HE4 (WFDC2) Promotes Tumor Growth in Endometrial Cancer Cell Lines
Abstract
:1. Introduction
2. Results
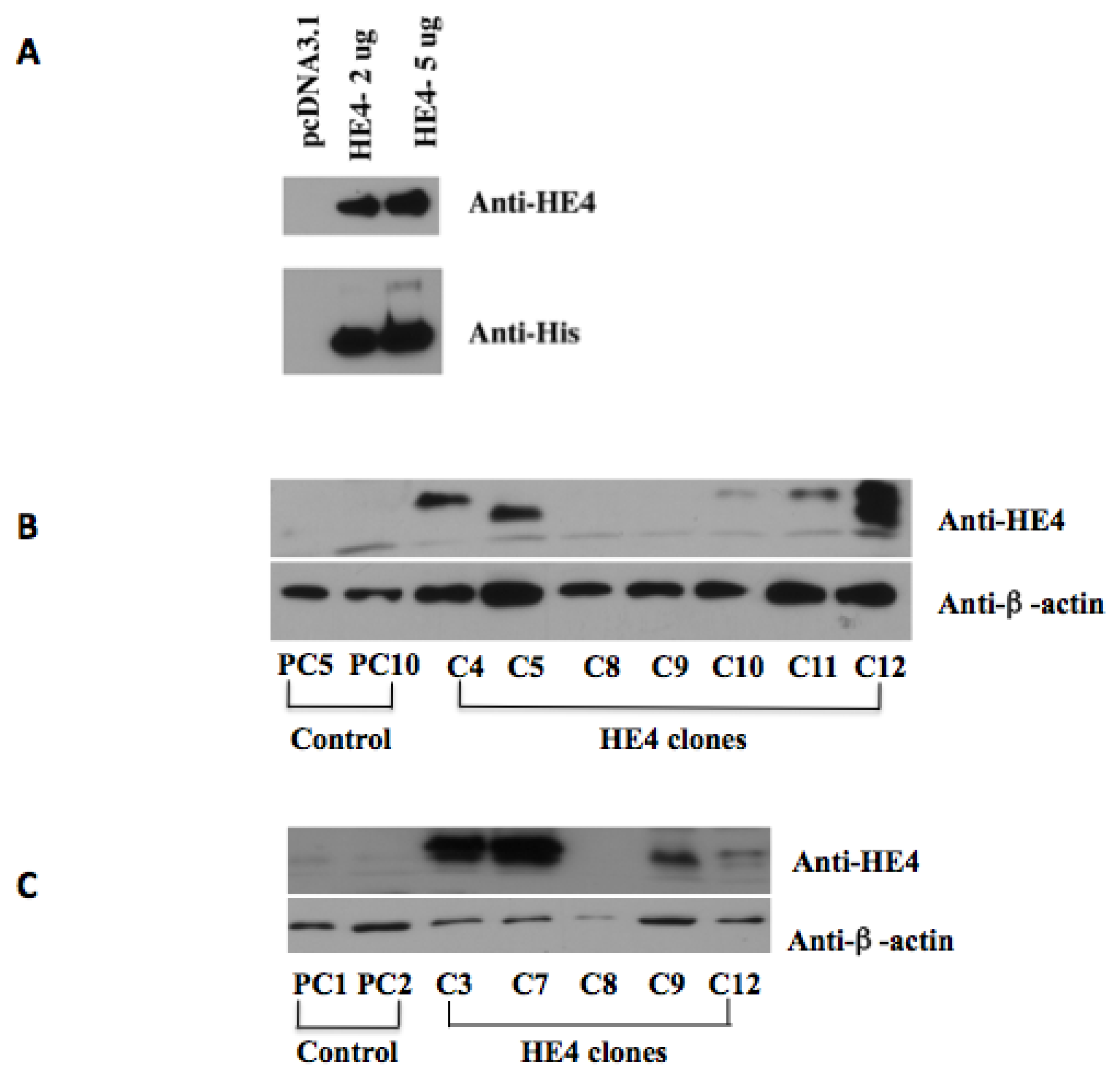
2.1. Overexpression of Human HE4 in Endometrial Cancer Cell Lines
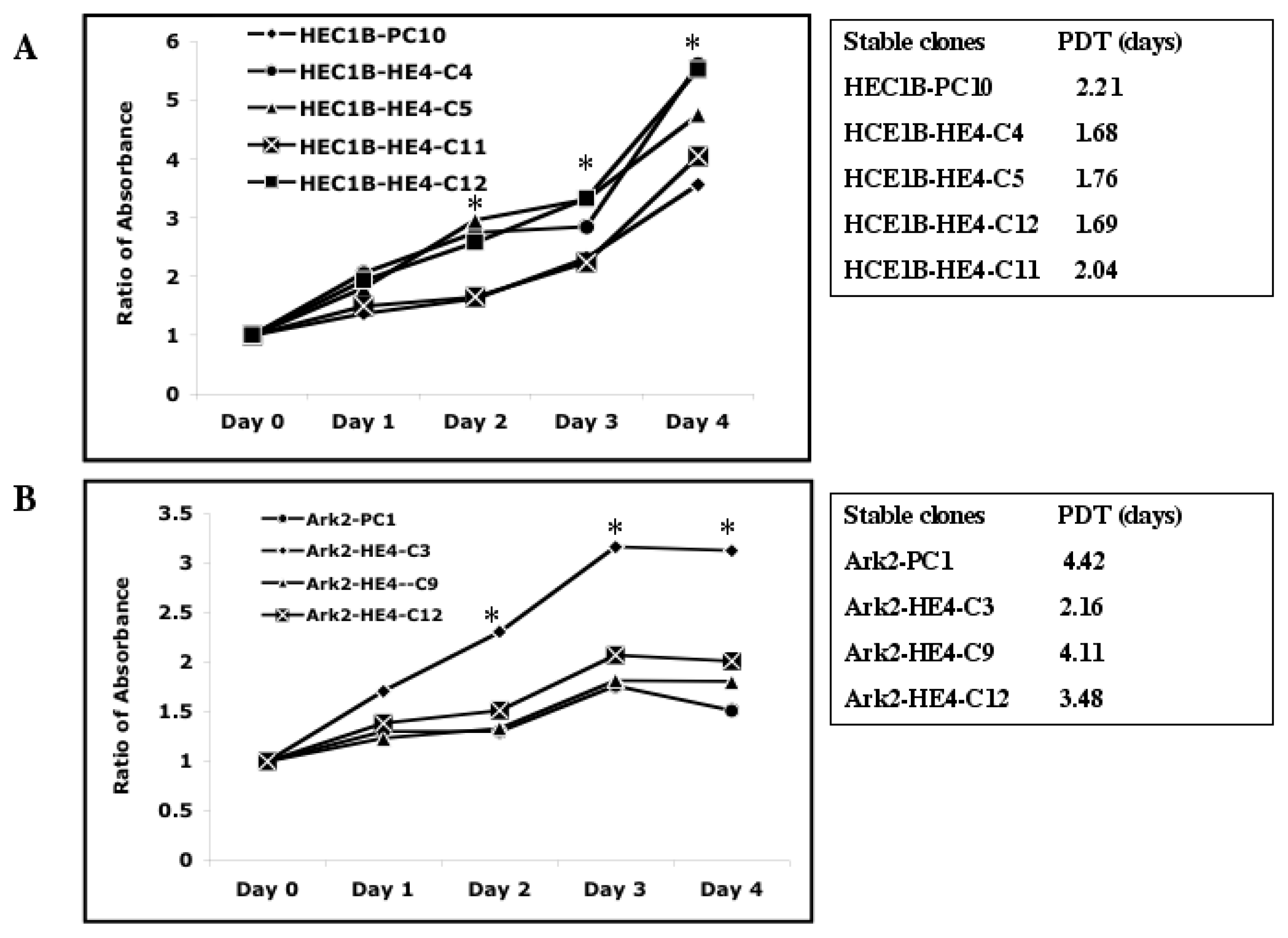
2.2. HE4 Overexpression Stimulates Endometrial Cancer Cell Growth
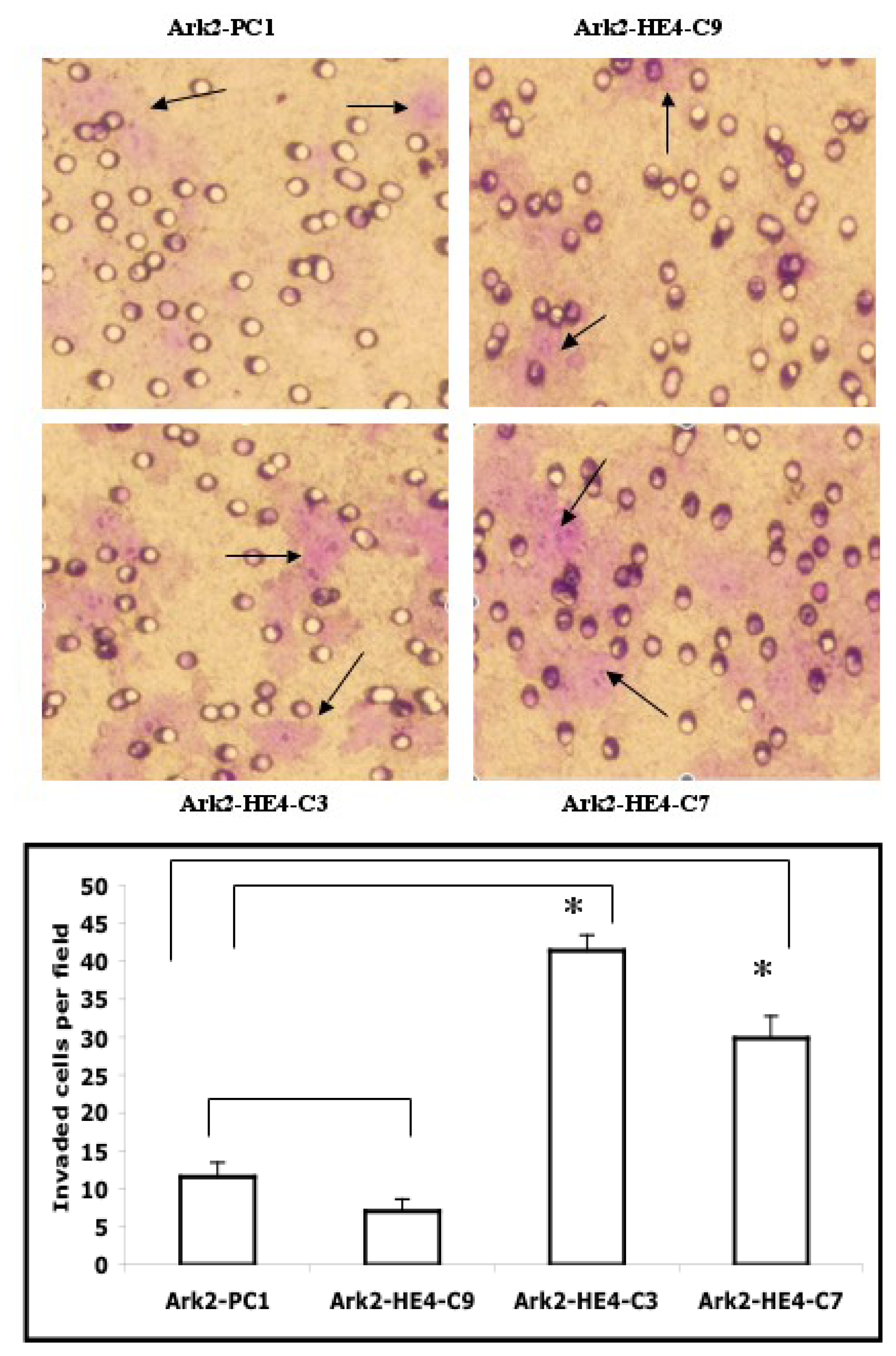
2.3. HE4 Overexpression Enhances Invasive Ability and Anchorage-Independent Growth in EC Cells
2.4. HE4 Overexpression Promotes EC Tumor Growth in Vivo
2.5. Determination of HE4 mRNA and Protein Levels in EC Tumor Tissues
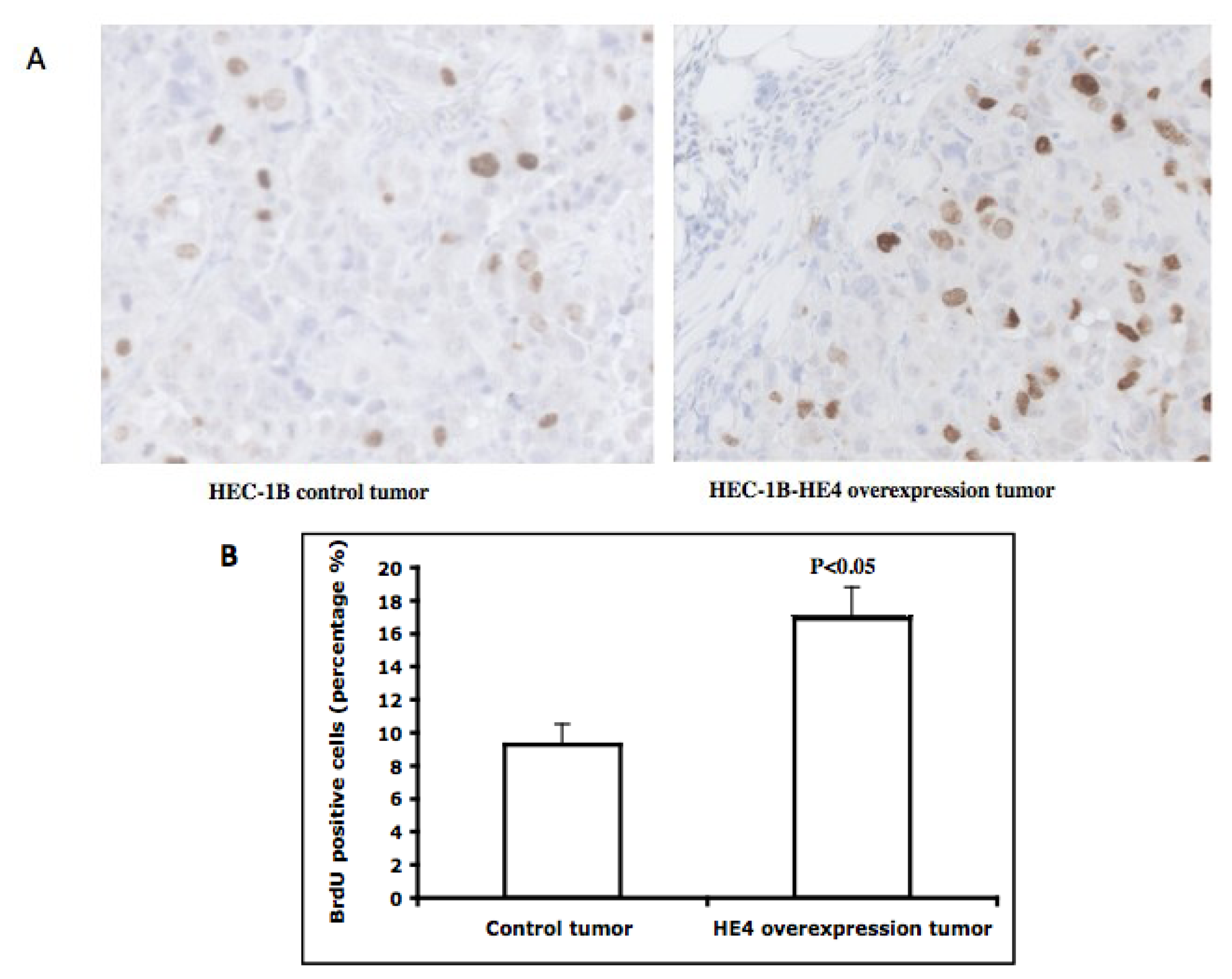
2.6. Tumors Overexpressing HE4 Have a Higher Number of Cells in the S-Phase
3. Discussion
4. Experimental Section
4.1. Cell Culture
4.2. Construction of Plasmid Vectors
4.3. RNA Isolation and Reverse Transcription
4.4. Real-Time PCR Analyses of mRNA Levels
4.5. In Vitro Proliferation Assay
4.6. Matrigel Invasion Assay
4.7. Soft Agar Colony Formation Assay
4.8. Mouse Xenograft Model
4.9. Detection of HE4 Expression in Tumor Tissues
4.10. Tumor Cell Proliferation Assay in Vivo
Supplementary Information
ijms-14-06026-s001.docxAcknowledgements
Conflict of Interest
References
- National Cancer Institute. SEER Stat Fact Sheets: Corpus and Uterus, NOS. Available online: http://seer.cancer.gov/statfacts/html/corp.html accessed on 12 March 2013.
- Bingle, L.; Singleton, V.; Bingle, C.D. The putative ovarian tumour marker gene HE4 (WFDC2), is expressed in normal tissues and undergoes complex alternative splicing to yield multiple protein isoforms. Oncogene 2002, 21, 2768–2773. [Google Scholar]
- Hellstrom, I.; Raycraft, J.; Hayden-Ledbetter, M.; Ledbetter, J.A.; Schummer, M.; McIntosh, M.; Drescher, C.; Urban, N.; Hellstrom, K.E. The HE4 (WFDC2) protein is a biomarker for ovarian carcinoma. Cancer Res 2003, 63, 3695–3700. [Google Scholar]
- Drapkin, R.; von Horsten, H.H.; Lin, Y.; Mok, S.C.; Crum, C.P.; Welch, W.R.; Hecht, J.L. Human epididymis protein 4 (HE4) is a secreted glycoprotein that is overexpressed by serous and endometrioid ovarian carcinomas. Cancer Res 2005, 65, 2162–2169. [Google Scholar]
- Jiang, S.-W.; Dowdy, S.C.; Fu, A.; Attewell, J.R.; Kalogera, E.; Drapkin, R.; Podratz, K.C.; Broaddus, R.; Li, J. HE4 Transcription- and splice variants-specific expression in 8 endometrial cancer and correlation with patient survival. Int. J. Mol. Sci 2013. submitted for publication. [Google Scholar]
- Galgano, M.T.; Hampton, G.M.; Frierson, H.F., Jr. Comprehensive analysis of HE4 expression in normal and malignant human tissues. Mod. Pathol. 2006, 19, 847–853. [Google Scholar]
- DeSouza, L.V.; Grigull, J.; Ghanny, S.; Dube, V.; Romaschin, A.D.; Colgan, T.J.; Siu, K.W. Endometrial carcinoma biomarker discovery and verification using differentially tagged clinical samples with multidimensional liquid chromatography and tandem mass spectrometry. Mol. Cell Proteomics 2007, 6, 1170–1182. [Google Scholar]
- Li, H.; DeSouza, L.V.; Ghanny, S.; Li, W.; Romaschin, A.D.; Colgan, T.J.; Siu, K.W. Identification of candidate biomarker proteins released by human endometrial and cervical cancer cells using two-dimensional liquid chromatography/tandem mass spectrometry. J. Proteome Res 2007, 6, 2615–2622. [Google Scholar]
- Moore, R.G.; Brown, A.K.; Miller, M.C.; Badgwell, D.; Lu, Z.; Allard, W.J.; Granai, C.O.; Bast, R.C., Jr; Lu, K. Utility of a novel serum tumor biomarker HE4 in patients with endometrioid adenocarcinoma of the uterus. Gynecol. Oncol. 2008, 110, 196–201. [Google Scholar]
- Bignotti, E.; Ragnoli, M.; Zanotti, L.; Calza, S.; Falchetti, M.; Lonardi, S.; Bergamelli, S.; Bandiera, E.; Tassi, R.A.; Romani, C.; et al. Diagnostic and prognostic impact of serum HE4 detection in endometrial carcinoma patients. Br. J. Cancer 2011, 104, 1418–1425. [Google Scholar] [Green Version]
- Kamei, M.; Yamashita, S.; Tokuishi, K.; Hashioto, T.; Moroga, T.; Suehiro, S.; Ono, K.; Miyawaki, M.; Takeno, S.; Yamamoto, S.; Kawahara, K. HE4 expression can be associated with lymph node metastases and disease-free survival in breast cancer. Anticancer Res 2010, 30, 4779–4783. [Google Scholar]
- Bouchard, D.; Morisset, D.; Bourbonnais, Y.; Tremblay, G.M. Proteins with whey-acidic-protein motifs and cancer. Lancet Oncol 2006, 7, 167–174. [Google Scholar]
- Micci, F.; Teixeira, M.R.; Haugom, L.; Kristensen, G.; Abeler, V.M.; Heim, S. Genomic aberrations in carcinomas of the uterine corpus. Genes Chromosomes Cancer 2004, 40, 229–246. [Google Scholar]
- Scholler, N.; Crawford, M.; Sato, A.; Drescher, C.W.; O’Briant, K.C.; Kiviat, N.; Anderson, G.L.; Urban, N. Bead-based ELISA for validation of ovarian cancer early detection markers. Clin. Cancer Res 2006, 12, 2117–2124. [Google Scholar]
- Scholler, N.; Lowe, K.A.; Bergan, L.A.; Kampani, A.V.; Ng, V.; Forrest, R.M.; Thorpe, J.D.; Gross, J.A.; Garvik, B.M.; Drapkin, R.; et al. Use of yeast-secreted in vivo biotinylated recombinant antibodies (Biobodies) in bead-based ELISA. Clin. Cancer Res 2008, 14, 2647–2655. [Google Scholar]
- Moore, R.G.; Brown, A.K.; Miller, M.C.; Skates, S.; Allard, W.J.; Verch, T.; Steinhoff, M.; Messerlian, G.; DiSilvestro, P.; Granai, C.O.; Bast, R.C., Jr. The use of multiple novel tumor biomarkers for the detection of ovarian carcinoma in patients with a pelvic mass. Gynecol. Oncol 2008, 108, 402–408. [Google Scholar]
- Hellstrom, I.; Hellstrom, K.E. SMRP and HE4 as biomarkers for ovarian carcinoma when used alone and in combination with CA125 and/or each other. Adv. Exp. Med. Biol 2008, 622, 15–22. [Google Scholar]
- Moore, R.G.; Jabre-Raughley, M.; Brown, A.K.; Robison, K.M.; Miller, M.C.; Allard, W.J.; Kurman, R.J.; Bast, R.C.; Skates, S.J. Comparison of a novel multiple marker assay vs the risk of Malignancy Index for the prediction of epithelial ovarian cancer in patients with a pelvic mass. Am. J. Obstet. Gynecol 2010, 203, 228e1–6. [Google Scholar]
- Havrilesky, L.J.; Whitehead, C.M.; Rubatt, J.M.; Cheek, R.L.; Groelke, J.; He, Q.; Malinowski, D.P.; Fischer, T.J.; Berchuck, A. Evaluation of biomarker panels for early stage ovarian cancer detection and monitoring for disease recurrence. Gynecol. Oncol 2008, 110, 374–382. [Google Scholar]
- Yurkovetsky, Z.; Skates, S.; Lomakin, A.; Nolen, B.; Pulsipher, T.; Modugno, F.; Marks, J.; Godwin, A.; Gorelik, E.; Jacobs, I.; et al. Development of a multimarker assay for early detection of ovarian cancer. J. Clin. Oncol 2010, 28, 2159–2166. [Google Scholar]
- Huhtinen, K.; Suvitie, P.; Hiissa, J.; Junnila, J.; Huvila, J.; Kujari, H.; Setala, M.; Harkki, P.; Jalkanen, J.; Fraser, J.; et al. Serum HE4 concentration differentiates malignant ovarian tumours from ovarian endometriotic cysts. Br. J. Cancer 2009, 100, 1315–1319. [Google Scholar]
- Park, Y.; Kim, Y.; Lee, E.Y.; Lee, J.H.; Kim, H.S. Reference ranges for HE4 and CA125 in a large Asian population by automated assays and diagnostic performances for ovarian cancer. Int. J. Cancer 2012, 130, 1136–1144. [Google Scholar]
- Ruggeri, G.; Bandiera, E.; Zanotti, L.; Belloli, S.; Ravaggi, A.; Romani, C.; Bignotti, E.; Tassi, R.A.; Tognon, G.; Galli, C.; et al. HE4 and epithelial ovarian cancer: Comparison and clinical evaluation of two immunoassays and a combination algorithm. Clin. Chim. Acta 2011, 412, 1447–1453. [Google Scholar]
- Shah, C.A.; Lowe, K.A.; Paley, P.; Wallace, E.; Anderson, G.L.; McIntosh, M.W.; Andersen, M.R.; Scholler, N.; Bergan, L.A.; Thorpe, J.D.; et al. Influence of ovarian cancer risk status on the diagnostic performance of the serum biomarkers mesothelin, HE4, and CA125. Cancer Epidemiol. Biomarkers Prev 2009, 18, 1365–1372. [Google Scholar]
- Anastasi, E.; Granato, T.; Marchei, G.G.; Viggiani, V.; Colaprisca, B.; Comploj, S.; Reale, M.G.; Frati, L.; Midulla, C. Ovarian tumor marker HE4 is differently expressed during the phases of the menstrual cycle in healthy young women. Tumour Biol 2010, 31, 411–415. [Google Scholar]
- Anderson, G.L.; McIntosh, M.; Wu, L.; Barnett, M.; Goodman, G.; Thorpe, J.D.; Bergan, L.; Thornquist, M.D.; Scholler, N.; Kim, N.; et al. Assessing lead time of selected ovarian cancer biomarkers: A nested case-control study. J. Natl. Cancer Inst 2010, 102, 26–38. [Google Scholar]
- Granato, T.; Midulla, C.; Longo, F.; Colaprisca, B.; Frati, L.; Anastasi, E. Role of HE4, CA72.4, and CA125 in monitoring ovarian cancer. Tumour Biol 2012, 33, 1335–1339. [Google Scholar]
- Yoshida, N.; Egami, H.; Yamashita, J.; Takai, E.; Tamori, Y.; Fujino, N.; Kitaoka, M.; Schalkwijk, J.; Ogawa, M. Immunohistochemical expression of SKALP/elafin in squamous cell carcinoma of human lung. Oncol. Rep 2002, 9, 495–501. [Google Scholar]
- Westin, U.; Nystrom, M.; Ljungcrantz, I.; Eriksson, B.; Ohlsson, K. The presence of elafin, SLPI, IL1-RA and STNFalpha RI in head and neck squamous cell carcinomas and their relation to the degree of tumour differentiation. Mediators Inflamm 2002, 11, 7–12. [Google Scholar]
- Clauss, A.; Ng, V.; Liu, J.; Piao, H.; Russo, M.; Vena, N.; Sheng, Q.; Hirsch, M.S.; Bonome, T.; Matulonis, U.; et al. Overexpression of elafin in ovarian carcinoma is driven by genomic gains and activation of the nuclear factor kappaB pathway and is associated with poor overall survival. Neoplasia 2010, 12, 161–172. [Google Scholar]
- Kluger, H.M.; Chelouche Lev, D.; Kluger, Y.; McCarthy, M.M.; Kiriakova, G.; Camp, R.L.; Rimm, D.L.; Price, J.E. Using a xenograft model of human breast cancer metastasis to find genes associated with clinically aggressive disease. Cancer Res 2005, 65, 5578–5587. [Google Scholar]
- Devoogdt, N.; Hassanzadeh Ghassabeh, G.; Zhang, J.; Brys, L.; de Baetselier, P.; Revets, H. Secretory leukocyte protease inhibitor promotes the tumorigenic and metastatic potential of cancer cells. Proc. Natl. Acad. Sci. USA 2003, 100, 5778–5782. [Google Scholar]
- Idoji, Y.; Watanabe, Y.; Yamashita, A.; Yamanishi, K.; Nishiguchi, S.; Shimada, K.; Yasunaga, T.; Yamanishi, H. In silico study of whey-acidic-protein domain containing oral protease inhibitors. Int. J. Mol. Med 2008, 21, 461–468. [Google Scholar]
- Kalogera, E.; Scholler, N.; Powless, C.; Weaver, A.; Drapkin, R.; Li, J.; Jiang, S.W.; Podratz, K.; Urban, N.; Dowdy, S.C. Correlation of serum HE4 with tumor size and myometrial invasion in endometrial cancer. Gynecol. Oncol 2012, 124, 270–275. [Google Scholar]
- Lu, K.H.; Patterson, A.P.; Wang, L.; Marquez, R.T.; Atkinson, E.N.; Baggerly, K.A.; Ramoth, L.R.; Rosen, D.G.; Liu, J.; Hellstrom, I.; et al. Selection of potential markers for epithelial ovarian cancer with gene expression arrays and recursive descent partition analysis. Clin. Cancer Res 2004, 10, 3291–3300. [Google Scholar]
- Li, J.; Dowdy, S.; Tipton, T.; Podratz, K.; Lu, W.G.; Xie, X.; Jiang, S.W. HE4 as a biomarker for ovarian and endometrial cancer management. Expert Rev. Mol. Diagn 2009, 9, 555–566. [Google Scholar]
- Scott, R.E.; Wille, J.J., Jr; Wier, M.L. Mechanisms for the initiation and promotion of carcinogenesis: A review and a new concept. Mayo Clin. Proc. 1984, 59, 107–117. [Google Scholar]
- Sanchez-Tillo, E.; Liu, Y.; de Barrios, O.; Siles, L.; Fanlo, L.; Cuatrecasas, M.; Darling, D.S.; Dean, D.C.; Castells, A.; Postigo, A. EMT-activating transcription factors in cancer: Beyond EMT and tumor invasiveness. Cell Mol. Life Sci 2012, 69, 3429–3456. [Google Scholar]
- Yang, J.; Zhu, J.; Sun, D.; Ding, A. Suppression of macrophage responses to bacterial lipopolysaccharide (LPS) by secretory leukocyte protease inhibitor (SLPI) is independent of its anti-protease function. Biochim. Biophys. Acta 2005, 1745, 310–317. [Google Scholar]
- McNeely, T.B.; Shugars, D.C.; Rosendahl, M.; Tucker, C.; Eisenberg, S.P.; Wahl, S.M. Inhibition of human immunodeficiency virus type 1 infectivity by secretory leukocyte protease inhibitor occurs prior to viral reverse transcription. Blood 1997, 90, 1141–1149. [Google Scholar]
- Van Dekken, H.; Alers, J.C.; Riegman, P.H.; Rosenberg, C.; Tilanus, H.W.; Vissers, K. Molecular cytogenetic evaluation of gastric cardia adenocarcinoma and precursor lesions. Am. J. Pathol 2001, 158, 1961–1967. [Google Scholar]
- Jin, F.; Dowdy, S.C.; Xiong, Y.; Eberhardt, N.L.; Podratz, K.C.; Jiang, S.W. Up-regulation of DNA methyltransferase 3B expression in endometrial cancers. Gynecol. Oncol 2005, 96, 531–538. [Google Scholar]
- Zeng, H.; Datta, K.; Neid, M.; Li, J.; Parangi, S.; Mukhopadhyay, D. Requirement of different signaling pathways mediated by insulin-like growth factor-I receptor for proliferation, invasion, and VPF/VEGF expression in a pancreatic carcinoma cell line. Biochem. Biophys. Res. Commun 2003, 302, 46–55. [Google Scholar]


© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, J.; Chen, H.; Mariani, A.; Chen, D.; Klatt, E.; Podratz, K.; Drapkin, R.; Broaddus, R.; Dowdy, S.; Jiang, S.-W. HE4 (WFDC2) Promotes Tumor Growth in Endometrial Cancer Cell Lines. Int. J. Mol. Sci. 2013, 14, 6026-6043. https://doi.org/10.3390/ijms14036026
Li J, Chen H, Mariani A, Chen D, Klatt E, Podratz K, Drapkin R, Broaddus R, Dowdy S, Jiang S-W. HE4 (WFDC2) Promotes Tumor Growth in Endometrial Cancer Cell Lines. International Journal of Molecular Sciences. 2013; 14(3):6026-6043. https://doi.org/10.3390/ijms14036026
Chicago/Turabian StyleLi, Jinping, Haibin Chen, Andrea Mariani, Dong Chen, Edward Klatt, Karl Podratz, Ronny Drapkin, Russell Broaddus, Sean Dowdy, and Shi-Wen Jiang. 2013. "HE4 (WFDC2) Promotes Tumor Growth in Endometrial Cancer Cell Lines" International Journal of Molecular Sciences 14, no. 3: 6026-6043. https://doi.org/10.3390/ijms14036026
APA StyleLi, J., Chen, H., Mariani, A., Chen, D., Klatt, E., Podratz, K., Drapkin, R., Broaddus, R., Dowdy, S., & Jiang, S.-W. (2013). HE4 (WFDC2) Promotes Tumor Growth in Endometrial Cancer Cell Lines. International Journal of Molecular Sciences, 14(3), 6026-6043. https://doi.org/10.3390/ijms14036026




