Identification and Characterization of a Bursaphelenchus xylophilus (Aphelenchida: Aphelenchoididae) Thermotolerance-Related Gene: Bx-HSP90
Abstract
:1. Introduction
2. Results and Discussion
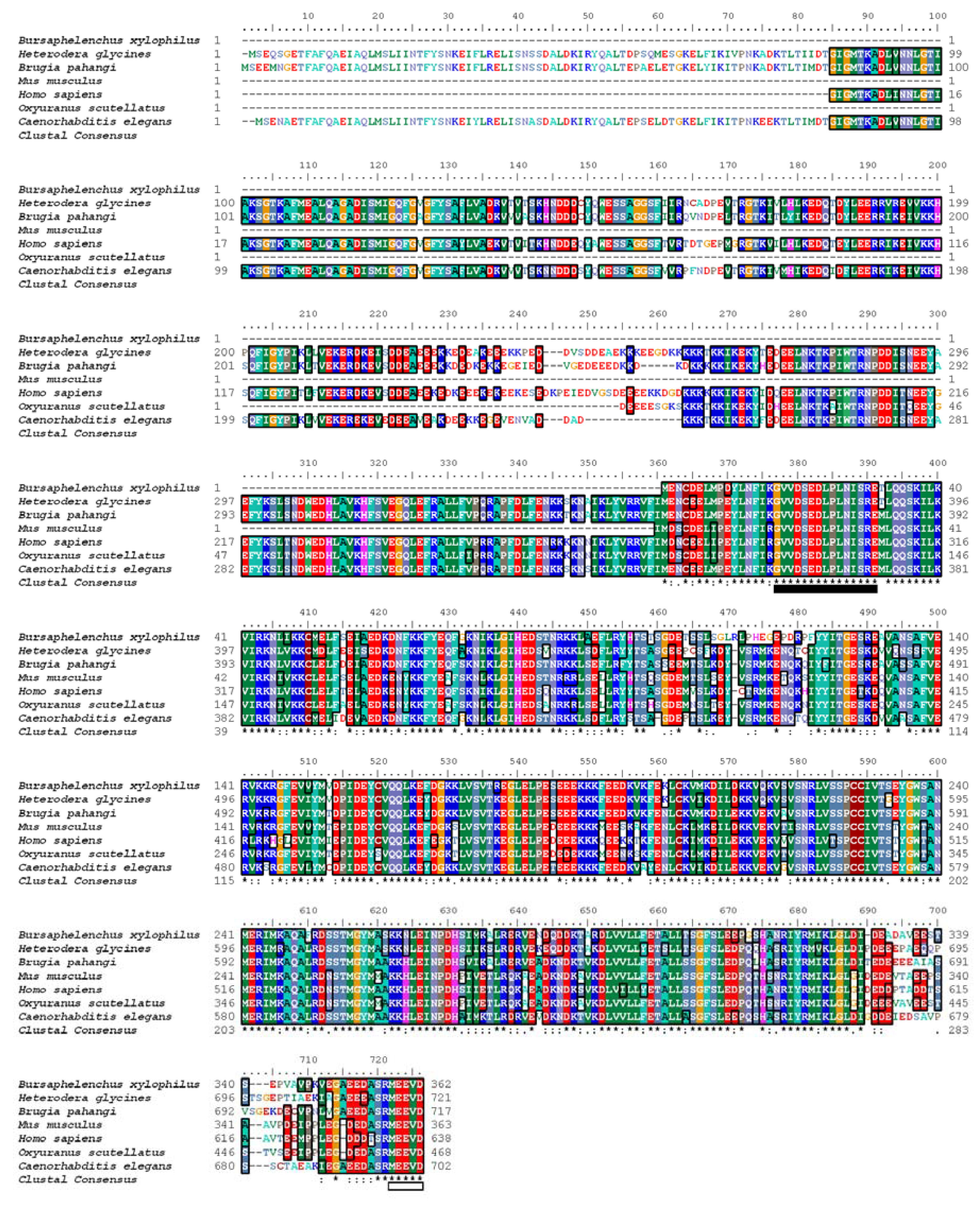
2.1. cDNA Cloning and Homology Analysis
2.2. In Situ Hybridization
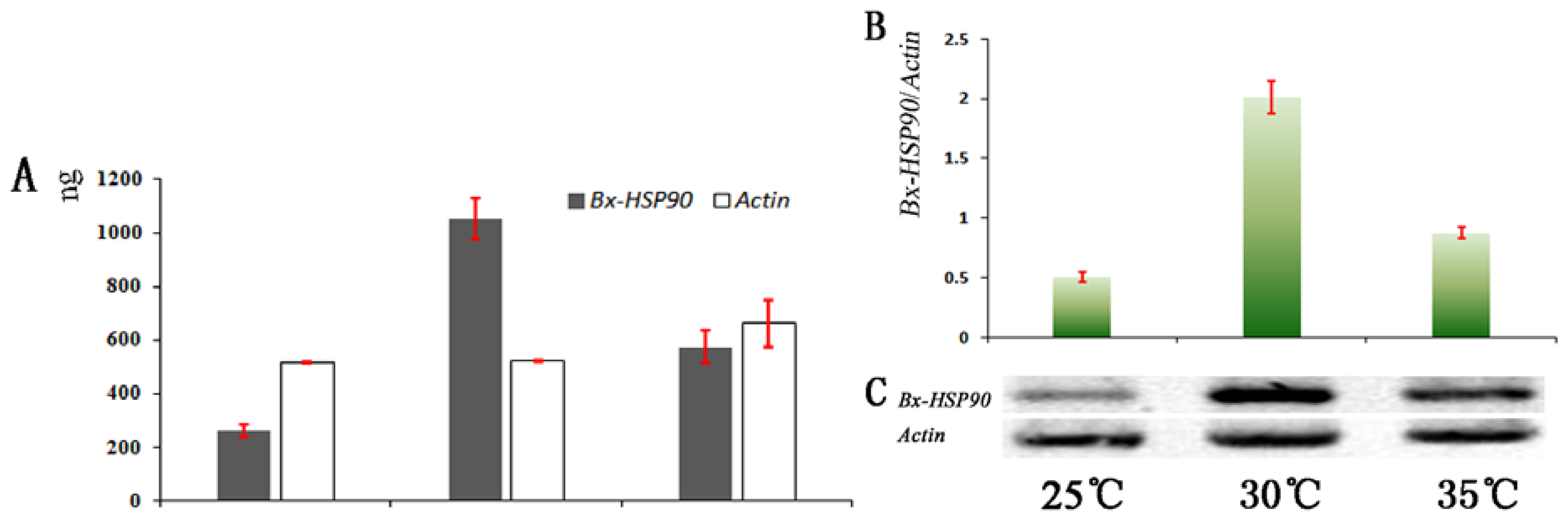
2.3. Analysis of Transcript Abundance
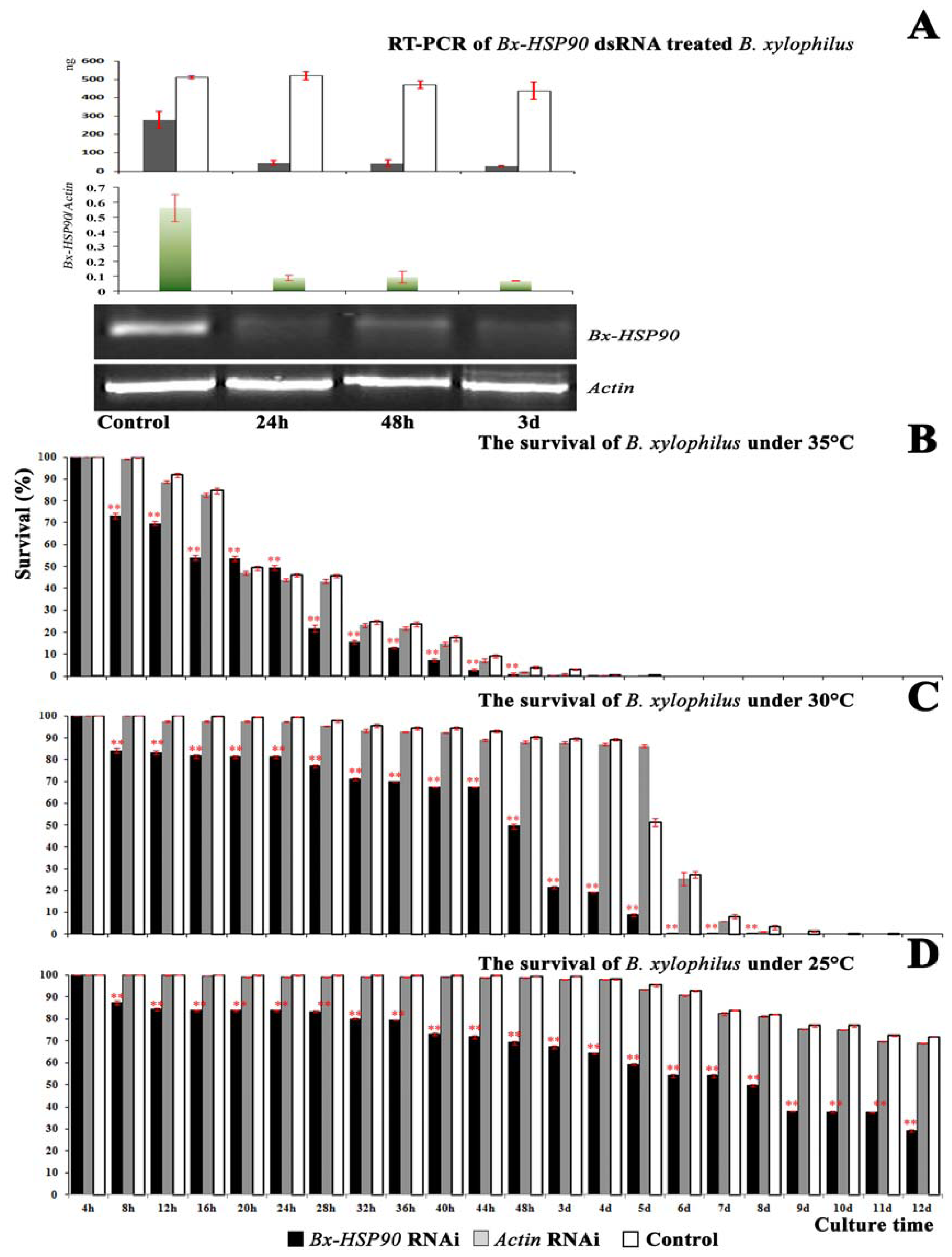
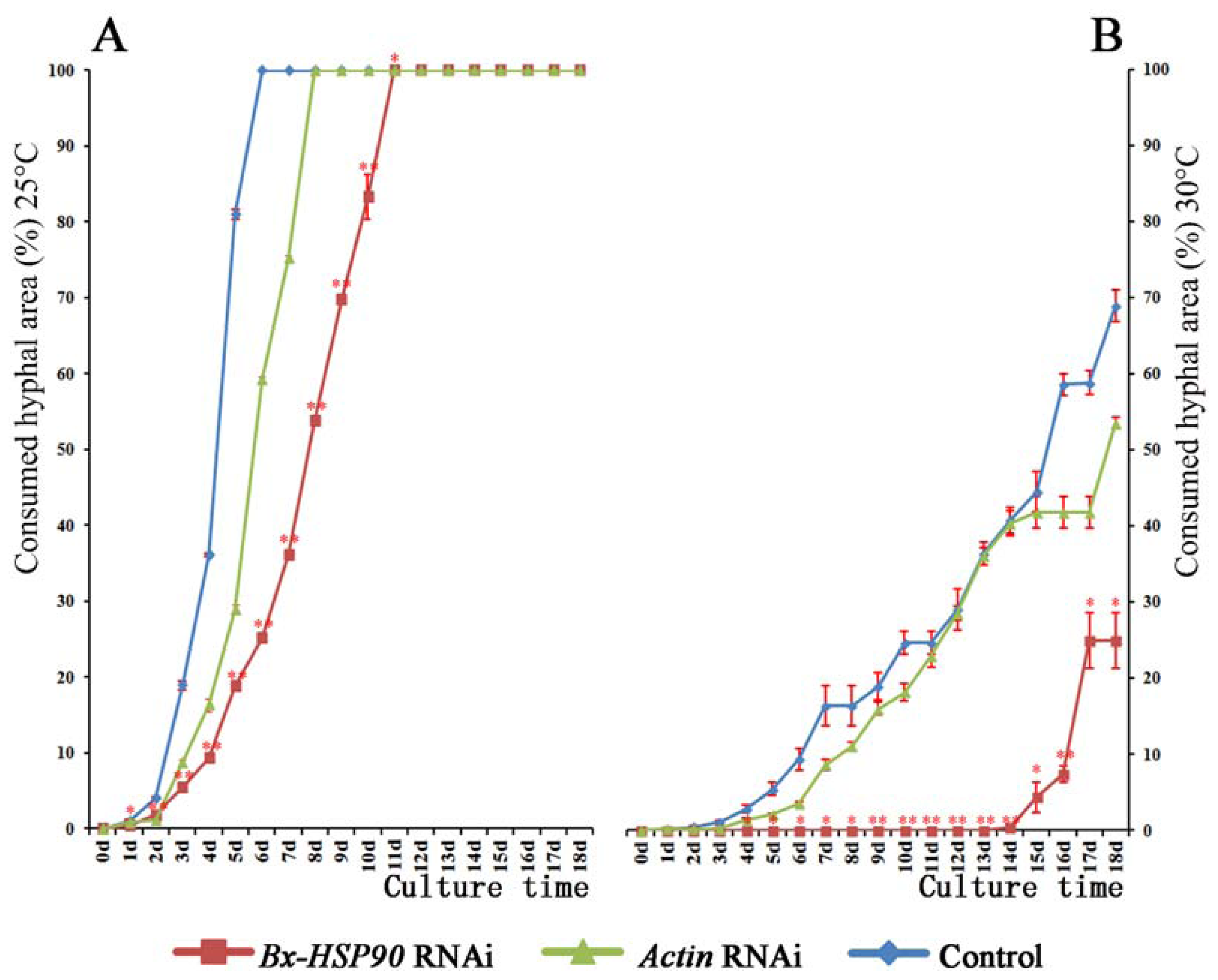
2.4. RNAi of Bx-HSP90
3. Experimental Section
3.1. Nematode Culture and Heat Shock
3.2. Construction of the Subtracted cDNA Libraries and Sequence Analyses
3.3. Cloning of Bx-HSP90
3.4. Homology Analysis
3.5. In Situ Hybridization
3.6. Analysis of Transcript Abundance
3.7. RNAi of Bx-HSP90
4. Conclusions
Acknowledgments
References
- Mamiya, Y. History of pine wilt disease in Japan. J. Nematol 1988, 20, 219–226. [Google Scholar]
- Rutherford, T.A.; Webster, J.M. Distribution of pine wilt disease with respect to temperature in North America, Japan, and Europe. Can. J. For. Res 1987, 17, 1050–1059. [Google Scholar]
- Bo, G.Z.; Kazuyoshi, F.; Jack, R.S.; Yuko, T. Pine Wilt Disease; Springer: Tokyo, Japan, 2008; pp. 2–40. [Google Scholar]
- Enda, N. Current status of pine wilt disease in Korea. For. Pests 1989, 38, 148–152. [Google Scholar]
- Mota, M.M.; Braasch, H.; Bravo, M.A.; Penas, A.C.; Burgermeister, W.; Metge, K.; Sousa, E. First report of Bursaphelenchus xylophilus in Portugal and in Europe. Nematology 1999, 8, 727–734. [Google Scholar]
- Devaney, E. Thermoregulation in the life cycle of nematodes. Int. J. Parasitol 2006, 36, 641–649. [Google Scholar]
- Mamiya, Y. Pathology of pine wilt disease caused by Bursaphelenchus xylophilus. Annu. Rev. Phytopathol 1983, 21, 201–220. [Google Scholar]
- Nik, A.H.; Victoria, G.; Richard, D.E.; Kirsty, M.; Eileen, D. Hsp-90 and the biology of nematodes. BMC Evol. Biol 2009, 9, 254. [Google Scholar]
- Veronika, H.; Christoph, J.O.K.; Klaus, R. Hsp90 in non-mammalian metazoan model systems. BBA Mol. Cell Res 2012, 1823, 712–721. [Google Scholar]
- Guangbo, C.; William, D.B.; Chris, W.S.; Rong, L. Hsp90 stress potentiates rapid cellular adaptation through induction of aneuploidy. Nature 2012, 482, 246–250. [Google Scholar]
- Mamiya, Y. The life history of the pine wood nematode, Bursaphelenchus lignicolus. Jpn. J. Nematol 1975, 5, 16–25. [Google Scholar]
- Kikuchi, T.; Cotton, J.A.; Dalzell, J.J.; Hasegawa, K.; Kanzaki, N.; McVeigh, P.; Takanashi, T.; Tsai, I.J.; Assefa, S.A.; Cock, P.J.A.; et al. Genomic insights into the origin of parasitism in the emerging plant pathogen Bursaphelenchus xylophilus. PLoS Pathog 2011, 7, 1–17. [Google Scholar]
- Thompson, F.J.; Cockroft, A.C.; Wheatley, I.; Britton, C.; Devaney, E. Heat shock and developmental expression of hsp83 in the filarial nematode Brugia pahangi. Eur. J. Biochem 2001, 268, 5808–5815. [Google Scholar]
- Devaney, E.; O’neill, K.; Harnett, W.; Whitesell, L.; Kinnaird, J.H. Hsp90 is essential in the filarial nematode Brugia pahangi. Int. J. Parasitol 2005, 35, 627–636. [Google Scholar]
- Urwin, P.E.; Lilley, C.J.; Atkinson, H.J. Ingestion of double–stranded RNA by preparasitic juvenile cyst nematodes leads to RNA interference. Mol. Plant Microbe Interaction 2002, 15, 747–752. [Google Scholar]
- Butov, A.; Johnson, T.; Cypser, J.; Sannikov, I.; Volkov, M.; Sehl, M.; Yashin, A. Hormesis and debilitation effects in stress experiments using the nematode worm Caenorhabditis elegans: the model of balance between cell damage and HSP levels. Exp. Gerontol 2001, 37, 57–66. [Google Scholar]
- Atkinson, H.J.; Urwin, P.E.; McPherson, M.J. Engineering plants for nematode resistance. Annu. Rev. Phytopathol 2003, 41, 615–639. [Google Scholar]
- Diatchenko, L.; Lau, Y.F.; Campbell, A.P.; Chenchik, A.; Moqadam, F.; Huang, B.; Lukyanov, S.; Lukyanov, K.; Gurskaya, N.; Sverdlov, E.D.; et al. Suppression subtractive hybridization: A method for generating differentially regulated or tissue-specific cDNA probes and libraries. Proc. Nat. Acad. Sci. USA 1996, 93, 6025–6030. [Google Scholar]
- Martin, S.A.; Thompson, F.J.; Devaney, E. The construction of spliced leader cDNA libraries from the filarial nematode Brugia pahangi. Mol. Biochem. Parasitol 1995, 70, 241–245. [Google Scholar]
- Blaxter, M.; Liu, L. Nematode spliced leader–ubiquity Evolution and Utility. Int. J. Parasitol 1996, 26, 1025–1033. [Google Scholar]
- NCBI. BLAST. Available online: http://www.ncbi.nlm.nih.gov/BLAST/ accessed on 20 July 2008.
- EBI. ClustalW. Available online: http://www.ebi.ac.uk/clustalw/ accessed on 20 July 2008.
- Bio-soft. Identity and Similarity Analysis program. Available online: http://www.bio-soft.net/sms/index.html/ accessed on 20 July 2008.
- Jannick, D.B.; Henrik, N.; von Gunnar, H.; Søren, B. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol 2004, 340, 783–795. [Google Scholar]
- De Boer, J.M.; Yan, Y.; Smant, G.; Davis, E.L.; Baum, T.J. In situ hybridization to messenger RNA in Heterodera glycines. J. Nematol 1998, 30, 309–312. [Google Scholar]
- Joachim, J.; Bartel, V.; Annelies, H. Four transthyretin–like genes of the migratory plant–parasitic nematode Radopholus similis: Members of an extensive nematode–specific family. Gene 2007, 402, 9–19. [Google Scholar]
- Rasband, W.S. ImageJ. Available online: http://imagej.nih.gov/ij/ accessed on 5 March 2012.
- Abràmoff, M.D.; Magalhães, P.J.; Ram, S.J. Image processing with ImageJ. Biophotonic. Int 2004, 11, 36–42. [Google Scholar]


| Primer Name | Primer Sequence | Reference |
|---|---|---|
| SL1 | 5′-GGTTTAATTACCCAAGTTTGAG-3′ | Blaxter et al. [20] |
| SLHSP90-A2L | 5′-GTCAAGAGGGCGGTCTCGA-3′ | This study |
| SLHSP90-S1L | 5′-CACCTCCGGAGACGAGACC-3′ | This study |
| EHSP90S2L | 5′-ACTTCATCATGGAGAACTGCG-3′ | This study |
| EHSP90A1L | 5′-ATCATTAGGAGGACAACAGA-3′ | This study |
| HSP90rtR1L | 5′-GCGATGAACTGATGCCCGACTAC-3′ | This study |
| HSP90rtF1L | 5′-CGAAGGCAGAGTTGGCGACG-3′ | This study |
| RsACT_F | 5′-GAAAGAGGGCCGGAAGAG-3′ | Joachim et al. [26] |
| RsACT_R | 5′-AGATCGTCCGCGACATAAAG-3′ | Joachim et al. [26] |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, F.; Wang, Z.; Li, D.; Chen, Q. Identification and Characterization of a Bursaphelenchus xylophilus (Aphelenchida: Aphelenchoididae) Thermotolerance-Related Gene: Bx-HSP90. Int. J. Mol. Sci. 2012, 13, 8819-8833. https://doi.org/10.3390/ijms13078819
Wang F, Wang Z, Li D, Chen Q. Identification and Characterization of a Bursaphelenchus xylophilus (Aphelenchida: Aphelenchoididae) Thermotolerance-Related Gene: Bx-HSP90. International Journal of Molecular Sciences. 2012; 13(7):8819-8833. https://doi.org/10.3390/ijms13078819
Chicago/Turabian StyleWang, Feng, Zhiying Wang, Danlei Li, and Qiaoli Chen. 2012. "Identification and Characterization of a Bursaphelenchus xylophilus (Aphelenchida: Aphelenchoididae) Thermotolerance-Related Gene: Bx-HSP90" International Journal of Molecular Sciences 13, no. 7: 8819-8833. https://doi.org/10.3390/ijms13078819
APA StyleWang, F., Wang, Z., Li, D., & Chen, Q. (2012). Identification and Characterization of a Bursaphelenchus xylophilus (Aphelenchida: Aphelenchoididae) Thermotolerance-Related Gene: Bx-HSP90. International Journal of Molecular Sciences, 13(7), 8819-8833. https://doi.org/10.3390/ijms13078819




