Modeling Human Liver Biology Using Stem Cell-Derived Hepatocytes
Abstract
:1. Introduction
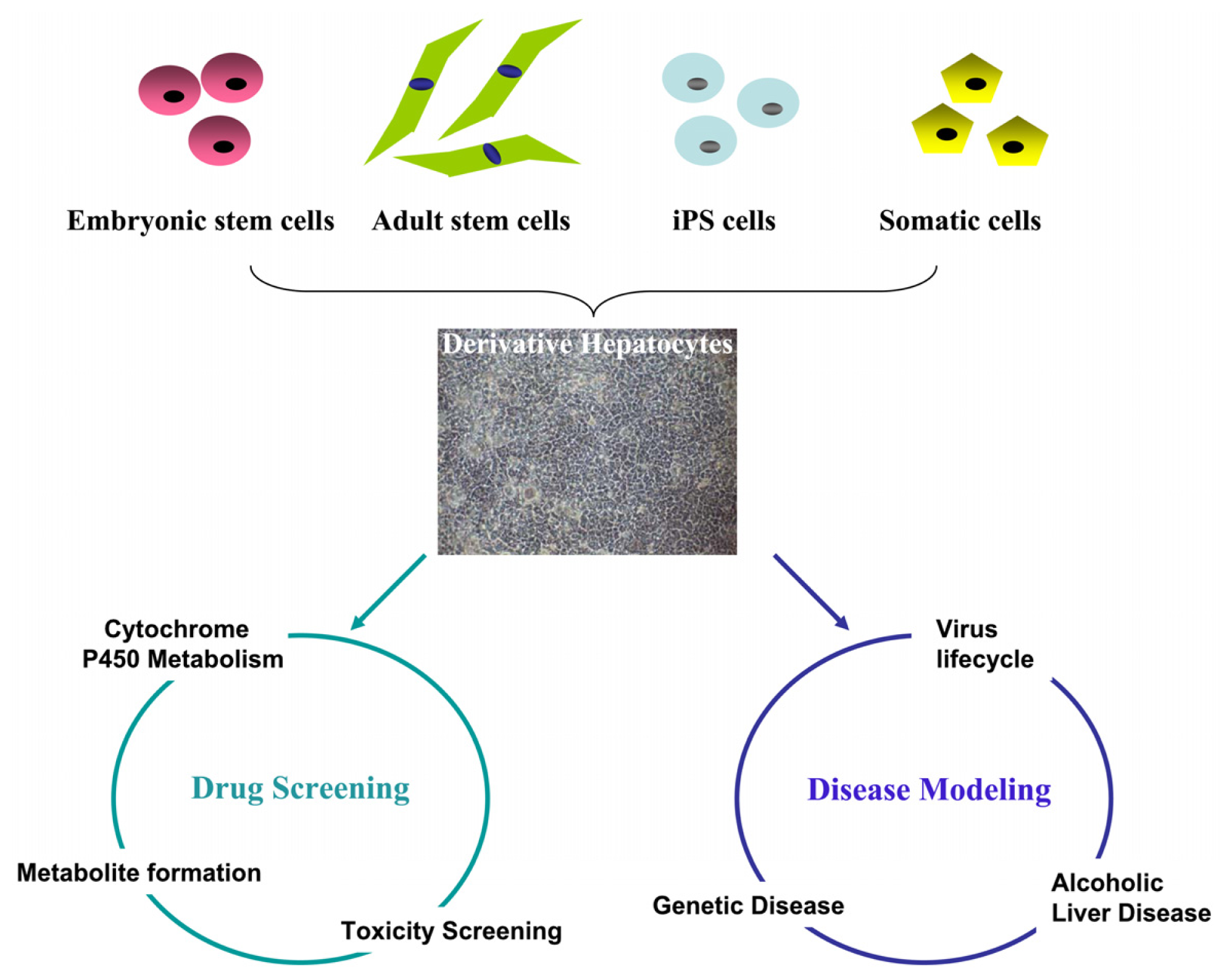
2. Generation of In Vitro-Derived Hepatocytes from Stem Cell or Somatic Cells
2.1. Hepatocytes Derived from Human Embryonic Stem Cells (hESCs)
2.2. Hepatocytes Derived from Induced Pluripotent Stem Cells (iPSCs)
2.3. Hepatocytes from Direct Reprogramming Somatic Cells
3. Modeling Human Drug Metabolism Using Stem Cell-Derived Hepatocytes
3.1. Drug Attrition
3.2. Stem Cell-Derived Hepatocytes
4. Modeling Human Disease Using Stem Cell-Derived Hepatocytes
4.1. Modeling Human Genetic Liver Disorders
4.2. Hepatitis Virus Infection and Replication
4.3. Alcoholic Liver Disease
5. Challenge and Prospect of Stem Cell-Derived Hepatocytes
Acknowledgments
Conflicts of Interest
References
- Kmiec, Z. Cooperation of liver cells in health and disease. Adv. Anat. Embryol. Cell Biol 2001, 161, 1–151. [Google Scholar]
- Espanol-Suner, R.; Carpentier, R.; van Hul, N.; Legry, V.; Achouri, Y.; Cordi, S.; Jacquemin, P.; Lemaigre, F.; Leclercq, I.A. Liver progenitor cells yield functional hepatocytes in response to chronic liver injury in mice. Gastroenterology 2012. [Google Scholar] [CrossRef]
- Thiele, G.M.; Klassen, L.W.; Tuma, D.J. Formation and immunological properties of aldehyde-derived protein adducts following alcohol consumption. Methods Mol. Biol 2008, 447, 235–257. [Google Scholar]
- Lecluyse, E.L.; Alexandre, E. Isolation and culture of primary hepatocytes from resected human liver tissue. Methods Mol. Biol 2010, 640, 57–82. [Google Scholar]
- Gomez-Lechon, M.J.; Donato, M.T.; Castell, J.V.; Jover, R. Human hepatocytes in primary culture: The choice to investigate drug metabolism in man. Curr. Drug Metab 2004, 5, 443–462. [Google Scholar]
- Hewitt, N.J.; Lechon, M.J.; Houston, J.B.; Hallifax, D.; Brown, H.S.; Maurel, P.; Kenna, J.G.; Gustavsson, L.; Lohmann, C.; Skonberg, C.; et al. Primary hepatocytes: Current understanding of the regulation of metabolic enzymes and transporter proteins, and pharmaceutical practice for the use of hepatocytes in metabolism, enzyme induction, transporter, clearance, and hepatotoxicity studies. Drug Metab. Rev 2007, 39, 159–234. [Google Scholar]
- Donato, M.T.; Lahoz, A.; Castell, J.V.; Gomez-Lechon, M.J. Cell lines: A tool for in vitro drug metabolism studies. Curr. Drug Metab 2008, 9, 1–11. [Google Scholar]
- Meylan, E.; Curran, J.; Hofmann, K.; Moradpour, D.; Binder, M.; Bartenschlager, R.; Tschopp, J. Cardif is an adaptor protein in the RIG-I antiviral pathway and is targeted by hepatitis C virus. Nature 2005, 437, 1167–1172. [Google Scholar]
- Jover, R.; Bort, R.; Gomez-Lechon, M.J.; Castell, J.V. Re-expression of C/EBP alpha induces CYP2B6, CYP2C9 and CYP2D6 genes in HepG2 cells. FEBS Lett 1998, 431, 227–230. [Google Scholar]
- Schrem, H.; Klempnauer, J.; Borlak, J. Liver-enriched transcription factors in liver function and development. Part II: The C/EBPs and D site-binding protein in cell cycle control, carcinogenesis, circadian gene regulation, liver regeneration, apoptosis, and liver-specific gene regulation. Pharmacol. Rev 2004, 56, 291–330. [Google Scholar]
- Wong, N.; Lai, P.; Pang, E.; Leung, T.W.; Lau, J.W.; Johnson, P.J. A comprehensive karyotypic study on human hepatocellular carcinoma by spectral karyotyping. Hepatology 2000, 32, 1060–1068. [Google Scholar]
- Thomson, J.A.; Itskovitz-Eldor, J.; Shapiro, S.S.; Waknitz, M.A.; Swiergiel, J.J.; Marshall, V.S.; Jones, J.M. Embryonic stem cell lines derived from human blastocysts. Science 1998, 282, 1145–1147. [Google Scholar]
- Reubinoff, B.E.; Pera, M.F.; Fong, C.Y.; Trounson, A.; Bongso, A. Embryonic stem cell lines from human blastocysts: Somatic differentiation in vitro. Nat. Biotechnol. 2000, 18, 399–404. [Google Scholar]
- Zhou, W.; Hannoun, Z.; Jaffray, E.; Medine, C.N.; Black, J.R.; Greenhough, S.; Zhu, L.; Ross, J.A.; Forbes, S.; Wilmut, I.; et al. SUMOylation of HNF4α regulates protein stability and hepatocyte function. J. Cell Sci 2012, 125, 3630–3635. [Google Scholar]
- Lavon, N.; Benvenisty, N. Study of hepatocyte differentiation using embryonic stem cells. J. Cell. Biochem 2005, 96, 1193–1202. [Google Scholar]
- Basma, H.; Soto-Gutierrez, A.; Yannam, G.R.; Liu, L.; Ito, R.; Yamamoto, T.; Ellis, E.; Carson, S.D.; Sato, S.; Chen, Y.; et al. Differentiation and transplantation of human embryonic stem cell-derived hepatocytes. Gastroenterology 2009, 136, 990–999. [Google Scholar]
- Duan, Y.; Catana, A.; Meng, Y.; Yamamoto, N.; He, S.; Gupta, S.; Gambhir, S.S.; Zern, M.A. Differentiation and enrichment of hepatocyte-like cells from human embryonic stem cells in vitro and in vivo. Stem Cells 2007, 25, 3058–3068. [Google Scholar]
- Hay, D.C.; Fletcher, J.; Payne, C.; Terrace, J.D.; Gallagher, R.C.; Snoeys, J.; Black, J.R.; Wojtacha, D.; Samuel, K.; Hannoun, Z.; et al. Highly efficient differentiation of hESCs to functional hepatic endoderm requires ActivinA and Wnt3a signaling. Proc. Natl. Acad. Sci. USA 2008, 105, 12301–12306. [Google Scholar]
- Itskovitz-Eldor, J.; Schuldiner, M.; Karsenti, D.; Eden, A.; Yanuka, O.; Amit, M.; Soreq, H.; Benvenisty, N. Differentiation of human embryonic stem cells into embryoid bodies compromising the three embryonic germ layers. Mol. Med 2000, 6, 88–95. [Google Scholar]
- Imamura, T.; Cui, L.; Teng, R.; Johkura, K.; Okouchi, Y.; Asanuma, K.; Ogiwara, N.; Sasaki, K. Embryonic stem cell-derived embryoid bodies in three-dimensional culture system form hepatocyte-like cells in vitro and in vivo. Tissue Eng. 2004, 10, 1716–1724. [Google Scholar]
- Vosough, M.; Omidinia, E.; Kadivar, M.; Shokrgozar, M.A.; Pournasr, B.; Aghdami, N.; Baharvand, H. Generation of functional hepatocyte-like cells from human pluripotent stem cells in a scalable suspension culture. Stem Cells Dev 2013, 22, 2693–2705. [Google Scholar]
- Hay, D.C.; Zhao, D.; Ross, A.; Mandalam, R.; Lebkowski, J.; Cui, W. Direct differentiation of human embryonic stem cells to hepatocyte-like cells exhibiting functional activities. Cloning Stem Cells 2007, 9, 51–62. [Google Scholar]
- Agarwal, S.; Holton, K.L.; Lanza, R. Efficient differentiation of functional hepatocytes from human embryonic stem cells. Stem Cells 2008, 26, 1117–1127. [Google Scholar]
- Hannoun, Z.; Fletcher, J.; Greenhough, S.; Medine, C.; Samuel, K.; Sharma, R.; Pryde, A.; Black, J.R.; Ross, J.A.; Wilmut, I.; et al. The comparison between conditioned media and serum-free media in human embryonic stem cell culture and differentiation. Cell Reprogram 2010, 12, 133–140. [Google Scholar]
- Hay, D.C.; Pernagallo, S.; Diaz-Mochon, J.J.; Medine, C.N.; Greenhough, S.; Hannoun, Z.; Schrader, J.; Black, J.R.; Fletcher, J.; Dalgetty, D.; et al. Unbiased screening of polymer libraries to define novel substrates for functional hepatocytes with inducible drug metabolism. Stem Cell Res 2011, 6, 92–102. [Google Scholar]
- Hay, D.C. Rapid and scalable human stem cell differentiation: Now in 3D. Stem Cells Dev 2013, 22, 2691–2692. [Google Scholar]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar]
- Dalgetty, D.M.; Medine, C.N.; Iredale, J.P.; Hay, D.C. Progress and future challenges in stem cell-derived liver technologies. Am. J. Physiol. Gastrointest. Liver Physiol 2009, 297, G241–G248. [Google Scholar]
- Sullivan, G.J.; Hay, D.C.; Park, I.H.; Fletcher, J.; Hannoun, Z.; Payne, C.M.; Dalgetty, D.; Black, J.R.; Ross, J.A.; Samuel, K.; et al. Generation of functional human hepatic endoderm from human induced pluripotent stem cells. Hepatology 2010, 51, 329–335. [Google Scholar]
- Kajiwara, M.; Aoi, T.; Okita, K.; Takahashi, R.; Inoue, H.; Takayama, N.; Endo, H.; Eto, K.; Toguchida, J.; Uemoto, S.; et al. Donor-dependent variations in hepatic differentiation from human-induced pluripotent stem cells. Proc. Natl. Acad. Sci. USA 2012, 109, 12538–12543. [Google Scholar]
- Takahashi, K.; Tanabe, K.; Ohnuki, M.; Narita, M.; Ichisaka, T.; Tomoda, K.; Yamanaka, S. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 2007, 131, 861–872. [Google Scholar]
- Yu, J.; Vodyanik, M.A.; Smuga-Otto, K.; Antosiewicz-Bourget, J.; Frane, J.L.; Tian, S.; Nie, J.; Jonsdottir, G.A.; Ruotti, V.; Stewart, R.; et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 2007, 318, 1917–1920. [Google Scholar]
- Huang, P.; He, Z.; Ji, S.; Sun, H.; Xiang, D.; Liu, C.; Hu, Y.; Wang, X.; Hui, L. Induction of functional hepatocyte-like cells from mouse fibroblasts by defined factors. Nature 2011, 475, 386–389. [Google Scholar]
- Sekiya, S.; Suzuki, A. Direct conversion of mouse fibroblasts to hepatocyte-like cells by defined factors. Nature 2011, 475, 390–393. [Google Scholar]
- Yu, B.; He, Z.-Y.; You, P.; Han, Q.-W.; Xiang, D.; Chen, F.; Wang, M.-J.; Liu, C.-C.; Lin, X.-W.; Borjigin, U. Reprogramming fibroblasts into bipotential hepatic stem cells by defined factors. Cell Stem Cell 2013, 13, 328–340. [Google Scholar]
- Vierbuchen, T.; Ostermeier, A.; Pang, Z.P.; Kokubu, Y.; Sudhof, T.C.; Wernig, M. Direct conversion of fibroblasts to functional neurons by defined factors. Nature 2010, 463, 1035–1041. [Google Scholar]
- Ieda, M.; Fu, J.D.; Delgado-Olguin, P.; Vedantham, V.; Hayashi, Y.; Bruneau, B.G.; Srivastava, D. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell 2010, 142, 375–386. [Google Scholar]
- Szabo, E.; Rampalli, S.; Risueno, R.M.; Schnerch, A.; Mitchell, R.; Fiebig-Comyn, A.; Levadoux-Martin, M.; Bhatia, M. Direct conversion of human fibroblasts to multilineage blood progenitors. Nature 2010, 468, 521–526. [Google Scholar]
- Kola, I.; Landis, J. Can the pharmaceutical industry reduce attrition rates? Nat. Rev. Drug Discov 2004, 3, 711–715. [Google Scholar]
- Orloff, J.; Douglas, F.; Pinheiro, J.; Levinson, S.; Branson, M.; Chaturvedi, P.; Ette, E.; Gallo, P.; Hirsch, G.; Mehta, C.; et al. The future of drug development: Advancing clinical trial design. Nat. Rev. Drug Discov 2009, 8, 949–957. [Google Scholar]
- Gomez-Lechon, M.J.; Lahoz, A.; Gombau, L.; Castell, J.V.; Donato, M.T. In vitro evaluation of potential hepatotoxicity induced by drugs. Curr. Pharm. Des 2010, 16, 1963–1977. [Google Scholar]
- Baranczewski, P.; Stanczak, A.; Sundberg, K.; Svensson, R.; Wallin, A.; Jansson, J.; Garberg, P.; Postlind, H. Introduction to in vitro estimation of metabolic stability and drug interactions of new chemical entities in drug discovery and development. Pharmacol. Rep 2006, 58, 453–472. [Google Scholar]
- Medine, C.N.; Lucendo-Villarin, B.; Storck, C.; Wang, F.; Szkolnicka, D.; Khan, F.; Pernagallo, S.; Black, J.R.; Marriage, H.M.; Ross, J.A.; et al. Developing high-fidelity hepatotoxicity models from pluripotent stem cells. Stem Cells Transl. Med 2013, 2, 505–509. [Google Scholar]
- Guengerich, F.P. Cytochrome p450 and chemical toxicology. Chem. Res. Toxicol 2008, 21, 70–83. [Google Scholar]
- Rodrigues, A.D. Integrated cytochrome P450 reaction phenotyping: Attempting to bridge the gap between cDNA-expressed cytochromes P450 and native human liver microsomes. Biochem. Pharmacol 1999, 57, 465–480. [Google Scholar]
- Thummel, K.E.; Wilkinson, G.R. In vitro and in vivo drug interactions involving human CYP3A. Annu. Rev. Pharmacol. Toxicol 1998, 38, 389–430. [Google Scholar]
- Giacomini, K.M.; Huang, S.M.; Tweedie, D.J.; Benet, L.Z.; Brouwer, K.L.; Chu, X.; Dahlin, A.; Evers, R.; Fischer, V.; Hillgren, K.M.; et al. Membrane transporters in drug development. Nat. Rev. Drug Discov 2010, 9, 215–236. [Google Scholar]
- Greenhough, S.; Medine, C.N.; Hay, D.C. Pluripotent stem cell derived hepatocyte like cells and their potential in toxicity screening. Toxicology 2010, 278, 250–255. [Google Scholar]
- Yildirimman, R.; Brolen, G.; Vilardell, M.; Eriksson, G.; Synnergren, J.; Gmuender, H.; Kamburov, A.; Ingelman-Sundberg, M.; Castell, J.; Lahoz, A.; et al. Human embryonic stem cell derived hepatocyte-like cells as a tool for in vitro hazard assessment of chemical carcinogenicity. Toxicol. Sci 2011, 124, 278–290. [Google Scholar]
- Ghodsizadeh, A.; Taei, A.; Totonchi, M.; Seifinejad, A.; Gourabi, H.; Pournasr, B.; Aghdami, N.; Malekzadeh, R.; Almadani, N.; Salekdeh, G.H.; et al. Generation of liver disease-specific induced pluripotent stem cells along with efficient differentiation to functional hepatocyte-like cells. Stem Cell Rev 2010, 6, 622–632. [Google Scholar]
- Rashid, S.T.; Corbineau, S.; Hannan, N.; Marciniak, S.J.; Miranda, E.; Alexander, G.; Huang-Doran, I.; Griffin, J.; Ahrlund-Richter, L.; Skepper, J.; et al. Modeling inherited metabolic disorders of the liver using human induced pluripotent stem cells. J. Clin. Investig 2010, 120, 3127–3136. [Google Scholar]
- Zhang, S.; Chen, S.; Li, W.; Guo, X.; Zhao, P.; Xu, J.; Chen, Y.; Pan, Q.; Liu, X.; Zychlinski, D.; et al. Rescue of ATP7B function in hepatocyte-like cells from Wilson’s disease induced pluripotent stem cells using gene therapy or the chaperone drug curcumin. Hum. Mol. Genet 2011, 20, 3176–3187. [Google Scholar]
- Wu, X.; Robotham, J.M.; Lee, E.; Dalton, S.; Kneteman, N.M.; Gilbert, D.M.; Tang, H. Productive hepatitis C virus infection of stem cell-derived hepatocytes reveals a critical transition to viral permissiveness during differentiation. PLoS Pathog 2012, 8, e1002617. [Google Scholar]
- Roelandt, P.; Obeid, S.; Paeshuyse, J.; Vanhove, J.; van Lommel, A.; Nahmias, Y.; Nevens, F.; Neyts, J.; Verfaillie, C.M. Human pluripotent stem cell-derived hepatocytes support complete replication of hepatitis C virus. J. Hepatol 2012, 57, 246–251. [Google Scholar]
- Schwartz, R.E.; Trehan, K.; Andrus, L.; Sheahan, T.P.; Ploss, A.; Duncan, S.A.; Rice, C.M.; Bhatia, S.N. Modeling hepatitis C virus infection using human induced pluripotent stem cells. Proc. Natl. Acad. Sci. USA 2012, 109, 2544–2548. [Google Scholar]
- Ganem, D.; Prince, A.M. Hepatitis B virus infection--natural history and clinical consequences. N. Engl. J. Med 2004, 350, 1118–1129. [Google Scholar]
- Paganelli, M.; Dallmeier, K.; Nyabi, O.; Scheers, I.; Kabamba, B.; Neyts, J.; Goubau, P.; Najimi, M.; Sokal, E.M. Differentiated umbilical cord matrix stem cells as a newin vitromodel to study early events during hepatitis B virus infection. Hepatology 2013, 57, 59–69. [Google Scholar]
- Taylor, J.M. Hepatitis delta virus. Virology 2006, 344, 71–76. [Google Scholar]
- Yan, H.; Zhong, G.; Xu, G.; He, W.; Jing, Z.; Gao, Z.; Huang, Y.; Qi, Y.; Peng, B.; Wang, H.; et al. Sodium taurocholate cotransporting polypeptide is a functional receptor for human hepatitis B and D virus. Elife 2012, 1, e00049. [Google Scholar]
- O’Shea, R.S.; Dasarathy, S.; McCullough, A.J. Alcoholic liver disease. Hepatology 2010, 51, 307–328. [Google Scholar]
- D’Souza El-Guindy, N.B.; Kovacs, E.J.; de Witte, P.; Spies, C.; Littleton, J.M.; de Villiers, W.J.; Lott, A.J.; Plackett, T.P.; Lanzke, N.; Meadows, G.G. Laboratory models available to study alcohol-induced organ damage and immune variations: Choosing the appropriate model. Alcohol. Clin. Exp. Res 2010, 34, 1489–1511. [Google Scholar]
- Seitz, H.K.; Stickel, F. Acetaldehyde as an underestimated risk factor for cancer development: Role of genetics in ethanol metabolism. Genes Nutr 2010, 5, 121–128. [Google Scholar]
- Pal, R.; Mamidi, M.K.; Das, A.K.; Gupta, P.K.; Bhonde, R. A simple and economical route to generate functional hepatocyte-like cells from hESCs and their application in evaluating alcohol induced liver damage. J. Cell. Biochem 2012, 113, 19–30. [Google Scholar]
- Szkolnicka, D.; Zhou, W.; Lucendo-Villarin, B.; Hay, D.C. Pluripotent stem cell-derived hepatocytes: Potential and challenges in pharmacology. Annu. Rev. Pharmacol. Toxicol 2013, 53, 147–159. [Google Scholar]
- Szkolnicka, D.; Farnworth, S.; Lucendo-Villarin, B.; Storck, C.; Zhou, W.; Iredale, J.P.; Flint, O.; Hay, D.C. Accurate prediction of potential liver injury using stem cell derived populations. Stem Cells Transl. Med 2014. accepted for publication. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Sun, P.; Zhou, X.; Farnworth, S.L.; Patel, A.H.; Hay, D.C. Modeling Human Liver Biology Using Stem Cell-Derived Hepatocytes. Int. J. Mol. Sci. 2013, 14, 22011-22021. https://doi.org/10.3390/ijms141122011
Sun P, Zhou X, Farnworth SL, Patel AH, Hay DC. Modeling Human Liver Biology Using Stem Cell-Derived Hepatocytes. International Journal of Molecular Sciences. 2013; 14(11):22011-22021. https://doi.org/10.3390/ijms141122011
Chicago/Turabian StyleSun, Pingnan, Xiaoling Zhou, Sarah L. Farnworth, Arvind H. Patel, and David C. Hay. 2013. "Modeling Human Liver Biology Using Stem Cell-Derived Hepatocytes" International Journal of Molecular Sciences 14, no. 11: 22011-22021. https://doi.org/10.3390/ijms141122011



