Role of Endoplasmic Reticulum Aminopeptidases in Health and Disease: from Infection to Cancer
Abstract
:1. Introduction
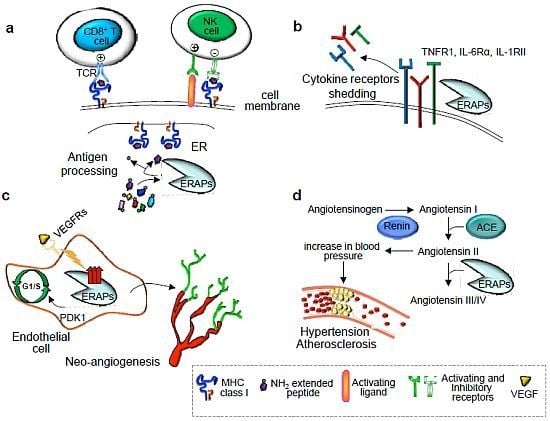
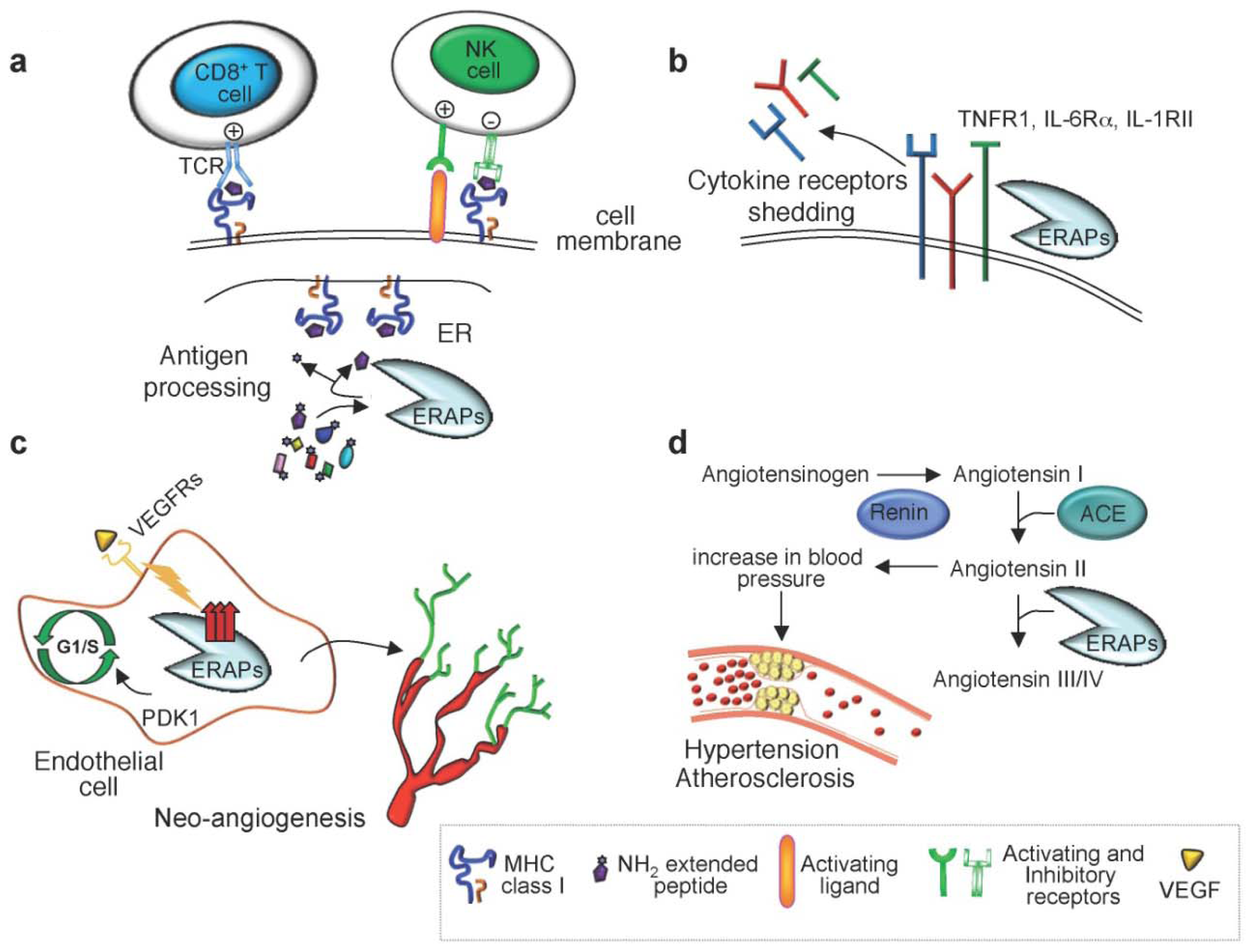
2. Immunological Functions of ER Aminopeptidases
2.1. Trimming of Antigenic Peptides by ER Aminopeptidases
2.2. Cytokine Receptor Shedding
3. Nonimmunological Functions of ER Aminopeptidases
4. Alteration of ERAP Functions in Human Diseases
4.1. Hypertension
4.2. Bacterial and Viral Infections
4.3. Autoimmune Diseases
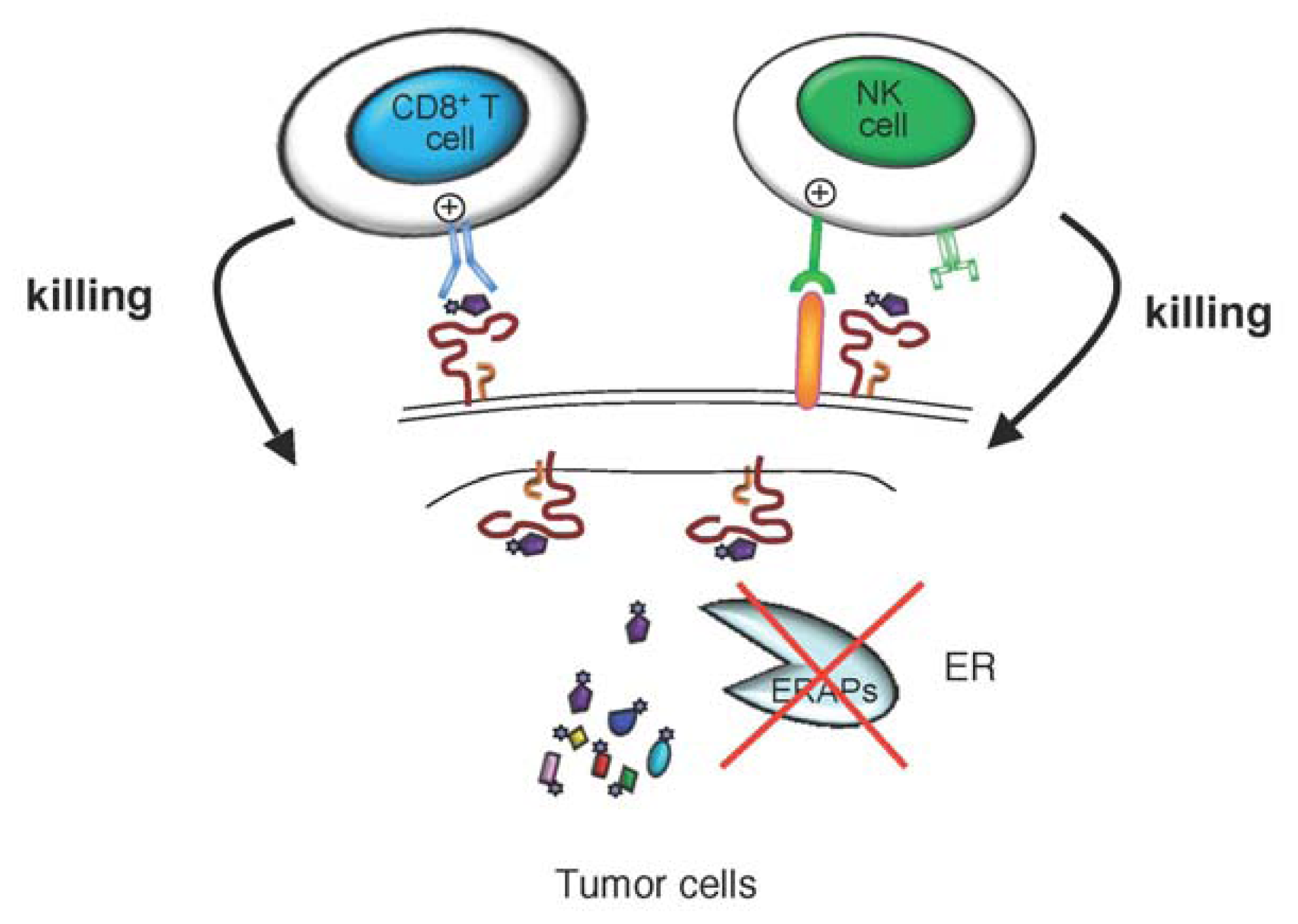
4.4. Cancer
5. Conclusions
Acknowledgments
References
- Hattori, A.; Matsumoto, H.; Mizutani, S.; Tsujimoto, M. Molecular cloning of adipocyte-derived leucine aminopeptidase highly related to placental leucine aminopeptidase/oxytocinase. J. Biochem 1999, 125, 931–938. [Google Scholar]
- Andres, A.M.; Dennis, M.Y.; Kretzschmar, W.W.; Cannons, J.L.; Lee-Lin, S.Q.; Hurle, B.; Schwartzberg, P.L.; Williamson, S.H.; Bustamante, C.D.; Nielsen, R.; et al. Balancing selection maintains a form of ERAP2 that undergoes nonsense-mediated decay and affects antigen presentation. PLoS Genet. 2010, 6, p. e1001157. Available online: http://www.plosgenetics.org/article/info%3Adoi%2F10.1371%2Fjournal.pgen.1001157 accessed on 29 June 2012.
- Saric, T.; Chang, S.C.; Hattori, A.; York, I.A.; Markant, S.; Rock, K.L.; Tsujimoto, M.; Goldberg, A.L. An IFN-gamma-induced aminopeptidase in the ER, ERAP1, trims precursors to MHC class I-presented peptides. Nat. Immunol 2002, 3, 1169–1176. [Google Scholar]
- Shin, E.C.; Seifert, U.; Urban, S.; Truong, K.T.; Feinstone, S.M.; Rice, C.M.; Kloetzel, P.M.; Rehermann, B. Proteasome activator and antigen-processing aminopeptidases are regulated by virus-induced type I interferon in the hepatitis C virus-infected liver. J. Interferon Cytokine Res 2007, 27, 985–990. [Google Scholar]
- Serwold, T.; Gonzalez, F.; Kim, J.; Jacob, R.; Shastri, N. ERAAP customizes peptides for MHC class I molecules in the endoplasmic reticulum. Nature 2002, 419, 480–483. [Google Scholar]
- Tanioka, T.; Hattori, A.; Masuda, S.; Nomura, Y.; Nakayama, H.; Mizutani, S.; Tsujimoto, M. Human leukocyte-derived arginine aminopeptidase. The third member of the oxytocinase subfamily of aminopeptidases. J. Biol. Chem 2003, 278, 32275–32283. [Google Scholar]
- Forloni, M.; Albini, S.; Limongi, M.Z.; Cifaldi, L.; Boldrini, R.; Nicotra, M.R.; Giannini, G.; Natali, P.G.; Giacomini, P.; Fruci, D. NF-kappaB, and not MYCN, regulates MHC class I and endoplasmic reticulum aminopeptidases in human neuroblastoma cells. Cancer Res 2010, 70, 916–924. [Google Scholar]
- York, I.A.; Chang, S.C.; Saric, T.; Keys, J.A.; Favreau, J.M.; Goldberg, A.L.; Rock, K.L. The ER aminopeptidase ERAP1 enhances or limits antigen presentation by trimming epitopes to 8–9 residues. Nat. Immunol 2002, 3, 1177–1184. [Google Scholar]
- Saveanu, L.; Carroll, O.; Lindo, V.; Del Val, M.; Lopez, D.; Lepelletier, Y.; Greer, F.; Schomburg, L.; Fruci, D.; Niedermann, G.; et al. Concerted peptide trimming by human ERAP1 and ERAP2 aminopeptidase complexes in the endoplasmic reticulum. Nat. Immunol 2005, 6, 689–697. [Google Scholar]
- Cui, X.; Hawari, F.; Alsaaty, S.; Lawrence, M.; Combs, C.A.; Geng, W.; Rouhani, F.N.; Miskinis, D.; Levine, S.J. Identification of ARTS-1 as a novel TNFR1-binding protein that promotes TNFR1 ectodomain shedding. J. Clin. Invest 2002, 110, 515–526. [Google Scholar]
- Cui, X.; Rouhani, F.N.; Hawari, F.; Levine, S.J. Shedding of the type II IL-1 decoy receptor requires a multifunctional aminopeptidase, aminopeptidase regulator of TNF receptor type 1 shedding. J. Immunol 2003, 171, 6814–6819. [Google Scholar]
- Cui, X.; Rouhani, F.N.; Hawari, F.; Levine, S.J. An aminopeptidase, ARTS-1, is required for interleukin-6 receptor shedding. J. Biol. Chem 2003, 278, 28677–28685. [Google Scholar]
- Sato, Y. Role of aminopeptidase in angiogenesis. Biol. Pharm. Bull 2004, 27, 772–776. [Google Scholar]
- Hattori, A.; Kitatani, K.; Matsumoto, H.; Miyazawa, S.; Rogi, T.; Tsuruoka, N.; Mizutani, S.; Natori, Y.; Tsujimoto, M. Characterization of recombinant human adipocyte-derived leucine aminopeptidase expressed in Chinese hamster ovary cells. J. Biochem 2000, 128, 755–762. [Google Scholar]
- Neefjes, J.; Jongsma, M.L.; Paul, P.; Bakke, O. Towards a systems understanding of MHC class I and MHC class II antigen presentation. Nat. Rev. Immunol 2011, 11, 823–836. [Google Scholar]
- Kloetzel, P.M. Generation of major histocompatibility complex class I antigens: Functional interplay between proteasomes and TPPII. Nat. Immunol 2004, 5, 661–669. [Google Scholar]
- Reits, E.; Neijssen, J.; Herberts, C.; Benckhuijsen, W.; Janssen, L.; Drijfhout, J.W.; Neefjes, J. A major role for TPPII in trimming proteasomal degradation products for MHC class I antigen presentation. Immunity 2004, 20, 495–506. [Google Scholar]
- Seifert, U.; Maranon, C.; Shmueli, A.; Desoutter, J.F.; Wesoloski, L.; Janek, K.; Henklein, P.; Diescher, S.; Andrieu, M.; de la Salle, H.; et al. An essential role for tripeptidyl peptidase in the generation of an MHC class I epitope. Nat. Immunol 2003, 4, 375–379. [Google Scholar]
- Beninga, J.; Rock, K.L.; Goldberg, A.L. Interferon-gamma can stimulate post-proteasomal trimming of the N terminus of an antigenic peptide by inducing leucine aminopeptidase. J. Biol. Chem 1998, 273, 18734–18742. [Google Scholar]
- Paz, P.; Brouwenstijn, N.; Perry, R.; Shastri, N. Discrete proteolytic intermediates in the MHC class I antigen processing pathway and MHC I-dependent peptide trimming in the ER. Immunity 1999, 11, 241–251. [Google Scholar]
- Stoltze, L.; Nussbaum, A.K.; Sijts, A.; Emmerich, N.P.; Kloetzel, P.M.; Schild, H. The function of the proteasome system in MHC class I antigen processing. Immunol. Today 2000, 21, 317–319. [Google Scholar]
- Stoltze, L.; Schirle, M.; Schwarz, G.; Schroter, C.; Thompson, M.W.; Hersh, L.B.; Kalbacher, H.; Stevanovic, S.; Rammensee, H.G.; Schild, H. Two new proteases in the MHC class I processing pathway. Nat. Immunol 2000, 1, 413–418. [Google Scholar]
- Lauvau, G.; Kakimi, K.; Niedermann, G.; Ostankovitch, M.; Yotnda, P.; Firat, H.; Chisari, F.V.; van Endert, P.M. Human transporters associated with antigen processing (TAPs) select epitope precursor peptides for processing in the endoplasmic reticulum and presentation to T cells. J. Exp. Med 1999, 190, 1227–1240. [Google Scholar]
- Chang, S.C.; Momburg, F.; Bhutani, N.; Goldberg, A.L. The ER aminopeptidase, ERAP1, trims precursors to lengths of MHC class I peptides by a “molecular ruler” mechanism. Proc. Natl. Acad. Sci. USA 2005, 102, 17107–17112. [Google Scholar]
- Serwold, T.; Gaw, S.; Shastri, N. ER aminopeptidases generate a unique pool of peptides for MHC class I molecules. Nat. Immunol 2001, 2, 644–651. [Google Scholar]
- Evnouchidou, I.; Momburg, F.; Papakyriakou, A.; Chroni, A.; Leondiadis, L.; Chang, S.C.; Goldberg, A.L.; Stratikos, E. The internal sequence of the peptide-substrate determines its N-terminus trimming by ERAP1. PLoS One 2008, 3, e3658. [Google Scholar]
- Hammer, G.E.; Gonzalez, F.; James, E.; Nolla, H.; Shastri, N. In the absence of aminopeptidase ERAAP, MHC class I molecules present many unstable and highly immunogenic peptides. Nat. Immunol 2007, 8, 101–108. [Google Scholar]
- Yan, J.; Parekh, V.V.; Mendez-Fernandez, Y.; Olivares-Villagomez, D.; Dragovic, S.; Hill, T.; Roopenian, D.C.; Joyce, S.; van Kaer, L. In vivo role of ER-associated peptidase activity in tailoring peptides for presentation by MHC class Ia and class Ib molecules. J. Exp. Med 2006, 203, 647–659. [Google Scholar]
- Firat, E.; Saveanu, L.; Aichele, P.; Staeheli, P.; Huai, J.; Gaedicke, S.; Nil, A.; Besin, G.; Kanzler, B.; van Endert, P.; et al. The role of endoplasmic reticulum-associated aminopeptidase 1 in immunity to infection and in cross-presentation. J. Immunol 2007, 178, 2241–2248. [Google Scholar]
- York, I.A.; Brehm, M.A.; Zendzian, S.; Towne, C.F.; Rock, K.L. Endoplasmic reticulum aminopeptidase 1 (ERAP1) trims MHC class I-presented peptides in vivo and plays an important role in immunodominance. Proc. Natl. Acad. Sci. USA 2006, 103, 9202–9207. [Google Scholar]
- Hammer, G.E.; Gonzalez, F.; Champsaur, M.; Cado, D.; Shastri, N. The aminopeptidase ERAAP shapes the peptide repertoire displayed by major histocompatibility complex class I molecules. Nat. Immunol 2006, 7, 103–112. [Google Scholar]
- Blanchard, N.; Shastri, N. Coping with loss of perfection in the MHC class I peptide repertoire. Curr. Opin. Immunol 2008, 20, 82–88. [Google Scholar]
- Blanchard, N.; Kanaseki, T.; Escobar, H.; Delebecque, F.; Nagarajan, N.A.; Reyes-Vargas, E.; Crockett, D.K.; Raulet, D.H.; Delgado, J.C.; Shastri, N. Endoplasmic reticulum aminopeptidase associated with antigen processing defines the composition and structure of MHC class I peptide repertoire in normal and virus-infected cells. J. Immunol 2010, 184, 3033–3042. [Google Scholar]
- Nagarajan, N.A.; Gonzalez, F.; Shastri, N. Nonclassical MHC class Ib-restricted cytotoxic T cells monitor antigen processing in the endoplasmic reticulum. Nat. Immunol 2012, 13, 579–586. [Google Scholar]
- Miyashita, H.; Yamazaki, T.; Akada, T.; Niizeki, O.; Ogawa, M.; Nishikawa, S.; Sato, Y. A mouse orthologue of puromycin-insensitive leucyl-specific aminopeptidase is expressed in endothelial cells and plays an important role in angiogenesis. Blood 2002, 99, 3241–3249. [Google Scholar]
- Akada, T.; Yamazaki, T.; Miyashita, H.; Niizeki, O.; Abe, M.; Sato, A.; Satomi, S.; Sato, Y. Puromycin insensitive leucyl-specific aminopeptidase (PILSAP) is involved in the activation of endothelial integrins. J. Cell Physiol 2002, 193, 253–262. [Google Scholar]
- Yamazaki, T.; Akada, T.; Niizeki, O.; Suzuki, T.; Miyashita, H.; Sato, Y. Puromycin-insensitive leucyl-specific aminopeptidase (PILSAP) binds and catalyzes PDK1, allowing VEGF-stimulated activation of S6K for endothelial cell proliferation and angiogenesis. Blood 2004, 104, 2345–52. [Google Scholar]
- Suzuki, T.; Abe, M.; Miyashita, H.; Kobayashi, T.; Sato, Y. Puromycin insensitive leucyl-specific aminopeptidase (PILSAP) affects RhoA activation in endothelial cells. J. Cell Physiol 2007, 211, 708–715. [Google Scholar]
- Yoshida, T.; Sato, Y.; Morita, I.; Abe, M. Pigpen, a nuclear coiled body component protein, is involved in angiogenesis. Cancer Sci 2010, 101, 1170–1176. [Google Scholar]
- Abe, M.; Sato, Y. Puromycin insensitive leucyl-specific aminopeptidase (PILSAP) is required for the development of vascular as well as hematopoietic system in embryoid bodies. Genes. Cells 2006, 11, 719–729. [Google Scholar]
- Fierabracci, A.; Milillo, A.; Locatelli, F.; Fruci, D. The putative role of endoplasmic reticulum aminopeptidases in autoimmunity: Insights from genomic-wide association studies. Autoimmun. Rev 2012, in press. [Google Scholar]
- Taranta, A.; Gianviti, A.; Palma, A.; De Luca, V.; Mannucci, L.; Procaccino, M.A.; Ghiggeri, G.M.; Caridi, G.; Fruci, D. Genetic risk factors in typical haemolytic uraemic syndrome. Nephrol. Dial. Transplant 2009, 24, 1851–1857. [Google Scholar]
- Yamamoto, N.; Nakayama, J.; Yamakawa-Kobayashi, K.; Hamaguchi, H.; Miyazaki, R.; Arinami, T. Identification of 33 polymorphisms in the adipocyte-derived leucine aminopeptidase (ALAP) gene and possible association with hypertension. Hum. Mutat 2002, 19, 251–257. [Google Scholar]
- Goto, Y.; Hattori, A.; Ishii, Y.; Mizutani, S.; Tsujimoto, M. Enzymatic properties of human aminopeptidase A. Regulation of its enzymatic activity by calcium and angiotensin IV. J. Biol. Chem 2006, 281, 23503–23513. [Google Scholar]
- Hallberg, P.; Lind, L.; Michaelsson, K.; Kurland, L.; Kahan, T.; Malmqvist, K.; Ohman, K.P.; Nystrom, F.; Liljedahl, U.; Syvanen, A.C.; et al. Adipocyte-derived leucine aminopeptidase genotype and response to antihypertensive therapy. BMC Cardiovasc. Disord 2003, 3, 11. [Google Scholar]
- Hill, L.D.; Hilliard, D.D.; York, T.P.; Srinivas, S.; Kusanovic, J.P.; Gomez, R.; Elovitz, M.A.; Romero, R.; Strauss, J.F. Fetal ERAP2 variation is associated with preeclampsia in African Americans in a case-control study. BMC Med. Genet 2011, 12, 64. [Google Scholar]
- Johnson, M.P.; Roten, L.T.; Dyer, T.D.; East, C.E.; Forsmo, S.; Blangero, J.; Brennecke, S.P.; Austgulen, R.; Moses, E.K. The ERAP2 gene is associated with preeclampsia in Australian and Norwegian populations. Hum. Genet 2009, 126, 655–666. [Google Scholar]
- Founds, S.A.; Conley, Y.P.; Lyons-Weiler, J.F.; Jeyabalan, A.; Hogge, W.A.; Conrad, K.P. Altered global gene expression in first trimester placentas of women destined to develop preeclampsia. Placenta 2009, 30, 15–24. [Google Scholar]
- Blanchard, N.; Gonzalez, F.; Schaeffer, M.; Joncker, N.T.; Cheng, T.; Shastri, A.J.; Robey, E.A.; Shastri, N. Immunodominant, protective response to the parasite Toxoplasma gondii requires antigen processing in the endoplasmic reticulum. Nat. Immunol 2008, 9, 937–944. [Google Scholar]
- Draenert, R.; Le Gall, S.; Pfafferott, K.J.; Leslie, A.J.; Chetty, P.; Brander, C.; Holmes, E.C.; Chang, S.C.; Feeney, M.E.; Addo, M.M.; et al. Immune selection for altered antigen processing leads to cytotoxic T lymphocyte escape in chronic HIV-1 infection. J. Exp. Med 2004, 199, 905–915. [Google Scholar]
- Tenzer, S.; Wee, E.; Burgevin, A.; Stewart-Jones, G.; Friis, L.; Lamberth, K.; Chang, C.H.; Harndahl, M.; Weimershaus, M.; Gerstoft, J. Antigen processing influences HIV-specific cytotoxic T lymphocyte immunodominance. Nat. Immunol 2009, 10, 636–646. [Google Scholar]
- Cagliani, R.; Riva, S.; Biasin, M.; Fumagalli, M.; Pozzoli, U.; Lo Caputo, S.; Mazzotta, F.; Piacentini, L.; Bresolin, N.; Clerici, M.; et al. Genetic diversity at endoplasmic reticulum aminopeptidases is maintained by balancing selection and is associated with natural resistance to HIV-1 infection. Hum. Mol. Genet 2010, 19, 4705–4714. [Google Scholar]
- Mehta, A.M.; Jordanova, E.S.; Corver, W.E.; van Wezel, T.; Uh, H.W.; Kenter, G.G.; Fleuren, G.J. Single nucleotide polymorphisms in antigen processing machinery component ERAP1 significantly associate with clinical outcome in cervical carcinoma. Genes. Chromosomes Cancer 2009, 48, 410–418. [Google Scholar]
- Kim, S.; Lee, S.; Shin, J.; Kim, Y.; Evnouchidou, I.; Kim, D.; Kim, Y.K.; Kim, Y.E.; Ahn, J.H.; Riddell, S.R.; et al. Human cytomegalovirus microRNA miR-US4–1 inhibits CD8(+) T cell responses by targeting the aminopeptidase ERAP1. Nat. Immunol 2011, 12, 984–991. [Google Scholar]
- Burton, P.R.; Clayton, D.G.; Cardon, L.R.; Craddock, N.; Deloukas, P.; Duncanson, A.; Kwiatkowski, D.P.; McCarthy, M.I.; Ouwehand, W.H.; Samani, N.J.; et al. Association scan of 14,500 nonsynonymous SNPs in four diseases identifies autoimmunity variants. Nat. Genet 2007, 39, 1329–1337. [Google Scholar]
- Maksymowych, W.P.; Inman, R.D.; Gladman, D.D.; Reeve, J.P.; Pope, A.; Rahman, P. Association of a specific ERAP1/ARTS1 haplotype with disease susceptibility in ankylosing spondylitis. Arthritis Rheum 2009, 60, 1317–1323. [Google Scholar]
- Davidson, S.I.; Wu, X.; Liu, Y.; Wei, M.; Danoy, P.A.; Thomas, G.; Cai, Q.; Sun, L.; Duncan, E.; Wang, N. Association of ERAP1, but not IL23R, with ankylosing spondylitis in a Han Chinese population. Arthritis Rheum 2009, 60, 3263–3268. [Google Scholar]
- Choi, C.B.; Kim, T.H.; Jun, J.B.; Lee, H.S.; Shim, S.C.; Lee, B.; Pope, A.; Uddin, M.; Rahman, P.; Inman, R.D. ARTS1 polymorphisms are associated with ankylosing spondylitis in Koreans. Ann. Rheum. Dis 2010, 69, 582–584. [Google Scholar]
- Harvey, D.; Pointon, J.J.; Evans, D.M.; Karaderi, T.; Farrar, C.; Appleton, L.H.; Sturrock, R.D.; Stone, M.A.; Oppermann, U.; Brown, M.A.; Wordsworth, B.P.; et al. Investigating the genetic association between ERAP1 and ankylosing spondylitis. Hum. Mol. Genet 2009, 18, 4204–4212. [Google Scholar]
- Evans, D.M.; Spencer, C.C.; Pointon, J.J.; Su, Z.; Harvey, D.; Kochan, G.; Oppermann, U.; Dilthey, A.; Pirinen, M.; Stone, M.A. Interaction between ERAP1 and HLA-B27 in ankylosing spondylitis implicates peptide handling in the mechanism for HLA-B27 in disease susceptibility. Nat. Genet 2011, 43, 761–767. [Google Scholar]
- Pazar, B.; Safrany, E.; Gergely, P.; Szanto, S.; Szekanecz, Z.; Poor, G. Association of ARTS1 gene polymorphisms with ankylosing spondylitis in the Hungarian population: the rs27044 variant is associated with HLA-B*2705 subtype in Hungarian patients with ankylosing spondylitis. J. Rheumatol 2010, 37, 379–384. [Google Scholar]
- Evnouchidou, I.; Kamal, R.P.; Seregin, S.S.; Goto, Y.; Tsujimoto, M.; Hattori, A.; Voulgari, P.V.; Drosos, A.A.; Amalfitano, A.; York, I.A.; et al. Cutting Edge: Coding single nucleotide polymorphisms of endoplasmic reticulum aminopeptidase 1 can affect antigenic peptide generation in vitro by influencing basic enzymatic properties of the enzyme. J. Immunol 2011, 186, 1909–1913. [Google Scholar]
- Fung, E.Y.; Smyth, D.J.; Howson, J.M.; Cooper, J.D.; Walker, N.M.; Stevens, H.; Wicker, L.S.; Todd, J.A. Analysis of 17 autoimmune disease-associated variants in type 1 diabetes identifies 6q23/TNFAIP3 as a susceptibility locus. Genes. Immun 2009, 10, 188–191. [Google Scholar]
- Guerini, F.R.; Cagliani, R.; Forni, D.; Agliardi, C.; Caputo, D.; Cassinotti, A.; Galimberti, D.; Fenoglio, C.; Biasin, M.; Asselta, R.; et al. A functional variant in ERAP1 predisposes to multiple sclerosis. PLoS One 2012, 7, e29931. [Google Scholar]
- Strange, A.; Capon, F.; Spencer, C.C.; Knight, J.; Weale, M.E.; Allen, M.H.; Barton, A.; Band, G.; Bellenguez, C.; Bergboer, J.G.; et al. A genome-wide association study identifies new psoriasis susceptibility loci and an interaction between HLA-C and ERAP1. Nat. Genet 2010, 42, 985–990. [Google Scholar]
- Sun, L.D.; Cheng, H.; Wang, Z.X.; Zhang, A.P.; Wang, P.G.; Xu, J.H.; Zhu, Q.X.; Zhou, H.S.; Ellinghaus, E.; Zhang, F.R.; et al. Association analyses identify six new psoriasis susceptibility loci in the Chinese population. Nat. Genet 2010, 42, 1005–1009. [Google Scholar]
- Franke, A.; McGovern, D.P.; Barrett, J.C.; Wang, K.; Radford-Smith, G.L.; Ahmad, T.; Lees, C.W.; Balschun, T.; Lee, J.; Roberts, R.; et al. Genome-wide meta-analysis increases to 71 the number of confirmed Crohn’s disease susceptibility loci. Nat. Genet 2010, 42, 1118–1125. [Google Scholar]
- Fruci, D.; Ferracuti, S.; Limongi, M.Z.; Cunsolo, V.; Giorda, E.; Fraioli, R.; Sibilio, L.; Carroll, O.; Hattori, A.; van Endert, P.M.; et al. Expression of endoplasmic reticulum aminopeptidases in EBV-B cell lines from healthy donors and in leukemia/lymphoma, carcinoma, and melanoma cell lines. J. Immunol 2006, 176, 4869–4879. [Google Scholar]
- Fruci, D.; Giacomini, P.; Nicotra, M.R.; Forloni, M.; Fraioli, R.; Saveanu, L.; van Endert, P.; Natali, P.G. Altered expression of endoplasmic reticulum aminopeptidases ERAP1 and ERAP2 in transformed non-lymphoid human tissues. J. Cell Physiol 2008, 216, 742–749. [Google Scholar]
- Kamphausen, E.; Kellert, C.; Abbas, T.; Akkad, N.; Tenzer, S.; Pawelec, G.; Schild, H.; van Endert, P.; Seliger, B. Distinct molecular mechanisms leading to deficient expression of ER-resident aminopeptidases in melanoma. Cancer Immunol. Immunother 2010, 59, 1273–1284. [Google Scholar]
- Kazeto, H.; Nomura, S.; Ito, N.; Ito, T.; Watanabe, Y.; Kajiyama, H.; Shibata, K.; Ino, K.; Tamakoshi, K.; Hattori, A.; et al. Expression of adipocyte-derived leucine aminopeptidase in endometrial cancer. Association with tumor grade and CA-125. Tumour. Biol 2003, 24, 203–208. [Google Scholar]
- Watanabe, Y.; Shibata, K.; Kikkawa, F.; Kajiyama, H.; Ino, K.; Hattori, A.; Tsujimoto, M.; Mizutani, S. Adipocyte-derived leucine aminopeptidase suppresses angiogenesis in human endometrial carcinoma via renin-angiotensin system. Clin. Cancer Res 2003, 9, 6497–6503. [Google Scholar]
- Shibata, K.; Kikkawa, F.; Mizokami, Y.; Kajiyama, H.; Ino, K.; Nomura, S.; Mizutani, S. Possible involvement of adipocyte-derived leucine aminopeptidase via angiotensin II in endometrial carcinoma. Tumour. Biol 2005, 26, 9–16. [Google Scholar]
- Mehta, A.M.; Jordanova, E.S.; Kenter, G.G.; Ferrone, S.; Fleuren, G.J. Association of antigen processing machinery and HLA class I defects with clinicopathological outcome in cervical carcinoma. Cancer Immunol. Immunother 2008, 57, 197–206. [Google Scholar]
- Cifaldi, L.; Lo Monaco, E.; Forloni, M.; Giorda, E.; Lorenzi, S.; Petrini, S.; Tremante, E.; Pende, D.; Locatelli, F.; Giacomini, P.; et al. Natural killer cells efficiently reject lymphoma silenced for the endoplasmic reticulum aminopeptidase associated with antigen processing. Cancer Res 2011, 71, 1597–1606. [Google Scholar]
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Cifaldi, L.; Romania, P.; Lorenzi, S.; Locatelli, F.; Fruci, D. Role of Endoplasmic Reticulum Aminopeptidases in Health and Disease: from Infection to Cancer. Int. J. Mol. Sci. 2012, 13, 8338-8352. https://doi.org/10.3390/ijms13078338
Cifaldi L, Romania P, Lorenzi S, Locatelli F, Fruci D. Role of Endoplasmic Reticulum Aminopeptidases in Health and Disease: from Infection to Cancer. International Journal of Molecular Sciences. 2012; 13(7):8338-8352. https://doi.org/10.3390/ijms13078338
Chicago/Turabian StyleCifaldi, Loredana, Paolo Romania, Silvia Lorenzi, Franco Locatelli, and Doriana Fruci. 2012. "Role of Endoplasmic Reticulum Aminopeptidases in Health and Disease: from Infection to Cancer" International Journal of Molecular Sciences 13, no. 7: 8338-8352. https://doi.org/10.3390/ijms13078338