Analysis of Expressed Sequence Tags from Chinese Bayberry Fruit (Myrica rubra Sieb. and Zucc.) at Different Ripening Stages and Their Association with Fruit Quality Development
Abstract
:1. Introduction
2. Results and Discussion
2.1. Changes in Quality Attributes during Fruit Development and Ripening
2.2. Analysis of Bayberry Fruit cDNA Libraries and EST Library
2.3. Functional Annotation of Bayberry EST
2.4. Analysis of Highly Expressed Genes from “Biqi” Fruit ESTs
2.5. Identification of Genes Regulating Bayberry Fruit Ripening and Senescence
2.6. Identification of Genes Associated with Quality Formation in Ripening Bayberry Fruit
3. Experimental Section
3.1. Plant Materials
3.2. Fruit Surface Color Measurement
3.3. Total Soluble Solid (TSS) and Titratable Acidity (TA)
3.4. Determination of Total Anthocyanin Contents
3.5. Determination of Alcohol Contents
3.6. Construction of Bayberry Fruit cDNA Library
3.7. Bioinformatics Analysis
3.8. Statistic Analysis
4. Conclusions
Acknowledgments
Conflict of Interest
References
- Chen, K.S.; Xu, C.J.; Zhang, B.; Ferguson, I.B. Red bayberry: Botany and horticulture. Hort. Rev 2004, 30, 83–114. [Google Scholar]
- Zhang, S.M.; Gao, Z.S.; Xu, C.J.; Chen, K.S.; Wang, G.Y.; Zheng, J.T.; Lu, T. Genetic diversity of Chinese bayberry (Myrica rubra Sieb. et Zucc.) accessions revealed by amplified fragment length polymorphism. HortScience 2009, 44, 487–491. [Google Scholar]
- Chen, K.S.; Xu, C.J.; Zhang, B.; Ferguson, I.B. Myrica rubra (Red Bayberry). In The Encyclopedia of Fruit and Nuts; Janick, J., Paull, R.E., Eds.; Cambridge University Press: Cambridge, UK, 2008; pp. 522–526. [Google Scholar]
- Zhang, W.S.; Li, X.; Zheng, J.T.; Wang, G.Y.; Sun, C.D.; Ferguson, I.B.; Chen, K.S. Bioactive components and antioxidant capacity of Chinese bayberry (Myrica rubra Sieb. and Zucc.) fruit in relation to fruit maturity and postharvest storage. Eur. Food Res. Technol 2008, 227, 1091–1097. [Google Scholar]
- Zhang, B.; Kang, M.; Xie, Q.; Xu, B.; Sun, C.; Chen, K.; Wu, Y. Anthocyanins from Chinese bayberry extract protect β cells from oxidative stress-mediated injury via HO-1 upregulation. J. Agric. Food Chem 2011, 59, 537–545. [Google Scholar]
- Sun, C.D.; Zhang, B.; Zhang, J.K.; Xu, C.J.; Wu, Y.L.; Li, X.; Chen, K.S. Cyanidin-3-glucoside-rich extract from Chinese bayberry fruit protects pancreatic β cells and ameliorates hyperglycemia in streptozotocin-induced diabetic mice. J. Med. Food 2012, 15, 288–298. [Google Scholar]
- Sun, C.D.; Zheng, Y.X.; Chen, Q.J.; Tang, X.L.; Jiang, M.; Zhang, J.K.; Li, X.; Chen, K.S. Purification and anti-tumor activity of cyanidin-3-O-glucoside from Chinese bayberry fruit. Food Chem 2012, 131, 1287–1294. [Google Scholar]
- Joyce, D.C. Evaluation of fresh red bayberry (Myrica rubra) fruit acceptance. N. Z. J. Crop Hort 2007, 35, 125–128. [Google Scholar]
- Karp, D. From China, only in a bottle, a berry with an alluring name. The New York Times 2007. [Google Scholar]
- Karp, D. Myrica rubra, a fruit of many names. Fruit Gardener 2008, 40, 30–33. [Google Scholar]
- Zhang, W.S.; Chen, K.S.; Zhang, B.; Sun, C.D.; Cai, C.; Zhou, C.H.; Xu, W.P.; Zhang, W.Q.; Ferguson, I.B. Postharvest responses of Chinese bayberry fruit. Postharvest Biol. Technol 2005, 37, 241–251. [Google Scholar]
- Zhang, W.S.; Li, X.; Wang, X.X.; Wang, G.Y.; Zheng, J.T.; Abeysinghe, D.C.; Ferguson, I.B.; Chen, K.S. Ethanol vapour treatment alleviates postharvest decay and maintains fruit quality in Chinese bayberry fruit. Postharvest Biol. Technol 2007, 46, 195–198. [Google Scholar]
- Yang, Z.F.; Heng, Y.H.; Cao, S.F. Effect of high oxygen atmosphere storage on quality, antioxidant enzymes, and DPPH-radical scavenging activity of Chinese bayberry fruit. J. Agric. Food Chem 2009, 57, 176–181. [Google Scholar]
- Wang, K.T.; Jin, P.; Tang, S.S.; Shang, H.T.; Rui, H.J.; Di, H.T.; Cai, Y.; Zheng, Y.H. Improved control of postharvest decay in Chinese bayberries by a combination treatment of ethanol vapor with hot air. Food Control 2011, 22, 82–87. [Google Scholar]
- Alba, R.; Fei, Z.; Payton, P.; Liu, Y.; Moore, S.L.; Debbie, P.; Cohn, J.; Ascenzo, M.D.; Gordon, J.S.; Rose, J.K.C.; et al. ESTs, cDNA microarrays, and gene expression profiling: Tools for dissecting plant physiology and development. Plant J 2004, 39, 697–714. [Google Scholar]
- Pashley, C.H.; Ellis, J.R.; McCauley, D.E.; Burke, J.M. EST databases as a source for molecular markers: Lessons from Helianthus. J. Hered 2006, 97, 381–388. [Google Scholar]
- Rudd, S. Expressed sequence tags: Alternative or complement to whole genome sequences? Trends Plant Sci 2003, 8, 321–329. [Google Scholar]
- Cercós, M.; Soler, G.; Iglesias, D.J.; Gadea, J.; Forment, J.; Talón, M. Global analysis of gene expression during development and ripening of citrus fruit flesh. A proposed mechanism for citric acid utilization. Plant Mol. Biol 2006, 62, 513–527. [Google Scholar]
- Terol, J.; Conesa, A.; Colmenero, J.M.; Cercos, M.; Tadeo, F.; Agustí, J.; Alós, E.; Andres, F.; Soler, G.; Brumos, J.; et al. Analysis of 13,000 unique Citrus clusters associated with fruit quality, production and salinity tolerance. BMC Genomics 2007, 8, 31. [Google Scholar]
- Janssen, B.J.; Thodey, K.; Schaffer, R.J.; Alba, R.; Balakrishnan, L.; Bishop, R.; Bowen, J.H.; Crowhurst, R.N.; Gleave, A.P.; Ledger, S.; et al. Global gene expression analysis of apple fruit development from the floral bud to ripe fruit. BMC Plant Biol 2008, 8, 16. [Google Scholar]
- Newcomb, R.D.; Crowhurst, R.N.; Gleave, A.P.; Rikkerink, E.H.A.; Allan, A.C.; Beuning, L.L.; Bowen, J.H.; Gera, E.; Jamieson, K.R.; Janssen, B.J.; et al. Analyses of expressed sequence tags from apple. Plant Physiol 2006, 141, 147–166. [Google Scholar]
- Park, S.; Sugimoto, N.; Larson, M.D.; Beaudry, R.; Nocker, S. Identification of genes with potential roles in apple fruit development and biochemistry through large-scale statistical analysis of expressed sequence tags. Plant Physiol 2006, 141, 811–824. [Google Scholar]
- Da Silva, F.G.; Iandolino, A.; Al-Kayal, F.; Bohlmann, M.C.; Cushman, M.A.; Lim, H.; Ergul, A.; Figueroa, R.; Kabuloglu, E.K.; Osborne, C.; et al. Characterizing the grape transcriptome. Analysis of expressed sequence tags from multiple Vitis species and development of a compendium of gene expression during berry development. Plant Physiol 2005, 139, 574–597. [Google Scholar]
- Wang, X.C.; Guo, L.; Shangguan, L.F.; Wang, C.; Yang, G.; Qu, S.C.; Fang, J.G. Analysis of expressed sequence tags from grapevine flower and fruit and development of simple sequence repeat markers. Mol. Biol. Rep 2012, 39, 6825–6834. [Google Scholar]
- Crowhurst, R.N.; Gleave, A.P.; MacRae, E.A.; Ampomah-Dwamena, C.; Atkinson, R.G.; Beuning, L.L.; Bulley, S.M.; Chagne, D.; Marsh, K.B.; Matich, A.J.; et al. Analysis of expressed sequence tags from Actinidia: Applications of a cross species EST database for gene discovery in the areas of flavor, health, color and ripening. BMC Genomics 2008, 9, 351. [Google Scholar]
- Ogundiwin, E.A.; Martí, C.; Forment, J.; Pons, C.; Granell, A.; Gradziel, T.M.; Peace, C.P.; Crisosto, C.H. Development of ChillPeach genomic tools and identification of cold-responsive genes in peach fruit. Plant Mol. Biol 2008, 68, 379–397. [Google Scholar]
- Niu, S.S.; Xu, C.J.; Zhang, W.S.; Zhang, B.; Li, X.; Lin-Wang, K.; Ferguson, I.B.; Allan, A.C.; Chen, K.S. Coordinated regulation of anthocyanin biosynthesis in Chinese bayberry (Myrica rubra) fruit by a R2R3 MYB transcription factor. Planta 2010, 231, 887–899. [Google Scholar]
- Ke, D.; Kader, A. Tolerance of “Valencia” oranges to controlled atmospheres as determined by physiological responses and quality attributes. J. Am. Soc. Hortic. Sci 1990, 115, 779–783. [Google Scholar]
- Xu, C.J.; Zhu, C.Q.; Gao, Z.S.; Chen, K.S. Application of fruit crop ESTs in studies on fruit development and ripening. J. Fruit Sci 2009, 26, 353–360. [Google Scholar]
- Wang, W.J.; Chen, K.S.; Xu, C.J. DNA quantification using EvaGreen and a real-time PCR instrument. Anal. Biochem 2006, 356, 303–305. [Google Scholar]
- The UniProt Consortium. The universal protein resource (UniProt). Nucleic Acids Res 2008, 36, D190–D195.
- UniProt. Available online: http://beta.uniprot.org/?tab=blast accessed on 12 December 2012.
- Bevan, M.; Bancroft, I.; Bent, E.; Love, K.; Goodman, H.; Dean, C.; Bergkamp, R.; Dirkse, W.; van Staveren, M.; Stiekema, W.; et al. Analysis of 1.9 Mb of contiguous sequence from chromosome 4 of Arabidopsis thaliana. Nature 1998, 391, 485–488. [Google Scholar]
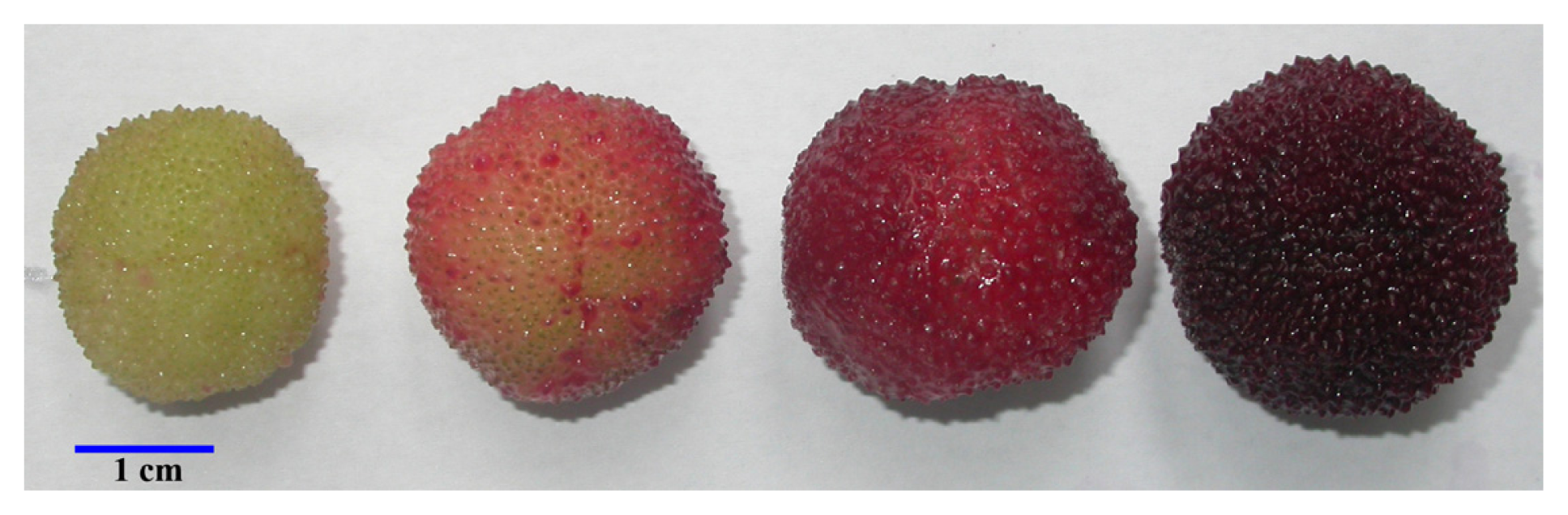

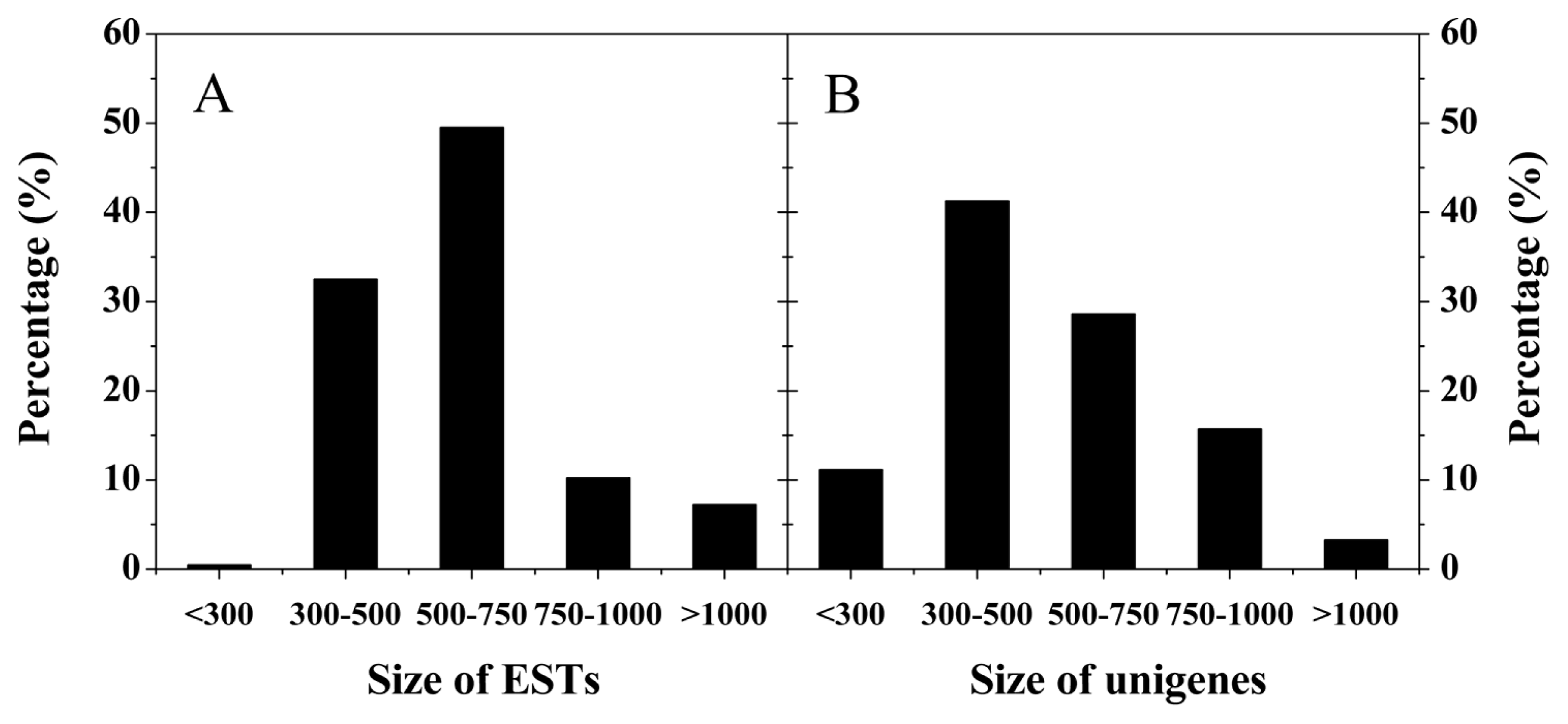
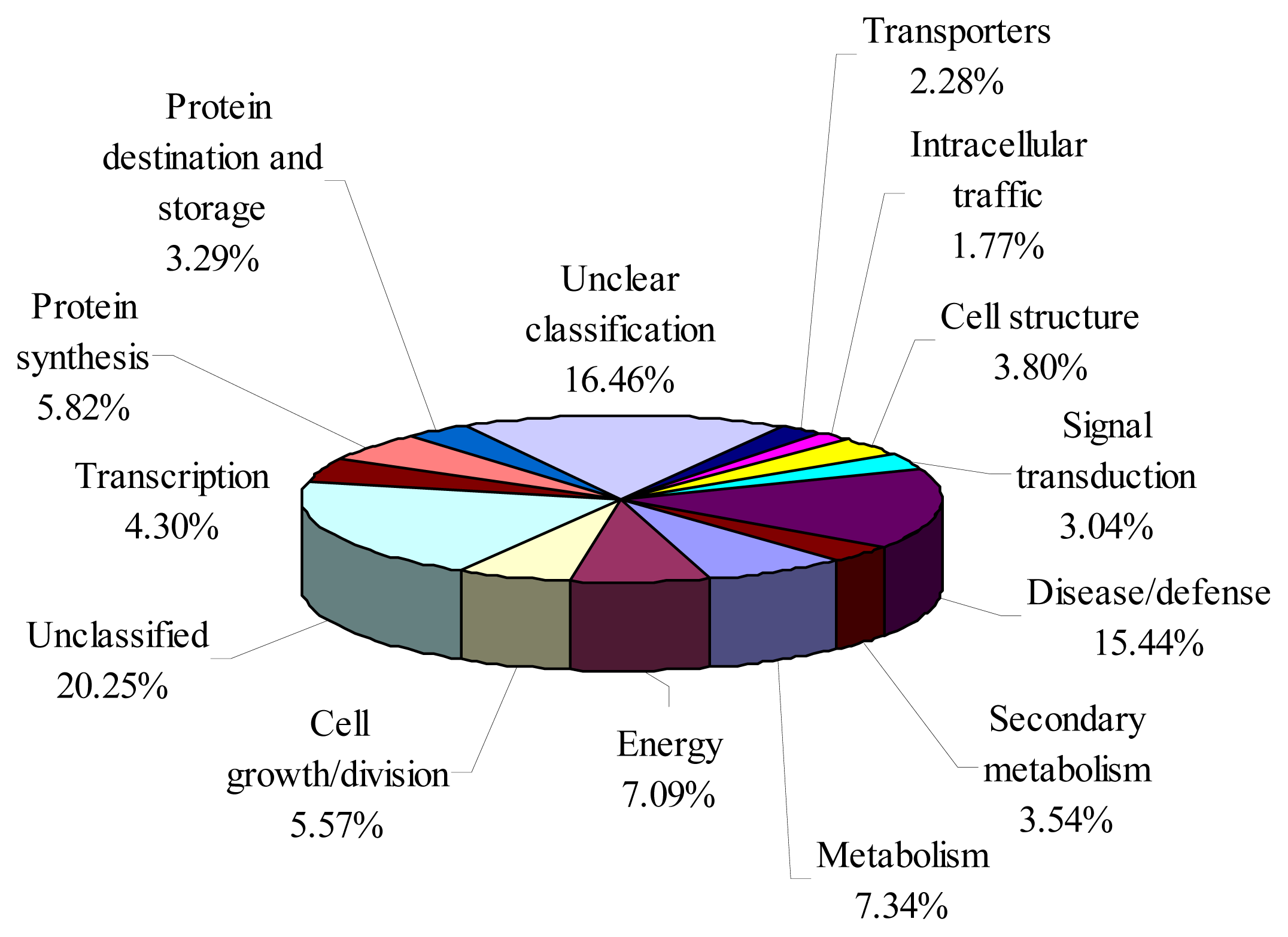

| Quality attributes | MG | PR | RR | FR |
|---|---|---|---|---|
| Length * (mm) | 18.86 ± 0.68 | 20.00 ± 0.90 | 25.44 ± 1.75 | 26.47 ± 1.19 |
| Diameter * (mm) | 19.92 ± 0.42 | 21.65 ± 1.12 | 27.19 ± 1.06 | 28.02 ± 1.16 |
| Weight * (g) | 3.91 ± 0.42 | 4.90 ± 0.63 | 9.45 ± 1.30 | 11.29 ± 1.47 |
| Color * (CIRG) | 1.18 ± 0.14 | 2.59 ± 0.35 | 3.61 ± 0.29 | 6.01 ± 1.03 |
| TSS * (∘Brix) | 8.76 ± 0.23 | 9.97 ± 0.44 | 11.80 ± 0.75 | 12.95 ± 0.71 |
| TA ** (mg/g FW) | 47.18 ± 0.55 | 30.87 ± 0.43 | 6.87 ± 0.95 | 6.65 ± 0.80 |
| Total anthocyanins ** (mg/100 g FW) | 0.25 ± 0.12 | 2.67 ± 0.37 | 63.06 ± 16.01 | 88.03 ± 9.24 |
| Ethanol ** (μmol/100 g FW) | 0.76 ± 0.46 | 3.21 ± 0.52 | 6.96 ± 2.53 | 62.92 ± 4.99 |
| Acetaldehyde ** (μmol/100 g FW) | 1.33 ± 0.05 | 1.46 ± 0.07 | 1.48 ± 0.10 | 3.38 ± 0.24 |
| Library parameter | MG | PR | RR | FR |
|---|---|---|---|---|
| Titer of original cDNA library | 2.01 × 105 | 1.49 × 105 | 3.92 × 105 | 1.18 × 106 |
| Titer of EST library | 3.13 × 109 | 3.04 × 109 | 3.53 × 109 | 3.95 × 109 |
| Recombinant rate | 94.74% | 100% | 94.74% | 94.74% |
| Average length of EST (bp) | 701 | 597 | 565 | 1272 |
| Number of TCs | 14 | 37 | 28 | 53 |
| Number of singletons | 45 | 202 | 36 | 79 |
| NO. | Unigene code | Highest BLAST match based on deduced amino acid sequences | Numbers of ESTs obtained | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Deduced protein | Plant | Identity (%) | MG | PR | RR | FR | Total | ||
| 1 | MRU00001 | Metallothionein-like protein type 3 (AF268393.1) | Musa acuminata | 77 | 150 | 64 | 117 | 77 | 408 |
| 2 | MRU00012 | Phytocystatin cysteine proteinase inhibitor 1 (AY390352.1) | Actinidia deliciosa | 74 | 19 | 11 | 48 | 65 | 143 |
| 3 | MRU00135 | Phase-change related protein (AJ271778.1) | Quercus robur | 59 | 122 | 2 | 1 | 0 | 125 |
| 4 | MRU00040 | Thaumatin-like protein (AJ871175.2) | Actinidia deliciosa | 79 | 65 | 25 | 11 | 13 | 114 |
| 5 | MRU00003 | GAST-like (cold-regulated LTCOR12)(EU717678.1) | Litchi chinensis | 71 | 3 | 8 | 45 | 20 | 76 |
| 6 | MRU00007 | Metallothionein-like protein type 2 (L02306.1) | Ricinus communis | 82 | 25 | 2 | 32 | 17 | 76 |
| 7 | MRU00008 | Flavonoid 3′-hydroxylase (F3′H) (DQ218417.1) | Gerbera hybrida | 82 | 1 | 24 | 12 | 20 | 57 |
| 8 | MRU00032 | Metallothionein-like protein class II (AJ130886.1) | Fagus sylvatica | 77 | 28 | 15 | 8 | 5 | 56 |
| 9 | MRU00026 | Putative tropinone reductase (AK175221.1) | Arabidopsis thaliana | 72 | 1 | 3 | 25 | 25 | 54 |
| 10 | MRU00028 | Small molecular heat shock protein 17.5 ( EF100453.1) | Nelumbo nucifera | 89 | 0 | 3 | 22 | 6 | 31 |
| Processes involved | Unigene code | Highest BLAST match based on deduced amino acid sequences | Numbers of ESTs obtained | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Deduced protein | Plant | Identity (%) | MG | PR | RR | FR | Total | ||
| Ethylene biosynthesis | MRU00059 | 1-aminocyclopropane-1-carboxylic acid oxidase (ACO) (AY211549.1) | Vitis vinifera | 89 | 1 | 1 | |||
| Ethylene signal transduction | MRU00110 | Ethylene response factor (ERF) (AJ606475.1) | Fagus sylvatica | 75 | 1 | 1 | |||
| MRU00129 | Ethylene-insensitive 3-like protein (EIL) (GU981740.1) | Citrus aurantium | 64 | 1 | 1 | ||||
| ABA biosynthesis | MRU00088 | 9-cis-epoxycarotenoid dioxygenase (NCED) (DQ309329.1) | Citrus clementina | 86 | 1 | 1 | |||
| Senescence and apoptosis | MRU00242 | Ascorbate peroxidase (HQ661034.1) | Dimocarpus longan | 94 | 1 | 1 | |||
| MRU00387 | Senescence-related protein (FJ014471.1) | Camellia sinensis | 77 | 1 | 1 | ||||
| MRU00062 | Senescence-associated protein (EU961590.1) | Zea mays | 71 | 1 | 1 | ||||
| MRU00050 | Cytochrome C reductase (8.0 KD subunit) (X79274.1) | Solanum tuberosum | 72 | 2 | 1 | 3 | 6 | ||
| Fruit quality attributes | Unigene code | Highest BLAST match based on deduced amino acid sequences | Numbers of ESTs obtained | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Deduced protein | Plant | Identity (%) | MG | PR | RR | FR | Total | ||
| Color | MRU00008 | Flavonoid 3′-hydroxylase (F3′H) (DQ218417.1) | Gerbera hybrida | 82 | 1 | 24 | 12 | 20 | 57 |
| MRU00113 | Myb transcription factor (HQ661039.1) | Dimocarpus longan | 76 | 3 | 2 | 5 | |||
| MRU00098 | UDP-glucose:flavonoid 3-O-glucosyltransferase (UFGT) (AY695815.1) | Fragaria ananassa | 59 | 1 | 1 | ||||
| Flavor | MRU00009 | Vacuolar ATP synthase subunit G1 (FJ906831.1) | Jatropha curcas | 81 | 3 | 5 | 8 | ||
| MRU00061 | 6-phosphogluconate dehydrogenase (EU815934.1) | Cucumis sativus | 87 | 1 | 1 | 1 | 3 | ||
| MRU00369 | Alpha glucosidase-like protein (AB240194.1) | Hordeum vulgare | 68 | 1 | 1 | ||||
| MRU00046 | β 1–3 glucanase (EU676805.1) | Vitis riparia | 75 | 2 | 2 | ||||
| Texture | MRU00130 | Polygalacturonase inhibitor-like protein (PGIP) (AJ515557.1) | Cicer arietinum | 88 | 1 | 1 | |||
| MRU00124 | Polygalacturonase (PG) (AY062697.1) | Arabidopsis thaliana | 63 | 1 | 1 | ||||
| MRU00022 | Peroxidase (POD) (FJ473421.1) | Litchi chinensis | 72 | 2 | 2 | ||||
| Aroma | MRU00010 | Alcohol dehydrogenase (ADH) (JF357957.1) | Diospyros kaki | 81 | 4 | 10 | 14 | ||
| MRU00108 | Alcohol dehydrogenase (ADH) (AM062702.1) | Alnus glutinosa | 94 | 1 | 1 | ||||
| MRU00126 | Alcohol dehydrogenase (ADH) (AF194173.1) | Vitis vinifera | 88 | 4 | 4 | ||||
| MRU00002 | Short chain alcohol dehydrogenase (AY084550.1) | Arabidopsis thaliana | 71 | 1 | 1 | ||||
| MRU00315 | Short chain alcohol dehydrogenase (AB192882.1) | Solanum tuberosum | 64 | 1 | 1 | ||||
© 2013 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhu, C.; Feng, C.; Li, X.; Xu, C.; Sun, C.; Chen, K. Analysis of Expressed Sequence Tags from Chinese Bayberry Fruit (Myrica rubra Sieb. and Zucc.) at Different Ripening Stages and Their Association with Fruit Quality Development. Int. J. Mol. Sci. 2013, 14, 3110-3123. https://doi.org/10.3390/ijms14023110
Zhu C, Feng C, Li X, Xu C, Sun C, Chen K. Analysis of Expressed Sequence Tags from Chinese Bayberry Fruit (Myrica rubra Sieb. and Zucc.) at Different Ripening Stages and Their Association with Fruit Quality Development. International Journal of Molecular Sciences. 2013; 14(2):3110-3123. https://doi.org/10.3390/ijms14023110
Chicago/Turabian StyleZhu, Changqing, Chao Feng, Xian Li, Changjie Xu, Chongde Sun, and Kunsong Chen. 2013. "Analysis of Expressed Sequence Tags from Chinese Bayberry Fruit (Myrica rubra Sieb. and Zucc.) at Different Ripening Stages and Their Association with Fruit Quality Development" International Journal of Molecular Sciences 14, no. 2: 3110-3123. https://doi.org/10.3390/ijms14023110
APA StyleZhu, C., Feng, C., Li, X., Xu, C., Sun, C., & Chen, K. (2013). Analysis of Expressed Sequence Tags from Chinese Bayberry Fruit (Myrica rubra Sieb. and Zucc.) at Different Ripening Stages and Their Association with Fruit Quality Development. International Journal of Molecular Sciences, 14(2), 3110-3123. https://doi.org/10.3390/ijms14023110






