1. Introduction
Rice researchers are currently focused on enhancing rice quality as well as improving yield. In Malaysia, 33 modern varieties of rice have been released for commercial planting, including waxy, non-waxy, aromatic and elongating varieties [
1]. Nevertheless, the most important components of rice quality are related to its cooking and eating qualities. Amylose content is considered to be the key determinant of the processing, cooking and eating characteristics of rice (
Oryza sativa L.), and it correlates directly with the volume expansion, water absorption and ultimate firmness of cooked rice [
2]. Generally, the amylose content of milled rice is categorized into five classes: waxy (0–2%), very low amylose (3–9%), low amylose (10–19%), intermediate amylose (20–24%) and high amylose (above 24%). Low amylose content is associated with cohesive, tender and glossy cooked rice. In contrast, high amylose content is associated with dry, firm, fluffy and well separated grains of cooked rice [
3]. Lower amylose rice is preferable to higher amylose rice (above 20%) because it does not become hard and dry when cooked.
Previous genetic studies have found that amylose content is governed mainly by an allelic series of genes at one locus and by one or several modifier genes with minor effects [
4]. However, its inheritance pattern is complex due to cytoplasmic effects, epistasis and the triploid nature of the endosperm [
5], as well as environmental effects [
6]. Environmental effects were found to have caused variations of up to 6% in amylose content for a given cultivar [
7]. Higher levels of amylose are reportedly controlled by either partial or complete dominance. Thus, heterozygotes cannot be identified using phenotypic measurements of amylose content, and this often leads to inefficiency in traditional rice breeding [
5,
8]. In the study conducted by Chen
et al. [
9], higher air temperature during grain development was found to be associated with a decrease in apparent amylose content of low and intermediate amylose content rice, but with an increase in apparent amylose content of high amylose content rice.
Over the past several decades, various methods have been reported for the determination of amylose content, including iodine binding, near infrared spectroscopy, size-exclusion chromatography and most recently, asymmetric flow field flow fractionation. However, none of these methods have been validated for routine use, and different values of amylose content have been reported using different methods for the same varieties [
10]. The utilization of microsatellites could overcome the shortcomings of amylose content measurements due to their co-dominant nature and their ability to reveal high levels of allelic diversity. Furthermore, microsatellites are easily assayed by polymerase chain reaction (PCR) and are amenable to marker-assisted selection (MAS).
Amylose synthesis in the rice kernel is catalyzed by the granule-bound starch synthase (GBSS) encoded by the
Wx gene on chromosome 6. There are two functional alleles present in the
Wx locus, namely
wxa and
wxb, which are found mainly in the indica and japonica varieties, respectively. These alleles produce different levels of
Wx protein in the endosperm, and consequently cause variation in amylose content. The
wxa allele synthesizes higher contents of GBSS and thus the grains have higher apparent amylose content than those expressing the
wxb allele [
11]. Previous studies have identified several nucleotide polymorphisms that are associated with the
Wx gene, including a polymorphic microsatellite (CT)
n and a G–T SNP located at the 5′-leader intron splice site [
12,
13]. Another two SNPs are present in exon 6 [A-C SNP] and exon 10 [C-T SNP], which exhibit a high correlation of amylose level with
Wx microsatellite and G-T SNPs [
14,
15].
The levels of waxy protein and amylose content were observed to be correlated with the excision of the leader intron of the
Wx transcripts. High amylose varieties contain high amounts of completely processed GBSS mRNA, while lower amylose varieties contain varying ratios of both completely processed GBSS and partially processed transcripts containing the leader intron [
12,
13]. The G to T mutation at the 5′-leader intron splice site was reported to be responsible for the characteristics of the
wxb allele; having low level of mature
Wx transcript, GBSS and apparent amylose content [
12,
15,
16]. Ayres
et al. confirmed the observation that this single-base change could interfere with mRNA processing in 89 US non-waxy varieties; all 30 low amylose varieties tested had the sequence AG
TTATA while 59 intermediate and high amylose varieties had AG
GTATA. They reported that the G-T mutation could explain 79.7% of the variation in the amylose content of the non-waxy varieties. Previous study has reported that the AG
GTATA and AG
TTATA sequences at the 5′-leader intron splice site coincided with the presence of the
wxa allele and
wxb allele, respectively [
16]. Various studies have shown similar results in which low amylose varieties had the AG
TTATA sequence, while the intermediate and high amylose varieties had the AG
GTATA sequence [
12,
17–
21].
Nevertheless, the G-T SNP alone is not sufficient to explain all of the observed genetic variations in amylose content among rice varieties. As an additional explanation on the variation in amylose content, Larkin and Park reported a sequence change in exon 6 of the
Wx gene, comprised of an A-C polymorphism, which resulted in an amino acid substitution and this was associated with intermediate apparent amylose content rice. This A-C SNP was genotyped together with the G-T SNP in the study conducted by Chen
et al. using 171 rice accessions, together explaining 86.7% of the variation in apparent amylose content, which characterized rice into low, intermediate and high apparent amylose content classes. Another identified SNP in the
Wx gene was found to be located in exon 10 and comprised of a C-T polymorphism. The C to T substitution was identified in high apparent amylose rice [
14]. All these SNPs could be used as tools to discriminate the different classes of amylose and thereby useful in breeding programs to develop rice varieties with desirable amylose levels.
The present study sought to analyze the Wx microsatellite and G-T SNPs at the 5′-leader intron splice site in 15 waxy and non-waxy rice varieties with a wide range of amylose contents. This is an essential first step towards the development of high quality local rice varieties with intermediate amylose contents using molecular approaches. Further, this study aimed to optimize a simple and reliable electrophoresis method, MAGE, which has not been extensively used for the detection of microsatellite alleles, particularly the Wx alleles.
3. Discussion
Previous studies have reported that a polymorphism in the
Wx microsatellite located 55-bp upstream of the putative 5′-leader intron splice site in the
Wx gene was responsible for more than 82% of the variation in the amylose content of non-waxy rice [
4,
14]. This indicates that the
Wx microsatellite can be used to distinguish most rice varieties with different amylose contents.
PAGE method and its limitations: The most common electrophoresis method employed in previous studies has been PAGE, due to its ability to separate many samples simultaneously with scoring accuracy [
4,
14,
21]. However, PAGE is technically challenging and the process can be cumbersome and time-consuming. Polyacrylamide gels are thin and fragile, and the glass plates used in casting them are difficult to handle. We worked on both 5% (data not shown) and 8% polyacrylamide gels and we found that 5% polyacrylamide gels are rather crumbly and easily broken. Besides, acrylamide is carcinogenic and is known to be a cumulative neurotoxin.
The advantages of MAGE over PAGE: In this study, MAGE was used instead of PAGE due to its ability to produce thicker and more stable gels and the fact that it uses a standard agarose gel electrophoresis system. We found these gels easier to prepare and free from any leakage problems during gel casting. This method was approximately 4 h shorter than the PAGE method employed by Jayamani
et al. [
21], and like PAGE, it is able to separate many samples simultaneously with good scoring accuracy. According to Ochsenreither
et al. [
22], PAGE and MAGE are both currently used for microsatellite analysis and they produce comparable and reproducible results. In our results, however, MAGE gives clearer single bands for the
Wx microsatellites with no shadow bands as compared with PAGE.
A total of 16
Wx microsatellite alleles, ranging in length from (CT)
4 to (CT)
22, have been found to be correlated with amylose content [
4,
14,
21,
23,
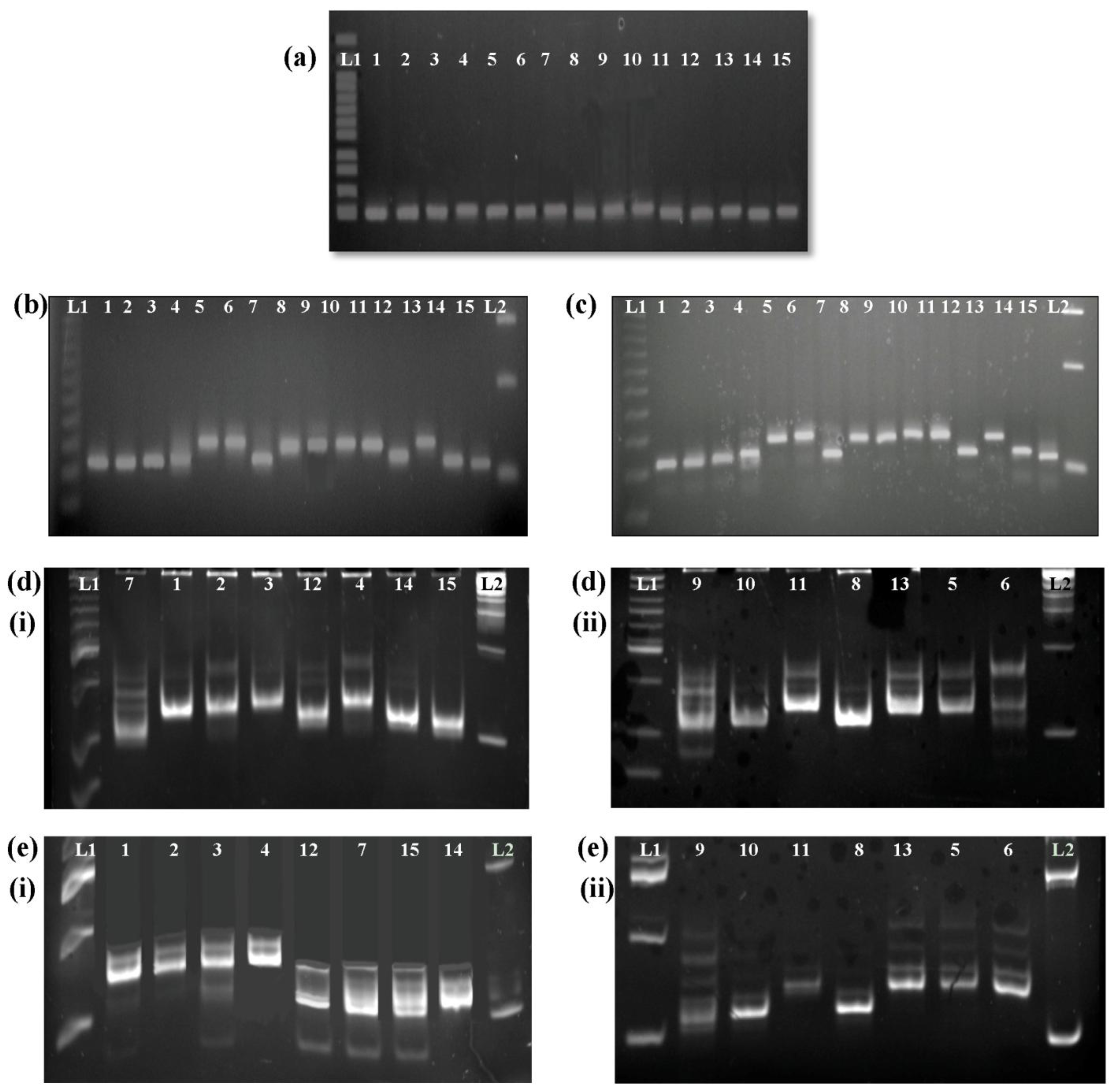
24]. In the present study, fifteen varieties that are commonly used in local breeding programs were characterized using
Wx microsatellite and G-T SNP analysis. Five microsatellite alleles of the
Wx gene were identified, namely (CT)
10, (CT)
11, (CT)
16, (CT)
17, and (CT)
18. All of the selected varieties with high amylose content (above 24%) were associated with shorter repeat alleles, namely (CT)
10 and (CT)
11, while longer repeat alleles, including (CT)
16, (CT)
17 and (CT)
18, were detected in the selected varieties with low and intermediate amylose contents (below 24%).
Of the five microsatellite alleles detected, (CT)
11 and (CT)
17 were found in the most tested varieties. Studies conducted by Ayres
et al. and Bergman
et al. found that varieties with the (CT)
11 allele had an amylose content of greater than 23%, consistent with the results obtained in this study, in which all varieties with the (CT)
11 allele were high amylose varieties with an apparent amylose content greater than 24%. According to Ayres
et al., the (CT)
17 allele is present in varieties with divergent amylose contents. In this study, the (CT)
17 allele was present in a waxy variety, Pulut Hitam 9, which has an amylose content of 0.34%, but it was also found in three varieties with apparent amylose contents of 14 to 19.6%, including MR263 and both the internationally accepted high quality varieties (Khaw Dawk Mali and Basmati 370). Our results were similar to those of previous studies. The (CT)
17 allele was reported to be prominent in the Thai rice germplasm; where 36 out of 68 Thai rice strains tested contained this allele, including Khaw Dawk Mali 105 [
25]. The (CT)
17 allele was also present in all eight Basmati accessions tested by Jayamani
et al. We also found that the
Wx microsatellites could not always be used to distinguish the waxy varieties, as concluded by Ayres
et al. The two waxy varieties tested, Pulut Siding and Pulut Hitam 9, had different alleles, (CT)
16 and (CT)
17, respectively. However, the waxy varieties were found to have larger repeat alleles, (CT)
n,
n > 15, as did other low amylose varieties.
To study the correlation between the
Wx microsatellite allele and the G-T SNP, we sequenced the region containing the microsatellite from all selected varieties representing all five detected microsatellite classes. The present results were consistent with those of previous studies, which found that all intermediate and high amylose varieties had the sequence AGGTATA, while waxy and low-amylose varieties had the sequence AGTTATA [
4,
15,
17,
19,
21,
25]. We found that all varieties with the (CT)
10, (CT)
11 and (CT)
18 alleles had the sequence AGGTATA, whereas all varieties with (CT)
16 had the sequence AG
TTATA. (CT)
17 was the only allele that could be associated with either the AG
GTATA or AG
TTATA sequences. The presence of both sequences in the (CT)
17 was also observed by Jayamani
et al. [
21] and Chen
et al. [
26].
The sequencing results of the G-T SNP at the 5′-leader intron splice site were further confirmed using the PCR-
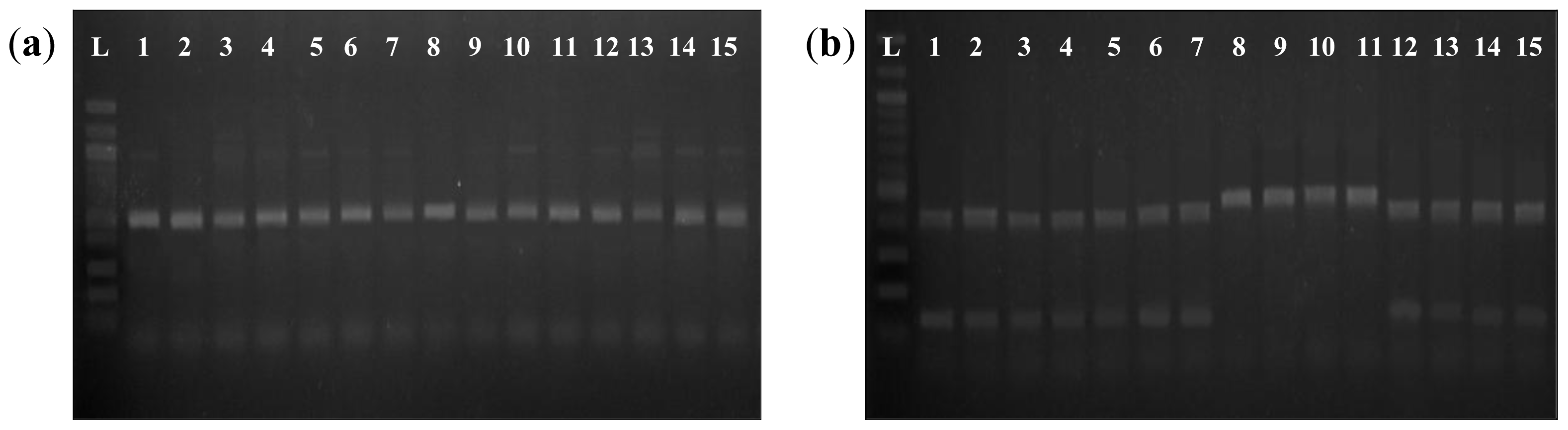
AccI CAPS method. In all cases, amplified fragments that contained the sequence AG
GTATA in the splice site were cleaved by
AccI, resulting in two fragments of the expected size on a 1.5% agarose gel, whereas amplified fragments containing the sequence AG
TTATA were not cleaved by
AccI and remained as a single fragment (
Figure 2). We noted that the PCR-
AccI CAPS method is effective in the detection of the G-T SNP and took less time than the sequencing reactions to discriminate waxy and low amylose varieties from varieties intermediate and high amylose varieties. G-T SNP analysis has been applied in breeding programs to produce indica varieties with lower amylose contents [
6]. Ayres
et al. reported that the G-T SNP could explain approximately 80% of the variation in amylose content in non-waxy rice varieties. In this study, the G-T SNP could distinguish varieties with 18% or less amylose from those with higher than 18% amylose with 100% accuracy. However, this G-T SNP could not be used to discriminate between intermediate and high amylose varieties.
Based on the present study, the combination of Wx microsatellite allele and G-T SNP identification could be used to differentiate high amylose (above 24%), intermediate amylose (20–24%) and low amylose or waxy varieties (below 19%). The Wx microsatellite allele could be used to discriminate high amylose varieties from intermediate and low amylose varieties. All of the selected varieties with high amylose contents had shorter repeat alleles, i.e., (CT)n; n = 10 and 11, whereas the selected varieties with 24% or less amylose contents had longer repeat alleles, i.e., (CT)n; n = 16, 17 and 18. Meanwhile, the G-T SNP could be used to discriminate intermediate and high amylose varieties from waxy and low amylose varieties. Hence, the Wx microsatellite and G-T SNP analysis can be used together as tools to develop rice varieties with the desired amylose levels through breeding work supported by MAS.
4. Experimental Section
Fifteen varieties were used in the study, including two internationally accepted quality rice varieties: Basmati 370 and Khaw Dawk Mali. Seeds were obtained from the Malaysian Agricultural Research and Development Institute (MARDI). The selected varieties were classified as waxy (0–2%); low amylose (below 19%); intermediate amylose (20–24%) and high amylose (above 24%) based on the data provided by MARDI (
Table 1). Apparent amylose content was estimated using the method described by Juliano (1971).The apparent amylose content (%) was estimated according to Juliano [
27]. The seeds were sown and grown in the greenhouse and leaves of 2- to 4-week-old seedlings were harvested for DNA isolation. Genomic DNA was extracted using the cetyltrimethyl ammonium bromide (CTAB) method described by Murray and Thompson [
28] with minor modifications. The quality of DNA samples was examined using agarose gel electrophoresis.
For the
Wx microsatellite assay, PCR was performed in 25 μL reaction mixtures, containing the respective 0.25 μg of template DNA, 1× Green GoTaq flexi buffer, 1.5 mM MgCl
2, 0.2 mM deoxynucleotides, 0.5 μM oligo 484 (forward primer: 5′-CTTTGTCTATCTCAAGACAC-3′), 0.5 μM oligo 485 (reverse primer: 5′-TTGCAGATGTTCTTCCTGATG-3′) [
4] and 2.5 units of GoTaq DNA polymerase (Promega, Madison, WI). The targeted fragments were amplified using a Mastercycler Gradient (Eppendorf, Hamburg) with the following steps: Denaturation at 95 °C for 4 min, followed by 35 cycles of 94 °C for 45 s, 55 °C for 30 s and 72 °C for 60 s, and 5 min at 72 °C for the final extension. After PCR, 5 μL of the PCR products was evaluated on a standard 1% agarose gel to confirm successful amplification. The remaining 20 μL was separated on 4% MetaPhor agarose gels and 8% polyacrylamide gels.
For the 4% MetaPhor gel preparation, 2% MetaPhor agarose (Lonza, Walkersville, MD, USA) and 2% standard agarose (First Base, Singapore, SG) were slowly sprinkled into 1× TAE buffer with rapid stirring. The agarose was soaked in the buffer for 15 min before heating to prevent the solution from foaming during heating. The solution was heated in a microwave at medium power for 2 min before reheating at high power for 1 min. Once the molten gel was cast and solidified, it was placed at 4 °C for 20 min prior to use for optimal resolution. The Wx microsatellite PCR products were electrophoresed for 2 or 4 h at 80 V on ice and stained with an ethidium bromide solution for 10 min followed by 20 min of destaining with water. The gels were observed under UV transillumination (Alpha-Imager). The size of the amplification products was estimated by comparisons to 25-bp and 100-bp DNA ladders (New England Biolabs, Ipswich, MA, USA).
PAGE protocol: For the 8% polyacrylamide gel preparation, 2.4 mL of Acrylamide/Bis 19:1 40% (Ambion, Woodward Austin, TX) were added to 2.4 mL of 5× TAE buffer and 7.2 mL of distilled water to give a total volume of 12 mL. Two hundred microliters of 10% ammonium persulfate (Sigma, Saint Louis, MO, USA) and 10 μL of N,N,N′,N′-tetramethylethylenediamine (Sigma, Saint Louis, MO, USA) were then added to the mixture to complete the gel before the acrylamide polymerized. Once the molten gel was cast and solidified, it was placed at 4 °C until needed prior to use for optimal resolution. The Wx microsatellite PCR products were electrophoresed for 2 or 4 h at 80 V and stained with an ethidium bromide solution for 5 min followed by 15 min of destaining with 1× TAE buffer. The gels were observed under UV transillumination (Alpha-Imager). The size of the amplification products was estimated by comparisons to 25-bp and 100-bp DNA ladders (New England Biolabs, Ipswich, MA, USA).
For sequencing analysis, the same PCR reaction conditions were used to obtain a larger amplified fragment from a subset of all tested varieties, using oligo 484 and W2R (reverse primer: 5′-TTTCCA GCCCAACACCTTAC-3′) [
4]. The PCR products were electrophoresed on a 1.5% agarose gel and the resulting bands were excised. The amplified products were purified using a QIAquick Gel Extraction Kit (Qiagen, Valencia, CA) according to the manufacturer’s protocol and were sequenced (Macrogen, Rockville, MD). To further confirm the presence or absence of the G-T polymorphism at the 5′-leader intron splice site, 10 μL of each amplified product obtained from PCR-
AccI CAPS (forward primer: 5′-ACCATTCCTTCAGTTCTTTG-3′; reverse primer: 5′-ATGATTTAACGAGAGTTGAA-3′) [
29] was digested with 2 units of
AccI (New England Biolabs, Ipswich, MA, USA) in a total volume of 15 μL for 2 h at 37 °C. The samples were electrophoresed for 1.5 h at 80 V in a standard 1.5% agarose gel.