Bacterial Motility Measured by a Miniature Chamber for High-Pressure Microscopy
Abstract
:1. Introduction
2. Results and Discussion
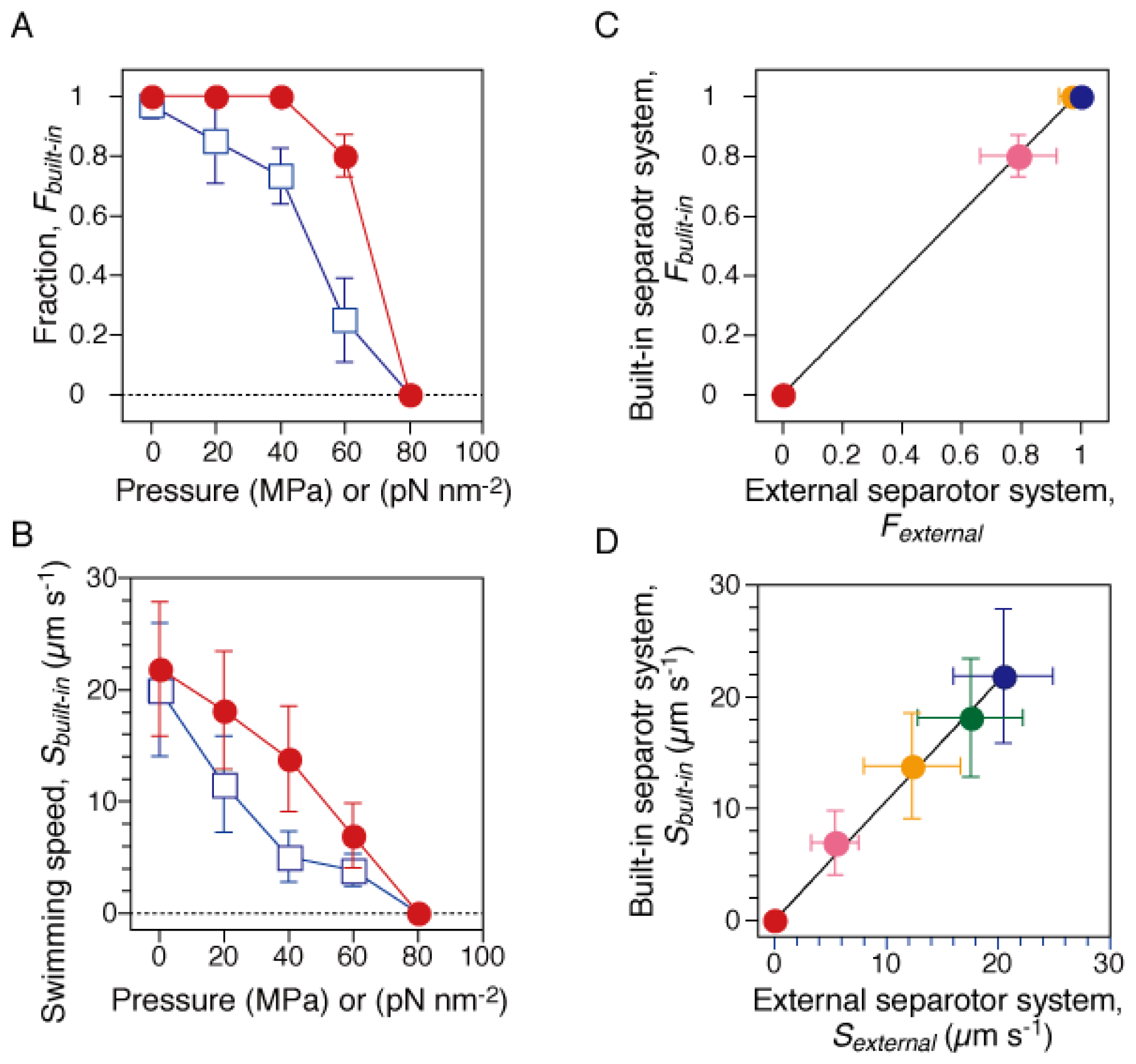
2.1. Pressure Dependence of Motility of Swimming E. coli
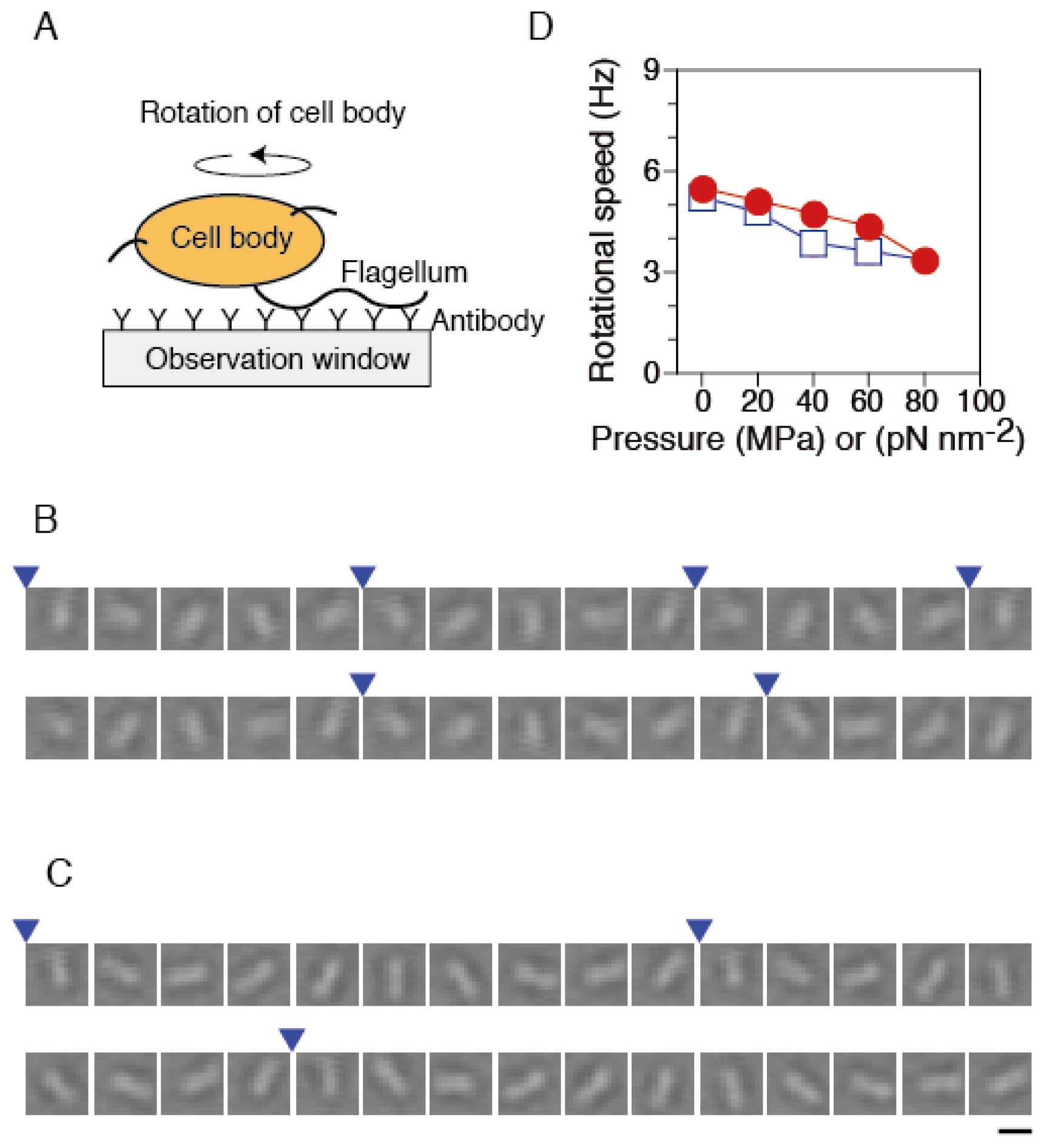
2.2. Torque Generation of Single Flagellar Motors in Tethered Cells
3. Experimental Section
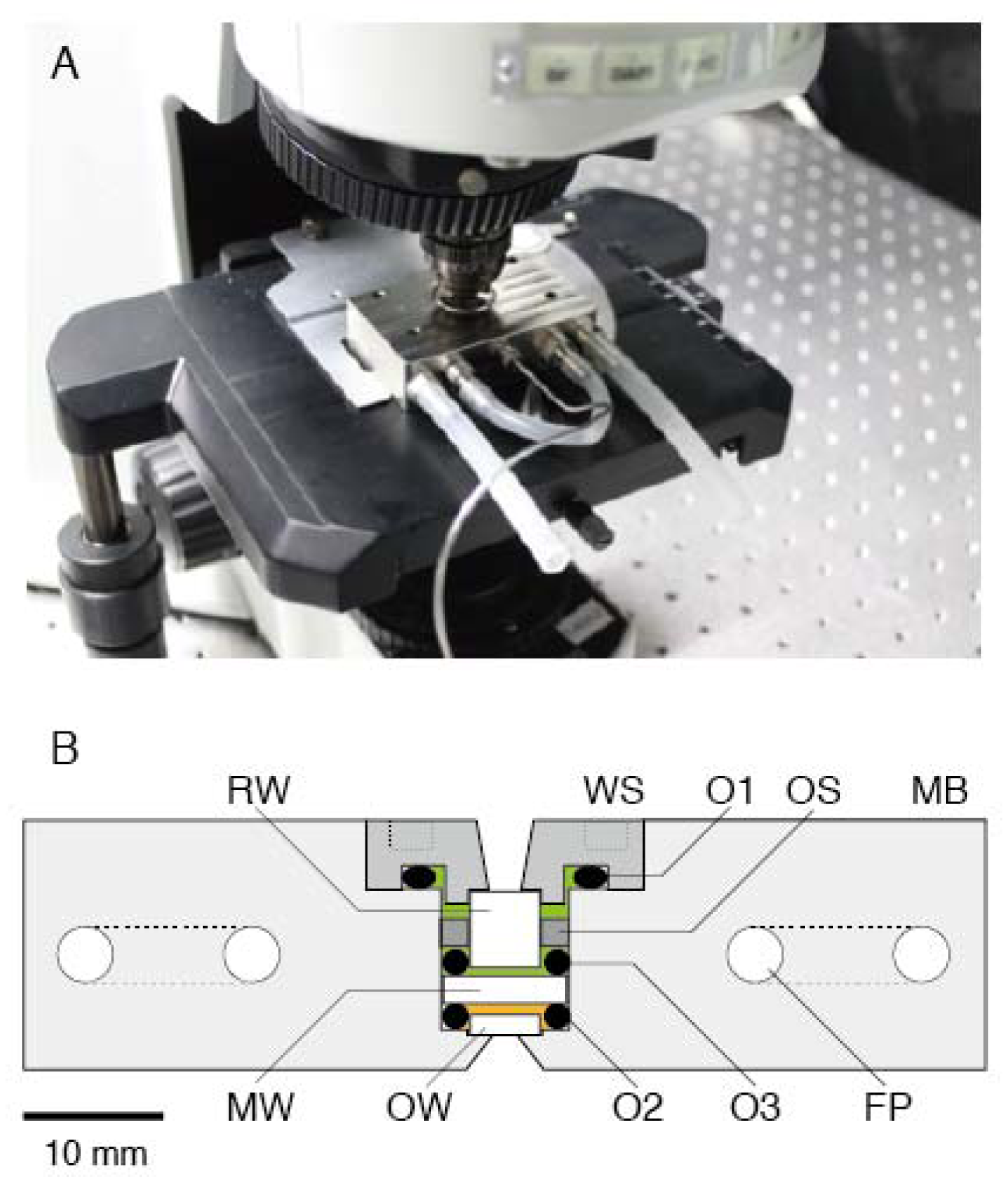
3.1. High-Pressure Chamber and Microscope
3.2. Bacterial Strains
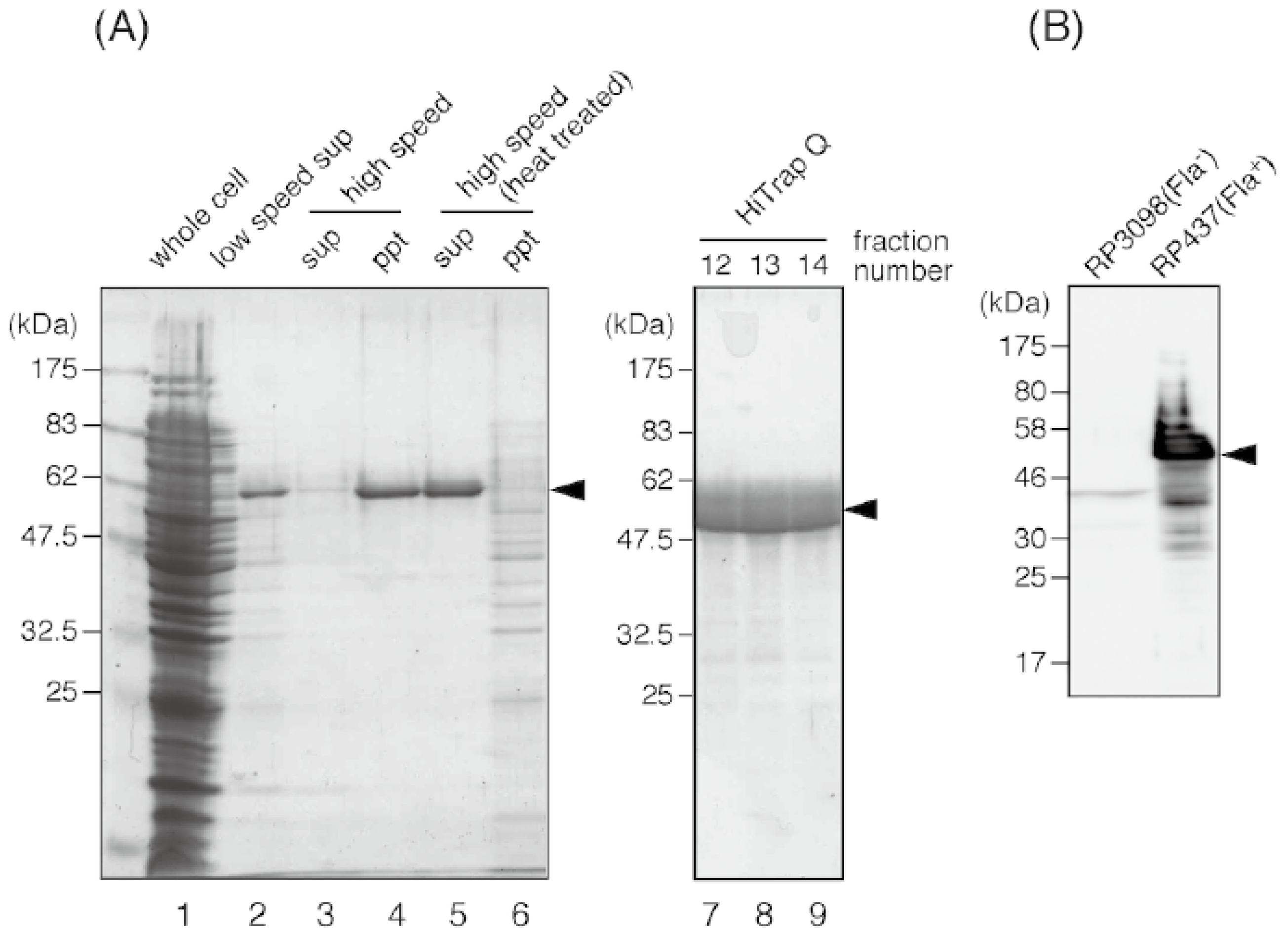
3.3. Purification of E. coli Flagellin and Preparation of the Anti-Flagellin Antibody
3.4. Motility Assays
4. Conclusions
Acknowledgements
References
- Kato, C.; Horikoshi, K. Gene expression under high pressure. Prog. Biotechnol 1996, 13, 59–66. [Google Scholar]
- Merrin, J.; Kumar, P.; Libchaber, A. Effects of pressure and temperature on the binding of RecA protein to single-stranded DNA. Proc. Natl. Acad. Sci. USA 2011, 108, 19913–19918. [Google Scholar]
- Kato, C.; Li, L.; Nogi, Y.; Nakamura, Y.; Tamaoka, J.; Horikoshi, K. Extremely barophilic bacteria isolated from the Mariana Trench, Challenger Deep, at a depth of 11,000 meters. Appl. Environ. Microbiol 1998, 64, 1510–1513. [Google Scholar]
- Bartlett, D.H. Pressure effects on in vivo microbial processes. BBA-Protein Struct M 2002, 1595, 367–381. [Google Scholar]
- Otter, T.; Salmon, E.D. Pressure-induced changes in Ca2+-channel excitability in Paramecium. J. exp. biol 1985, 117, 29–43. [Google Scholar]
- Abe, F. Exploration of the effects of high hydrostatic pressure on microbial growth, physiology and survival: perspectives from piezophysiology. Biosci. Biotechnol. Biochem 2007, 71, 2347–2357. [Google Scholar]
- Inoue, S.; Fuseler, J.; Salmon, E.D.; Ellis, G.W. Functional organization of mitotic microtubules. Physical chemistry of the in vivo equilibrium system. Biophys. J 1975, 15, 725–744. [Google Scholar]
- Frey, B.; Hartmann, M.; Herrmann, M.; Meyer-Pittroff, R.; Sommer, K.; Bluemelhuber, G. Microscopy under pressure—an optical chamber system for fluorescence microscopic analysis of living cells under high hydrostatic pressure. Microsc. Res. Tech 2006, 69, 65–72. [Google Scholar]
- Ranatunga, K.W.; Fortune, N.S.; Geeves, M.A. Hydrostatic compression in glycerinated rabbit muscle fibers. Biophys. J 1990, 58, 1401–1410. [Google Scholar]
- Friedrich, O.; Kress, K.R.; Hartmann, M.; Frey, B.; Sommer, K.; Ludwig, H.; Fink, R.H. Prolonged high-pressure treatments in mammalian skeletal muscle result in loss of functional sodium channels and altered calcium channel kinetics. Cell Biochem. Biophys 2006, 45, 71–83. [Google Scholar]
- Zobell, C.E.; Cobet, A.B. Growth, reproduction, and death rates of Escherichia coli at increased hydrostatic pressures. J. Bacteriol 1962, 84, 1228–1236. [Google Scholar]
- Welch, T.J.; Farewell, A.; Neidhardt, F.C.; Bartlett, D.H. Stress response of Escherichia coli to elevated hydrostatic pressure. J. Bacteriol 1993, 175, 7170–7177. [Google Scholar]
- Ishii, A.; Sato, T.; Wachi, M.; Nagai, K.; Kato, C. Effects of high hydrostatic pressure on bacterial cytoskeleton FtsZ polymers in vivo and in vitro. Microbiology 2004, 150, 1965–1972. [Google Scholar]
- Vass, H.; Black, S.L.; Herzig, E.M.; Ward, F.B.; Clegg, P.S.; Allen, R.J. A multipurpose modular system for high-resolution microscopy at high hydrostatic pressure. Rev. Sci. Instrum 2010, 81, 053710. [Google Scholar]
- Manas, P.; Mackey, B.M. Morphological and physiological changes induced by high hydrostatic pressure in exponential- and stationary-phase cells of Escherichia coli: Relationship with cell death. Appl. Environ. Microbiol 2004, 70, 1545–1554. [Google Scholar]
- Kawarai, T.; Wachi, M.; Ogino, H.; Furukawa, S.; Suzuki, K.; Ogihara, H.; Yamasaki, M. SulA-independent filamentation of Escherichia coli during growth after release from high hydrostatic pressure treatment. Appl. Microbiol. Biotechnol 2004, 64, 255–262. [Google Scholar]
- Moussa, M.; Perrier-Cornet, J.M.; Gervais, P. Damage in Escherichia coli cells treated with a combination of high hydrostatic pressure and subzero temperature. Appl. Environ. Microbiol 2007, 73, 6508–6518. [Google Scholar]
- Dzwolak, W.; Kato, M.; Taniguchi, Y. Fourier transform infrared spectroscopy in high-pressure studies on proteins. BBA-Protein Struct M 2002, 1595, 131–144. [Google Scholar]
- Kamatari, Y.O.; Kitahara, R.; Yamada, H.; Yokoyama, S.; Akasaka, K. High-pressure NMR spectroscopy for characterizing folding intermediates and denatured states of proteins. Methods 2004, 34, 133–143. [Google Scholar]
- Winter, R. Synchrotron X-ray and neutron small-angle scattering of lyotropic lipid mesophases, model biomembranes and proteins in solution at high pressure. BBA-Protein Struct M 2002, 1595, 160–184. [Google Scholar]
- Brooks, N.J.; Gauthe, B.L.L.E.; Terrill, N.J.; Rogers, S.E.; Templer, R.H.; Ces, O.; Seddon, J.M. Automated high pressure cell for pressure jump X-ray diffraction. Rev. Sci. Instrum 2010, 81, 064103. [Google Scholar]
- Filabozzi, A.; Deriu, A.; di Bari, M.T.; Russo, D.; Croci, S.; di Venere, A. Elastic incoherent neutron scattering as a probe of high pressure induced changes in protein flexibility. Biochim. Biophys. Acta 2010, 1804, 63–67. [Google Scholar] [Green Version]
- Colloc’h, N.; Girard, E.; Dhaussy, A.C.; Kahn, R.; Ascone, I.; Mezouar, M.; Fourme, R. High pressure macromolecular crystallography: The 140-MPa crystal structure at 2.3 angstrom resolution of urate oxidase, a 135-kDa tetrameric assembly. BBA-Proteins Proteom 2006, 1764, 391–397. [Google Scholar]
- Usui, K.; Hiraki, T.; Kawamoto, J.; Kurihara, T.; Nogi, Y.; Kato, C.; Abe, F. Eicosapentaenoic acid plays a role in stabilizing dynamic membrane structure in the deep-sea piezophile Shewanella violacea: A study employing high-pressure time-resolved fluorescence anisotropy measurement. BBA-Biomembranes 2012, 1818, 574–583. [Google Scholar]
- Salmon, E.D.; Ellis, G.W. A new miniature hydrostatic pressure chamber for microscopy. Strainfree optical glass windows facilitate phase-contrast and polarized-light microscopy of living cells. Optional fixture permits simultaneous control of pressure and temperature. J. Cell Biol 1975, 65, 587–602. [Google Scholar]
- Mukai, S.A.; Deguchi, S.; Tsujii, K. A high-temperature and -pressure microscope cell to observe colloidal behaviors in subcritical and supercritical water: Brownian motion of colloids near a wall. Colloid Surface A 2006, 282, 483–488. [Google Scholar]
- Nishiyama, M.; Kimura, Y.; Nishiyama, Y.; Terazima, M. Pressure-induced changes in the structure and function of the kinesin-microtubule complex. Biophys. J 2009, 96, 1142–1150. [Google Scholar]
- Nishiyama, M.; Shimoda, Y.; Hasumi, M.; Kimura, Y.; Terazima, M. Microtubule depolymerization at high pressure. Ann. N. Y. Acad. Sci 2010, 1189, 86–90. [Google Scholar]
- Nishiyama, M.; Sowa, M. Microscopic analysis of bacterial motility at high pressure. Biophys. J 2012, 102, 1872–1880. [Google Scholar]
- Hazelbauer, G.L.; Falke, J.J.; Parkinson, J.S. Bacterial chemoreceptors: high-performance signaling in networked arrays. Trends Biochem. Sci 2008, 33, 9–19. [Google Scholar]
- Porter, S.L.; Wadhams, G.H.; Armitage, J.P. Signal processing in complex chemotaxis pathways. Nat. Rev. Microbiol 2011, 9, 153–165. [Google Scholar]
- Taylor, B.L. Aer on the inside looking out: paradigm for a PAS-HAMP role in sensing oxygen, redox and energy. Mol. Microbiol 2007, 65, 1415–1424. [Google Scholar]
- Yang, H.; Inokuchi, H.; Adler, J. Phototaxis away from blue light by an Escherichia coli mutant accumulating protoporphyrin IX. Proc. Natl. Acad. Sci. USA 1995, 92, 7332–7336. [Google Scholar]
- Maeda, K.; Imae, Y.; Shioi, J.I.; Oosawa, F. Effect of temperature on motility and chemotaxis of Escherichia coli. J. Bacteriol 1976, 127, 1039–1046. [Google Scholar]
- Nishiyama, S.I.; Umemura, T.; Nara, T.; Homma, M.; Kawagishi, I. Conversion of a bacterial warm sensor to a cold sensor by methylation of a single residue in the presence of an attractant. Mol. Microbiol 1999, 32, 357–365. [Google Scholar]
- Nishiyama, S.; Maruyama, I.N.; Homma, M.; Kawagishi, I. Inversion of thermosensing property of the bacterial receptor Tar by mutations in the second transmembrane region. J. Mol. Biol 1999, 286, 1275–1284. [Google Scholar]
- Salman, H.; Libchaber, A. A concentration-dependent switch in the bacterial response to temperature. Nat. Cell Biol 2007, 9, 1098–1100. [Google Scholar]
- Paster, E.; Ryu, W.S. The thermal impulse response of Escherichia coli. Proc. Natl. Acad. Sci. USA 2008, 105, 5373–5377. [Google Scholar]
- Nishiyama, S.; Ohno, S.; Ohta, N.; Inoue, Y.; Fukuoka, H.; Ishijima, A.; Kawagishi, I. Thermosensing function of the Escherichia coli redox sensor Aer. J. Bacteriol 2010, 192, 1740–1743. [Google Scholar]
- Frank, V.; Koler, M.; Furst, S.; Vaknin, A. The physical and functional thermal sensitivity of bacterial chemoreceptors. J. Mol. Biol 2011, 411, 554–566. [Google Scholar]
- Namba, K.; Vonderviszt, F. Molecular architecture of bacterial flagellum. Q. Rev. Biophys 1997, 30, 1–65. [Google Scholar]
- Berry, R.M.; Armitage, J.P. The bacterial flagella motor. Adv. Microb. Physiol 1999, 41, 291–337. [Google Scholar]
- Berg, H.C. The rotary motor of bacterial flagella. Annu. Rev. Biochem 2003, 72, 19–54. [Google Scholar]
- Kojima, S.; Blair, D.F. The bacterial flagellar motor: structure and function of a complex molecular machine. Int. Rev. Cytol 2004, 233, 93–134. [Google Scholar]
- Minamino, T.; Imada, K.; Namba, K. Molecular motors of the bacterial flagella. Curr. Opin. Struct. Biol 2008, 18, 693–701. [Google Scholar]
- Sowa, Y.; Berry, R.M. Bacterial flagellar motor. Q. Rev. Biophys 2008, 41, 103–132. [Google Scholar]
- Terashima, H.; Kojima, S.; Homma, M. Flagellar motility in bacteria structure and function of flagellar motor. Int. Rev. Cell Mol. Biol 2008, 270, 39–85. [Google Scholar]
- Silverman, M.; Simon, M. Flagellar rotation and the mechanism of bacterial motility. Nature 1974, 249, 73–74. [Google Scholar]
- Meganathan, R.; Marquis, R.E. Loss of bacterial motility under pressure. Nature 1973, 246, 525–527. [Google Scholar]
- Eloe, E.A.; Lauro, F.M.; Vogel, R.F.; Bartlett, D.H. The deep-sea bacterium Photobacterium profundum SS9 utilizes separate flagellar systems for swimming and swarming under high-pressure conditions. Appl. Environ. Microbiol 2008, 74, 6298–6305. [Google Scholar]
- Sowa, Y.; Rowe, A.D.; Leake, M.C.; Yakushi, T.; Homma, M.; Ishijima, A.; Berry, R.M. Direct observation of steps in rotation of the bacterial flagellar motor. Nature 2005, 437, 916–919. [Google Scholar]
- Nakamura, S.; Kami-ike, N.; Yokota, J.P.; Minamino, T.; Namba, K. Evidence for symmetry in the elementary process of bidirectional torque generation by the bacterial flagellar motor. Proc. Natl. Acad. Sci. USA 2010, 107, 17616–17620. [Google Scholar]
- Manson, M.D.; Tedesco, P.; Berg, H.C.; Harold, F.M.; van der Drift, C. A protonmotive force drives bacterial flagella. Proc. Natl. Acad. Sci. USA 1977, 74, 3060–3064. [Google Scholar]
- Hirota, N.; Kitada, M.; Imae, Y. Flagellar Motors of Alkalophilic Bacillus Are Powered by an Electrochemical Potential Gradient of Na+. FEBS Lett 1981, 132, 278–280. [Google Scholar]
- Sowa, Y.; Hotta, H.; Homma, M.; Ishijima, A. Torque-speed relationship of the Na+-driven flagellar motor of Vibrio alginolyticus. J. Mol. Biol 2003, 327, 1043–1051. [Google Scholar]
- Ryu, W.S.; Berry, R.M.; Berg, H.C. Torque-generating units of the flagellar motor of Escherichia coli have a high duty ratio. Nature 2000, 403, 444–447. [Google Scholar]
- Inoue, Y.; Lo, C.J.; Fukuoka, H.; Takahashi, H.; Sowa, Y.; Pilizota, T.; Wadhams, G.H.; Homma, M.; Berry, R.M.; Ishijima, A. Torque-speed relationships of Na+-driven chimeric flagellar motors in Escherichia coli. J. Mol. Biol 2008, 376, 1251–1259. [Google Scholar]
- Iwazawa, J.; Imae, Y.; Kobayasi, S. Study of the torque of the bacterial flagellar motor using a rotating electric field. Biophys. J 1993, 64, 925–933. [Google Scholar]
- Lowe, G.; Meister, M.; Berg, H.C. Rapid Rotation of Flagellar Bundles in Swimming Bacteria. Nature 1987, 325, 637–640. [Google Scholar]
- Chen, X.; Berg, H.C. Torque-speed relationship of the flagellar rotary motor of Escherichia coli. Biophys. J 2000, 78, 1036–1041. [Google Scholar]
- Yuan, J.; Berg, H.C. Thermal and solvent-isotope effects on the flagellar rotary motor near zero load. Biophys. J 2010, 98, 2121–2126. [Google Scholar]
- Baker, M.A.; Inoue, Y.; Takeda, K.; Ishijima, A.; Berry, R.M. Two methods of temperature control for single-molecule measurements. Eur. Biophys. J 2011, 40, 651–660. [Google Scholar]
- Manson, M.D.; Tedesco, P.M.; Berg, H.C. Energetics of Flagellar Rotation in Bacteria. J. Mol. Biol 1980, 138, 541–561. [Google Scholar]
- Shioi, J.I.; Matsuura, S.; Imae, Y. Quantitative measurements of proton motive force and motility in Bacillus subtilis. J. Bacteriol 1980, 144, 891–897. [Google Scholar]
- Blair, D.F.; Berg, H.C. The MotA Protein of Escherichia-coli is a proton-conducting component of the flagellar motor. Cell 1990, 60, 439–449. [Google Scholar]
- Kojima, S.; Blair, D.F. Solubilization and purification of the MotA/MotB complex of Escherichia coli. Biochemistry 2004, 43, 26–34. [Google Scholar]
- Braun, T.F.; Al-Mawsawi, L.Q.; Kojima, S.; Blair, D.F. Arrangement of core membrane segments in the MotA/MotB proton-channel complex of Escherichia coli. Biochemistry 2004, 43, 35–45. [Google Scholar]
- Hosking, E.R.; Manson, M.D. Clusters of charged residues at the C terminus of MotA and N terminus of MotB are important for function of the Escherichia coli flagellar motor. J. Bacteriol 2008, 190, 5517–5521. [Google Scholar]
- Suzuki, H.; Yonekura, K.; Namba, K. Structure of the rotor of the bacterial flagellar motor revealed by electron cryomicroscopy and single-particle image analysis. J. Mol. Biol 2004, 337, 105–113. [Google Scholar]
- Lee, L.K.; Ginsburg, M.A.; Crovace, C.; Donohoe, M.; Stock, D. Structure of the torque ring of the flagellar motor and the molecular basis for rotational switching. Nature 2010, 466, 996–1000. [Google Scholar]
- Minamino, T.; Imada, K.; Kinoshita, M.; Nakamura, S.; Morimoto, Y.V.; Namba, K. Structural insight into the rotational switching mechanism of the bacterial flagellar motor. PLoS Biol 2011, 9, e1000616. [Google Scholar]
- Garza, A.G.; Harris-Haller, L.W.; Stoebner, R.A.; Manson, M.D. Motility protein interactions in the bacterial flagellar motor. Proc. Natl. Acad. Sci. USA 1995, 92, 1970–1974. [Google Scholar]
- Zhou, J.; Lloyd, S.A.; Blair, D.F. Electrostatic interactions between rotor and stator in the bacterial flagellar motor. Proc. Natl. Acad. Sci. USA 1998, 95, 6436–6441. [Google Scholar]
- Lloyd, S.A.; Blair, D.F. Charged residues of the rotor protein FliG essential for torque generation in the flagellar motor of Escherichia coli. J. Mol. Biol 1997, 266, 733–744. [Google Scholar]
- Kojima, S.; Blair, D.F. Conformational change in the stator of the bacterial flagellar motor. Biochemistry 2001, 40, 13041–13050. [Google Scholar]
- Che, Y.S.; Nakamura, S.; Kojima, S.; Kami-ike, N.; Namba, K.; Minamino, T. Suppressor analysis of the MotB(D33E) mutation to probe bacterial flagellar motor dynamics coupled with proton translocation. J. Bacteriol 2008, 190, 6660–6667. [Google Scholar]
- Nakamura, S.; Kami-ike, N.; Yokota, J.P.; Kudo, S.; Minamino, T.; Namba, K. Effect of intracellular pH on the torque-speed relationship of bacterial proton-driven flagellar motor. J. Mol. Biol 2009, 386, 332–338. [Google Scholar]
- Kudo, S.; Magariyama, Y.; Aizawa, S. Abrupt changes in flagellar rotation observed by laser dark-field microscopy. Nature 1990, 346, 677–680. [Google Scholar]
- Sowa, Y.; Steel, B.C.; Berry, R.M. A simple backscattering microscope for fast tracking of biological molecules. Rev. Sci. Instrum 2010, 81, 113704. [Google Scholar]
- Bai, F.; Branch, R.W.; Nicolau, D.V., Jr; Pilizota, T.; Steel, B.C.; Maini, P.K.; Berry, R.M. Conformational spread as a mechanism for cooperativity in the bacterial flagellar switch. Science 2010, 327, 685–689. [Google Scholar]
- Terasawa, S.; Fukuoka, H.; Inoue, Y.; Sagawa, T.; Takahashi, H.; Ishijima, A. Coordinated reversal of flagellar motors on a single Escherichia coli cell. Biophys. J 2011, 100, 2193–2200. [Google Scholar]
- Fukuoka, H.; Inoue, Y.; Terasawa, S.; Takahashi, H.; Ishijima, A. Exchange of rotor components in functioning bacterial flagellar motor. Biochem. Biophys. Res. Commun 2010, 394, 130–135. [Google Scholar]
- Delalez, N.J.; Wadhams, G.H.; Rosser, G.; Xue, Q.; Brown, M.T.; Dobbie, I.M.; Berry, R.M.; Leake, M.C.; Armitage, J.P. Signal-dependent turnover of the bacterial flagellar switch protein FliM. Proc. Natl. Acad. Sci. USA 2010, 107, 11347–11351. [Google Scholar]
- Lazova, M.D.; Ahmed, T.; Bellomo, D.; Stocker, R.; Shimizu, T.S. Response rescaling in bacterial chemotaxis. Proc. Natl. Acad. Sci. USA 2011, 108, 13870–13875. [Google Scholar]
- Chiu, S.W.; Leake, M.C. Functioning nanomachines seen in real-time in living bacteria using single-molecule and super-resolution fluorescence imaging. Int. J. Mol. Sci 2011, 12, 2518–2542. [Google Scholar]
- Taniguchi, Y.; Choi, P.J.; Li, G.W.; Chen, H.; Babu, M.; Hearn, J.; Emili, A.; Xie, X.S. Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 2010, 329, 533–538. [Google Scholar]
- Wang, S.; Furchtgott, L.; Huang, K.C.; Shaevitz, J.W. Helical insertion of peptidoglycan produces chiral ordering of the bacterial cell wall. Proc. Natl. Acad. Sci. USA 2012, 109, E595–E604. [Google Scholar]
- Parkinson, J.S. Complementation analysis and deletion mapping of Escherichia coli mutants defective in chemotaxis. J. Bacteriol 1978, 135, 45–53. [Google Scholar]
- Scharf, B.E.; Fahrner, K.A.; Turner, L.; Berg, H.C. Control of direction of flagellar rotation in bacterial chemotaxis. Proc. Natl. Acad. Sci. USA 1998, 95, 201–206. [Google Scholar]
- Li, N.; Kojima, S.; Homma, M. Characterization of the periplasmic region of PomB, a Na+-driven flagellar stator protein in Vibrio alginolyticus. J. Bacteriol 2011, 193, 3773–3784. [Google Scholar]
- Muramoto, K.; Macnab, R.M. Deletion analysis of MotA and MotB, components of the force-generating unit in the flagellar motor of Salmonella. Mol. Microbiol 1998, 29, 1191–1202. [Google Scholar]
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Nishiyama, M.; Kojima, S. Bacterial Motility Measured by a Miniature Chamber for High-Pressure Microscopy. Int. J. Mol. Sci. 2012, 13, 9225-9239. https://doi.org/10.3390/ijms13079225
Nishiyama M, Kojima S. Bacterial Motility Measured by a Miniature Chamber for High-Pressure Microscopy. International Journal of Molecular Sciences. 2012; 13(7):9225-9239. https://doi.org/10.3390/ijms13079225
Chicago/Turabian StyleNishiyama, Masayoshi, and Seiji Kojima. 2012. "Bacterial Motility Measured by a Miniature Chamber for High-Pressure Microscopy" International Journal of Molecular Sciences 13, no. 7: 9225-9239. https://doi.org/10.3390/ijms13079225



