Genome-Wide Analysis of Respiratory Burst Oxidase Homologs in Grape (Vitis vinifera L.)
Abstract
:1. Introduction
2. Results
2.1. Identification of Vvrboh Genes in the Grape Genome
2.2. Sequence Analysis and Domain Organization in Grape Rboh Homologs
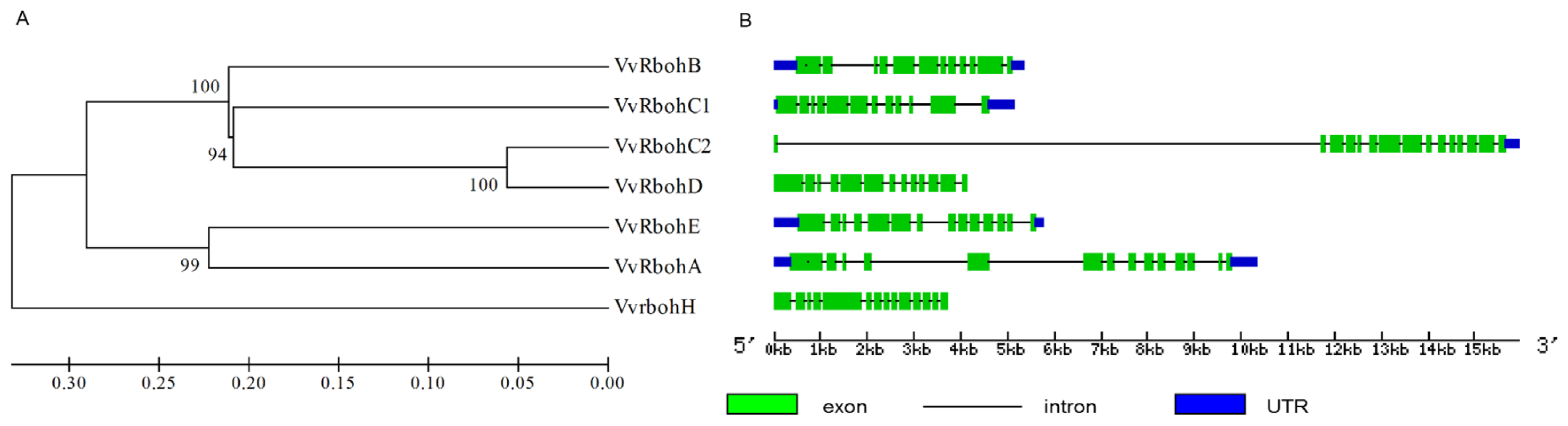
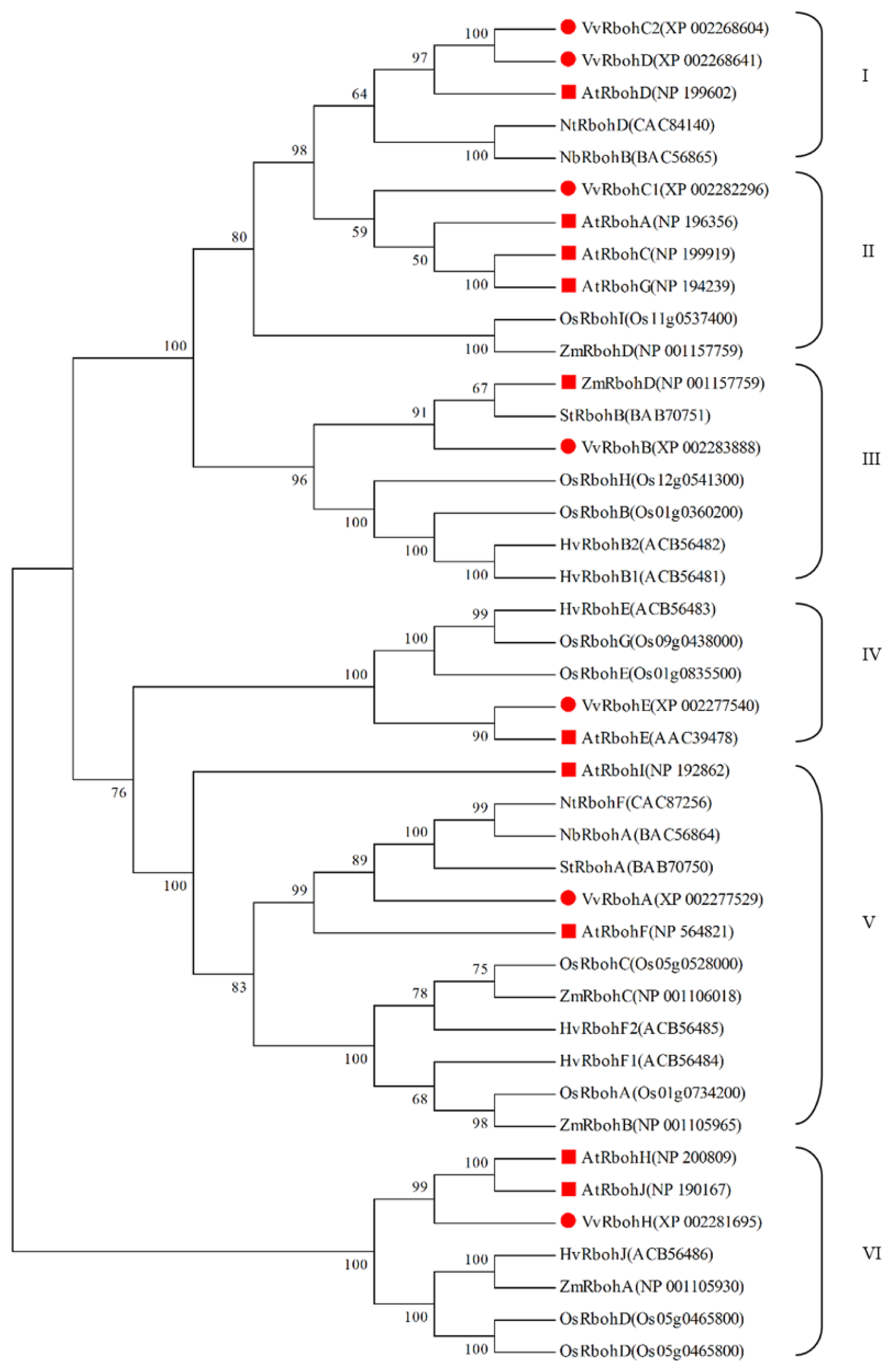
2.3. Phylogenetic Analysis of Vvrboh Genes
2.4. Gene Structure Analysis of Vvrboh Genes
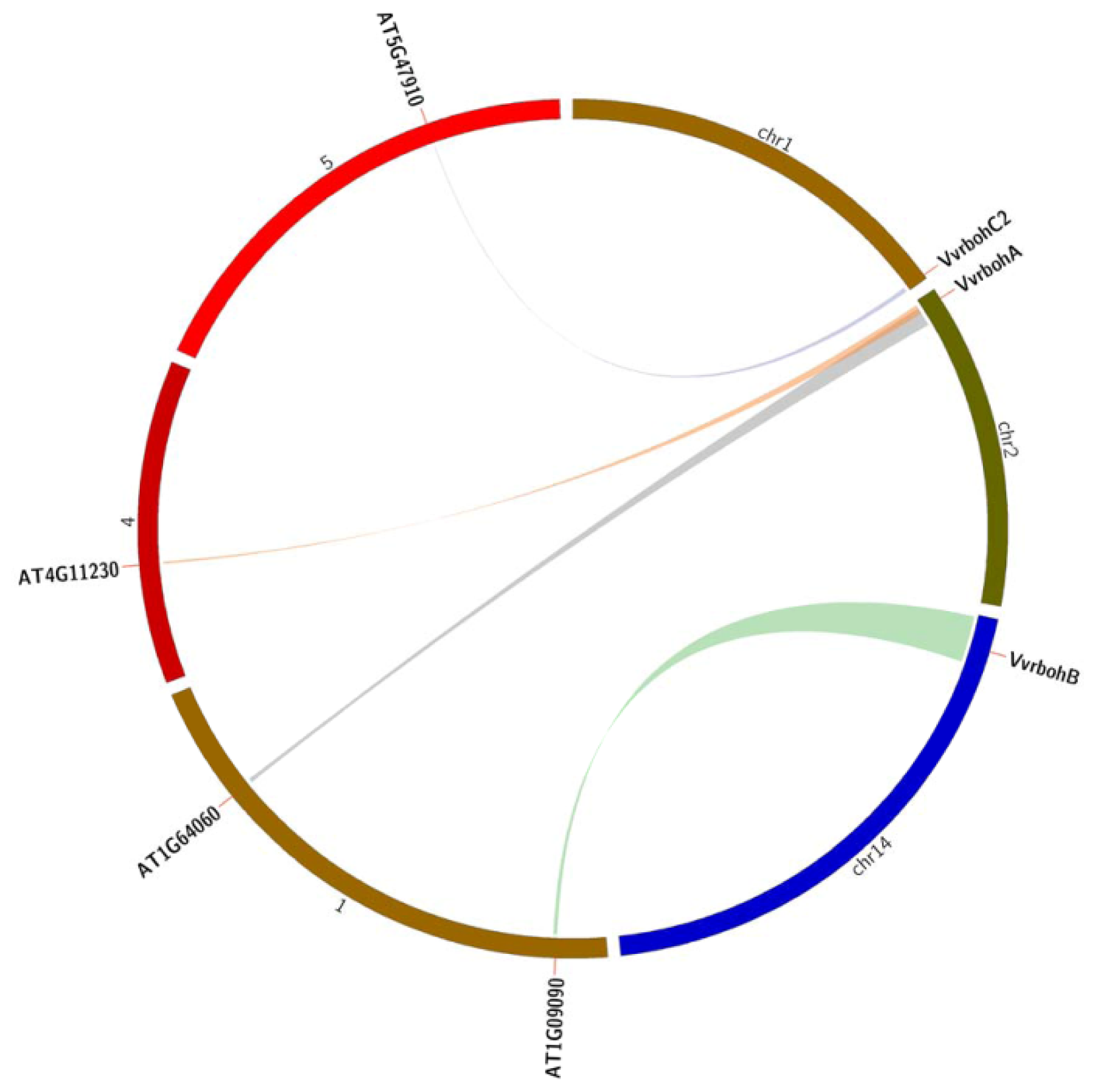
2.5. Evolutionary Relationships between Grape and Arabidopsis rboh Genes
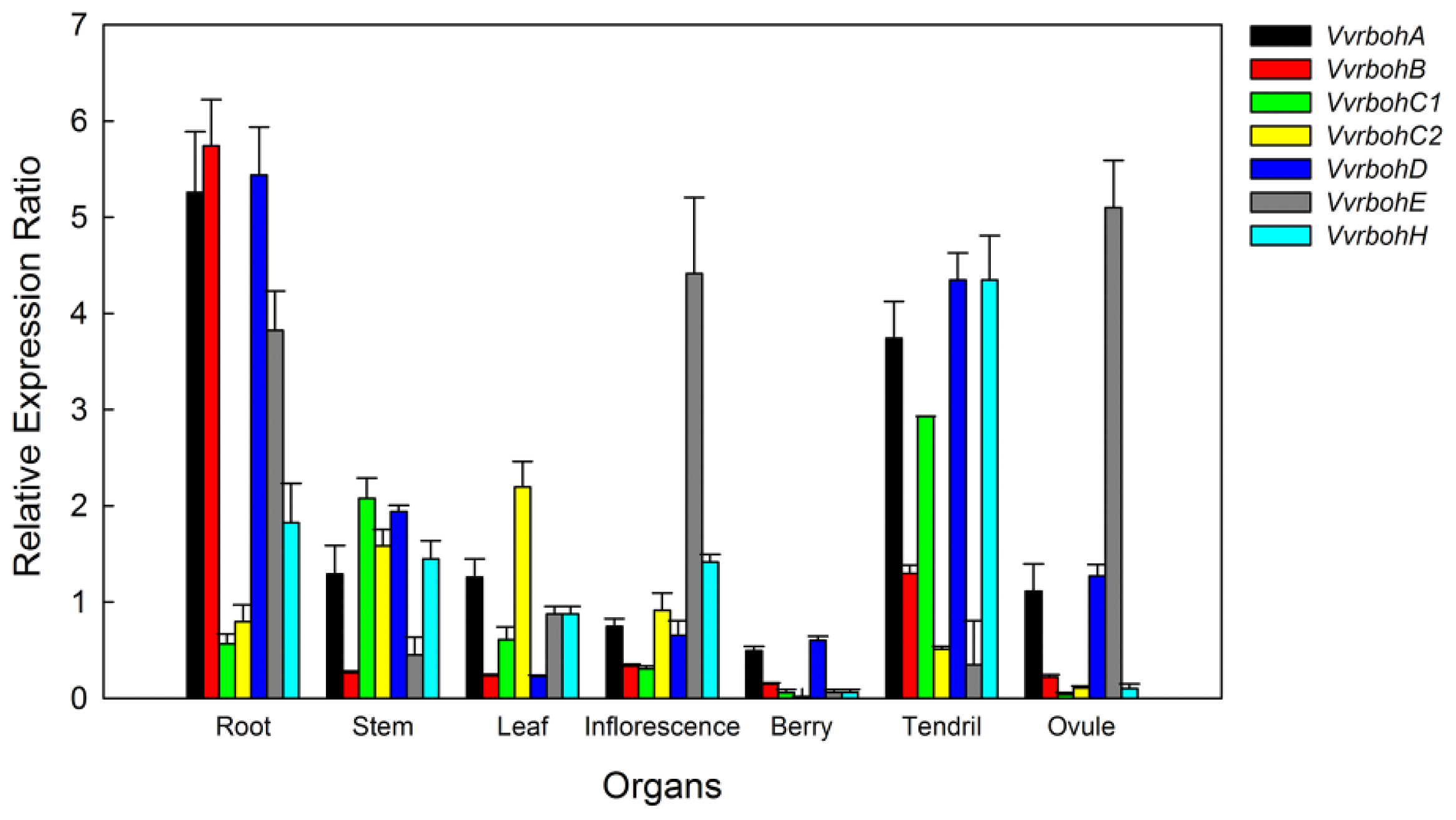
2.6. Expression Patterns of Vvrboh Genes in Different Tissues and Organs
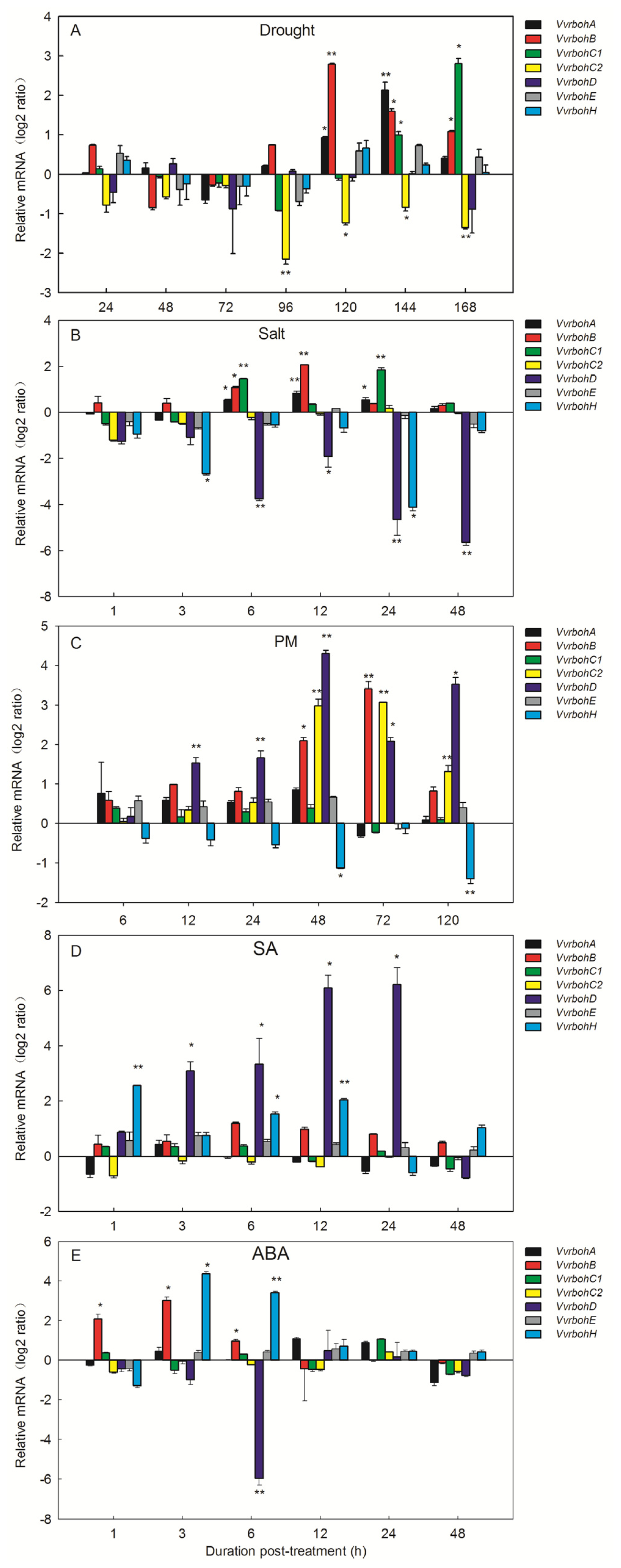
2.7. Expression Profiles of Vvrboh Genes Following Various Stress Hormone Treatments
3. Discussion
3.1. Identification and Sequence Analysis of Vvrboh Genes
3.2. The Evolution of Rboh Proteins in Grape and Arabidopsis and Functional Prediction of Vvrboh Genes
3.3. Spatial Expression Patterns of Vvrboh Genes in Various Grape Tissues
3.4. Vvrboh Genes Respond to a Range of Biological Stresses
4. Experimental Section
4.1. Identification and Annotation of Grape Respiratory Burst Oxidase Homolog (Vvrboh) Genes
4.2. Amino Acid Sequence Alignment and Phylogenetic Analysis
4.3. Exon/Intron Structure Analysis of Vvrboh Genes
4.4. Synteny Analysis
4.5. Targeting Signal Prediction
4.6. Plant Materials
4.7. Abiotic and Biotic Stress Treatments and Hormone Applications
4.8. Quantitative RT-PCR Analysis
5. Conclusions
Supplementary Information
ijms-14-24169-s001.pdfAcknowledgments
Abbreviations
| ABA | abscisic acid |
| At | Arabidopsis thaliana |
| HR | hypersensitive response |
| ORF | open reading frame |
| qRT-PCR | quantitative reverse transcription PCR |
| SA | salicylic acid |
| Vv | Vitis vinifera |
Conflicts of Interest
References
- Mittler, R.; Vanderauwera, S.; Gollery, M.; van Breusegem, F. Reactive oxygen gene network of plants. Trends Plant Sci 2004, 9, 490–498. [Google Scholar]
- Foreman, J.; Demidchik, V.; Bothwell, J.H.; Mylona, P.; Miedema, H.; Torres, M.A.; Linstead, P.; Costa, S.; Brownlee, C.; Jones, J.D. Reactive oxygen species produced by NADPH oxidase regulate plant cell growth. Nature 2003, 422, 442–446. [Google Scholar]
- Torres, M.A.; Dangl, J.L. Functions of the respiratory burst oxidase in biotic interactions, abiotic stress and development. Curr. Opin. Plant Biol 2005, 8, 397–403. [Google Scholar]
- Pei, Z.-M.; Murata, Y.; Benning, G.; Thomine, S.; Klüsener, B.; Allen, G.J.; Grill, E.; Schroeder, J.I. Calcium channels activated by hydrogen peroxide mediate abscisic acid signalling in guard cells. Nature 2000, 406, 731–734. [Google Scholar]
- Apel, K.; Hirt, H. Reactive oxygen species: Metabolism, oxidative stress, and signal transduction. Annu. Rev. Plant Biol 2004, 55, 373–399. [Google Scholar]
- Mittler, R.; Vanderauwera, S.; Suzuki, N.; Miller, G.; Tognetti, V.B.; Vandepoele, K.; Gollery, M.; Shulaev, V.; van Breusegem, F. ROS signaling: The new wave? Trends Plant Sci 2011, 16, 300–309. [Google Scholar]
- Sagi, M.; Fluhr, R. Superoxide production by plant homologues of the gp91phox NADPH oxidase. Modulation of activity by calcium and by tobacco mosaic virus infection. Plant Physiol 2001, 126, 1281–1290. [Google Scholar]
- Groom, Q.J.; Torres, M.A.; Fordham-Skelton, A.P.; Hammond-Kosack, K.E.; Robinson, N.J.; Jones, J.D. RbohA, a rice homologue of the mammalian gp91phox respiratory burst oxidase gene. Plant J 1996, 10, 515–522. [Google Scholar]
- Torres, M.A.; Onouchi, H.; Hamada, S.; Machida, C.; Hammond-Kosack, K.E.; Jones, J.D. Six Arabidopsis thaliana homologues of the human respiratory burst oxidase (gp91phox). Plant J 1998, 14, 365–370. [Google Scholar]
- Sagi, M.; Davydov, O.; Orazova, S.; Yesbergenova, Z.; Ophir, R.; Stratmann, J.W.; Fluhr, R. Plant respiratory burst oxidase homologs impinge on wound responsiveness and development in Lycopersicon esculentum. Plant Cell 2004, 16, 616–628. [Google Scholar]
- Orozco-Cárdenas, M.L.; Narváez-Vásquez, J.; Ryan, C.A. Hydrogen peroxide acts as a second messenger for the induction of defense genes in tomato plants in response to wounding, systemin, and methyl jasmonate. Plant Cell 2001, 13, 179–191. [Google Scholar]
- Zhang, H.; Fang, Q.; Zhang, Z.; Wang, Y.; Zheng, X. The role of respiratory burst oxidase homologues in elicitor-induced stomatal closure and hypersensitive response in Nicotiana benthamiana. J. Exp. Bot 2009, 60, 3109–3122. [Google Scholar]
- Asai, S.; Ohta, K.; Yoshioka, H. MAPK signaling regulates nitric oxide and NADPH oxidase-dependent oxidative bursts in Nicotiana benthamiana. Plant Cell 2008, 20, 1390–1406. [Google Scholar]
- Yoshioka, H.; Numata, N.; Nakajima, K.; Katou, S.; Kawakita, K.; Rowland, O.; Jones, J.D.; Doke, N. Nicotiana benthamiana gp91phox homologs NbrbohA and NbrbohB participate in H2O2 accumulation and resistance to Phytophthora infestans. Plant Cell 2003, 15, 706–718. [Google Scholar]
- Simon-Plas, F.; Elmayan, T.; Blein, J.P. The plasma membrane oxidase NtrbohD is responsible for AOS production in elicited tobacco cells. Plant J 2002, 31, 137–147. [Google Scholar]
- Yoshioka, H.; Sugie, K.; Park, H.-J.; Maeda, H.; Tsuda, N.; Kawakita, K.; Doke, N. Induction of plant gp91phox homolog by fungal cell wall, arachidonic acid, and salicylic acid in potato. Mol. Plant Microbe Inter 2001, 14, 725–736. [Google Scholar]
- Kobayashi, M.; Ohura, I.; Kawakita, K.; Yokota, N.; Fujiwara, M.; Shimamoto, K.; Doke, N.; Yoshioka, H. Calcium-dependent protein kinases regulate the production of reactive oxygen species by potato NADPH oxidase. Plant Cell 2007, 19, 1065–1080. [Google Scholar]
- Lin, F.; Ding, H.; Wang, J.; Zhang, H.; Zhang, A.; Zhang, Y.; Tan, M.; Dong, W.; Jiang, M. Positive feedback regulation of maize NADPH oxidase by mitogen-activated protein kinase cascade in abscisic acid signalling. J. Exp. Bot 2009, 60, 3221–3238. [Google Scholar]
- Si, Y.; Dane, F.; Rashotte, A.; Kang, K.; Singh, N.K. Cloning and expression analysis of the Ccrboh gene encoding respiratory burst oxidase in Citrullus colocynthis and grafting onto Citrullus lanatus (watermelon). J. Exp. Bot 2010, 61, 1635–1642. [Google Scholar]
- Trujillo, M.; Altschmied, L.; Schweizer, P.; Kogel, K.-H.; Hückelhoven, R. Respiratory burst oxidase homologue A of barley contributes to penetration by the powdery mildew fungus Blumeria graminis f. sp. hordei. J. Exp. Bot 2006, 57, 3781–3791. [Google Scholar]
- Lightfoot, D.J.; Boettcher, A.; Little, A.; Shirley, N.; Able, A.J. Identification and characterisation of barley (Hordeum vulgare) respiratory burst oxidase homologue family members. Funct. Plant Biol 2008, 35, 347–359. [Google Scholar]
- Marino, D.; Andrio, E.; Danchin, E.G.; Oger, E.; Gucciardo, S.; Lambert, A.; Puppo, A.; Pauly, N. A Medicago truncatula NADPH oxidase is involved in symbiotic nodule functioning. New Phytol 2011, 189, 580–592. [Google Scholar]
- Müller, K.; Linkies, A.; Leubner-Metzger, G.; Kermode, A.R. Role of a respiratory burst oxidase of Lepidium sativum (cress) seedlings in root development and auxin signalling. J. Exp. Bot 2012, 63, 6325–6334. [Google Scholar]
- Sagi, M.; Fluhr, R. Production of reactive oxygen species by plant NADPH oxidases. Plant Physiol 2006, 141, 336–340. [Google Scholar]
- Zimmermann, P.; Hirsch-Hoffmann, M.; Hennig, L.; Gruissem, W. Genevestigator. Arabidopsis microarray database and analysis toolbox. Plant Physiol 2004, 136, 2621–2632. [Google Scholar]
- Kwak, J.M.; Mori, I.C.; Pei, Z.-M.; Leonhardt, N.; Torres, M.A.; Dangl, J.L.; Bloom, R.E.; Bodde, S.; Jones, J.D.; Schroeder, J.I. NADPH oxidase AtrbohD and AtrbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EMBO J 2003, 22, 2623–2633. [Google Scholar]
- Torres, M.A.; Dangl, J.L.; Jones, J.D. Arabidopsis gp91phox homologues AtrbohD and AtrbohF are required for accumulation of reactive oxygen intermediates in the plant defense response. Proc. Natl. Acad. Sci. USA 2002, 99, 517–522. [Google Scholar]
- Miller, G.; Schlauch, K.; Tam, R.; Cortes, D.; Torres, M.A.; Shulaev, V.; Dangl, J.L.; Mittler, R. The plant NADPH oxidase RBOHD mediates rapid systemic signaling in response to diverse stimuli. Sci. Signal 2009, 2. [Google Scholar] [CrossRef]
- Vignais, P. The superoxide-generating NADPH oxidase: Structural aspects and activation mechanism. Cell Mol. Life Sci 2002, 59, 1428–1459. [Google Scholar]
- Yoshida, L.S.; Saruta, F.; Yoshikawa, K.; Tatsuzawa, O.; Tsunawaki, S. Mutation at histidine 338 of gp91phox depletes FAD and affects expression of cytochrome b558 of the human NADPH oxidase. J. Biol. Chem 1998, 273, 27879–27886. [Google Scholar]
- Wong, H.L.; Pinontoan, R.; Hayashi, K.; Tabata, R.; Yaeno, T.; Hasegawa, K.; Kojima, C.; Yoshioka, H.; Iba, K.; Kawasaki, T. Regulation of rice NADPH oxidase by binding of Rac GTPase to its N-terminal extension. Plant Cell 2007, 19, 4022–4034. [Google Scholar]
- Takeda, S.; Gapper, C.; Kaya, H.; Bell, E.; Kuchitsu, K.; Dolan, L. Local positive feedback regulation determines cell shape in root hair cells. Science 2008, 319, 1241–1244. [Google Scholar]
- Finegold, A.A.; Shatwell, K.P.; Segal, A.W.; Klausner, R.D.; Dancis, A. Intramembrane bis-heme motif for transmembrane electron transport conserved in a yeast iron reductase and the human NADPH oxidase. J. Biol. Chem 1996, 271, 31021–31024. [Google Scholar]
- Wang, G.-F.; Li, W.-Q.; Li, W.-Y.; Wu, G.-L.; Zhou, C.-Y.; Chen, K.-M. Characterization of rice NADPH oxidase genes and their expression under various environmental conditions. Int. J. Mol. Sci 2013, 14, 9440–9458. [Google Scholar]
- Keller, T.; Damude, H.G.; Werner, D.; Doerner, P.; Dixon, R.A.; Lamb, C. A plant homolog of the neutrophil NADPH oxidase gp91phox subunit gene encodes a plasma membrane protein with Ca2+ binding motifs. Plant Cell 1998, 10, 255–266. [Google Scholar]
- Lyons, E.; Pedersen, B.; Kane, J.; Alam, M.; Ming, R.; Tang, H.; Wang, X.; Bowers, J.; Paterson, A.; Lisch, D. Finding and comparing syntenic regions among Arabidopsis and the outgroups papaya, poplar, and grape: CoGe with rosids. Plant Physiol 2008, 148, 1772–1781. [Google Scholar]
- Müller, K.; Carstens, A.C.; Linkies, A.; Torres, M.A.; Leubner-Metzger, G. The NADPH oxidase AtrbohB plays a role in Arabidopsis seed after ripening. N. Phytol 2009, 184, 885–897. [Google Scholar]
- Desikan, R.; Reynolds, A.; Hancock, J.; Neill, S. Harpin and hydrogen peroxide both initiate programmed cell death but have differential effects on defence gene expression in Arabidopsis suspension cultures. Biochem. J 1998, 330, 115–120. [Google Scholar]
- Shinozaki, K.; Yamaguchi-Shinozaki, K. Gene networks involved in drought stress response and tolerance. J. Exp. Bot 2007, 58, 221–227. [Google Scholar]
- Shirasu, K.; Nakajima, H.; Rajasekhar, V.K.; Dixon, R.A.; Lamb, C. Salicylic acid potentiates an agonist-dependent gain control that amplifies pathogen signals in the activation of defense mechanisms. Plant Cell 1997, 9, 261–270. [Google Scholar]
- Rao, M.V.; Davis, K.R. Ozone-induced cell death occurs via two distinct mechanisms in Arabidopsis: The role of salicylic acid. Plant J 1999, 17, 603–614. [Google Scholar]
- Draper, J. Salicylate, superoxide synthesis and cell suicide in plant defence. Trends Plant Sci 1997, 2, 162–165. [Google Scholar]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Higgins, D.G. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar]
- Hervé, C.; Tonon, T.; Collén, J.; Corre, E.; Boyen, C. NADPH oxidases in Eukaryotes: Red algae provide new hints! Curr. Genet 2006, 49, 190–204. [Google Scholar]
- Letunic, I.; Doerks, T.; Bork, P. SMART 7: Recent updates to the protein domain annotation resource. Nucleic Acids Res 2012, 40, D302–D305. [Google Scholar]
- Ren, J.; Wen, L.; Gao, X.; Jin, C.; Xue, Y.; Yao, X. DOG 1.0: Illustrator of protein domain structures. Cell Res 2009, 19, 271–273. [Google Scholar]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol 2011, 28, 2731–2739. [Google Scholar]
- Rice, P.; Longden, I.; Bleasby, A. EMBOSS: The European molecular biology open software suite. Trends Genet 2000, 16, 276–277. [Google Scholar]
- Guo, A.Y.; Zhu, Q.H.; Chen, X.; Luo, J.C. GSDS: A gene structure display server. Yi Chuan 2007, 29, 1023–1026. [Google Scholar]
- Krzywinski, M.; Schein, J.; Birol, İ.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res 2009, 19, 1639–1645. [Google Scholar]
- Nakai, K.; Kanehisa, M. A knowledge base for predicting protein localization sites in eukaryotic cells. Genomics 1992, 14, 897–911. [Google Scholar]
- Xu, X.; Guo, R.; Cheng, C.; Zhang, H.; Zhang, Y.; Wang, X. Overexpression of ALDH2B8, an aldehyde dehydrogenase gene from grapevine, sustains Arabidopsis growth upon salt stress and protects plants against oxidative stress. Plant Cell Tiss. Org 2013, 114, 187–196. [Google Scholar]
- Peng, S.; Zhu, Z.; Zhao, K.; Shi, J.; Yang, Y.; He, M.; Wang, Y. A novel heat shock transcription factor, VpHsf1, from Chinese wild Vitis pseudoreticulata is involved in biotic and abiotic stresses. Plant Mol. Biol. Rep 2013, 31, 240–247. [Google Scholar]
- Hou, H.; Yan, Q.; Wang, X.; Xu, H. A SBP-box gene VpSBP5 from Chinese wild Vitis species responds to Erysiphe necator and defense signaling molecules. Plant Mol. Biol. Rep 2013, 31, 1261–1270. [Google Scholar]
- Xiao, H.; Nassuth, A. Stress-and development-induced expression of spliced and unspliced transcripts from two highly similar dehydrin 1 genes in V. riparia and V. vinifera. Plant Cell Rep 2006, 25, 968–977. [Google Scholar]
- Wang, Y.; Liu, Y.; He, P.; Chen, J.; Lamikanra, O.; Lu, J. Evaluation of foliar resistance to Uncinula necator in Chinese wild Vitis species. Vitis 1995, 34, 159–164. [Google Scholar]


| Gene ID | Gene Locus ID | Accession No. | Putative function | Chromosome | Start | End | Predicted gene length (kb) | Predicted ORF length (bp) |
|---|---|---|---|---|---|---|---|---|
| VvrbohA | GSVIVT01019429001 | XP_002277529.1 | plasma membrane | chr2 | 621477 | 631791 | 10.310 | 2769 |
| VvrbohB | GSVIVT01031128001 | XP_002283888.1 | chloroplast thylakoid membrane | chr14 | 1892425 | 1897786 | 5.362 | 2622 |
| VvrbohC1 | GSVIVT01014350001 | XP_002282296.2 | plasma membrane | chr19 | 2924506 | 2929619 | 5.114 | 2523 |
| VvrbohC2 | GSVIVT01001122001 | XP_002268604.1 | chloroplast thylakoid membrane | chr1 | 22798594 | 22814521 | 15.930 | 2484 |
| VvrbohD | GSVIVT01001123001 | XP_002268641.1 | plasma membrane | chr1 | 22815076 | 22819209 | 4.134 | 2721 |
| VvrbohE | GSVIVT01015025001 | XP_002277540.1 | chloroplast thylakoid membrane | chr11 | 542212 | 547987 | 5.776 | 2754 |
| VvrbohH | GSVIVT01025074001 | XP_002281695.1 | chloroplast thylakoid membrane | chr6 | 4762503 | 4766209 | 3.707 | 2559 |
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Cheng, C.; Xu, X.; Gao, M.; Li, J.; Guo, C.; Song, J.; Wang, X. Genome-Wide Analysis of Respiratory Burst Oxidase Homologs in Grape (Vitis vinifera L.). Int. J. Mol. Sci. 2013, 14, 24169-24186. https://doi.org/10.3390/ijms141224169
Cheng C, Xu X, Gao M, Li J, Guo C, Song J, Wang X. Genome-Wide Analysis of Respiratory Burst Oxidase Homologs in Grape (Vitis vinifera L.). International Journal of Molecular Sciences. 2013; 14(12):24169-24186. https://doi.org/10.3390/ijms141224169
Chicago/Turabian StyleCheng, Chenxia, Xiaozhao Xu, Min Gao, Jun Li, Chunlei Guo, Junyang Song, and Xiping Wang. 2013. "Genome-Wide Analysis of Respiratory Burst Oxidase Homologs in Grape (Vitis vinifera L.)" International Journal of Molecular Sciences 14, no. 12: 24169-24186. https://doi.org/10.3390/ijms141224169




