comparing the experiments done with the human [
70] and the Drosophila proteins [
52,
53]. First, and as expected, it was found that NAD(P)H interfered with fluorescence monitoring of pyrene-labeled actin polymerization state (
Figure 13). At the wavelength used to excite actin-bound pyrene (λ
ex, 365 nm), NAD(P)H absorbs a significant fraction of light causing a drastic decrease of the intensity of the light emitted by pyrene at 407 nm in F-actin. Thus, the fluorescence intensity values alone may not fully reflect the rate and extent of pyrenyl-actin polymerization/depolymerization. Secondly, human MICAL-MO appeared to be more active than the Drosophila forms (compare
Figure 12 and
Figures 13,
14,
15). Almost stoichiometric amounts of Drosophila MICAL-MO or MICAL-MOCH were routinely used to carry out the depolymerization reaction, which was completed in ~60 min, namely 600 nM MICAL form and 1–5 μM actin monomer. With a similar concentration of human MICAL-MO, the reaction was completed within 2–5 min from MICAL addition even in the absence of actin (
i.e., in the NADPH oxidase activity). As a consequence of this observation, all experiments with the human protein were carried out with 10–100 nM MICAL-MO (
i.e., with catalytic amounts of MICAL) in the presence of pre-polymerized 2–5 μM actin and in buffer stabilizing the filamentous actin form. Furthermore, in several of the experiments with the Drosophila protein, it appears that G-actin was mixed with MICAL forms and NADPH prior to inducing polymerization. With this assay format, it may be difficult to know how much residual NADPH is present in solution by the time actin polymerization is initiated.
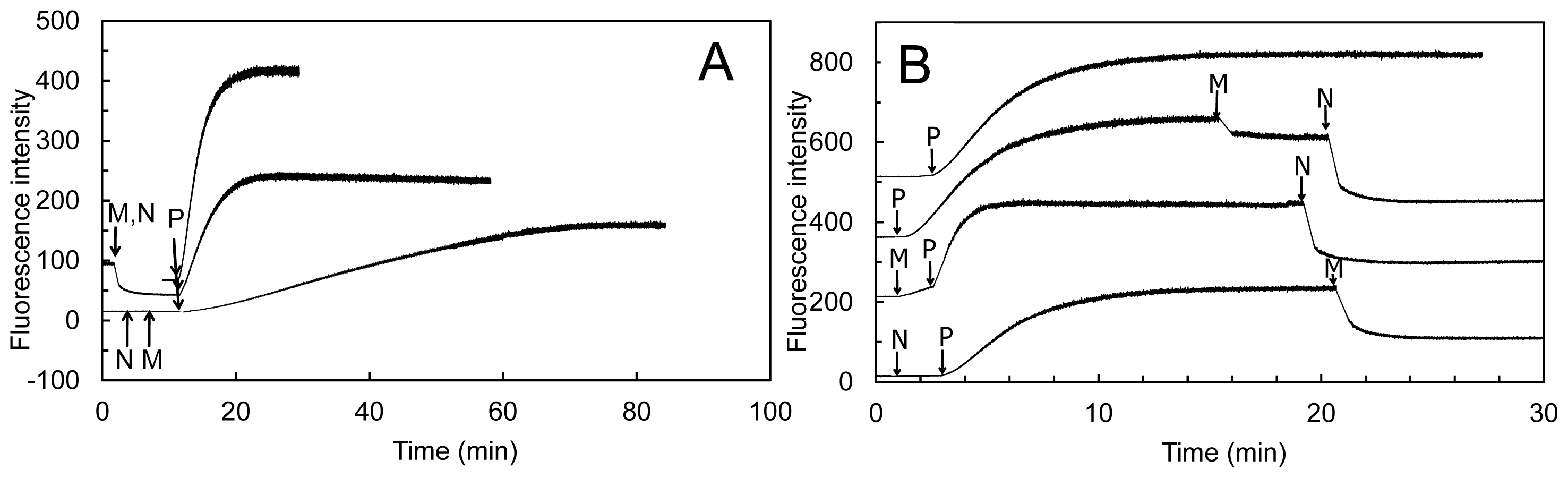
More importantly, inspection of the time-course and extent of NADPH consumption in the presence of human MICAL-MO (
Figures 14 and
15, [
70]) revealed an as yet unsolved puzzling aspect. When experiments like those shown in
Figure 14 and
15 were carried out, the extent and time-course of NADPH oxidation was different from those expected. Assuming that MICAL-MO catalyzes the hydroxylation of one specific actin residue, with a 1:1 stoichiometry of NADPH being oxidized per actin monomer being modified, one would have predicted biphasic traces. Initially, MICAL would behave as a monooxygenase. The rate of NADPH consumption would be fast and the extent of the phase should be lower or equal to the total actin monomer concentration. A second slower phase of NADPH oxidation should follow due to the MICAL NADPH oxidase reaction with residual oxygen.
On the contrary, although traces showed some biphasicity, which could not be clearly correlated with the total actin concentration present, the second phase was much faster than that measured in the NADPH oxidase activity under the same conditions (
Figure 14) and led to oxidation of all NADPH present. The observed extent and time-course of NADPH consumption are consistent with a case of substrate recycling suggesting that the effect of MICAL (and NADPH) on F-actin may be reversible. In support of this hypothesis is the fact that when the reaction was monitored by DLS, actin was never fully converted to the monomeric form, as judged by the average size of the particles in solution (
Figure 15). An allusion to this possibility is made also by Hung
et al. [
52], perhaps taking into account the results obtained with the human MICAL-MO [
70].
The same extent and time-course of NADPH oxidation were obtained when assays were done in buffers that do not stabilize F-actin, and in the absence of dithiothreitol (DTT). The latter, which is included in the buffers to protect actin from non specific oxidation, efficiently scavenges any hydrogen peroxide present, but may also reverse oxidation of Cys residues. How one could reconcile the extent and rate of NADPH consumption observed with the human MICAL-MO [
69] and the irreversible depolymerization of actin due to quantitative chemical modification of actin monomers reported by Hung
et al. [
52] with the Drosophila protein is not clear. Thorough studies of the reaction are indeed required. However, it is unlikely that the effect on actin is due to hydrogen peroxide generation on the basis of independent observations. Only high H
2O
2 concentrations lead to actin depolymerization as judged from decrease of fluorescence of pyrenyl-labeled actin [
52,
53] and DLS (unpublished data). Direct contact between Drosophila MICAL forms and F-actin was required to cause depolymerization [
52]. Control experiments also ruled out that NADP
+ alone or in combination with H
2O
2 may be responsible for actin depolymerization [
53]. However, as pointed out before, a high local hydrogen peroxide concentration produced by MICAL oxidase activity when bound to F-actin, may be sufficient to chemically modify one or more actin residues and cause filaments’ depolymerization.