Development and Characterization of New Single Nucleotide Polymorphism Markers from Expressed Sequence Tags in Common Carp (Cyprinus carpio)
Abstract
:1. Introduction
2. Results and Discussion
3. Experimental Section
3.1. Detection and Annotation of Putative SNPs
3.2. Validation and Characterization of SNPs
3.3. Data Analysis
4. Conclusions
Supplementary Materials
Acknowledgements
References
- Wohlfarth, G.W. Heterosis for growth rate in common carp. Aquaculture 1993, 113, 31–46. [Google Scholar]
- Food and Agriculture Organization of the United Nations Fisheries and Aquaculture Department. Fishery and Aquaculture Statistics, 2009; FAO: Rome, Italy, 2011. Available online: www.fao.org/fishery/statistics/global-aquaculture-production/en accessed on 25 May 2012.
- Crooijmans, R.; Bierbooms, V.; Komen, J.; Van, D.P.; Groenen, M. Microsatellite markers in common carp (Cyprinus carpio L.). Anim. Genet 1997, 28, 129–134. [Google Scholar]
- Wang, D.; Liao, X.; Cheng, L.; Yu, X.; Tong, J. Development of novel EST-SSR markers in common carp by data mining from public EST sequences. Aquaculture 2007, 271, 558–574. [Google Scholar]
- Yue, G.H.; Ho, M.Y.; Orban, L.; Komen, J. Microsatellites within genes and ESTs of common carp and their applicability in silver crucian carp. Aquaculture 2004, 234, 85–98. [Google Scholar]
- Liu, Z.J.; Cordes, J.F. DNA marker technologies and their applications in aquaculture genetics. Aquaculture 2004, 238, 1–37. [Google Scholar]
- Garvin, M.R.; Saitoh, K.; Gharrett, A.J. Application of single nucleotide polymorphisms to non-model species: A technical review. Mol. Ecol. Resour 2010, 10, 915–934. [Google Scholar]
- Picoult-Newberg, L.; Ideker, T.E.; Pohl, M.G.; Taylor, S.L.; Donaldson, M.A.; Nickerson, D.A.; Boyce-Jacino1, M. Mining SNPs from EST databases. Genome Res 1999, 9, 167–174. [Google Scholar]
- Cenadelli, S.; Maran, V.; Bongioni, G.; Fusetti, L.; Parma, P.; Aleandri, R. Identification of nuclear SNPs in gilthead seabream. J. Fish Biol 2007, 70, 399–405. [Google Scholar]
- Hayes, B.; Laerdahl, J.K.; Lien, S.; Moen, T.; Berg, P.; Hindar, K.; Davidson, W.S.; Koop, B.F.; Adzhubei, A.; Hoyheim, B. An extensive resource of single nucleotide polymorphism markers associated with Atlantic salmon (Salmo salar) expressed sequences. Aquaculture 2007, 265, 82–90. [Google Scholar]
- Hubert, S.; Bussey, J.T.; Higgins, B.; Curtis, B.A.; Bowman, S. Development of single nucleotide polymorphism markers for Atlantic cod (Gadus morhua) using expressed sequences. Aquaculture 2009, 296, 7–14. [Google Scholar]
- Vera, M.; Álvarez-Dios, J.A.; Millán, A.; Pardo, B.G.; Bouza, C.; Hermida, M.; Fernández, C.; Herrán, R.; Molina-Luzón, M.J.; Martínez, P. Validation of single nucleotide polymorphism (SNP) markers from an immune expressed sequence tag (EST) turbot, Scophthalmus maximus, database. Aquaculture 2011, 313, 31–41. [Google Scholar]
- Zheng, X.; Kuang, Y.; Zhang, X.; Lu, C.; Cao, D.; Li, C.; Sun, X. A genetic linkage map and comparative genome analysis of common carp (Cyprinus carpio L.) using microsatellites and SNPs. Mol. Genet. Genomics 2011. [Google Scholar] [CrossRef]
- Nicod, J.C.; Largiadèr, C.R. SNPs by AFLP (SBA): A rapid SNP isolation strategy for non-model organisms. Nucleic Acids Res 2003, 31. [Google Scholar] [CrossRef]
- Smith, C.T.; Elfstrom, C.M.; Seeb, L.W.; Seeb, J.E. Use of sequence data from rainbow trout and Atlantic salmon for SNP detection in Pacific salmon. Mol. Ecol 2005, 14, 4193–4203. [Google Scholar]
- He, C.; Chen, L.; Simmons, M.; Li, P.; Kim, S.; Liu, Z.J. Putative SNP discovery in interspecific hybrids of catfish by comparative EST analysis. Anim. Genet 2003, 34, 445–448. [Google Scholar]
- Larhammar, D.; Risinger, C. Molecular genetic aspects of tetraploidy in the common carp Cyprinus carpio. Mol. Phylogenet. Evol 1994, 3, 59–68. [Google Scholar]
- David, L.; Blum, S.; Feldman, M.W.; Lavi, U.; Hillel, J. Recent duplication of the common carp (Cyprinus carpio L.) genome as revealed by analyses of microsatellite loci. Mol. Biolo. Evol 2003, 20, 1425–1434. [Google Scholar]
- Stickney, H.L.; Schmutz, J.; Woods, I.G.; Holtzer, C.C.; Dickson, M.C.; Kelly, P.D.; Myers, R.M.; Talbot, W.S. Rapid mapping of zebrafish mutations with SNPs and oligonucleotide microarrays. Genome Res 2002, 12, 1929–1934. [Google Scholar]
- Andolfatto, P. Adaptive evolution of non-coding DNA in Drosophila. Nature 2005, 437, 1149–1152. [Google Scholar]
- Lynch, M.; Koskella, B.; Schaack, S. Mutation pressure and the evolution of organelle genomic architecture. Science 2006, 311, 1727–1730. [Google Scholar]
- Pedretti, K.; Scheetz, T.; Braun, T.; Roberts, C.; Robinson, N.; Casavant, T. A Parallel Expressed Sequence Tag (EST) Clustering Program. In Proceedings of the 6th International Conference on Parallel Computing Technologies; Malyshkin, V., Ed.; Springer-Verlag Berlin: Novosibirsk, Russia, 2001; pp. 490–497. [Google Scholar]
- Gordon, D.; Abajian, C.; Green, P. Consed: A graphical tool for sequence finishing. Genome Res 1998, 8, 195–202. [Google Scholar]
- Barker, G.; Batley, J.; O’Sullivan, H.; Edwards, K.J.; Edwards, D. Redundancy based detection of sequence polymorphisms in expressed sequence tag data using autoSNP. Bioinformatics 2003, 19, 421–422. [Google Scholar]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory: Cold Spring Harbor, NY, USA, 2001; pp. 463–465. [Google Scholar]
- Gallagher, P.M.; Lowe, G.; Fitzgerald, T.; Bella, A.; Greene, C.M.; McElvaney, N.G.; O’Neill, S.J. Association of IL-10 polymorphism with severity of illness in community acquired pneumonia. Thorax 2003, 58, 154–156. [Google Scholar]
- Gaudet, M.; Fara, A.-G.; Sabatti, M.; Kuzminsky, E.; Mugnozza, G.S. Single-reaction for SNP genotyping on agarose gel by allele-specific PCR in black poplar (Populus nigra L.). Plant Mol. Biol. Rep 2007, 25, 1–9. [Google Scholar]
- Thompson, J.; Gibson, T.; Plewniak, F.; Jeanmougin, F.; Higgins, D. The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 1997, 25, 4876–4882. [Google Scholar]
- Yeh, F.C.; Boyle, T. POPGENE: Microsoft Window-based Freeware for Population Genetic Analysis, version 1.31; University of Alberta: Edmonton, Canada, 1999. [Google Scholar]
- Park, S.D.E. Trypanotolerance in West African Cattle and the Population Genetic Effects of Selection. Ph.D. Dissertation, University of Dublin, Dublin, Ireland, 2001. [Google Scholar]
- Excoffier, L.; Laval, G.; Schneider, S. ARLEQUIN: An integrated software package for population genetics data analysis. Version 3. Evol. Bioinform. Online 2005, 1, 47–50. [Google Scholar]
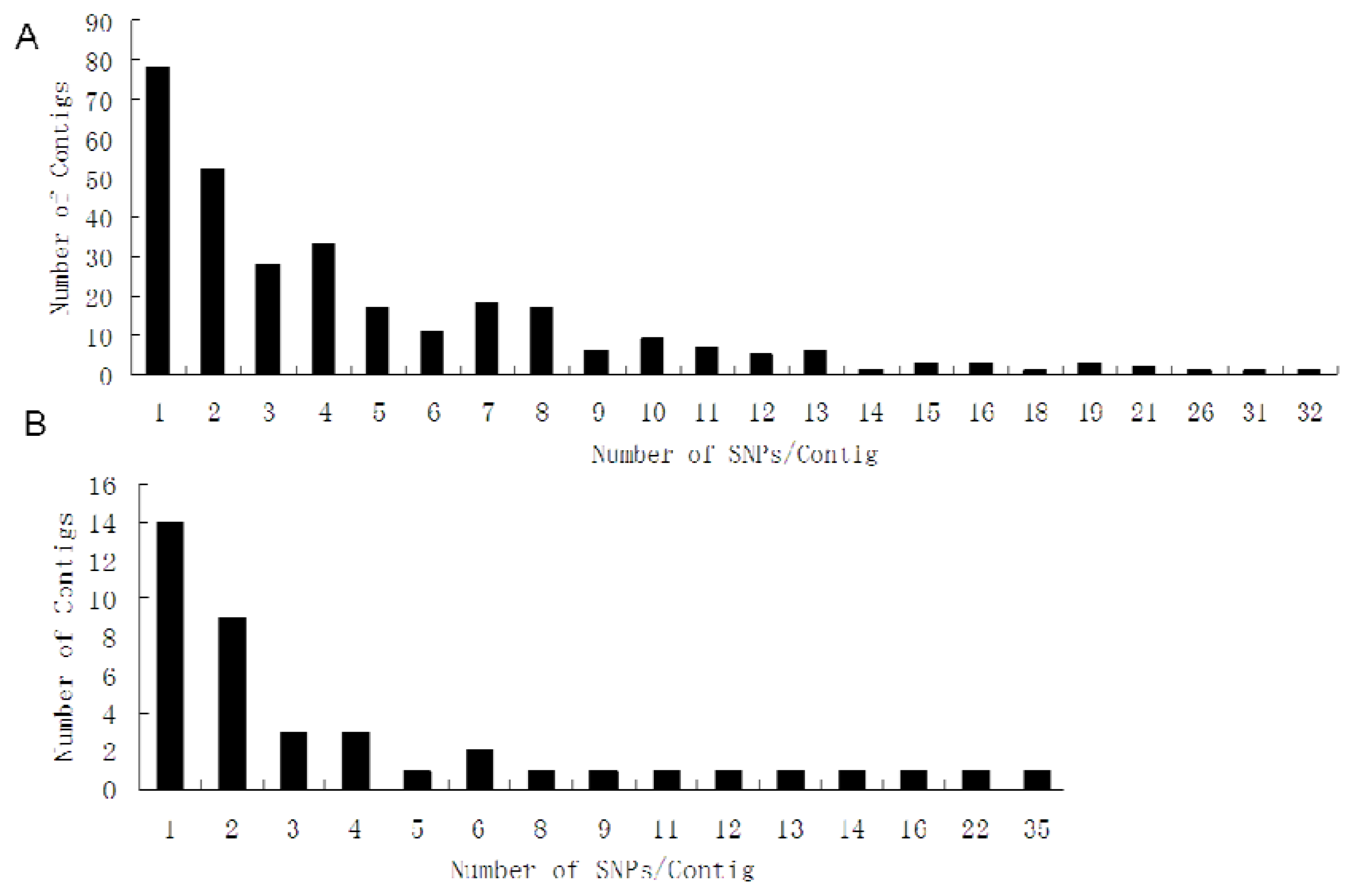
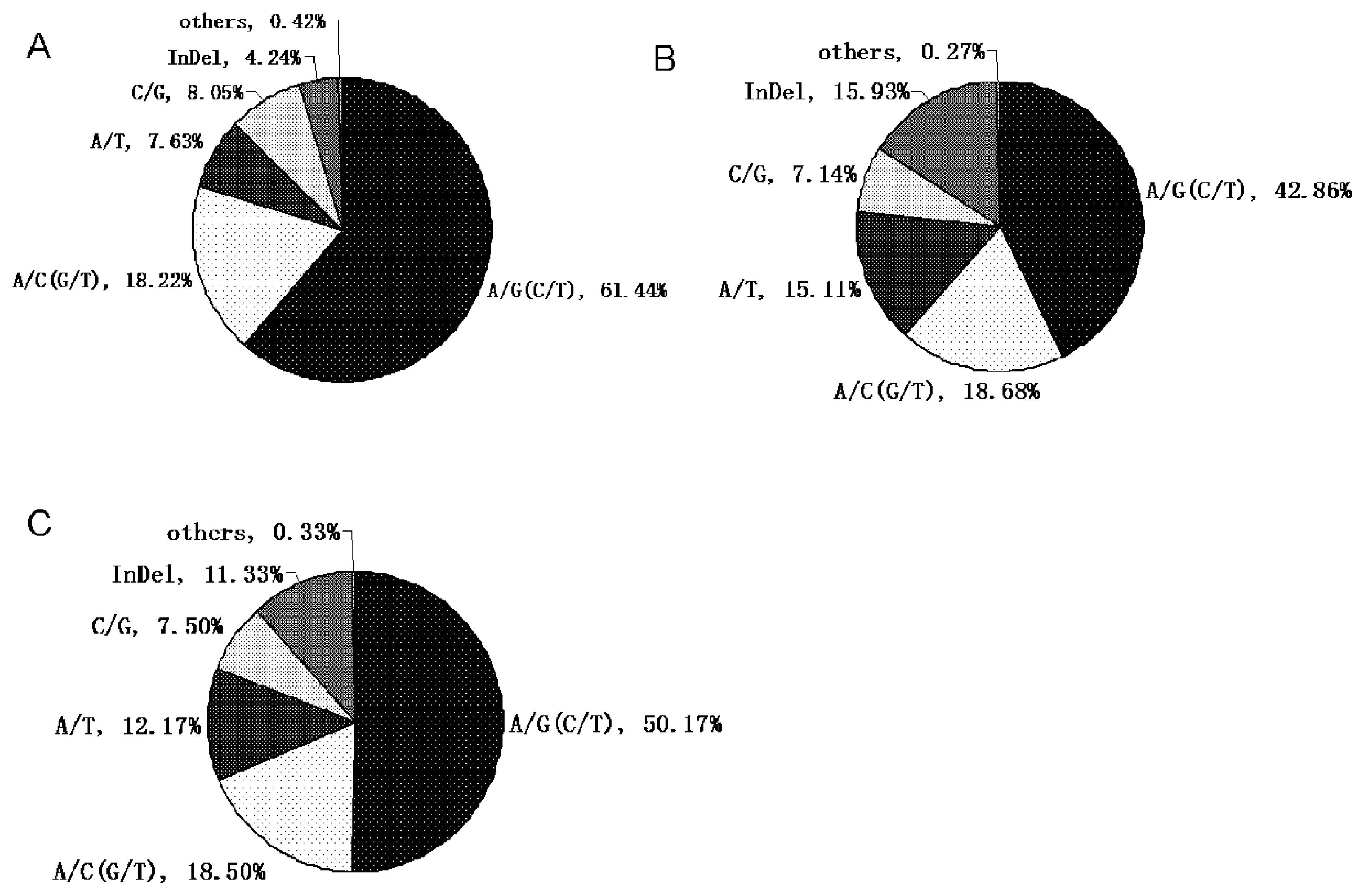
| Cluster ID | Primers designed | Length (bp, anticipative/actual) | Number of SNPs (putative/validated) | New SNPs in exon/intron | Annotation |
|---|---|---|---|---|---|
| Cyprinus_Cluster7873.seq.Contig1 | 7873-1 | 180/388 | 1/0 | 14/32(5) a | extracellular space |
| 7873-2 | 196/313 | 1/0 | 4/0 | ||
| Cyprinus_Cluster7877.seq.Contig1 | 7877 | 163/250 | 6/1 | 0/0 | malate dehydrogenase (oxaloacetate-decarboxylating) (NADP+) activity |
| Cyprinus_Cluster7885.seq.Contig1 | 7885 | 215/215 | 1/0 | 1(2)/− | actin binding |
| Cyprinus_Cluster7889.seq.Contig1 | 7889 | 319/822 | 1/0 | 0/0 | cornified envelope |
| Cyprinus_Cluster7892.seq.Contig1 | 7892 | 253/377 | 4/1 | 0/0 | structural constituent of ribosome |
| Cyprinus_Cluster7895.seq.Contig1 | 7895-2 | 157/954 | 1/0 | 2/11(5) | cerebroside-sulfatase activity |
| Cyprinus_Cluster7896.seq.Contig1 | 7896 | 448/450 | 1/0 | 1/− | nucleus |
| Cyprinus_Cluster7898.seq.Contig1 | 7898 | 238/238 | 0/0 | 13/− | creatine kinase activity |
| Cyprinus_Cluster7909.seq.Contig1 | 7909 | 533/533 | 2/0 | 13/− | N.A. b |
| Cyprinus_Cluster7918.seq.Contig1 | 7918 | 278/550 | 1/1 | 1/0 | cathepsin H activity |
| Cyprinus_Cluster7921.seq.Contig1 | 7921 | 164/319 | 1/0 | 1/2 | N.A. |
| Cyprinus_Cluster7927.seq.Contig1 | 7927 | 322/322 | 1/1 | 1/− | N.A. |
| Cyprinus_Cluster7929.seq.Contig1 | 7929 | 222/1433 | 1/0 | 1/13(3) | 3-hydroxyanthranilate 3,4-dioxygenase activity |
| Cyprinus_Cluster7933.seq.Contig1 | 7933 | 429/1016 | 1/1 | 34(1)/55(18) | glutathione transferase activity |
| Cyprinus_Cluster7943.seq.Contig1 | 7943 | 402/405 | 4/0 | 1(2)/− | N.A. |
| Cyprinus_Cluster7944.seq.Contig1 | 7944 | 298/298 | 4/0 | 5/− | GTPase activity |
| Cyprinus_Cluster7947.seq.Contig1 | 7947 | 236/419 | 2/1 | 2/3 | lysozyme activity |
| Cyprinus_Cluster7953.seq.Contig1 | 7953 | 347/347 | 1/1 | 7/− | serine-type endopeptidase inhibitor activity |
| Cyprinus_Cluster7955.seq.Contig1 | 7955 | 240/357 | 3/0 | 0/0 | actin binding |
| Cyprinus_Cluster7957.seq.Contig1 | 7957 | 354/1428 | 5/0 | 2/17(2) | fructose-bisphosphate ldolase activity |
| Cyprinus_Cluster7961.seq.Contig1 | 7961 | 222/418 | 2/1 | 21/26(5) | external side of plasma membrane |
| Cyprinus_Cluster7969.seq.Contig1 | 7969 | 217/216 | 4/4 | 7/− | antigen binding |
| Cyprinus_Cluster7974.seq.Contig1 | 7974 | 215/443 | 6/0 | 1/5 | N.A. |
| Cyprinus_Cluster7980.seq.Contig1 | 7980-2 | 211/529 | 7/0 | 1/13(1) | l-lactate dehydrogenase activity |
| Cyprinus_Cluster7986.seq.Contig1 | 7986-2 | 191/404 | 1/0 | 4/17 | binding |
| Cyprinus_Cluster7988.seq.Contig1 | 7988 | 184/324 | 1/0 | 16/30(2) | receptor activity |
| Cyprinus_Cluster7990.seq.Contig1 | 7990 | 273/1301 | 2/1 | (2)/3 | ubiquitin-protein ligase activity |
| Cyprinus_Cluster7992.seq.Contig1 | 7992 | 139/139 | 1/0 | 1/− | adenyl-nucleotide exchange factor activity |
| Cyprinus_Cluster8000.seq.Contig1 | 8000 | 419/709 | 2/0 | 2/3 | glucose-6-phosphatase activity |
| Cyprinus_Cluster8001.seq.Contig1 | 8001 | 376/1412 | 2/1 | 2/4(2) | N.A. |
| Cyprinus_Cluster8008.seq.Contig1 | 8008 | 233/1715 | 2/1 | 0/0 | integral to membrane |
| Cyprinus_Cluster8009.seq.Contig1 | 8009-2 | 213/412 | 3/1 | 3/3 | calcium ion binding |
| Cyprinus_Cluster8011.seq.Contig1 | 8011 | 303/302 | 2/0 | 2/− | thyroxine 5′-deiodinase activity |
| Cyprinus_Cluster8012.seq.Contig1 | 8012 | 203/203 | 2/0 | 1(1)/0 | N.A. |
| Cyprinus_Cluster8013.seq.Contig1 | 8013 | 260/257 | 5/1 | 1/0 | N.A. |
| Cyprinus_Cluster8017.seq.Contig1 | 8017 | 313/1384 | 3/0 | (1)/6 | regulation of progression through cell cycle |
| Cyprinus_Cluster8021.seq.Contig1 | 8021 | 447/656 | 8/0 | 6/1 | N.A. |
| Cyprinus_Cluster8025.seq.Contig1 | 8025 | 318/875 | 2/2 | 0/7 | steroid binding |
| Cyprinus_Cluster8034.seq.Contig1 | 8034 | 327/327 | 2/0 | 2/− | bisphosphoglycerate mutase activity |
| Cyprinus_Cluster8041.seq.Contig1 | 8041 | 173/329 | 1/0 | 0/2 | cytoplasm |
| Cyprinus_Cluster8048.seq.Contig1 | 8048 | 256/989 | 2/0 | 1/0 | dolichyl-diphosphooligosaccharide-protein glycotransferase activity |
| Cyprinus_Cluster8050.seq.Contig1 | 8050-1 | 179/179 | 1/0 | 6(1)/− | signal transducer activity |
| Cyprinus_Cluster8052.seq.Contig2 | 8052cg2 | 286/792 | 1/1 | 13/53(15) | l-lactate dehydrogenase activity |
| Cyprinus_Cluster8123.seq.Contig1 | 8123 | 428/428 | 7/3 | 6/0 | mitochondrion |
| Cyprinus_Cluster8142.seq.Contig1 | 8142 | 420/420 | 7/3 | 1/0 | N.A. |
| Cyprinus_Cluster8184.seq.Contig1 | 8184 | 413/413 | 3/0 | 12/0 | N.A. |
| Total | 47 | 13,213/27,010 | 121/26 | 202(10)/306(58) | - |
| Transitions | Transversions | Multiple | |||||
|---|---|---|---|---|---|---|---|
| A/G | C/T | A/C | A/T | G/C | GT | ||
| In exons | 64 | 81 | 27 | 18 | 19 | 18 | 1C/G/T |
| In introns | 88 | 68 | 29 | 55 | 26 | 39 | 1A/G/T |
| In complete sequences | 152 | 149 | 56 | 73 | 45 | 55 | 2 |
| Loci | Allele Frequencies | He | Ho | PIC | p-value |
|---|---|---|---|---|---|
| CC a 7892G > A | 0.355(A)/0.645(G) | 0.464 | 0.342 | 0.403 | 0.15446 |
| CC7953C > G | 0.263(G)/0.737(C) | 0.393 | 0.526 | 0.313 | 0.04006 |
| CC7943G > A | 0.316(A)/0.684(G) | 0.438 | 0.474 | 0.339 | 0.71678 |
| CC7909-1G > A | 0.355(A)/0.645(G) | 0.464 | 0.500 | 0.353 | 0.73065 |
| CC7909-2T > C | 0.145(C)/0.855(T) | 0.251 | 0.237 | 0.217 | 0.57020 |
| CC7909-3A > G | 0.25(G)/0.75(A) | 0.380 | 0.395 | 0.305 | 1.00000 |
| CC7909-4A > T | 0.145(T)/0.855(A) | 0.251 | 0.237 | 0.217 | 0.56984 |
| CC7909-5T > C | 0.145(C)/0.856(T) | 0.251 | 0.237 | 0.217 | 0.56926 |
| CC7909-6G > C | 0.145(C)/0.857(G) | 0.251 | 0.237 | 0.217 | 0.56908 |
| CC7909-7T > A | 0.092(A)/0.908(T) | 0.169 | 0.079 | 0.153 | 0.01852 |
| CC7909-8A > G | 0.079(G)/0.921(A) | 0.147 | 0.105 | 0.135 | 0.19207 |
| CC7909-9T > A | 0.066(A)/0.934(T) | 0.125 | 0.079 | 0.115 | 0.13050 |
| CC7909-10T > C | 0.224(C)/0.776(T) | 0.352 | 0.184 | 0.287 | 0.00768 |
| CC7909-11T > C | 0.092(C)/0.908(T) | 0.169 | 0.079 | 0.153 | 0.01862 |
| CC7909-12T > A | 0.092(A)/0.909(T) | 0.169 | 0.079 | 0.153 | 0.01853 |
| CC7909-13T > C | 0.368(C)/0.632(T) | 0.472 | 0.158 | 0.357 | 0.00003 |
| CC7969-1G > T | 0.053(T)/0.947(G) | 0.101 | 0.053 | 0.095 | 0.07912 |
| CC7969-2C > A | 0.290(A)/0.710(C) | 0.417 | 0.158 | 0.327 | 0.00030 |
| CC7969-3C > G | 0.111(G)/0.889(C) | 0.200 | 0.111 | 0.178 | 0.04048 |
| CC7969-4A > G | 0.167(G)/0.833(A) | 0.282 | 0.333 | 0.239 | 0.55952 |
| CC7969-5A > C | 0.236(C)/0.764(A) | 0.366 | 0.472 | 0.296 | 0.15560 |
| CC7969-6A > G | 0.236(G)/0.764(A) | 0.366 | 0.472 | 0.296 | 0.15479 |
| CC7969-7T > G | 0.236(G)/0.764(T) | 0.366 | 0.472 | 0.296 | 0.15639 |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhu, C.; Cheng, L.; Tong, J.; Yu, X. Development and Characterization of New Single Nucleotide Polymorphism Markers from Expressed Sequence Tags in Common Carp (Cyprinus carpio). Int. J. Mol. Sci. 2012, 13, 7343-7353. https://doi.org/10.3390/ijms13067343
Zhu C, Cheng L, Tong J, Yu X. Development and Characterization of New Single Nucleotide Polymorphism Markers from Expressed Sequence Tags in Common Carp (Cyprinus carpio). International Journal of Molecular Sciences. 2012; 13(6):7343-7353. https://doi.org/10.3390/ijms13067343
Chicago/Turabian StyleZhu, Chuankun, Lei Cheng, Jingou Tong, and Xiaomu Yu. 2012. "Development and Characterization of New Single Nucleotide Polymorphism Markers from Expressed Sequence Tags in Common Carp (Cyprinus carpio)" International Journal of Molecular Sciences 13, no. 6: 7343-7353. https://doi.org/10.3390/ijms13067343





