Arabidopsis Non-Coding RNA Regulation in Abiotic Stress Responses
Abstract
:1. Introduction
2. Small RNAs (sRNAs)
2.1. Micro RNAs (miRNAs)
2.2. Small Interfering RNAs (siRNAs)
3. Long Non-Coding RNAs (lncRNAs)
3.1. Long Antisense Non-Coding RNAs (lancRNAs)
3.2. Long Intergenic Non-Coding RNAs (lincRNAs)
4. Conclusions and Perspectives
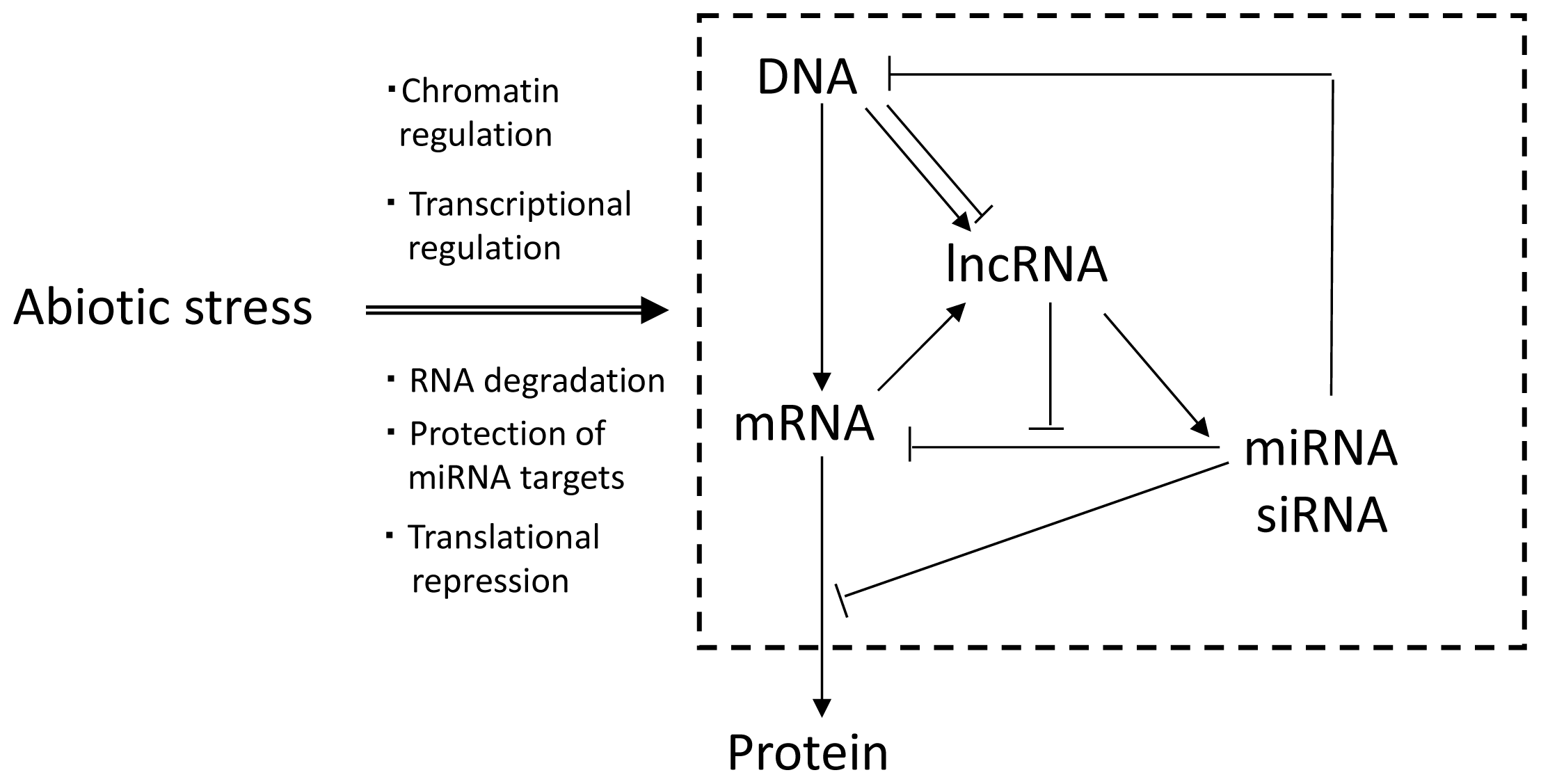
| Small ncRNAs (sRNAs) |
|
| Long ncRNAs (lncRNAs) |
|
Acknowledgments
Conflicts of Interest
References
- Hirayama, T.; Shinozaki, K. Research on plant abiotic stress responses in the postgenome era: Past, present and future. Plant J 2010, 61, 1041–1052. [Google Scholar]
- Laporte, P.; Merchan, F.; Amor, B.B.; Wirth, S.; Crespi, M. Riboregulators in plant development. Biochem. Soc. Trans 2007, 35, 1638–1642. [Google Scholar]
- Rymarquis, L.A.; Kastenmayer, J.P.; Huttenhofer, A.G.; Green, P.J. Diamonds in the rough: mRNA-like non-coding RNAs. Trends Plant Sci 2008, 13, 329–334. [Google Scholar]
- Matsui, A.; Ishida, J.; Morosawa, T.; Mochizuki, Y.; Kaminuma, E.; Endo, T.A.; Okamoto, M.; Nambara, E.; Nakajima, M.; Kawashima, M.; et al. Arabidopsis transcriptome analysis under drought, cold, high-salinity and ABA treatment conditions using a tiling array. Plant Cell Physiol 2008, 49, 1135–1149. [Google Scholar]
- Mazzucotelli, E.; Mastrangelo, A.M.; Crosatti, C.; Guerra, D.; Stanca, A.M.; Cattivelli, L. Abiotic stress response in plants: When post-transcriptional and post-translational regulations control transcription. Plant Sci 2008, 174, 420–431. [Google Scholar]
- Khraiwesh, B.; Zhu, J.K.; Zhu, J. Role of miRNAs and siRNAs in biotic and abiotic stress responses of plants. Biochim. Biophys. Acta 2012, 1819, 137–148. [Google Scholar]
- De Lima, J.C.; Loss-Morais, G.; Margis, R. microRNAs play critical roles during plant development and in response to abiotic stresses. Genet. Mol. Biol 2012, 35, 1069–1077. [Google Scholar]
- Sunkar, R.; Zhu, J.K. Novel and stress-regulated MicroRNAs and other small RNA fromArabidopsis. Plant Cell 2004, 16, 2001–2019. [Google Scholar]
- Voinnet, O. Origin, biogenesis, and activity of plant microRNAs. Cell 2009, 136, 669–687. [Google Scholar]
- Yamaguchi-Shinozaki, K.; Shinozaki, K. A novel cis-acting element in an Arabidopsis gene is involved in responsiveness. Plant Cell 1994, 6, 251–264. [Google Scholar]
- Mundy, J.; Yamaguchi-Shinozaki, K.; Chua, N.H. Nuclear proteins bind conserved elements in the abscisic acid-responsive promoter of a rice rab gene. Proc. Natl. Acad. Sci. USA 1990, 87, 1406–1410. [Google Scholar]
- Shang, Y.; Yan, L.; Liu, Z.Q.; Zheng, C.; Chao, M.; Xin, Q.; Wu, F.Q.; Wang, X.F.; Du, S.Y.; Jiang, T.; et al. The Mg-chelatase H subunit of Arabidopsis antagonizes a group of transcription repressors to relieve ABA-responsive genes of inhibition. Plant Cell 2010, 22, 1909–1935. [Google Scholar]
- Zhou, X.; Wang, G.; Sutoh, K.; Zhu, J.K.; Zhang, W. Identification of cold-inducible microRNAs in plants by transcriptome analysis. Biochim. Biophys. Acta 2008, 1779, 780–788. [Google Scholar]
- Liu, H.H.; Tian, X.; Li, Y.U.; Wu, C.A.; Zheng, C.C. Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA 2008, 14, 836–843. [Google Scholar]
- Zhou, L.; Liu, Y.; Liu, Z.; Kong, D.; Duan, M.; Luo, L. Genome-wide identification and analysis of drought-responsive microRNAs in Oryza sativa. J. Exp. Bot 2010, 61, 4157–4168. [Google Scholar]
- Fuji, H.; Chiou, T.J.; Lin, S.I.; Aung, K.; Zhu, J.K. A miRNA involved in phosphate starvation response in Arabidopsis. Curr. Biol 2005, 15, 2038–2043. [Google Scholar]
- Sunkar, R.; Kapoor, A.; Zhu, J.K. Posttranscriptional induction of two Cu/Zn superoxide dismutase genes in Arabidopsis is mediated by downregulation of miR398 and important for oxidative stress tolerance. Plant Cell 2006, 18, 2051–2065. [Google Scholar]
- Yamasaki, H.; Abdel-Ghany, S.E.; Cohu, C.M.; Kobayashi, Y.; Shikanai, T.; Pilon, M. Regulation of copper homeostasis by micro-RNA in Arabidopsis. J. Biol. Chem 2007, 282, 16369–16378. [Google Scholar]
- Brodersen, P.; Sakvarelidze-Achard, L.; Bruun-Rasmussen, M.; Dunoyer, P.; Yamamoto, Y.Y.; Sieburth, L.; Voinnet, O. Widespread translational inhibition by plant miRNAs and siRNAs. Science 2008, 320, 1185–1190. [Google Scholar]
- Li, W.; Cui, X.; Meng, Z.; Huang, X.; Xie, Q.; Wu, H.; Jin, H.; Zhang, D.; Liang, W. Transcriptional regulation of Arabidopsis miR168a and ARGONAUTE1 homeostasis in abscisic acid and abiotic stress responses. Plant Physiol 2013, 158, 1279–1292. [Google Scholar]
- Liu, P.P.; Montgomery, T.A.; Fahlgren, N.; Kasschau, K.D.; Nonogaki, H.; Carrington, J.C. Repression of AUXIN RESPONSE FACTOR10 by microRNA160 is critical for seed germination and post-germination stages. Plant J 2007, 52, 133–146. [Google Scholar]
- Blomster, T.; Salojarvi, J.; Sipari, N.; Brosche, M.; Ahlfors, R.; Keinanen, M.; Overmyer, K.; Kangasjarvi, J. Apoplastic reactive oxygen species transiently decrease auxin signaling and cause stress-induced morphogenic response inArabidopsis. Plant Physiol 2011, 157, 1866–1883. [Google Scholar]
- Wu, M.F.; Tian, Q.; Reed, J.W. Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 2006, 133, 4211–4218. [Google Scholar]
- Marin, E.; Jouannet, V.; Herz, A.; Lokerse, A.S.; Weijers, D.; Vaucheret, H.; Nussaume, L.; Crespi, M.D.; Maizel, A. miR390, Arabidopsis TAS3 tasiRNAs, and their AUXIN RESPONSE FACTOR targets define an autoregulatory network quantitatively regulating lateral root growth. Plant Cell 2010, 22, 1104–1117. [Google Scholar]
- Iglesias, M.J.; Terrile, M.C.; Bartoli, C.G.; D’Ippolito, S.; Casalongue, C.A. Auxin signaling participates in the adaptative response against oxidative stress and salinity by interacting with redox metabolism inArabidopsis. Plant Mol. Biol 2010, 74, 215–222. [Google Scholar]
- Jones-Rhoades, M.W.; Bartel, D.P. Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol. Cell 2004, 14, 787–799. [Google Scholar]
- Aung, K.; Lin, S.I.; Wu, C.C.; Huang, Y.T.; Su, C.L.; Chiou, T.J. pho2, a phosphate overaccumulator, is caused by a nonsense mutation in a microRNA399 target gene. Plant Physiol 2006, 141, 1000–1011. [Google Scholar]
- Franco-Zorrilla, J.M.; Valli, A.; Todesco, M.; Mateos, I.; Puga, M.I.; Rubio-Somoza, I.; Leyva, A.; Weigel, D.; Garcia, J.A.; Paz-Ares, J. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet 2007, 39, 1033–1037. [Google Scholar]
- Mallory, A.C.; Vaucheret, H. Functions of microRNAs and related small RNAs in plants. Nat. Genet 2006, 38, S31–S36. [Google Scholar]
- Ito, H.; Gaubert, H.; Bucher, E.; Mirouze, M.; Vaillant, I.; Paszkowski, J. An siRNA pathway prevents transgenerational retrotransposition in plants subjected to stress. Nature 2011, 472, 115–119. [Google Scholar]
- Boyko, A.; Blevins, T.; Yao, Y.; Golubov, A.; Bilichak, A.; Ilnytskyy, Y.; Hollunder, J.; Meins, F., Jr.; Kovalchuk, I. Transgenerational adaptation of Arabidopsis to stress requires DNA methylation and the function of Dicer-like proteins. PLoS One 2010, 5, e9514. [Google Scholar]
- Ronemus, M.; Vaughn, M.W.; Martienssen, R.A. MicroRNA-targeted and small interfering RNA-mediated mRNA degradation is regulated by argonaute, dicer, and RNA-dependent RNA polymerase in Arabidopsis. Plant Cell 2006, 18, 1559–1574. [Google Scholar]
- Allen, E.; Xie, Z.; Gustafson, A.M.; Carrington, J.C. MicroRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 2005, 121, 207–221. [Google Scholar]
- Ben Amor, B.; Wirth, S.; Merchan, F.; Laporte, P.; d’Aubenton-Carafa, Y.; Hirsch, J.; Maizel, A.; Mallory, A.; Lucas, A.; Deragon, J.M.; et al. Novel long non-protein coding RNAs involved in Arabidopsis differentiation and stress responses. Genome Res 2009, 19, 57–69. [Google Scholar]
- Yang, J.H.; Seo, H.H.; Han, S.J.; Yoon, E.K.; Yang, M.S.; Lee, W.S. Phytohormone abscisic acid control RNA-dependent RNA polymerase 6 gene expression and post-transcriptional gene silencing in rice cells. Nucleic Acids Res 2008, 36, 1220–1226. [Google Scholar]
- Bartels, D.; Sunkar, R. Drought and salt tolerance in plants. Crit. Rev. Plant. Sci 2005, 24, 23–58. [Google Scholar]
- Borsani, O.; Zhu, J.; Verslues, P.E.; Sunkar, R.; Zhu, J.K. Endogenous siRNAs derived from a pair of natural cis-antisense transcripts regulate salt tolerance in Arabidopsis. Cell 2005, 123, 1279–1291. [Google Scholar]
- Zhang, X.; Lii, Y.; Wu, Z.; Polishko, A.; Zhang, H.; Chinnusamy, V.; Lonardi, S.; Zhu, J.K.; Liu, R.; Jin, H. Mechanisms of small RNA generation from cis-NATs in response to environmental and developmental cues. Mol. Plant 2013, 6, 704–715. [Google Scholar]
- Henz, S.R.; Cumbie, J.S.; Kasschau, K.D.; Lohmann, J.U.; Carrington, J.C.; Weigel, D.; Schmid, M. Distinct expression patterns of natural antisense transcripts in Arabidopsis. Plant Physiol 2007, 144, 1247–1255. [Google Scholar]
- Zhan, S.; Lukens, L. Protein-coding cis-natural antisense transcripts have high and broad expression in Arabidopsis thaliana. Plant Physiol 2013, 161, 2171–2180. [Google Scholar]
- Yamada, K.; Lim, J.; Dale, J.M.; Chen, H.; Shinn, P.; Palm, C.J.; Southwick, A.M.; Wu, H.C.; Kim, C.; Nguyen, M.; et al. Empirical analysis of transcriptional activity in the Arabidopsis genome. Science 2003, 302, 842–846. [Google Scholar]
- Li, L.; Wang, X.F.; Stolc, V.; Li, X.Y.; Zhang, D.F.; Su, N.; Tongprasit, W.; Li, S.G.; Cheng, Z.K.; Wang, J.; et al. Genome-wide transcription analyses in rice using tiling microarrays. Nat. Genet 2006, 38, 124–129. [Google Scholar]
- Lister, R.; O’Malley, R.C.; Tonti-Filippini, J.; Gregory, B.D.; Berry, C.C.; Millar, A.H.; Ecker, J.R. Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell 2008, 133, 523–536. [Google Scholar]
- Laubinger, S.; Zeller, G.; Henz, S.R.; Sachsenberg, T.; Widmer, C.K.; Naouar, N.; Vuylsteke, M.; Scholkopf, B.; Ratsch, G.; Weigel, D. At-TAX: A whole genome tiling array resource for developmental expression analysis and transcript identification in Arabidopsis thaliana. Genome Biol 2008, 9, R112. [Google Scholar]
- Hazen, S.P.; Naef, F.; Quisel, T.; Gendron, J.M.; Chen, H.; Ecker, J.R.; Borevitz, J.O.; Kay, S.A. Exploring the transcriptional landscape of plant circadian rhythms using genome tiling arrays. Genome Biol 2009, 10, R17. [Google Scholar]
- Filichkin, S.A.; Priest, H.D.; Givan, S.A.; Shen, R.; Bryant, D.W.; Fox, S.E.; Wong, W.K.; Mockler, T.C. Genome-wide mapping of alternative splicing in Arabidopsis thaliana. Genome Res 2010, 20, 45–58. [Google Scholar]
- Swiezewski, S.; Liu, F.; Magusin, A.; Dean, C. Cold-induced silencing by long antisense transcripts of an Arabidopsis Polycomb target. Nature 2009, 462, 799–802. [Google Scholar]
- Luo, C.; Sidote, D.J.; Zhang, Y.; Kerstetter, R.A.; Michael, T.P.; Lam, E. Integrative analysis of chromatin states in Arabidopsis identified potential regulatory mechanisms for natural antisense transcript production. Plant J 2013, 73, 77–90. [Google Scholar]
- Rajeswaran, R.; Aregger, M.; Zvereva, A.S.; Borah, B.K.; Gubaeva, E.G.; Pooggin, M.M. Sequencing of RDR6-dependent double-stranded RNAs reveals novel features of plant siRNA biogenesis. Nucleic Acids Res 2012, 40, 6241–6254. [Google Scholar]
- Ner-Gaon, H.; Halachmi, R.; Savaldi-Goldstein, S.; Rubin, E.; Ophir, R.; Fluhr, R. Intron retention is a major phenomenon in alternative splicing in Arabidopsis. Plant J 2004, 39, 877–885. [Google Scholar]
- Hanada, K.; Higuchi-Takeuchi, M.; Okamoto, M.; Yoshizumi, T.; Shimizu, M.; Nakaminami, K.; Nishi, R.; Ohashi, C.; Iida, K.; Tanaka, M.; et al. Small open reading frames associated with morphogenesis are hidden in plant genomes. Proc. Natl. Acad. Sci. USA 2013, 110, 2395–2400. [Google Scholar]
- Moghe, G.D.; Lehti-Shiu, M.D.; Seddon, A.E.; Yin, S.; Chen, Y.; Juntawong, P.; Brandizzi, F.; Bailey-Serres, J.; Shiu, S.H. Characteristics and significance of intergenic polyadenylated RNA transcription in Arabidopsis. Plant Physiol 2013, 161, 1210–1224. [Google Scholar]
- Wu, J.; Okada, T.; Fukushima, T.; Tsudzuki, T.; Sugiura, M.; Yukawa, Y. A novel hypoxic stress-responsive long non-coding RNA transcribed by RNA polymerase III in Arabidopsis. RNA Biol 2012, 9, 302–313. [Google Scholar]
- Kapranov, P.; Willingham, A.T.; Gingeras, T.R. Genome-wide transcription and the implications for genomic organization. Nat. Rev. Genet 2007, 8, 413–423. [Google Scholar]
- Beclin, C.; Boutet, S.; Waterhouse, P.; Vaucheret, H. A branched pathway for transgene-induced RNA silencing in plants. Curr. Biol 2002, 12, 684–688. [Google Scholar]
- Yoshikawa, M.; Peragine, A.; Park, M.Y.; Poethig, R.S. A pathway for the biogenesis of trans-acting siRNAs in Arabidopsis. Genes Dev 2005, 19, 2164–2175. [Google Scholar]
- Zheng, Q.; Ryvkin, P.; Li, F.; Dragomir, I.; Valladares, O.; Yang, J.; Cao, K.; Wang, L.S.; Gregory, B.D. Genome-wide double-stranded RNA sequencing reveals the functional significance of base-paired RNAs in Arabidopsis. PLoS Genet 2010, 6, e1001141. [Google Scholar]
- Gregory, B.D.; O’Malley, R.C.; Lister, R.; Urich, M.A.; Tonti-Filippini, J.; Chen, H.; Millar, A.H.; Ecker, J.R. A link between RNA metabolism and silencing affecting Arabidopsis development. Dev. Cell 2008, 14, 854–866. [Google Scholar]
- Gazzani, S.; Lawrenson, T.; Woodward, C.; Headon, D.; Sablowski, R. A link between mRNA turnover and RNA interference in Arabidopsis. Science 2004, 306, 1046–1048. [Google Scholar]
- Dunoyer, P.; Himber, C.; Voinnet, O. DICER-LIKE 4 is required for RNA interference and produces the 21-nucleotide small interfering RNA component of the plant cell-to-cell silencing signal. Nat. Genet 2005, 37, 1356–1360. [Google Scholar]
- Xie, Z.; Allen, E.; Fahlgren, N.; Calamar, A.; Givan, S.A.; Carrington, J.C. Expression of Arabidopsis miRNA genes. Plant Physiol 2005, 138, 2145–2154. [Google Scholar]
- Jauvion, V.; Rivard, M.; Bouteiller, N.; Elmayan, T.; Vaucheret, H. RDR2 partially antagonizes the production of RDR6-dependent siRNA in sense transgene-mediated PTGS. PLoS One 2012, 7, e29785. [Google Scholar]
- Glazov, E.; Phillips, K.; Budziszewski, G.J.; Schob, H.; Meins, F., Jr.; Levin, J.Z. A gene encoding an RNase D exonuclease-like protein is required for post-transcriptional silencing in Arabidopsis. Plant J 2003, 35, 342–349. [Google Scholar]
- Herr, A.J.; Jensen, M.B.; Dalmay, T.; Baulcombe, D.C. RNA polymerase IV directs silencing of endogenous DNA. Science 2005, 308, 118–120. [Google Scholar]
- Dalmay, T.; Horsefield, R.; Braunstein, T.H.; Baulcombe, D.C. SDE3 encodes an RNA helicase required for post-transcriptional gene silencing in Arabidopsis. EMBO J 2001, 20, 2069–2078. [Google Scholar]
- Liu, F.; Marquardt, S.; Lister, C.; Swiezewski, S.; Dean, C. Targeted 3′ processing of antisense transcripts triggers Arabidopsis FLC chromatin silencing. Science 2010, 327, 94–97. [Google Scholar]
- Angel, A.; Song, J.; Dean, C.; Howard, M. A Polycomb-based switch underlying quantitative epigenetic memory. Nature 2011, 476, 105–108. [Google Scholar]
- De Lucia, F.; Crevillen, P.; Jones, A.M.; Greb, T.; Dean, C. A PHD-polycomb repressive complex 2 triggers the epigenetic silencing of FLC during vernalization. Proc. Natl. Acad. Sci. USA 2008, 105, 16831–16836. [Google Scholar]
- Helliwell, C.A.; Robertson, M.; Finnegan, E.J.; Buzas, D.M.; Dennis, E.S. Vernalization-repression of Arabidopsis FLC requires promoter sequences but not antisense transcripts. PLoS One 2011, 6, e2151. [Google Scholar]
- Sun, Q.; Csorba, T.; Skourti-Stathaki, K.; Proudfoot, N.J.; Dean, C. R-loop stabilization represses antisense transcription at the Arabidopsis FLC locus. Science 2013, 340, 619–621. [Google Scholar]
- Heo, J.B.; Sung, S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 2011, 331, 76–79. [Google Scholar]
- Wang, K.C.; Chang, H.Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar]
- Kramerov, D.A.; Vassetzky, N.S. Short retroposons in eukaryotic genomes. Int. Rev. Cytol 2005, 247, 165–221. [Google Scholar]
- Yukawa, Y.; Felis, M.; Englert, M.; Stojanov, M.; Matousek, J.; Beier, H.; Sugiura, M. Plant 7SL RNA genes belong to type 4 of RNA polymerase III- dependent genes that are composed of mixed promoters. Plant J 2005, 43, 97–106. [Google Scholar]
- Shin, H.; Shin, H.S.; Chen, R.; Harrison, M.J. Loss of At4 function impacts phosphate distribution between the roots and the shoots during phosphate starvation. Plant J 2006, 45, 712–726. [Google Scholar]
- Lin, S.I.; Chiang, S.F.; Lin, W.Y.; Chen, J.W.; Tseng, C.Y.; Wu, P.C.; Chiou, T.J. Regulatory network of microRNA399 and PHO2 by systemic signaling. Plant Physiol 2008, 147, 732–746. [Google Scholar]
- Pant, B.D.; Buhtz, A.; Kehr, J.; Scheible, W.R. MicroRNA399 is a long-distance signal for the regulation of plant phosphate homeostasis. Plant J 2008, 53, 731–738. [Google Scholar]
- Wu, H.J.; Wang, Z.M.; Wang, M.; Wang, X.J. Wide-spread long non-coding RNAs (lncRNAs) as endogenous target mimics (eTMs) for microRNAs in plants. Plant Physiol 2013, 161, 1875–1884. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Matsui, A.; Nguyen, A.H.; Nakaminami, K.; Seki, M. Arabidopsis Non-Coding RNA Regulation in Abiotic Stress Responses. Int. J. Mol. Sci. 2013, 14, 22642-22654. https://doi.org/10.3390/ijms141122642
Matsui A, Nguyen AH, Nakaminami K, Seki M. Arabidopsis Non-Coding RNA Regulation in Abiotic Stress Responses. International Journal of Molecular Sciences. 2013; 14(11):22642-22654. https://doi.org/10.3390/ijms141122642
Chicago/Turabian StyleMatsui, Akihiro, Anh Hai Nguyen, Kentaro Nakaminami, and Motoaki Seki. 2013. "Arabidopsis Non-Coding RNA Regulation in Abiotic Stress Responses" International Journal of Molecular Sciences 14, no. 11: 22642-22654. https://doi.org/10.3390/ijms141122642




