Characterization of a Polyamine Microsphere and Its Adsorption for Protein
Abstract
:1. Introduction
2. Results and Discussion
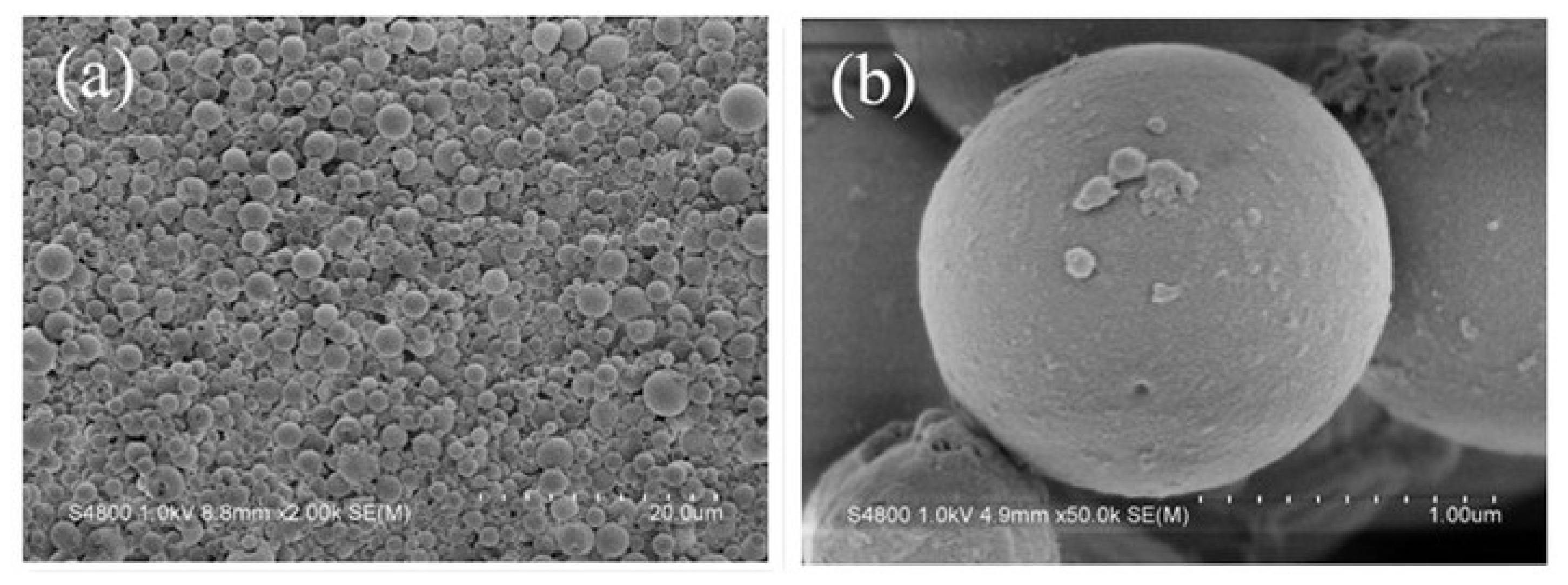
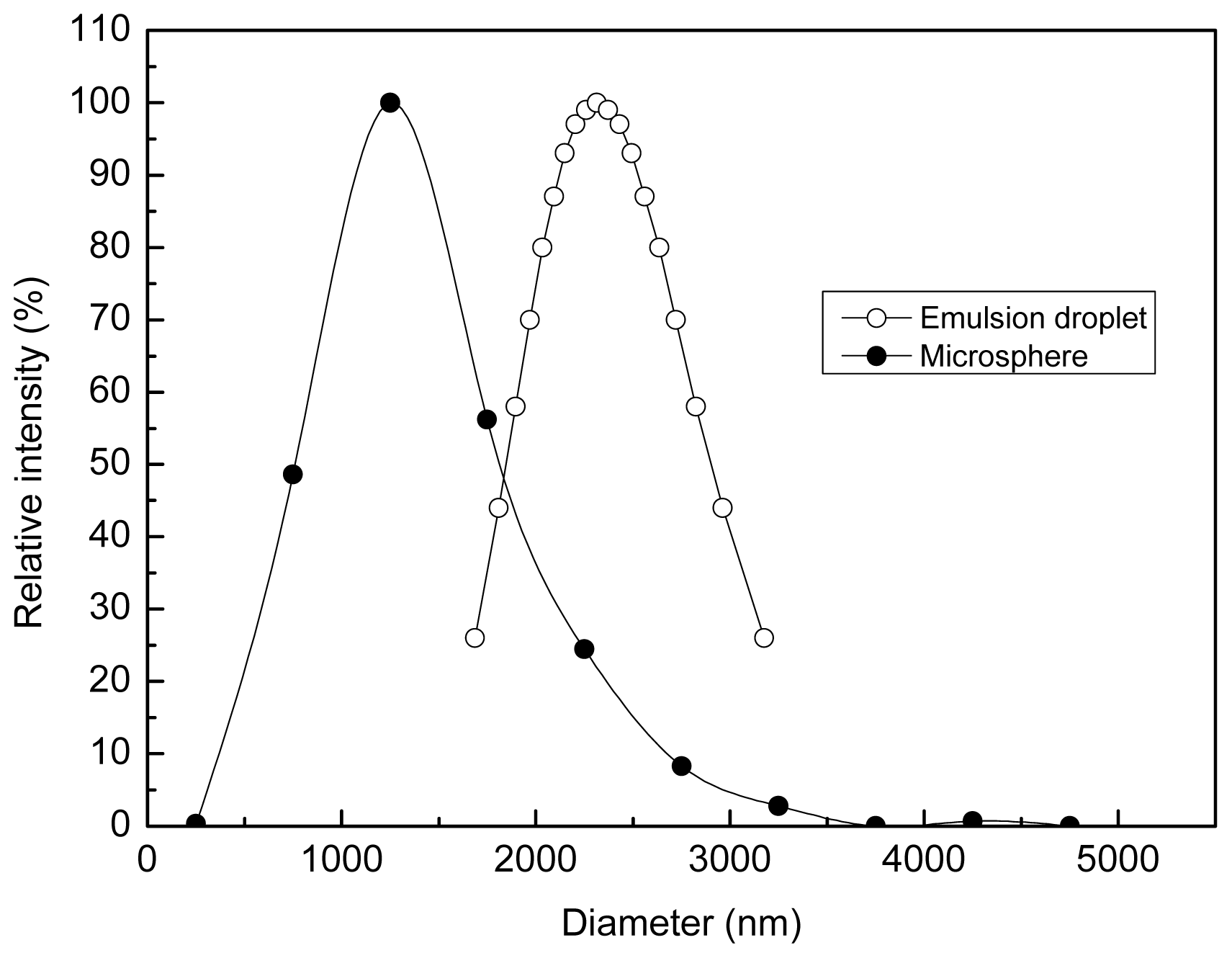
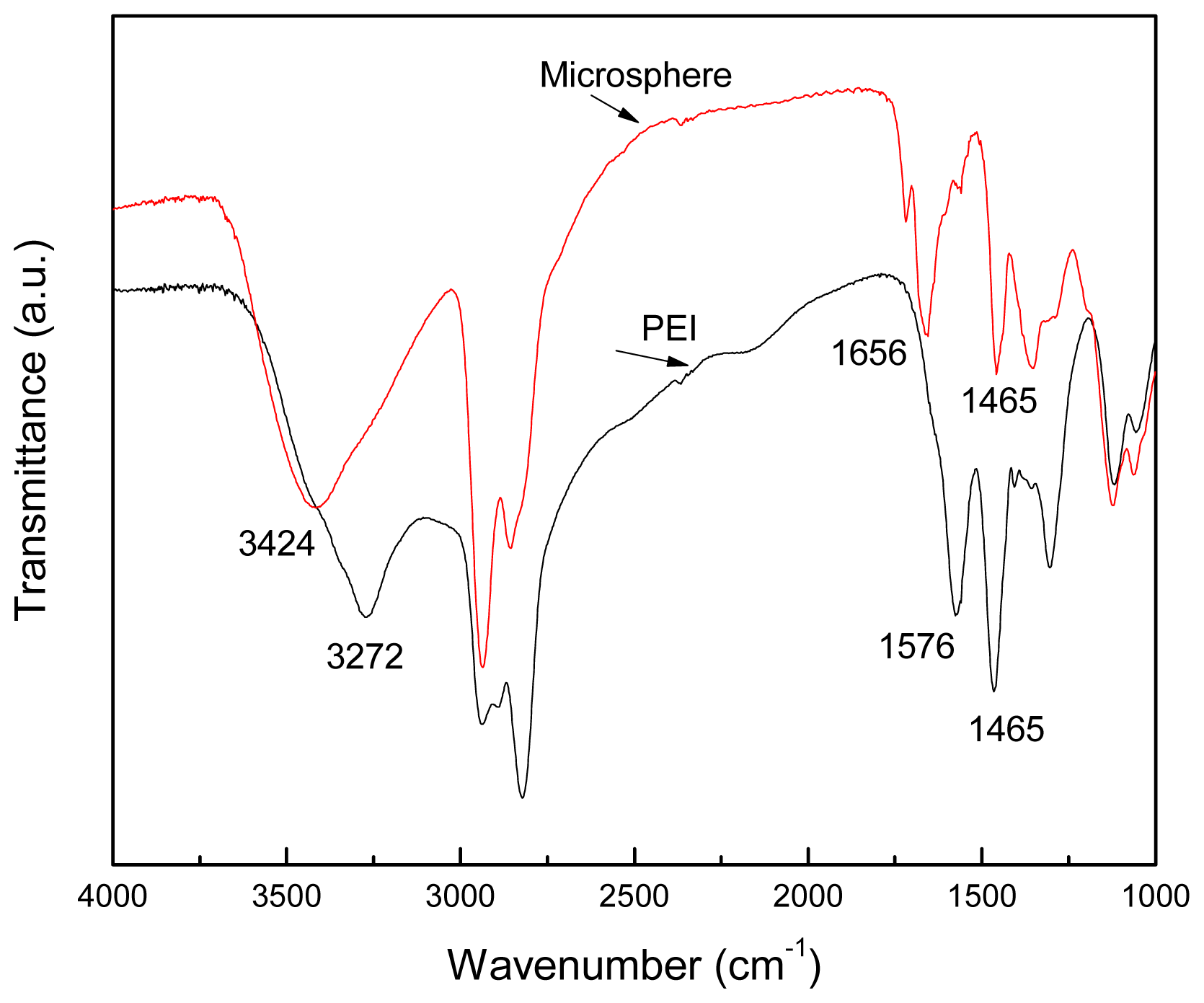
2.1. Structure of the Polyamine Microsphere
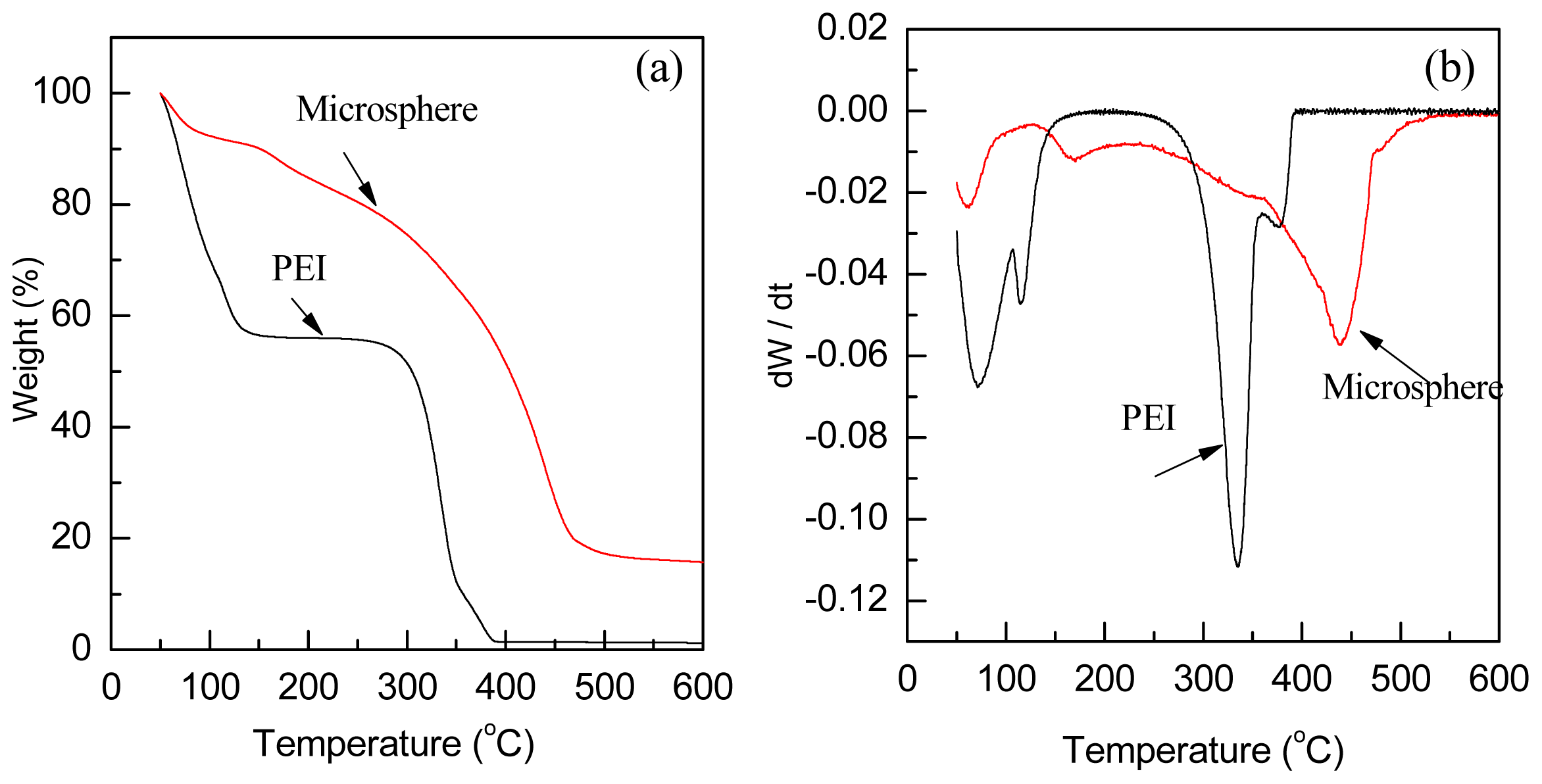
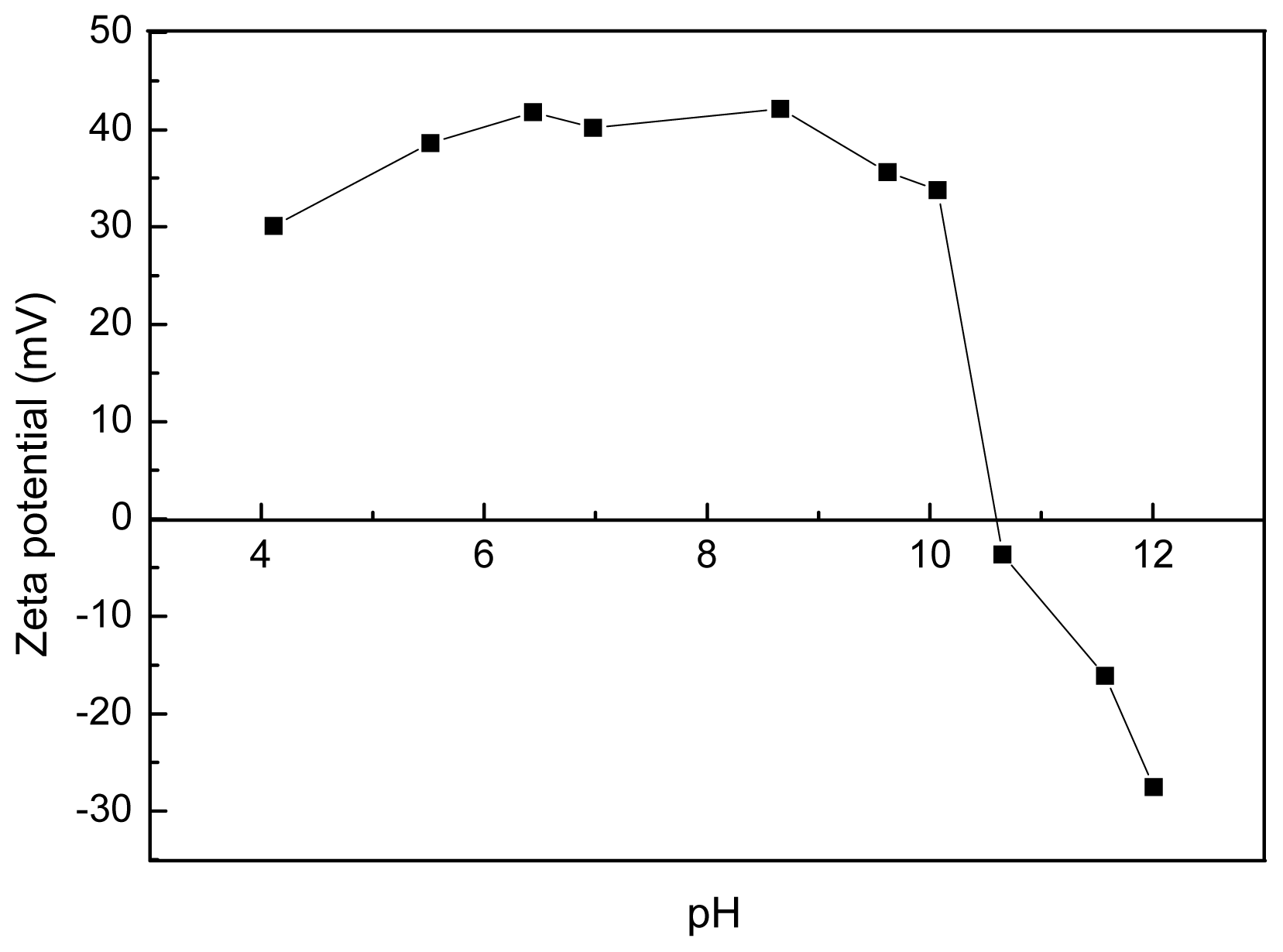
2.2. Thermalstability and Zeta Potential of the Polyamine Microsphere
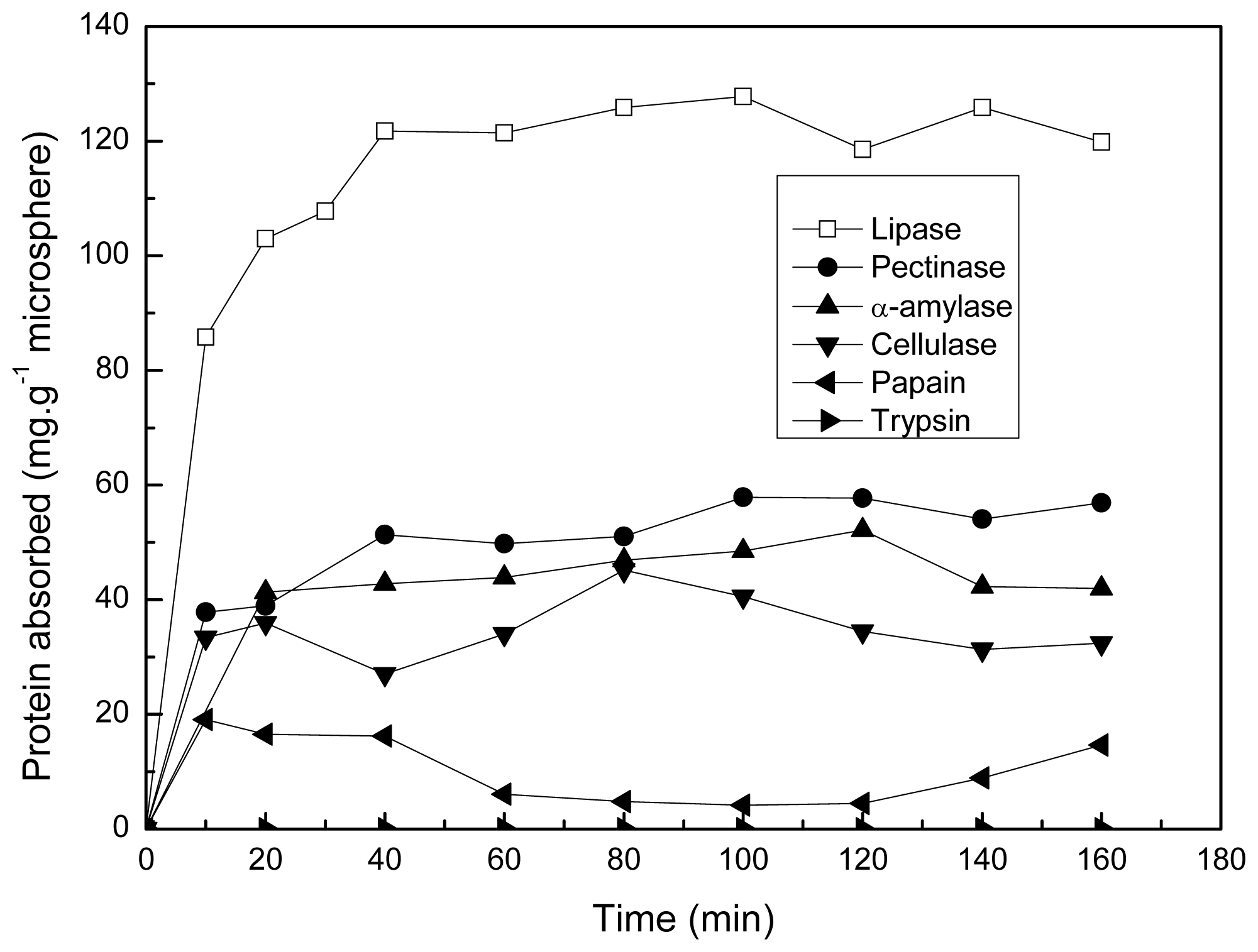
2.3. Protein Adsorption of the Microsphere
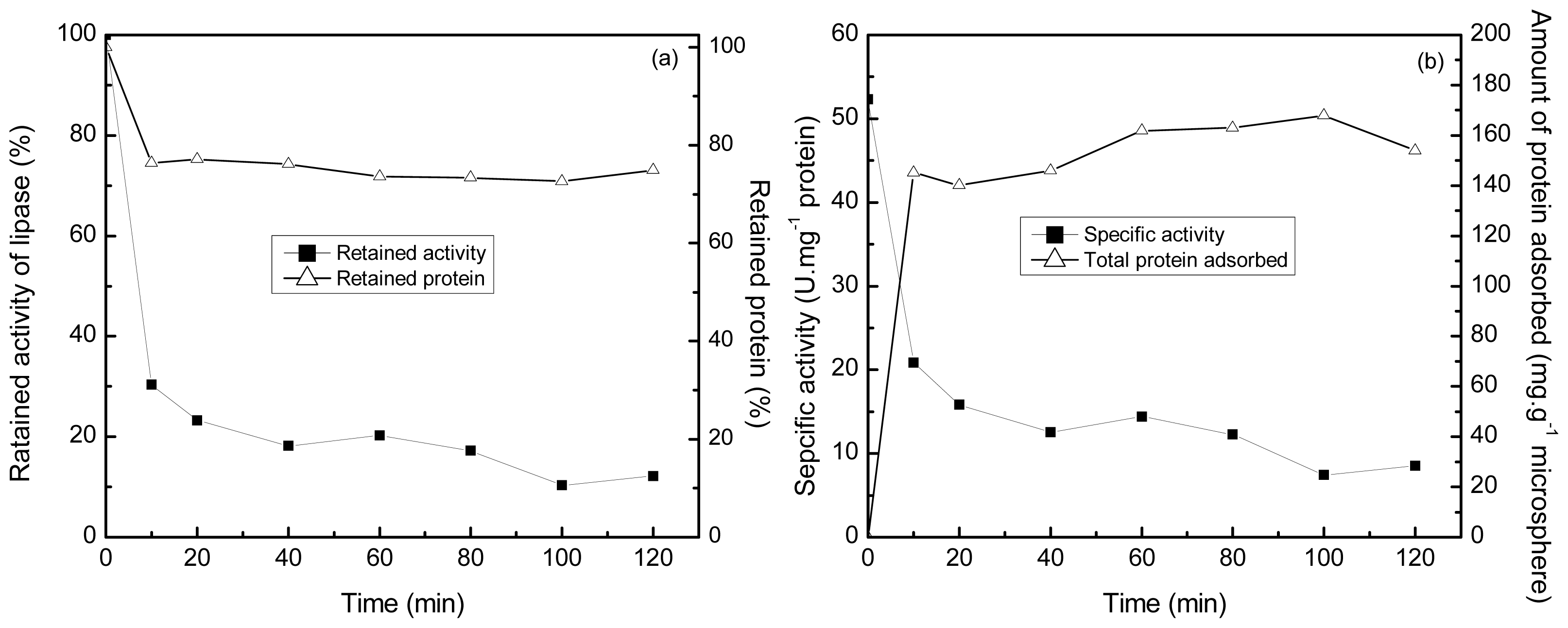
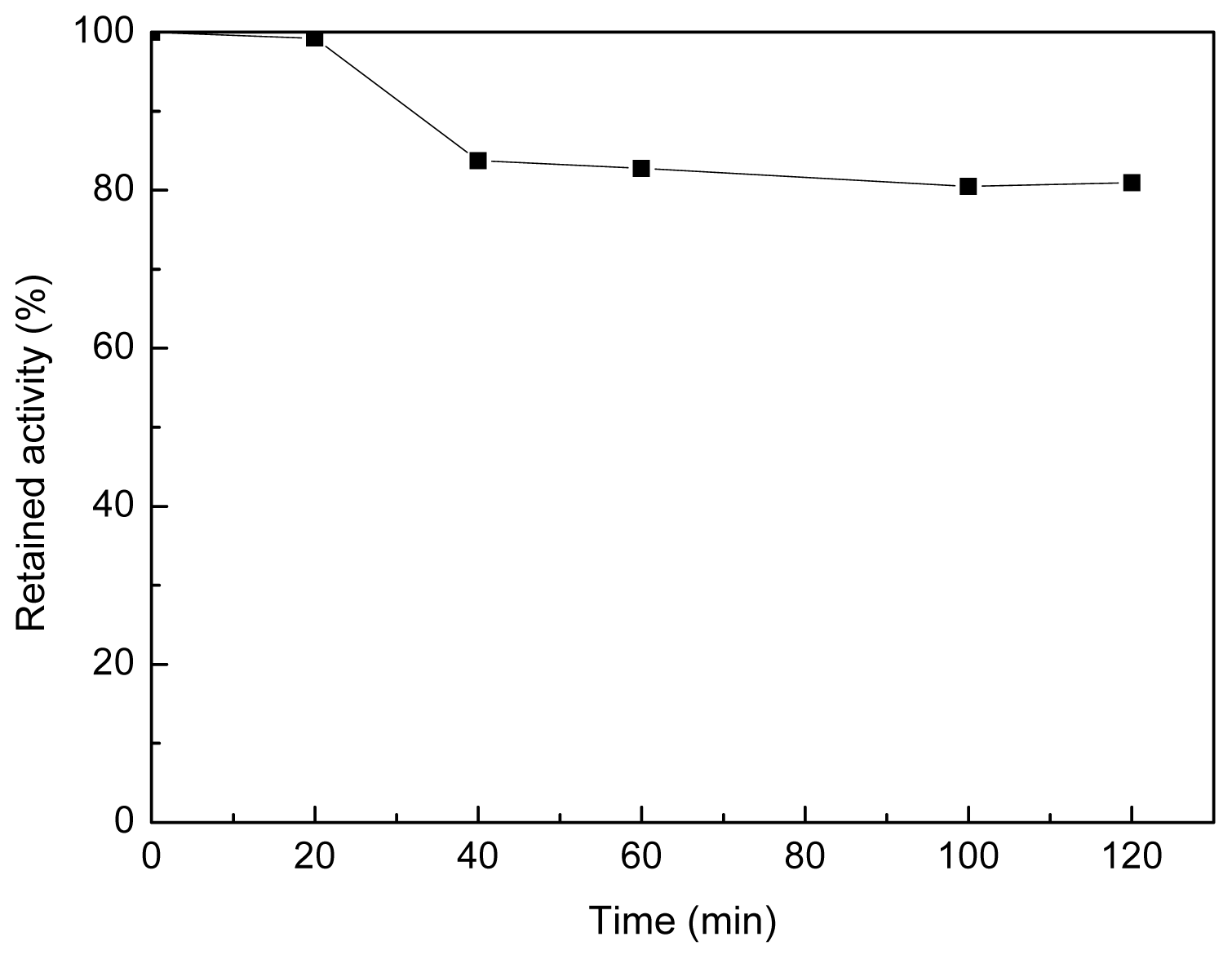
2.4. Selective Adsorption of Lipase onto the Polyamine Microsphere
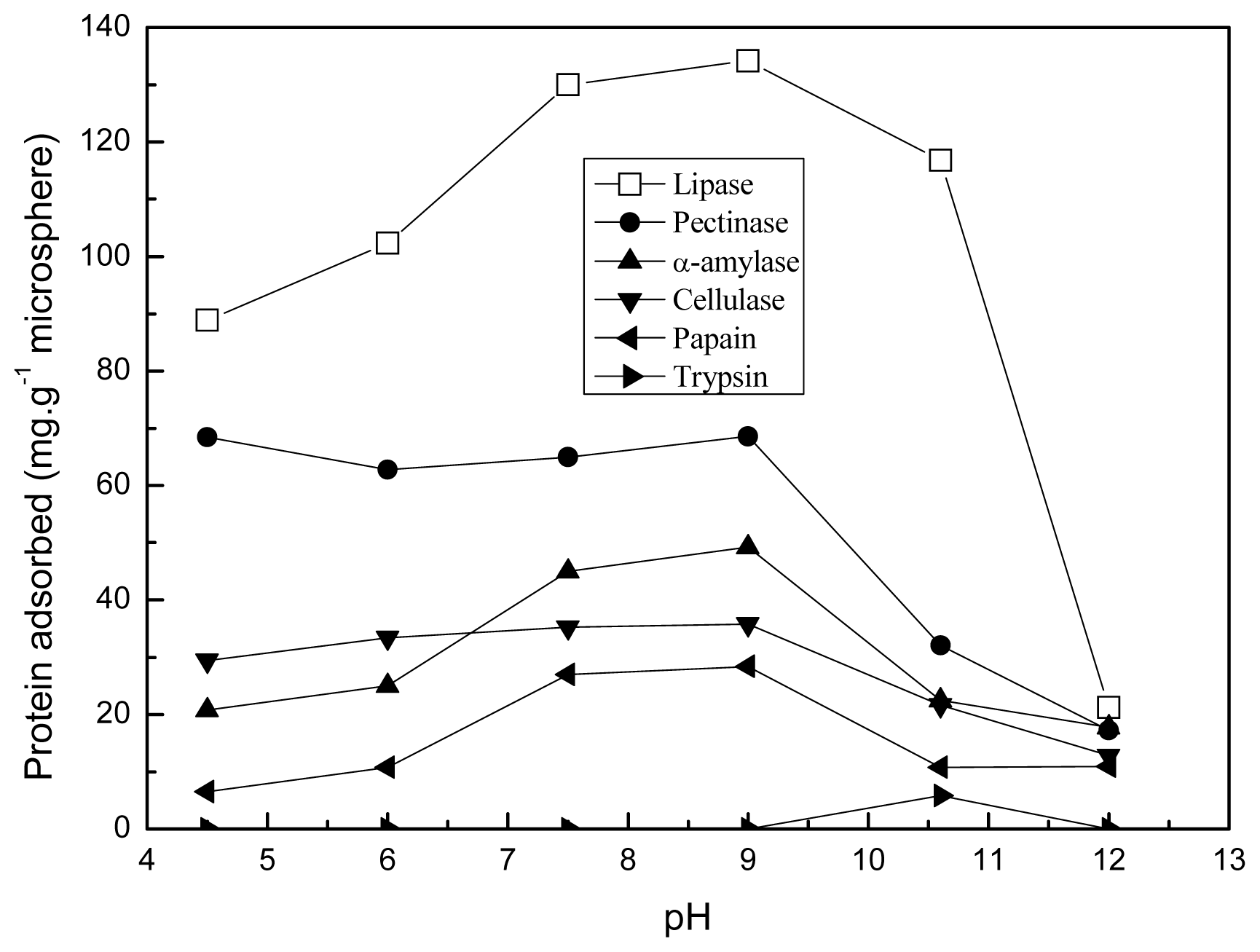
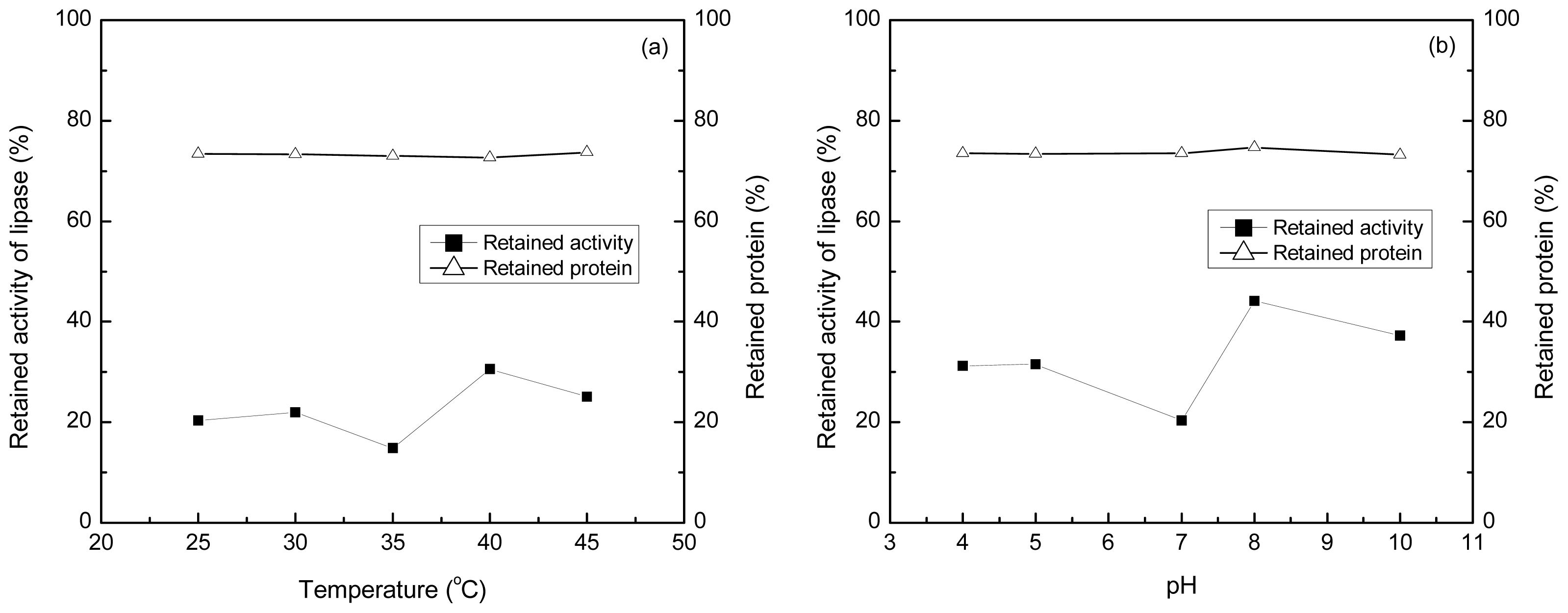
2.5. Effect of pH and Temperature on the Adsorption
3. Experimental Section
3.1. Materials
3.2. Preparation of Polyamine Microsphere in Inverse Emulsion
3.3. The Adsorption of Protein
3.4. Determination of Protein Content and the Activity of Lipase
3.5. Characterization of the Polyamine Microsphere
4. Conclusions
Acknowledgments
References
- Shin, J.; Anisur, R.M. Hollow manganese oxide nanoparticles as multifunctional agents for magnetic resonance imaging and drug delivery. Angew. Chem. Int. Ed 2009, 48, 321–324. [Google Scholar]
- Yang, X.; Chen, L.; Huang, B.; Bai, F.; Yang, X. Synthesis of pH-sensitive hollow polymer microspheres and their application as drug carriers. Polymer 2009, 50, 3556–3563. [Google Scholar]
- Yang, S.; Wang, C. Facile dicyandiamide-mediated fabrication of well-defined CuO hollow microspheres and their catalytic application. Mater. Chem. Phys 2010, 120, 296–301. [Google Scholar]
- McQuade, D.T.; Pullen, A.E. Conjugated polymer-based chemical sensors. Chem. Rev 2000, 100, 2537–2574. [Google Scholar]
- Hu, J.; Li, S.J.; Liu, B.L. Adsorption of BSA onto sulfonated microspheres. Biochem. Eng. J 2005, 23, 259–263. [Google Scholar]
- Shao, D.; Xu, K.; Song, X.; Hu, J.; Yang, W.; Wang, C. Effective adsorption and separation of lysozyme with PAA-modified Fe3O4@silica core/shell microspheres. J. Colloid Interface Sci 2009, 336, 526–532. [Google Scholar]
- Sari, M.M. Investigation of yeast invertase immobilization onto cupric ion-chelated, porous, and biocompatible poly(hydroxyethyl methacrylate-n-vinyl imidazole) microspheres. Appl. Biochem. Biotechnol 2011, 163, 1020–1037. [Google Scholar]
- Srisuwan, Y.; Srihanam, P.; Baimark, Y. Preparation of silk fibroin microspheres and its application to protein adsorption. J. Macromol. Sci. Part A 2009, 46, 521–525. [Google Scholar]
- Kawai, T.; Saito, K.; Lee, W. Protein binding to polymer brush, based on ion-exchange, hydrophobic, and affinity interactions. J. Chromatogr. B 2003, 790, 131–142. [Google Scholar]
- Patil, S.; Sandberg, A.; Heckert, E.; Self, W.; Seal, S. Protein adsorption and cellular uptake of cerium oxide nanoparticles as a function of zeta potential. Biomaterials 2007, 28, 4600–4607. [Google Scholar]
- Mateo, C.; Abian, O.; Fernandez-Lafuente, R.; Guisan, J.M. Reversible enzyme immobilization via a very strong and nondistorting ionic adsorption on support-polyethylenimine composites. Biotechnol. Bioeng 2000, 68, 98–105. [Google Scholar]
- Wang, F.; Gu, Z.G.; Cui, Z.G.; Liu, L.M. Comparison of covalent immobilization of amylase on polystyrene pellets with pentaethylenehexamine and pentaethylene glycol spacers. Bioresour. Technol 2011, 102, 9374–9379. [Google Scholar]
- Bolivar, J.M.; Rocha-Martin, J.; Mateo, C.; Cava, F.; Berenguer, J.; Fernandez-Lafuente, R.; Guisan, J.M. Coating of soluble and immobilized enzymes with ionic polymers: Full stabilization of the quaternary structure of multimeric enzymes. Biomacromolecules 2009, 10, 742–747. [Google Scholar]
- Murakami, Y.; Sugo, K.; Hirano, M.; Okuyama, T. Surface chemical analysis and chromatographic characterization of polyethylenimine-coated hydroxyapatite with various amount of polyethylenimine. Talanta 2011, 85, 1298–1303. [Google Scholar]
- Niepel, M.S.; Peschel, D.; Sisquella, X.; Planell, J.A.; Groth, T. pH-dependent modulation of fibroblast adhesion on multilayers composed of poly(ethylene imine) and heparin. Biomaterials 2009, 30, 4939–4947. [Google Scholar]
- Murugesan, R.; Orsat, V. Spray drying for the production of nutraceutical ingredients—A review. Food Bioprocess Technol 2012, 5, 3–14. [Google Scholar]
- Bose, S.; Schenck, D.; Ghosh, I.; Hollywood, A.; Maulit, E.; Ruegger, C. Application of spray granulation for conversion of a nanosuspension into a dry powder form. Eur. J. Pharm. Sci 2012, 47, 35–43. [Google Scholar]
- Grunlan, J.C.; Choi, J.K.; Lin, A. Antimicrobial behavior of polyelectrolyte multilayer films containing cetrimide and silver. Biomacromolecules 2005, 6, 1149–1153. [Google Scholar]
- Ji, J.; Tan, Q.; Fan, D.Z.; Sun, F.Y.; Barbosa, M.A.; Shen, J. Fabrication of alternating polycation and albumin multilayer coating onto stainless steel by electrostatic layer-by-layer adsorption. Colloids Surf. B 2004, 34, 185–190. [Google Scholar]
- Kassab, H.; Maksoud, M.; Aguado, S.; Pera-Titus, M.; Albela, B.; Bonneviot, L. Polyethylenimine covalently grafted on mesostructured porous silica for CO2 capture. RSC Adv 2012, 2, 2508–2516. [Google Scholar]
- Gao, B.; Li, Y.; Chen, Z. Adsorption behaviour of functional grafting particles based on polyethyleneimine for chromate anions. Chem. Eng. J 2009, 150, 337–343. [Google Scholar]
- Hu, X.; Ji, J. Covalent Layer-by-Layer assembly of hyperbranched polyether and polyethyleneimine: Multilayer films providing possibilities for surface functionalization and local drug delivery. Biomacromolecules 2011, 12, 4264–4271. [Google Scholar]
- Xia, B.; Dong, C.; Lu, Y.; Rong, M.; Lv, Y.-Z.; Shi, J. Preparation and characterization of chemically-crosslinked polyethyleneimine films on hydroxylated surfaces for stable bactericidal coatings. Thin Solid Films 2011, 520, 1120–1124. [Google Scholar]
- Sunintaboon, P.; Ho, K.M. Formation of nanostructured materials via coalescence of amphiphilic hollow particles. J. Am. Chem. Soc 2006, 128, 2168–2169. [Google Scholar]
- Tong, W.; Gao, C.; Mohwald, H. Poly(ethyleneimine) microcapsules: Glutaraldehyde-mediated assembly and the influence of molecular weight on their properties. Polym. Adv. Technol 2008, 19, 817–823. [Google Scholar]
- Schiller, R.; Weiss, C.K.; Geserick, J.; Husing, N.; Landfester, K. Synthesis of mesoporous silica particles and capsules by miniemulsion technique. Chem. Mater 2009, 21, 5088–5098. [Google Scholar]
- Rossier-Miranda, F.J.; Schroën, C.G.P.H.; Boom, R.M. Colloidosomes: Versatile microcapsules in perspective. Colloids Surf. A 2009, 343, 43–49. [Google Scholar]
- Son, W.J.; Choi, J.S. Adsorptive removal of carbon dioxide using polyethyleneimine-loaded mesoporous silica materials. Microporous Mesoporous Mater 2008, 113, 31–40. [Google Scholar]
- Fernandez-Lafuente, R.; Armisen, P.; Sabuquillo, P.; Fernandez-Lorente, G.; Guisan, J.M. Immobilization of lipases by selective adsorption on hydrophobic supports. Chem. Phys. Lipids 1998, 93, 185–197. [Google Scholar]
- Volpato, G.; Filice, M.; Rivas, B.D.L.; Rodrigues, R.C.; Heck, J.X.; Fernandez-Lafuente, R.; Guisan, J.M.; Mateo, C.; Ayub, M.A.Z. Purification, immobilization, and characterization of a specific lipase from Staphylococcus warneri EX17 by enzyme fractionating via adsorption on different hydrophobic supports. Biotechnol. Prog 2011, 27, 717–723. [Google Scholar]
- Zheng, M.M.; Lu, Y.; Dong, L.; Guo, P.M.; Deng, Q.C.; Li, W.-L.; Feng, Y.Q.; Huang, F.H. Immobilization of Candida rugosa lipase on hydrophobic/strong cation-exchange functional silica particles for biocatalytic synthesis of phytosterol esters. Bioresour. Technol 2012, 115, 141–146. [Google Scholar]
- Lowry, O.; Rosebrough, N.; Farr, A.; Rabdall, R. Protein measurement with folin-phenol reagent. J. Biol. Chem 1951, 193, 265–275. [Google Scholar]
- Abramic, M.; Lescic, I.; Korica, T.; Vitale, L.; Saenger, W.; Pgac, J. Purification and properties of extracellular lipase from Streptomyces rimosus. Enzym. Microb. Tech 1999, 25, 522–529. [Google Scholar]
- Cohen, J.L.; Schubert, S.; Wich, P.R.; Cui, L.; Cohen, J.A.; Mynar, J.L.; Frechet, J.M.J. Acid-degradable cationic dextran particles for the delivery of siRNA therapeutics. Bioconjug. Chem 2011, 22, 1056–1065. [Google Scholar]


© 2013 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, F.; Liu, P.; Nie, T.; Wei, H.; Cui, Z. Characterization of a Polyamine Microsphere and Its Adsorption for Protein. Int. J. Mol. Sci. 2013, 14, 17-29. https://doi.org/10.3390/ijms14010017
Wang F, Liu P, Nie T, Wei H, Cui Z. Characterization of a Polyamine Microsphere and Its Adsorption for Protein. International Journal of Molecular Sciences. 2013; 14(1):17-29. https://doi.org/10.3390/ijms14010017
Chicago/Turabian StyleWang, Feng, Pei Liu, Tingting Nie, Huixian Wei, and Zhenggang Cui. 2013. "Characterization of a Polyamine Microsphere and Its Adsorption for Protein" International Journal of Molecular Sciences 14, no. 1: 17-29. https://doi.org/10.3390/ijms14010017
APA StyleWang, F., Liu, P., Nie, T., Wei, H., & Cui, Z. (2013). Characterization of a Polyamine Microsphere and Its Adsorption for Protein. International Journal of Molecular Sciences, 14(1), 17-29. https://doi.org/10.3390/ijms14010017



