Species Differentiation of Chinese Mollitrichosiphum (Aphididae: Greenideinae) Driven by Geographical Isolation and Host Plant Acquirement
Abstract
:1. Introduction
2. Results and Discussion
2.1. Results
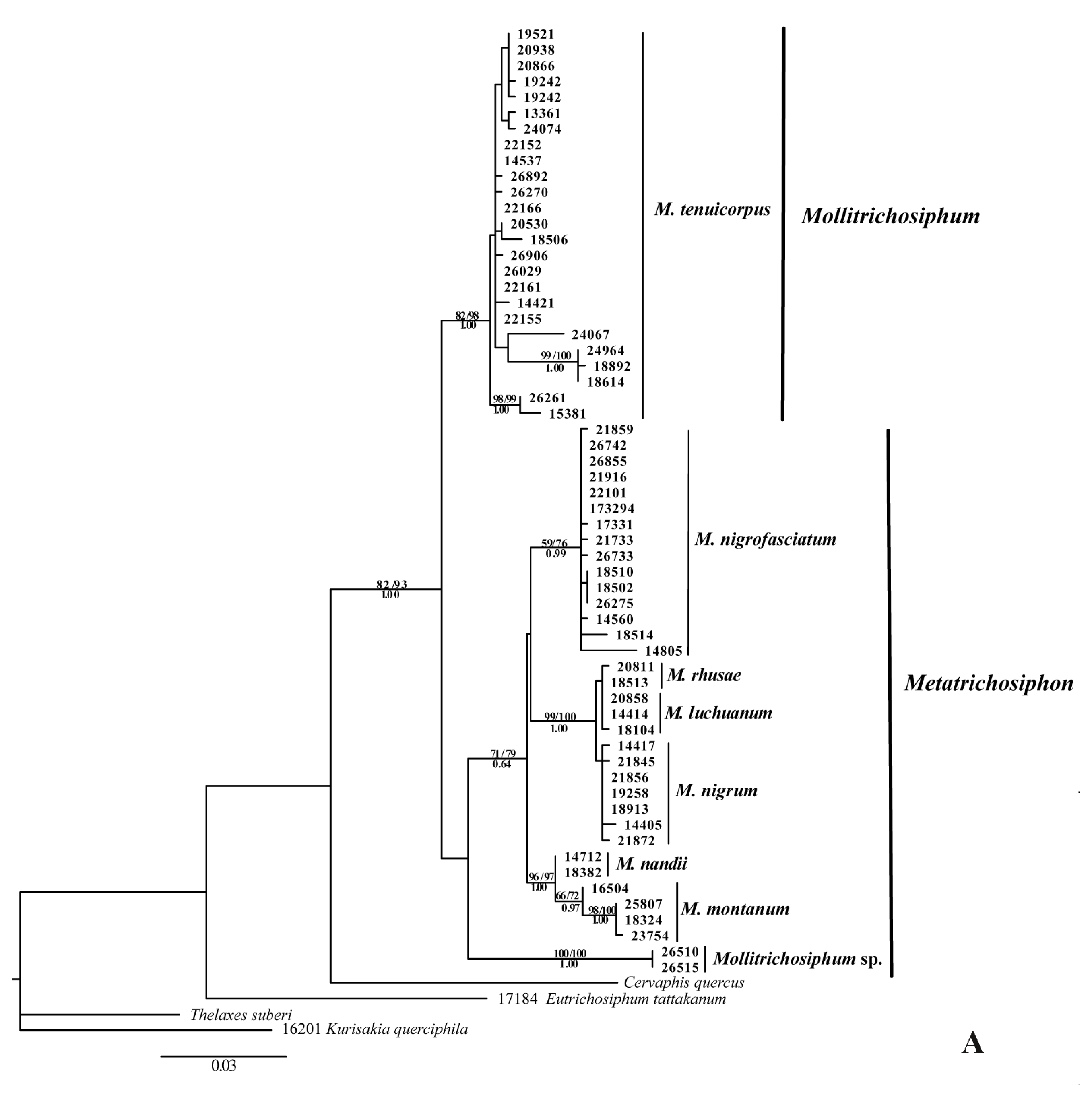
2.1.1. Phylogenetic Analyses
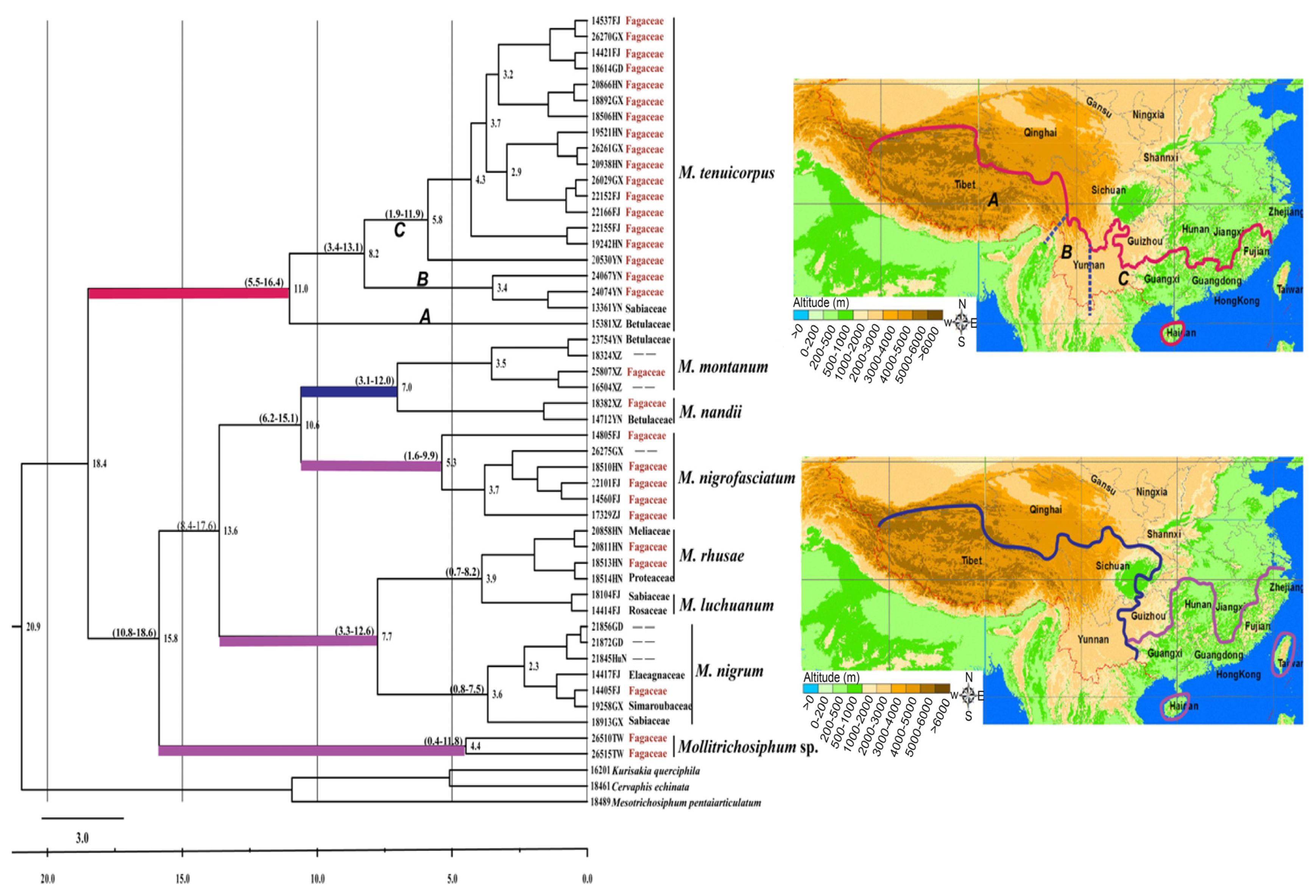
2.1.2. Divergence Time and Geographical Distribution
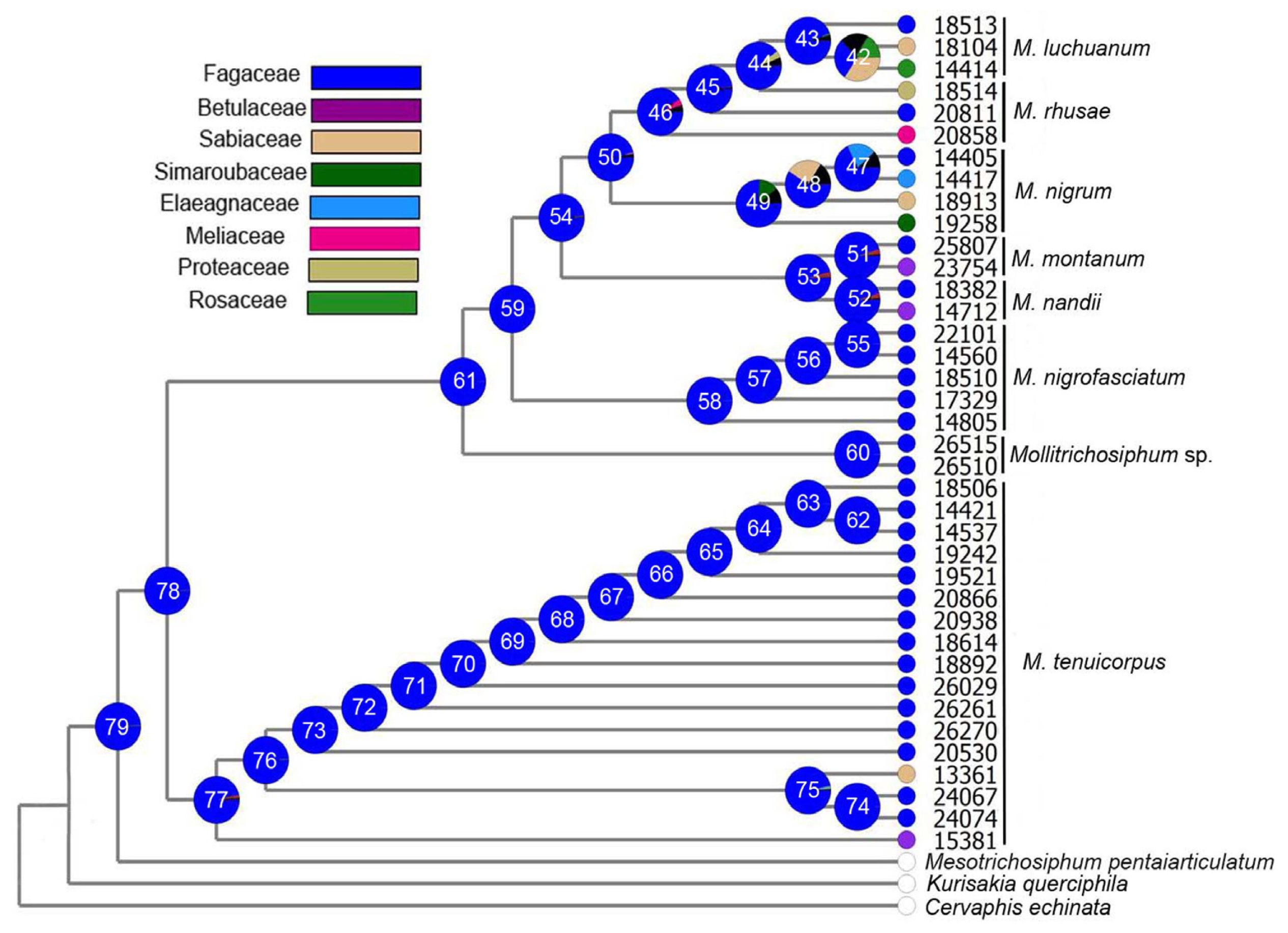
2.1.3. Ancestral State Reconstruction
2.2. Discussion
2.2.1. Phylogenetic Relationships
2.2.2. Divergence Times and Biogeography
2.2.3. Species Differentiation of Chinese Mollitrichosiphum
3. Experimental Section
3.1. Taxa Sampling
3.2. DNA Extraction, PCR and Sequencing
3.3. Sequence Alignment and Phylogenetic Analyses
3.4. Divergence Time Estimation and Ancestral State Reconstruction
4. Conclusions
Supplementary Materials
ijms-13-10441-s001.pdfAcknowledgments
References
- Myers, N.; Mittermeier, R.A.; Mittermeier, C.G.; Fonseca, G.A.B.; Kent, J. Biodiversity hotspots for conservation priorities. Nature 2000, 403, 854–858. [Google Scholar]
- Shi, Y.F.; Li, J.J.; Li, B.Y. Uplift and Environmental Change of Qinghai-Xizang (Tibetan) Plateau in the Late Cenozoic; Guangdong Science and Technology Press: Guangzhou, China, 1998. [Google Scholar]
- Shi, Y.F.; Li, J.J.; Li, B.Y.; Yao, T.D.; Wang, S.M.; Li, S.J. Uplift of the Qinghai-Xizang (Tibetan) Plateau and east Asia environmental change during late cenozoic. J. Geogr. Sci 1999, 54, 10–21. [Google Scholar]
- Li, B.Y.; Pan, B.T. Progress in paleogeographic study of the Tibetan Plateau. Geogr. Res 2002, 21, 62–70. [Google Scholar]
- Li, J.J.; Wen, S.X.; Zhang, Q.S.; Wang, F.B.; Zheng, B.X.; Li, B.Y. A discussion on the period, amplitude and type of the uplift of the Qinghai-Xizang Plateau. Sci. Sin 1979, 22, 1314–1328. [Google Scholar]
- Harrison, T.M.; Copeland, P.; Kidd, W.S.F.; Yin, A. Raising Tibet. Science 1992, 255, 1663–1670. [Google Scholar]
- Zhang, R.Z. Zoogeography of China; Science Press: Beijing, China, 1999. [Google Scholar]
- Cao, W.X.; Chen, Y.Y.; Wu, Y.F.; Zhu, S.Q. Origin and Evolution of Schizothoracine Fishes in Relation to the Upheaval of the Qinghai-Xizang Plateau. In Studies on the Period, Amplitude and Type of the Uplift of the Qinghai-Xizang Plateau; Li, Q.F., Ed.; Science Press: Beijing, China, 1981; pp. 118–129. [Google Scholar]
- Huang, F.S. Promontory of Tibet Plateau and Insect Fauna. In Insects of Xizang; Chen, S.X., Ed.; Science Press: Beijing, China, 1981; Volume 1, pp. 1–31. [Google Scholar]
- López-Pujol, J.; Zhang, F.M.; Ge, S. Plant biodiversity in China: Richly varied, endangered, and in need of conservation. Biodivers. Conserv 2006, 15, 3983–4026. [Google Scholar]
- Chen, Y.H.; Bi, J.F. Biogeography and hotspots of amphibian species of China: Implications to reserve selection and conservation. Curr. Sci 2007, 92, 480–489. [Google Scholar]
- Huang, X.L.; Qiao, G.X.; Lei, F.M. Diversity and distribution of aphids in the Qinghai-Tibetan Plateau-Himalayas. Ecol. Entomol 2006, 31, 608–615. [Google Scholar]
- Huang, X.L.; Lei, F.M.; Qiao, G.X. Areas of endemism and patterns of diversity for aphids of the Qinghai-Tibetan Plateau and the Himalayas. J. Biogeogr 2008, 35, 230–240. [Google Scholar]
- Lei, F.M.; Wei, G.A.; Zhao, H.F.; Yin, Z.H.; Lu, J.L. China subregional avian endemism and biodiversity conservation. Biodivers. Conserv 2007, 16, 1119–1130. [Google Scholar]
- Liu, Z.; Huang, X.L.; Jiang, L.Y.; Qiao, G.X. The species diversity and geographical distribution of aphids in China (Hemiptera, Aphidoidea). Acta Zootax. Sin 2009, 34, 277–291. [Google Scholar]
- Lu, L.; Fritsch, P.W.; Cruz, B.C.; Wang, H.; Li, D.Z. Reticulate evolution, cryptic species, and character convergence in the core East Asian clade of Gaultheria (Ericaceae). Mol. Phylogenet. Evol 2010, 57, 364–379. [Google Scholar]
- Wang, Y.H.; Chen, J.M.; Xu, C.; Liu, X.; Wang, Q.F.; Motley, T.J. Population genetic structure of an aquatic herb Batrachium bungei (Ranuculaceae) in the Hengduan Mountains of China. Aquat. Bot 2010, 92, 221–225. [Google Scholar]
- Li, Y.; Zhai, S.N.; Qiu, Y.X.; Guo, Y.P.; Ge, X.J.; Comes, H.P. Glacial survival east and west of the ‘Mekong-Salween-Divide’ in the Himalaya-Hengduan Mountains region as revealed by AFLP and cpDNA sequence variation in Sinopodophyllum hexandrum (Berberidaceae). Mol. Phylogenet. Evol 2011, 59, 412–424. [Google Scholar]
- Liu, Z.J.; Ren, B.P.; Wei, F.W.; Long, Y.C.; Hao, Y.L.; Li, M. Phylogeography and population structure of the Yunnan snub-nosed monkey (Rhinopithecus bieti) inferred from mitochondrial control region DNA sequence analyses. Mol. Ecol 2007, 16, 3334–3349. [Google Scholar]
- Song, G.; Qu, Y.H.; Yin, Z.H.; Li, S.H.; Liu, N.F.; Lei, F.M. Phylogeography of the Alcippe morrisonia (Aves: Timaliidae): Long population history beyond late Pleistocene glaciations. BMC Evol. Biol 2009, 9, 143. [Google Scholar]
- Huang, Z.H.; Liu, N.F.; Liang, W.; Zhang, Y.Y.; Liao, X.J.; Ruan, L.Z.; Yang, Z.S. Phylogeography of Chinese bamboo partridge, Bambusicola thoracica thoracica (Aves: Galliformes) in south China: Inference from mitochondrial DNA control-region sequences. Mol. Phylogenet. Evol 2010, 56, 273–280. [Google Scholar]
- Gao, B.; Yu, L.J.; Qu, Y.H.; Song, G.; Dai, C.Y.; Zhang, R.Y.; Yin, Z.H.; Wang, K.F.; Gao, X.B.; Li, S.H.; et al. An unstructured phylogeographic pattern with extensive gene flow in an endemic bird of South China: Collared finchbill (Spizixos semitorques). Int. J. Mol. Sci 2011, 12, 3635–3647. [Google Scholar]
- Che, J.; Zhou, W.W.; Hu, J.S.; Yan, F.; Papenfuss, T.J.; Wake, D.B.; Zhang, Y.P. Spiny frogs (Paini) illuminate the history of the Himalayan region and Southeast Asia. Proc. Natl. Acad. Sci. USA 2010, 31, 13765–13770. [Google Scholar]
- Remaudière, G.; Remaudiėr, M. Catalogue of the World’s Aphididae. In Homoptera Aphidoidea; Institut National de la Recherche Agronomique: Paris, France, 1997; pp. 1–475. [Google Scholar]
- Zhang, D.; Qiao, G.X. Mollitrichosiphum Suenaga from China (Hemiptera, Aphididae), with the description of one new species. Zootaxa 2010, 2608, 1–24. [Google Scholar]
- Zhang, G.X.; Zhong, T.S. Homoptera: Aphidinea. Economic Insect Fauna of China 1983, Part I, 123–135. [Google Scholar]
- Noordam, D. Greenideinae from Java (Homoptera, Aphididae). Zool. Verh 1994, 296, 1–284. [Google Scholar]
- Tao, C.C. Aphid Fauna of Taiwan Province, China; Taiwan Provincial Museum: Taipei, Taiwan, 1990. [Google Scholar]
- Tao, C.C. List of Aphidoidea (Homoptera) of China; Taiwan Agricultural Research Institute: Taipei, Taiwan, 1999. [Google Scholar]
- Zhang, R.L.; Huang, X.L.; Jiang, L.Y.; Qiao, G.X. Phylogeny and species differentiation of Mollitrichosiphum (Aphididae, Greenideinae) based on mitochondrial COI and Cytb genes. Curr. Zool 2011, 57, 806–815. [Google Scholar]
- Ghosh, A.K.; Agarwala, B.K. Subfamily: Greenideinae. The Fauna of India and the Adjacent Countries: Homoptera, Aphidoidea 1993, Part 6. [Google Scholar]
- Zhang, H.C.; Qiao, G.X. Application of gene sequences in molecular phylogenetic study on Aphididae (Hemiptera). Acta Entomol. Sin 2006, 49, 521–527. [Google Scholar]
- Spicer, R.A.; Harris, N.B.W.; Widdowson, M.; Herman, A.B.; Guo, S.; Valdes, P.J.; Wolfe, J.A.; Kelley, S.P. Constant elevation of southern Tibet over the past 15 million years. Nature 2003, 421, 622–624. [Google Scholar]
- Royden, L.H.; Burchfiel, B.C.; van der Hilst, R.D. The geological evolution of the Tibetan Plateau. Science 2008, 321, 1054–1058. [Google Scholar]
- Zhou, S.Z.; Wang, X.L.; Wang, J.; Xu, L.B. A preliminary study on timing of the oldest Pleistocene glaciation in Qinghai-Tibetan Plateau. Quat. Int 2006, 155, 44–51. [Google Scholar]
- Peng, Z.; Ho, S.Y.W.; Zhang, Y.; He, S. Uplift of the Tibetan plateau: Evidence from divergence times of glyptosternoid catfishes. Mol. Phylogenet. Evol 2006, 39, 568–572. [Google Scholar]
- Ward, F.K. The Mekong-Salween divide as a geographical barrier. Geogr. J 1921, 58, 49–56. [Google Scholar]
- Clark, M.K.; Schoenbohm, L.M.; Royden, L.H.; Whipple, K.X.; Burchfiel, B.C.; Zhang, X.; Tang, W.; Wang, E.; Chen, L. Surface uplift, tectonics, and erosion of eastern Tibet from large-scale drainage patterns. Tectonics 2004, 23, 1006–1029. [Google Scholar]
- Hall, R. The Plate Tectonics of Cenozoic SE Asia and the Distribution of Land and Sea. In Biogeography and Geological Evolution of SE Asia; Backhuys publishers: Leiden, The Netherlands, 1998. [Google Scholar]
- Harrison, S.P.; Yu, G.; Takahara, H.; Prentice, I.C. Palaeovegetation (communications arising): Diversity of temperate plants in East Asian. Nature 2001, 413, 129–130. [Google Scholar]
- Wegierek, P.; Peñalver, E. Fossil representatives of the family Greenideidae (Hemiptera, Aphidoidea) from the Miocene of Europe. Geobios-Lyon 2002, 35, 745–757. [Google Scholar]
- Penalver, E.; de Santisteban, C.; Barrón, E. Fossil Insects and Palaeobotany of the Rubielos de Mora Basin (Teruel). In The Geological and Paleontological Heritage of Central and Eastern Iberia (Iberian Range, Spain); Meléndez, G., Soria-Llop, C., Eds.; Publicaciones del Seminario de Paleontología: Zaragoza, Spain, 1999; pp. 95–116. [Google Scholar]
- Van Andel, T.H. New Views on an Old Planet: A History of Global Change; Cambridge University Press: Cambridge, UK, 1994; pp. 166–230. [Google Scholar]
- Ghosh, A.K. Biotaxonomy of Greenideinae (Homoptera: Aphidoidea). Proceedings of the International Smyposia on Population Structure, Genetics and Taxonomy of Aphids and Thysanoptera, Smolenice, Czechoslovakia, 9–14 September 1985.Holman, J.; Pelikán, J.; Dixon, A.F.G.; Weismann, L. (Eds.) SPB Academic Publisher: Hague, The Netherlands, 1987; pp. 273–292.
- Blackman, R.L.; Eastop, V.F. Aphids on the World’s Trees. In An Identification and Information Guide; CAB International: Wallingford, CT, USA; p. 1994.
- Dobzhansky, T. Genetics and the Origin of Species; Columbia University Press: New York, NY, USA; p. 1937.
- Mayr, E. Animal Species and Evolution; Belknap Press: Cambridge, UK; p. 1963.
- Funk, D.J.; Nosil, P.; Etges, W.J. Ecological divergence exhibits consistently positive associations with reproductive isolation across disparate taxa. Proc. Natl. Acad. Sci. USA 2006, 103, 3209–3213. [Google Scholar]
- Peccoud, J.; Ollivier, A.; Plantegenest, M.; Simon, J.C. A continuum of genetic divergence from sympatric host races to species in the pea aphid complex. Proc. Natl. Acad. Sci. USA 2009, 106, 7495–7500. [Google Scholar]
- Peccoud, J.; Simon, J.C.; McLaughlin, H.J.; Moran, N.A. Post-Pleistocene radiation of the pea aphid complex revealed by rapidly evolving endosymbionts. Proc. Natl. Acad. Sci. USA 2009, 106, 16315–16320. [Google Scholar]
- Heie, O.E. Paleontology and Phylogeny. In Aphids, Their Biology, Natural Enemies, and Control; Minks, A.K., Harrewijn, P., Eds.; Elsevier: Amsterdam, The Netherlands, 1987; Volume 2A, pp. 367–391. [Google Scholar]
- Ortiz-Rivas, B.; Moya, A.; Martinez-Torres, D. Combination of molecular data support the existence of three main lineages in the phylogeny of aphids (Hemiptera: Aphididae) and the basal position of the subfamily Lachninae. Mol. Phylogenet. Evol 2010, 55, 305–317. [Google Scholar]
- Vavre, F.; Girin, C.; Boulétreau, M. Phylogenetic status of a fecundity enhancing Wolbachia that does not induce thelytoky in Trichogramma. Insect Mol. Biol 1999, 8, 67–72. [Google Scholar]
- Foottit, R.G.; Maw, B.L.; von Dohlen, C.D.; Hebert, P.D.N. Species identification of aphids (Insecta: Hemiptera: Aphididae) through DNA barcodes. Mol. Ecol. Resour 2008, 8, 1189–1201. [Google Scholar]
- Harry, M.; Solignac, M.; Lachaise, D. Molecular evidence for parallel evolution of adaptive syndromes in fig-breeding Lissocephala (Drosophilidae). Mol. Phylogenet. Evol 1998, 9, 542–551. [Google Scholar]
- Palumbi, S.R. Nucleic Acids II: The Polymerase Chain Reaction. In Molecular Systematics; Hillis, D.M., Moritz, C., Mable, B.K., Eds.; Sinauer: Sunderland, MA, USA, 1996; pp. 205–247. [Google Scholar]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acid. Res 1997, 25, 4876–4882. [Google Scholar]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA 4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol 2007, 24, 1596–1599. [Google Scholar]
- Swofford, D.L. PAUP*, version 4.0; Phylogenetic Analysis Using Parsimony (and Other Methods); Sinauer Associates, Inc: Sunderland, MA, USA; p. 2002.
- Ronquist, F.; Huelsenbeck, J.P. Bayesian Analysis of Molecular Evolution Using MrBayes. In Statistical Methods in Molecular Evolution; Nielsen, R., Ed.; Springer: New York, NY, USA, 2005; pp. 183–232. [Google Scholar]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar]
- Posada, D.; Crandall, K.A. MODELTEST: Testing the model of DNA substitution. Bioinformatics 1998, 14, 817–818. [Google Scholar]
- Posada, D.; Buckley, T.R. Model selection and model averaging in phylogenetics: Advantages of akaike information criterion and Bayesian approaches over likelihood ratio tests. Syst. Biol 2004, 53, 793–808. [Google Scholar]
- Guindon, S.; Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol 2003, 52, 696–704. [Google Scholar]
- Drummond, A.J.; Suchard, M.A. Bayesian random local clocks or one rate to rule them all. BMC Boil 2010, 8. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A.J. Tracer, version 1.5; BioMed Central: London, UK, 2007. Available online: http://tree.bio.ed.ac.uk/software/tracer accessed on 30 November 2009.
- Rambaut, A. FigTree, version 1.3.1; BioMed Central: London, UK, 2009. Available online: http://tree.bio.ed.ac.uk/software/figtree accessed on 21 December 2009.
- Yu, Y.; Harris, A.J.; He, X.J. RASP, version 2.0b; Elsevier: Amsterdam, The Netherlands, 2010. Available online: http://mnh.scu.edu.cn/soft/blog/RASP accessed on 23 November 2011.

| Subgenera | Species | Host plants (Family) | |
|---|---|---|---|
| Mollitrichosiphum | M. godavariense | 1 | Fagaceae |
| M. nigriabdominalis | 1 | Fagaceae | |
| M. tenuicorpus | 3 | Fagaceae | |
| Betulaceae | |||
| Sabiaceae | |||
| M. trilokum | 1 | Fagaceae | |
| Metatrichosiphon | M. buddleiae | 2 | Buddlejaceae |
| Betulaceae | |||
| M. kazirangi | 1 | unknown | |
| M. montanum | 3 | Betulaceae | |
| Juglandaceae | |||
| Fagaceae | |||
| M. nandii | 4 | Betulaceae | |
| Lauraceae | |||
| Sapindaceae | |||
| Fagaceae | |||
| M. rhusae | 4 | Anacardiaceae | |
| Proteaceae | |||
| Meliaceae | |||
| Fagaceae | |||
| M. elongatum | 1 | Fagaceae | |
| M. syzygii | 1 | Myrtaceae | |
| M. luchuanum | 2 | Rosaceae | |
| Fagaceae | |||
| M. nigrofasciatum | 1 | Fagaceae | |
| M. nigrum | 3 | Sabiaceae | |
| Simaroubaceae | |||
| Elaeagnaceae | |||
| M. yamabiwae | 1 | Sabiaceae | |
| M. niitakaensis | 1 | Fagaceae | |
| M. glaucae | 1 | Fagaceae | |
| M. taiwanum | 1 | Sabiaceae | |
| Mollitrichosiphum sp. | 1 | Fagaceae | |
| Species | Voucher No. | Locality | Dates | Host plant | GenBank accession Nos: COI/Cytb/EF-1α |
|---|---|---|---|---|---|
| Ingroups | |||||
| M. tenuicorpus (Okajima) | 20938 | Hainan: Jianfengling Mt. | 25 March 2008 | Castanopsis fabri (Fagaceae) | JF969334/JF969383/JN645046 |
| 13361 | Yunnan: Baoshan City | 20 September 2002 | Meliosma rigida (Sabiaceae) | JF969308/JF969357/JN645020 | |
| 24074 | Yunnan: Ruili City | 2 December 2009 | Castanopsis calathiformis (Fagaceae) | JF969339/JF969388/JQ418332 | |
| 24067 | Yunnan: Ruili City | 3 December 2009 | Castanopsis calathiformis (Fagaceae) | JF969343/JF969392/JQ418331 | |
| 14421 | Fujian: Wuyishan Mt. | 6 July 2003 | Castanea sp. (Fagaceae) | JF969311/JF969360/JN645042 | |
| 14537 | Fujian: Wuyishan Mt. | 19 July 2003 | Castanopsis sclerophylla (Fagaceae) | JF969313/JF969362/JN645043 | |
| 18506 | Hainan: Diaoluoshan Mt. | 29 March 2006 | Cyclobalanopsis neglecta (Fagaceae) | JF969321/JF969370/JQ418327 | |
| 19242 | Hainan: Changjiang County | 26 March 2007 | Fagaceae | JF969327/JF969376/JN645045 | |
| 20866 | Hainan: Jianfengling Mt. | 15 December 2007 | Fagaceae | JF969333/JF969382/JN645030 | |
| 22152 | Fujian: Nanjing County | 23 November 2008 | unknown | JF969335/JF969384/JN645037 | |
| 22155 | Fujian: Huboliao District | 23 November 2008 | unknown | JF969336/JF969385/JN645038 | |
| 19521 | Hainan: Jianfengling Mt. | 16 November 2006 | Quercus sp. (Fagaceae) | JF969329/JF969378/JN645048 | |
| 22166 | Fujian: Huboliao District | 25 November 2008 | unknown | JF969337/JF969386/JN645039 | |
| 26029 | Guangxi: Huaping County | 1 November 2010 | Castanopsis eyrei (Fagaceae) | JN644999/JN645015/JN645054 | |
| 26261 | Guangxi: Shiwandashan Mt. | 19 November 2010 | Castanopsis fargesii Franch. (Fagaceae) | JN645000/JN645016/JN645050 | |
| 22161 | Fujian: Huboliao District | 24 November 2008 | unknown | JN644997/JN645013/JN645041 | |
| 24964 | Guangxi: Damin Mt. | 28 May 2011 | Fagaceae | JQ418333 | |
| 18614 | Guangdong: Chebaling District | 13 April 2006 | Castanopsis carlesii (Fagaceae) | JF969347/JF969396/JN645049 | |
| 20530 | Yunnan: Simao City | 27 July 2007 | Castanopsis ferox (Fagaceae) | JF969330/JF969379/JN645028 | |
| 15381 | Tibet: Motuo County | 14 August 2003 | Alnus cremastogyne (Betulaceae) | JF969317/JF969366/JN645044 | |
| 18892 | Guangxi: Guilin City | 18 May 2006 | Fagaceae | JF969348/JF969397/JN645025 | |
| 26892 | Fujian: Wuyishan Mt. | 15 June 2011 | Fagaceae | JQ418339 | |
| 26270 | Guangxi: Shangsi Country | 20 November 2010 | Fagaceae | JQ418313/JQ418317/JQ418334 | |
| 26906 | Fujian: Jiangle Country | 17 June 2011 | Fagaceae | JQ418340 | |
| M. nigrum (Zhang & Qiao) | 14417 | Fujian: Wuyishan Mt. | 6 July 2003 | Elaeagnus pungens (Elaeagnaceae) | JF969310/JF969359/JN645052 |
| 19258 | Guangxi: Xingan County | 2 July 2006 | Ailanthus altissima (Simaroubaceae) | JF969328/JF969377/JN645027 | |
| 21845 | Hunan: Mangshan Mt. | 17 July 2008 | unknown | JF969341/JF969390/JN645032 | |
| 21856 | Guangdong: Nanling Mt. | 18 July 2008 | unknown | JF969342/JF969391/JN645033 | |
| 18913 | Guangxi: Longsheng County | 21 May 2006 | Meliosma cuneifolia (Sabiaceae) | JF969326/JF969375/JN645026 | |
| 21872 | Guangdong: Nanling Mt. | 19 July 2008 | unknown | JN644995/JN645011/JN645035 | |
| 14405 | Fujian: Wuyishan Mt. | 4 July 2003 | Castanea sp.(Fagaceae) | JN644988/JN645004/JN645019 | |
| M. nandii (Busu) | 14712 | Yunnan: Baoshan City | 27 October 2003 | Alnus cremastogyne (Betulaceae) | JF969315/JF969364/JQ418318 |
| 18382 | Tibet: Tongmai District | 31 August 2005 | Fagus longipetiolata (Fagaceae) | JF969320/JF969369/JQ418323 | |
| M. nigrofasciatum (Maki) | 14560 | Fujian: Wuyishan Mt. | 20 July 2003 | Lithocarpus glaber (Fagaceae) | JF969314/JF969363/JN645022 |
| 14805 | Fujian: Wuyishan Mt. | 25 May 2004 | Cyclobalanopsis glauca (Fagaceae) | JF969346/JF969395/JQ418319 | |
| 22101 | Fujian: Liangyeshan Mt. | 14 November 2008 | Lithocarpus glaber (Fagaceae) | JF969351/JF969400/JN645036 | |
| 26742 | Zhejiang: Anji Country | 29 May 2011 | Fagaceae | JQ418338 | |
| 21916 | Guangdong: Nanling Mt. | 21 July 2008 | unknown | JN645047 | |
| 18510 | Hainan: Lingshui Mt. | 30 March 2006 | Lithocarpus elmerrillii (Fagaceae) | JN644994/JN645010/JQ418328 | |
| 26275 | Guangxi: Shiwandashan Mt. | 20 November 2010 | unknown | JN645001/JN645016/JN645051 | |
| 17329 | Zhejiang: Taishun County | 29 July 2005 | Quercus sp. (Fagaceae) | JN644990/JN645006/JQ418322 | |
| 26733 | Zhejiang: Anji Country | 29 May 2011 | Fagaceae | JQ418337 | |
| 18502 | Hainan: Lingshui County | 28 March 2006 | Castanopsis fabri Hance (Fagaceae) | JN644993/JN645008/JQ418326 | |
| 17331 | Zhejiang: Taishun County | 29 July 2005 | Fagaceae | JN645023 | |
| 21733 | Hunan: Leiling County | 7 July 2008 | unknown | JN645031 | |
| 21859 | Guangdong: Nanling Mt. | 19 July 2008 | unknown | JN645034 | |
| M. rhusae (Ghosh) | 18513 | Hainan: Diaoluoshan Mt. | 30 March 2006 | Fagaceae | JF969324/JF969373/JQ418329 |
| 20811 | Hainan: Wuzhishan Mt. | 7 December 2006 | Fagaceae | JF969331/JF969380/JQ418330 | |
| 20858 | Hainan: Diaoluoshan Mt. | 13 December 2007 | Meliaceae | JF969332/JF969381/JN645029 | |
| M. luchuanum (Takahashi) | 14414 | Fujian: Wuyishan Mt. | 6 July 2003 | Amygdalus persica (Rosaceae) | JF969309/JF969358/JN645021 |
| 18104 | Fujian: Wuyishan Mt. | 24 October 2005 | Meliosma rigida (Sabiaceae) | JF969319/JF969368/JN645024 | |
| M. montanum (van der Goot) | 16504 | Tibet: Zhangmu County | 27 July 2005 | unknown | JF969318/JF969367/JQ418321 |
| 23754 | Yunnan: Qingliang County | 10 November 2009 | Alnus nepalensis (Betulaceae) | JF969338/JF969387/JN645040 | |
| 18324 | Tibet: Linzhi City | 22 August 2005 | unknown | JF969344/JF969393/JN645018 | |
| 25807 | Tibet: Zhangmu County | 8 August 2010 | Fagaceae | JN644998/JN645012/JN645053 | |
| Mollitrichosiphum sp. | 26510 | Taiwan: Taman Mt. | 14 June 2011 | Fagaceae | JN645002/JQ418315/JQ418335 |
| 26515 | Taiwan: Hualian County | 20 June 2011 | Fagaceae | JN645003/JQ418316/JQ418336 | |
| Outgroups | |||||
| Eutrichosiphum tattakanum (Takahashi) | 17184 | Sichuan: Miyi County | 20 April 2005 | unknown | JN645055 |
| Kurisakia querciphila (Takahashi) | 16201 | Guizhou: Leigongshan Mt. | 31 May 2005 | Fagaceae | JF969355/JF969404/JQ418320 |
| Mesotrichosiphum pentaiarticulatum (Zhang & Qiao) | 18489 | Hainan: Lingshui Mt. | 27 March 2006 | Fagaceae | JQ418312/JQ418314/JQ418325 |
| Cervaphis echinata (Hille Ris Lambers) | 18461 | Hainan: Jianfengling Mt. | 21 March 2006 | Paulownia sp. (Scrophulariaceae) | JF969356/JF969405/JQ418324 |
| Individual datasets | Combined dataset | |||
|---|---|---|---|---|
| COI a | Cytb b | EF-1α c | ||
| Number of taxa | 72 | 57 | 59 | 50 |
| Aligned sequence length (bp) | 606 | 665 | 785 | 2056 |
| Conserved sites | 435 (473) | 450 (498) | 596 (656) | 1500 (1644) |
| Variable sites | 171 (133) | 215 (167) | 189 (129) | 556 (412) |
| Parsimony-informative sites | 147 (128) | 141 (134) | 111 (73) | 365 (328) |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhang, R.; Huang, X.; Jiang, L.; Lei, F.; Qiao, G. Species Differentiation of Chinese Mollitrichosiphum (Aphididae: Greenideinae) Driven by Geographical Isolation and Host Plant Acquirement. Int. J. Mol. Sci. 2012, 13, 10441-10460. https://doi.org/10.3390/ijms130810441
Zhang R, Huang X, Jiang L, Lei F, Qiao G. Species Differentiation of Chinese Mollitrichosiphum (Aphididae: Greenideinae) Driven by Geographical Isolation and Host Plant Acquirement. International Journal of Molecular Sciences. 2012; 13(8):10441-10460. https://doi.org/10.3390/ijms130810441
Chicago/Turabian StyleZhang, Ruiling, Xiaolei Huang, Liyun Jiang, Fumin Lei, and Gexia Qiao. 2012. "Species Differentiation of Chinese Mollitrichosiphum (Aphididae: Greenideinae) Driven by Geographical Isolation and Host Plant Acquirement" International Journal of Molecular Sciences 13, no. 8: 10441-10460. https://doi.org/10.3390/ijms130810441





