Efficient Agrobacterium-Mediated Transformation of Hybrid Poplar Populus davidiana Dode × Populus bollena Lauche
Abstract
:1. Introduction
2. Results and Discussion
2.1. Optimization of Poplar Shoot Regeneration
2.2. Improvement of Genetic Transformation of Poplar
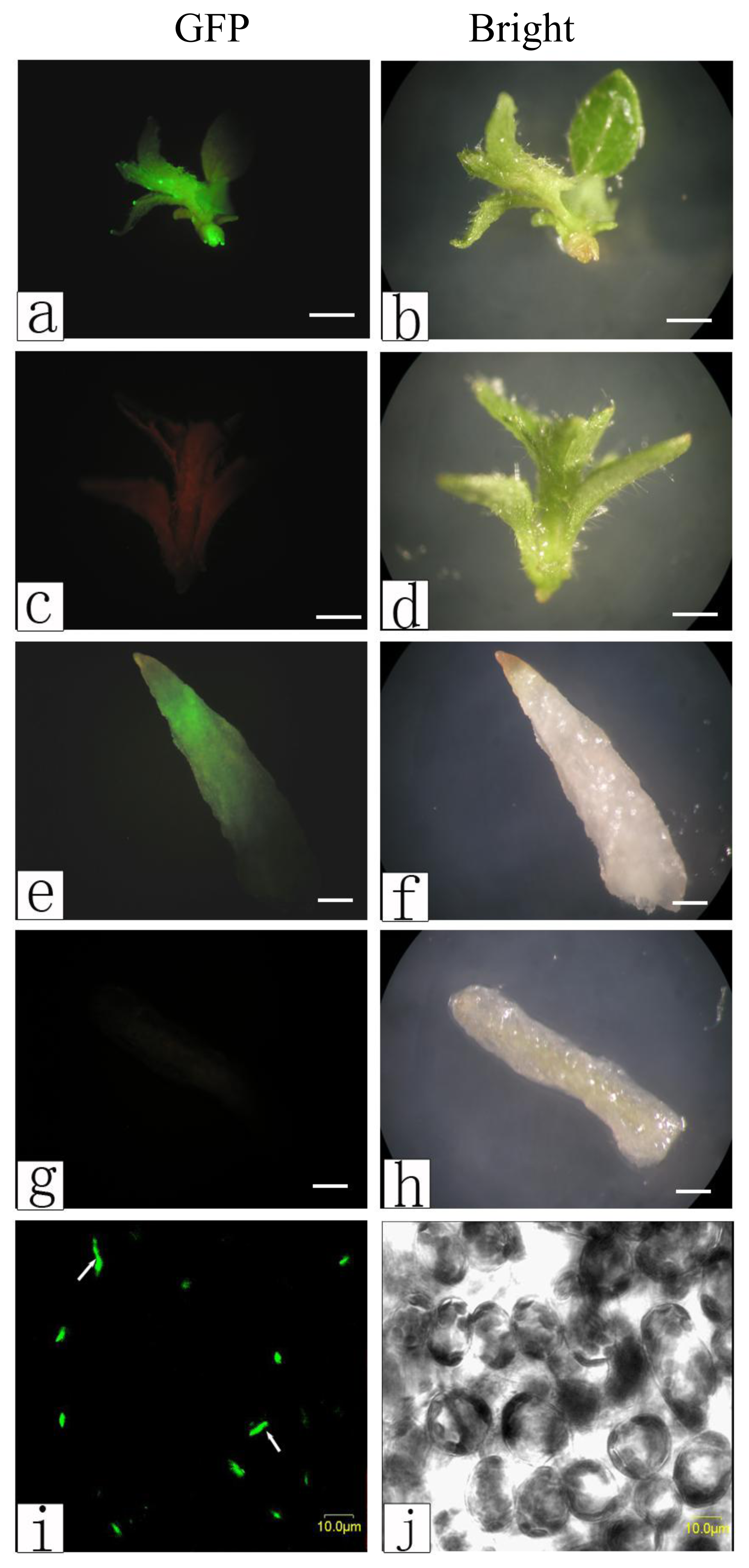
2.3. Observations of Genetic Transformation
3. Experimental Section
3.1. Plant Material, Agrobacterium Strain, Probe, and Plasmid
3.2. Establishment of Regeneration System for Poplar
3.3. Plant Transformation
3.3.1. Sample Cultivation
3.3.2. Effects of Agrobacterium Concentration and Infection Time
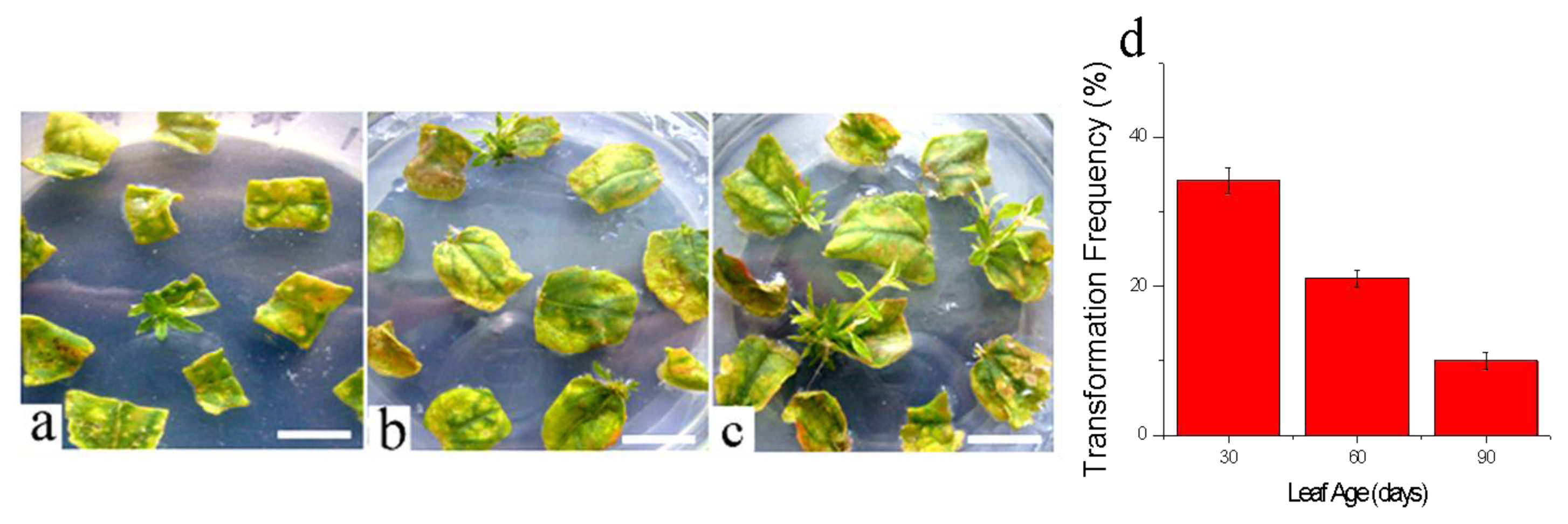
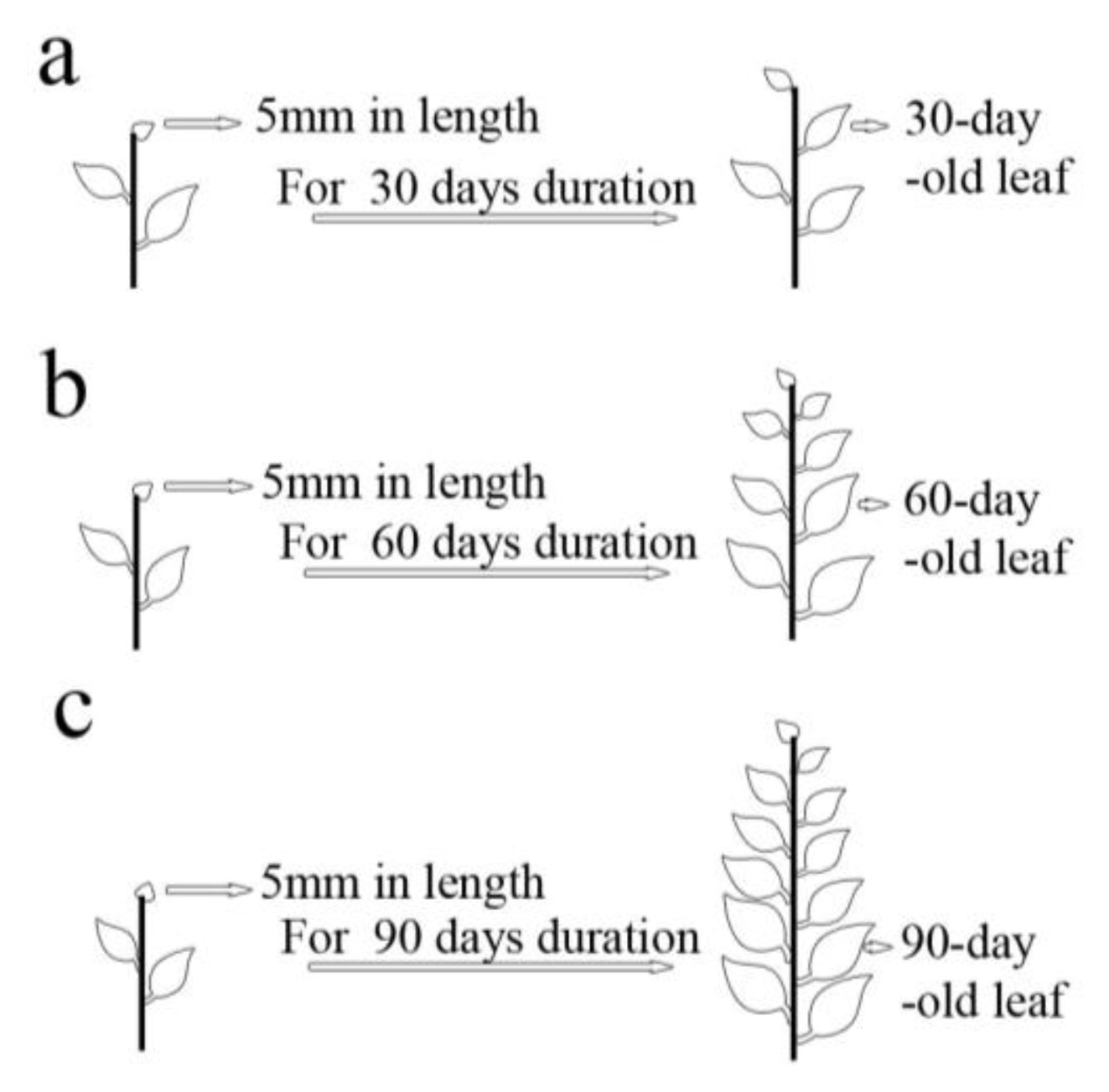
3.3.3. Effects of Leaf Ages
3.4. Visualization of Genetic Transformation
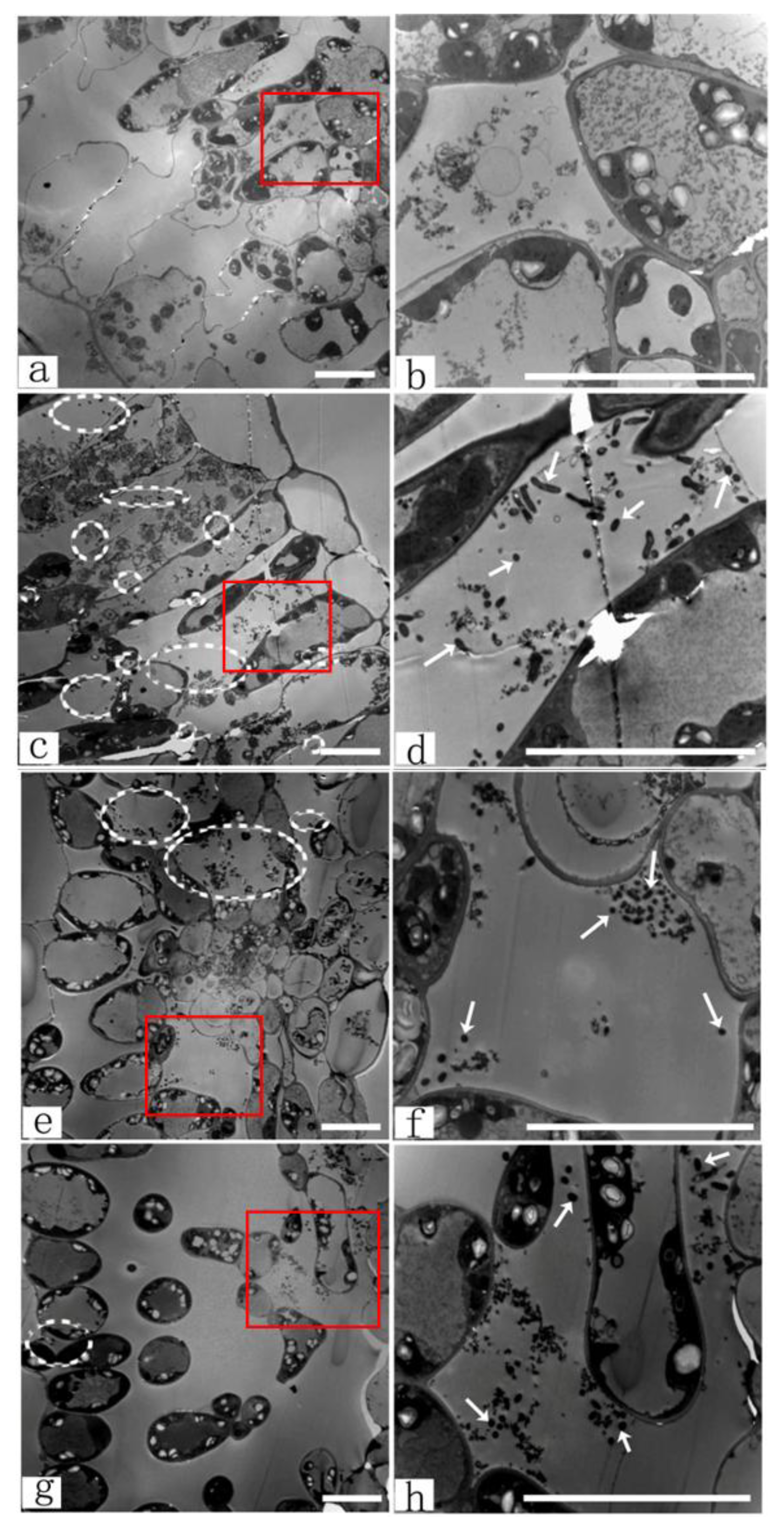
3.4.1. TEM Observations of Agrobacterium Infection
3.4.2. Green Fluorescent Protein Fluorescence Observations in Transgenic Poplar
3.4.3. Southern and Northern Blot Analysis
4. Conclusions
Acknowledgments
Abbreviations
| AS | Acetosyringone |
| BA | 6-Benzylaminopurine |
| Cef | Cefotaxime sodium |
| GFP | Green fluorescent protein |
| Kan | Kanamycin sulfate |
| MS | Murashige and Skoog |
| NAA | Naphthaleneacetic acid |
| Rif | Rifampicin |
| TEM | Transmission electron microscopy |
| FOX | Full-length cDNA overexpression |
Conflict of Interest
References
- Christie, C.B. Rapid propagation of aspens and silver poplars using tissue culture techniques. Proc. Int. Plant Prop. Soc 1978, 28, 255–260. [Google Scholar]
- De Block, M. Factors influencing the tissue culture and the Agrobacterium tumefaciens-mediated transformation of hybrid aspen and Poplar clones. Plant Physiol 1990, 93, 1110–1116. [Google Scholar]
- Son, S.H.; Hall, R.B. Multiple shoot regeneration from root organ cultures of Populus alba × P. grandidentata. Plant Cell Tiss. Organ. Cult 1990, 20, 53–57. [Google Scholar]
- Tsai, C.J.; Podila, G.K.; Chiang, V.L. Agrobacterium-mediated transformation of quaking aspen (Populus tremuloides) and regeneration of transgenic plants. Plant Cell Rep 1994, 14, 94–97. [Google Scholar]
- Gozukirmizi, N.; Bajrovic, K.; Ipekqi, Z. Genotype differencies in direct plant regeneration from stem explants of Populus tremula in Turkey. J. For. Res 1998, 3, 123–126. [Google Scholar]
- Noel, N.; Leple, J.C.; Pilate, G. Optimization of in vitro micropropagation and regeneration for Populus × interamericana and Populus × euramericana hybrids (P. deltoides, P. trichocarpa, and P. nigra). Plant Cell Rep 2002, 20, 1150–1155. [Google Scholar]
- Thakur, D.K.; Srivastava, D.K. High-efficiency plant regeneration from leaf explants of male Himalayan poplar (Populus ciliata Wall). In Vitro Cell Dev. Biol. Plant 2006, 42, 144–147. [Google Scholar]
- Yadav, R.; Arora, P.; Kumar, D.; Katyal, D.; Dilbaghi, N.; Chaudhury, A. High frequency direct plant regeneration from leaf, internode, and root segments of Eastern Cottonwood (Populus deltoides). Plant Biotechnol. Rep 2009, 3, 175–182. [Google Scholar]
- Huang, D.Q.; Dai, W.H. Direct regeneration from in vitro leaf and petiole tissues of Populus tremula ‘Erecta’. Plant Cell Tiss. Organ. Cult 2011, 107, 169–174. [Google Scholar]
- Harry, I.S.; Thrope, T.A. In vitro Culture of Forest Trees. In Plant Cell and Tissue Culture; Vasil, I.K., Thrope, T.A., Eds.; Kluwer Academic: Dordrecht, The Netherlands, 1994; pp. 539–560. [Google Scholar]
- Ahuja, M.R. Somatic cell differentiation and rapid clonal propagation of aspen. Silvae Genet 1983, 32, 131–135. [Google Scholar]
- Fillatti, J.J.; Sellmer, J.; McCown, B.; Haissig, B.; Comai, L. Agrobacterium mediated transformation and regeneration of Populus. Mol. Gen. Genet. 1987, 206, 192–199. [Google Scholar]
- Dai, W.H.; Cheng, Z.M.; Sargent, W. Plant regeneration and Agrobacterium-mediated transformation of two elite aspen hybrid clones from in vitro leaf tissues. In Vitro Cell Dev. Biol. Plant 2003, 39, 6–11. [Google Scholar]
- Ferreira, S.; Batista, D.; Serrazina, S.; Pais, M.S. Morphogenesis induction and organogenic nodule differentiation in Populus euphratica Oliv. leaf explants. Plant Cell Tiss. Organ. Cult 2009, 96, 35–43. [Google Scholar]
- Thakur, A.K.; Sarita, S.; Srivastava, D.K. Plant regeneration and genetic transformation studies in petiole tissue of Himalayan poplar (Populus ciliata Wall). Curr. Sci 2005, 89, 664–668. [Google Scholar]
- Tzfira, T.; Jensen, C.S.; Vainstein, A.; Altman, A. Transformation and regeneration of transgenic aspen plants via shoot formation from stem explants. Physiol. Plant 1997b, 99, 554–561. [Google Scholar]
- Kang, B.; Osburn, L.; Kopsell, D.; Tuskan, G.A.; Cheng, Z.M. Micropropagation of Populus trichocarpa ‘Nisqually-1’: The genotype deriving the Populus reference genome. Plant Cell Tiss. Organ. Cult 2009, 99, 251–257. [Google Scholar]
- Horsch, R.B.; Fry, J.E.; Hoffmann, N.L.; Eichholtz, D.; Rogers, S.G.; Fraley, R.T. A simple and general method for transferring genes into plants. Science 1985, 227, 1229–1231. [Google Scholar]
- Parsons, T.J.; Sinkar, V.P.; Stettler, R.F.; Nester, E.W.; Gordon, M.P. Transformation of poplar by Agrobacterium tumefaciens. Nat. Biotech 1986, 4, 533–536. [Google Scholar]
- Confalonieri, M.; Balestrazzi, A.; Bisoffi, S. Genetic transformation of Populus nigra by Agrobacterium tumefaciens. Plant Cell Rep 1994, 13, 256–261. [Google Scholar]
- Tzfira, T.; Jensen, C.S.; Wang, W.X.; Zuker, A.; Vinocur, B.; Altman, A.; Vainstein, A. Transgenic Populus tremula: A step-by-step protocol for its Agrobacterium-mediated transformation. Plant Mol. Biol. Rep 1997a, 15, 219–235. [Google Scholar]
- Han, K.H.; Gordon, M.P.; Strauss, S.H. High-frequency transformation of cottonwoods (genus Populus) by Agrobacterium rhizogenes. Can. J. For. Res 1997, 27, 464–470. [Google Scholar]
- Han, K.H.; Meilan, R.; Ma, C.; Strauss, S.H. An Agrobacterium tumefaciens transformation protocol effective on a variety of cottonwood hybrids (genus Populus). Plant Cell Rep 2000, 19, 315–320. [Google Scholar]
- Zhan, X.C.; Kawai, S.; Katayama, Y.; Morohoshi, N. A new approach based on the leaf disc method for Agrobacterium mediated transformation and regeneration of aspen. Plant Sci 1997, 123, 105–112. [Google Scholar]
- Okumura, S.; Sawada, M.; Park, Y.W.; Takahisa, H.Y.S.; Shimamura, H.; Takase, H.; Tomizawa, K.I. Transformation of poplar (Populus alba) plastids and expression of foreign proteins in tree chloroplasts. Transgenic Res 2006, 15, 637–646. [Google Scholar]
- Nishiguchi, M.; Yoshida, K.; Mohri, T.; Igasaki, T.; Shinohara, K. An improved transformation system for Lombardy poplar (Populus nigra var. italica). J. For. Res 2006, 11, 175–180. [Google Scholar]
- Cseke, L.J.; Cseke, S.B.; Podila, G.K. High efficiency poplar transformation. Plant Cell Rep 2007, 26, 1529–1538. [Google Scholar]
- Yevtushenko, D.P.; Santosh, M. Efficient Agrobacterium-mediated transformation of commercial hybrid poplar Populus nigra L. × P. maximowiczii A. Henry. Plant Cell Rep 2010, 29, 211–221. [Google Scholar]
- Meilan, R.; Ma, C. Protocols, 2nd ed; Wang, K., Ed.; Humana Press: Totowa, NJ, USA, 2006; Volume 344, pp. 143–151. [Google Scholar]
- Confalonieri, M.; Balestrazzi, A.; Bisoffi, S.; Cella, R. Factors affecting Agrobacterium tumefaciens-mediated transformation in several black poplar clones. Plant Cell Tiss. Organ. Cult 1995, 43, 215–222. [Google Scholar]
- Confalonieri, M.; Balestrazzi, A.; Cella, R. Genetic transformation of Populus deltoides and P.x euramericana clones using Agrobacterium tumefaciens. Plant Cell Tiss. Organ. Cult 1997, 48, 53–61. [Google Scholar]
- Li, X.Y.; Huang, F.H.; Gbur, E.E. Factors affecting transformation efficiency of poplar hybrid line NC5331 by Agrobacterium tumefaciens. J. Arkansas Acad. Sci 1997, 51, 116–120. [Google Scholar]
- Ma, C.P.; Strauss, S.H.; Meilan, R. Agrobacterium-mediated transformation of the genome sequenced poplar clone, Nisqually-1(Populus trichocarpa). Plant Mol. Biol. Rep 2004, 22, 1–9. [Google Scholar]
- Ichikawa, T.; Nakazawa, M.; Kawashima, M. The FOX hunting system: An alternative gain-of-function gene hunting technique. Plant J 2006, 45, 974–985. [Google Scholar]
- Kondou, Y.; Higuchi, M.; Takahashi, S. Systematic approaches to using the FOX hunting system to identify useful rice genes. Plant J 2009, 57, 883–894. [Google Scholar]
- Nakamura, H.; Hakata, M.; Amano, K.; Miyao, A. A genome-wide gain-of-function analysis of rice genes using the FOX-Hunting system. Plant Mol. Biol 2007, 65, 357–371. [Google Scholar]
- Busov, V.B.; Meilan, R.; Pearce, D.W.; Ma, C.; Rood, S.B.; Strauss, S.H. Activation tagging of a dominant gibberellin catabolism gene (GA 2-oxidase) from poplar that regulates tree stature. Plant Physiol 2003, 132, 1283–1291. [Google Scholar]
- Murashige, T.; Skoog, F. A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol. Plant 1962, 15, 473–497. [Google Scholar]
- Godwin, I.; Todd, G.; Brian, F.L.; Newbury, H.J. The effects of acetosyringone and pH on Agrobacterium-mediated transformation vary according to plant species. Plant Cell Rep 1991, 9, 671–675. [Google Scholar]
- Smith, M.A.L.; McCown, B.H. A comparison of source tissue for protoplast isolation from three woody plant species. Plant Sci. Lett 1983, 28, 149–156. [Google Scholar]
- Civinova, B.; Sladky, Z. Stimulation of the regeneration capacity of tree shoot segment explants in vitro. Biol. Plant 1990, 32, 407–413. [Google Scholar]
- Graves, A.E.; Goldman, S.L.; Banks, S.W.; Graves, A.C.F. Scanning electron microscope studies of Agrobacterium tumefaciens attachment to Zea mays, Gladiolus sp., and Triticum aestivum. J. Bactriol 1988, 170, 2395–2400. [Google Scholar]
- Verma, A.; Nain, V.; Kumari, C.; Singh, S.K.; Lakshmi, N.M.; Ananda, K.P. Tissue specific response of Agrobacterium tumefaciens attachment to Sorghum bicolor (L) Moench. Physiol. Mol. Biol. Plants 2008, 14, 307–313. [Google Scholar]
- Willians, D.; Carter, C.B. Transmission Electron Microscopy; Plenum Press: New York, NY, USA, 1996. [Google Scholar]
- Porebski, L.; Bailey, L.; Baum, B. Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol. Biol. Rep 1997, 15, 8–15. [Google Scholar]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed; Cold Spring Harbor Laboratory Press: New York, NY, USA, 1989. [Google Scholar]





| Medium | BA (mg/L) | NAA (mg/L) | Mean No. shoots formed per explant | Explant regeneration (%) |
|---|---|---|---|---|
| 1 | 0.1 | 0.01 | 0 | 0 |
| 2 | 0.1 | 0.03 | 0 | 0 |
| 3 | 0.1 | 0.05 | 0.18 ± 0.51 | 14.4 ± 1.93 |
| 4 | 0.1 | 0.08 | 0.08 ± 0.02 | 5.56 ± 1.93 |
| 5 | 0.1 | 0.1 | 0.02 ± 0.02 | 2.22 ± 1.92 |
| 6 | 0.3 | 0.01 | 3.61 ± 0.27 | 76.7 ± 6.65 |
| 7 | 0.3 | 0.03 | 1.82 ± 0.27 | 60.0 ± 10.0 |
| 8 | 0.3 | 0.05 | 0.42 ± 0.15 | 17.8 ± 5.09 |
| 9 | 0.3 | 0.08 | 5.87 ± 0.49 | 90.0 ± 8.81 |
| 10 | 0.3 | 0.1 | 2.08 ± 0.34 | 55.6 ± 5.09 |
| 11 | 0.5 | 0.01 | 0.59 ± 0.17 | 14.4 ± 3.85 |
| 12 | 0.5 | 0.03 | 1.54 ± 0.68 | 16.7 ± 5.77 |
| 13 | 0.5 | 0.05 | 0.78 ± 0.31 | 40.0 ± 10.0 |
| 14 | 0.5 | 0.08 | 0.03 ± 0.03 | 3.33 ± 3.34 |
| 15 | 0.5 | 0.1 | 0.53 ± 0.12 | 5.56 ± 3.85 |
© 2013 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Han, X.; Ma, S.; Kong, X.; Takano, T.; Liu, S. Efficient Agrobacterium-Mediated Transformation of Hybrid Poplar Populus davidiana Dode × Populus bollena Lauche. Int. J. Mol. Sci. 2013, 14, 2515-2528. https://doi.org/10.3390/ijms14022515
Han X, Ma S, Kong X, Takano T, Liu S. Efficient Agrobacterium-Mediated Transformation of Hybrid Poplar Populus davidiana Dode × Populus bollena Lauche. International Journal of Molecular Sciences. 2013; 14(2):2515-2528. https://doi.org/10.3390/ijms14022515
Chicago/Turabian StyleHan, Xue, Shurong Ma, Xianghui Kong, Tetsuo Takano, and Shenkui Liu. 2013. "Efficient Agrobacterium-Mediated Transformation of Hybrid Poplar Populus davidiana Dode × Populus bollena Lauche" International Journal of Molecular Sciences 14, no. 2: 2515-2528. https://doi.org/10.3390/ijms14022515




