Isolation and Expression of Glucosinolate Synthesis Genes CYP83A1 and CYP83B1 in Pak Choi (Brassica rapa L. ssp. chinensis var. communis (N. Tsen & S.H. Lee) Hanelt)
Abstract
:1. Introduction
2. Results and Discussion
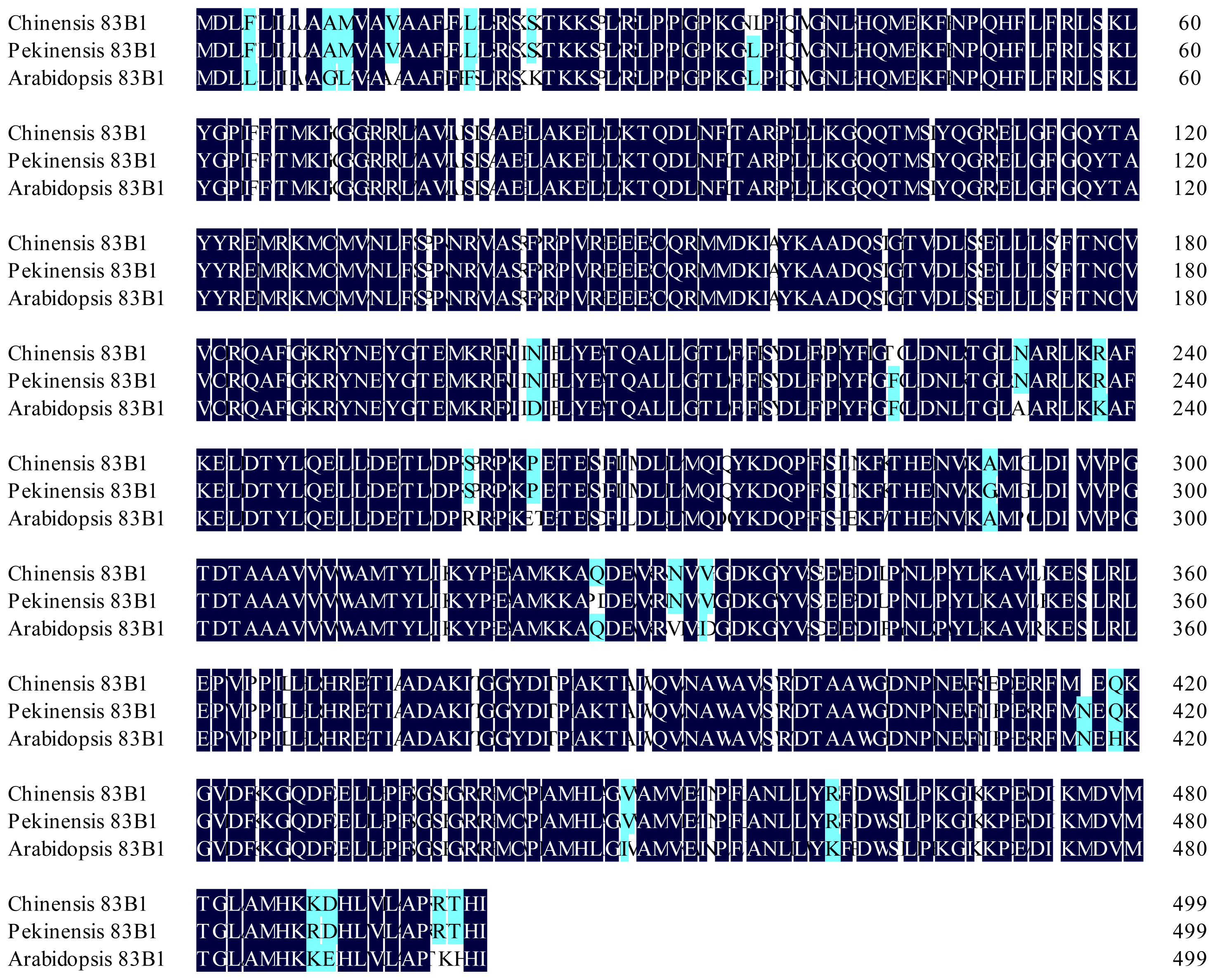
2.1. Identification and Sequence Analysis of CYP83A1 and CYP83B1
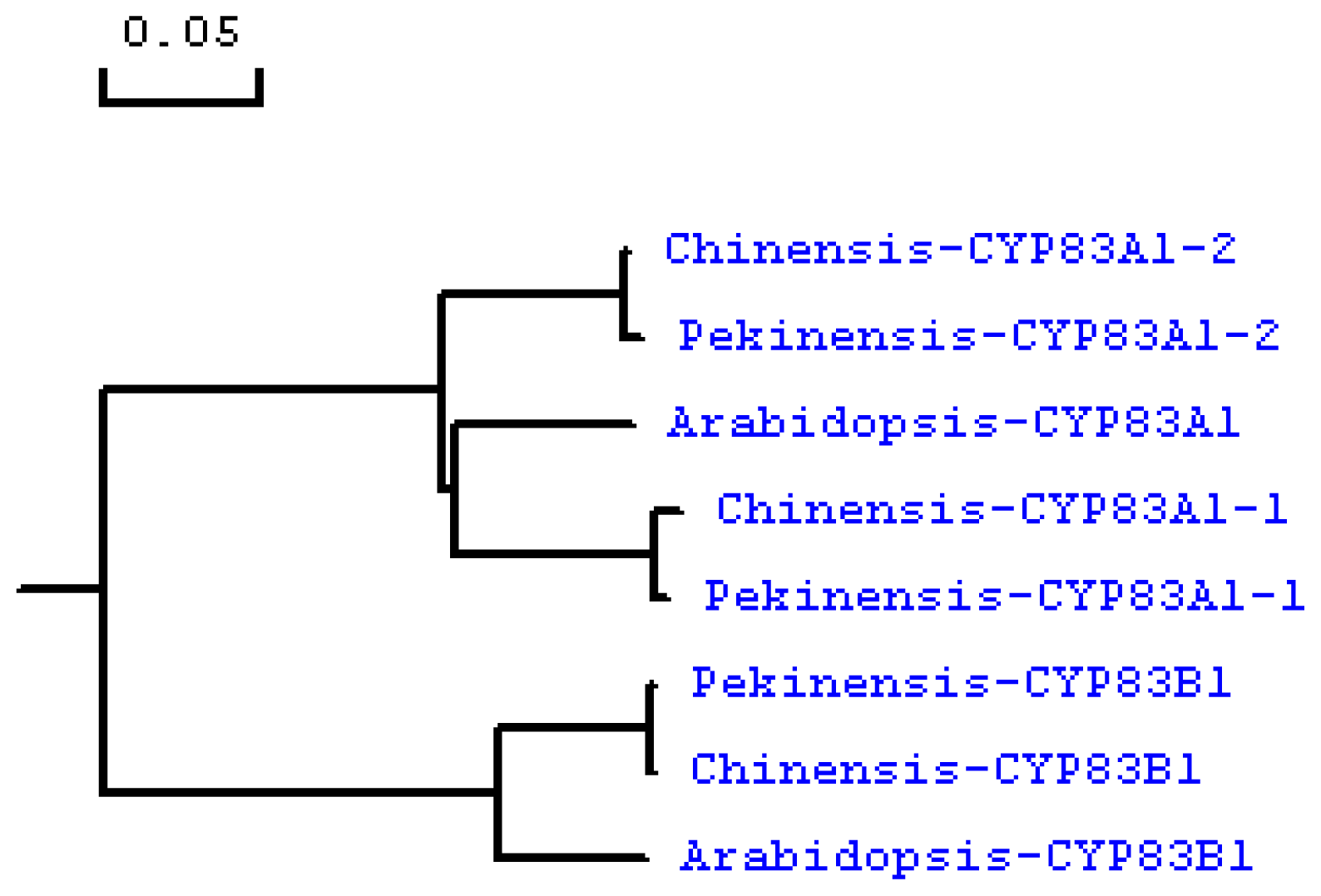
2.2. Phylogenetic Analysis of CYP83A1 and CYP83B1
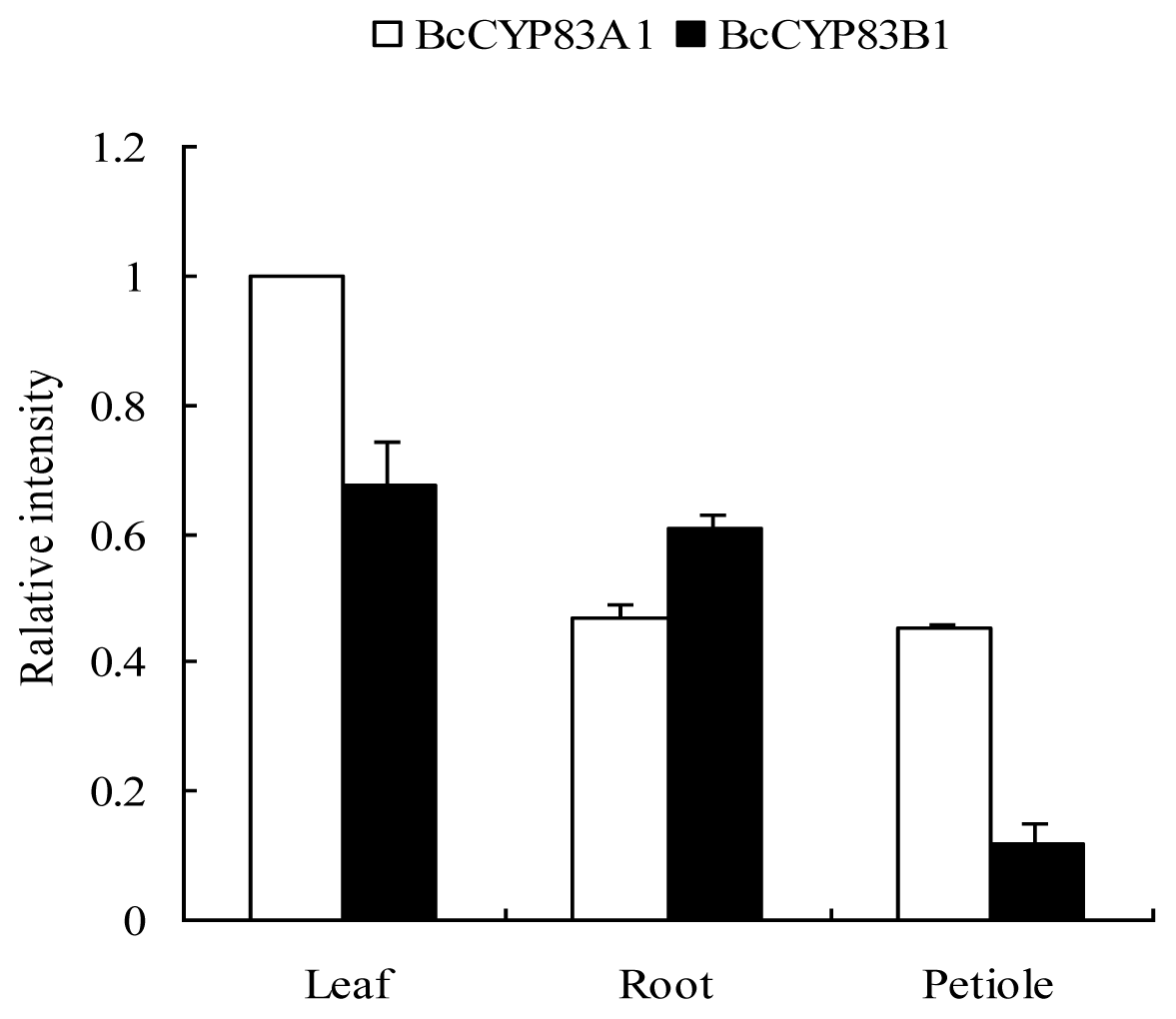
2.3. Expression of CYP83A1 and CYP83B1 in Different Cultivars and Various Organs
2.4. Glucosinolate Profile Concentrations in Different Cultivars and Various Organs
2.5. Discussion
3. Experimental Section
3.1. Plant Materials
3.2. Isolation of Total RNA and Synthesis of Full-Length cDNA Sequence
3.3. Phylogenetic Analysis
3.4. Quantitative Real-Time RT-PCR
| CYP83A1 |
| (F) 5′-TCCTCTCCTTATCCCTCGTGCTTGC-3′ |
| (R) 5′-CCAAGACGCATTCCAGGGCACATTC-3′ |
| CYP83B1 |
| (F) 5′-AGACTCTTGACCCTAGCCGTCCTA-3′ |
| (R) 5′-CCTTTGTCACCGACCACATTCCT-3′ |
| Actin |
| (F) 5′-CGCTTAACCCGAAAGCTAAC-3′ |
| (R) 5′-TACGCCCACTAGCGTAAAGA-3′ |
3.5. Glucosinolate Analysis
4. Conclusions
Acknowledgments
References
- Tay, D.C.S.; Toxopeus, H. Brassica rapa L. cv. Group Pak Choi. In Plant Resources of South-East Asia 8, Vegetables; Siemonsma, J.S., Piluek, K., Eds.; Pudoc Scientific Publishers: Wageningen, The Netherlands, 1993; pp. 130–134. [Google Scholar]
- He, H.J.; Fingerling, G.; Schnitzler, W.H. Seasonal variations of the glucosinolate amounts in Brassica campestris L. spp. chinensis. J. Appl. Bot 2000, 74, 198–202. [Google Scholar]
- Lewis, L.; Fenwick, G.R. Glucosinolate content of brassica vegetables: Analysis of twenty-four cultivars of calabrese (green sprouting broccoli, Brassica oleracea L. var. botrytis subvar. cymosa Lam.). Food Chem 1987, 25, 259–268. [Google Scholar]
- Kurilich, A.C.; Tsau, G.J.; Brown, A.; Howard, L.; Klein, B.P.; Jeffery, E.H.; Kushad, M.; Wallig, M.A.; Juvik, J.A. Carotene, tocopherol and ascorbate contents in subspecies of Brassica oleracea. J. Agric. Food Chem 1999, 47, 1576–1581. [Google Scholar]
- Mithen, R.F.; Dekker, M.; Verkerk, R.; Rabot, S.; Johnson, I.T. The nutritional significance, biosynthesis and bioavailability of glucosinolates in human foods. J. Sci. Food Agric 2000, 80, 967–984. [Google Scholar]
- Manici, L.M.; Lazzeri, L.; Palmieri, S. In vitro fungitoxicactivity of some glucosinolates and their enzyme-derived products towards plant pathogenic fungi. J. Agric. Food Chem 1997, 45, 2768–2773. [Google Scholar]
- Wittstock, U.; Gershenzon, J. Constitutive plant toxins and their role in defense against herbivores and pathogens. Curr. Opin. Plant Biol 2002, 5, 300–307. [Google Scholar]
- Wittstock, U.; Kliebenstein, D.J.; Lambrix, V.; Reichelt, M.; Gershenzon, J. Glucosinolate Hydrolysis and Its Impact on Generalist and Specialist Insect Herbivores. In Integrative Phytochemistry: From Ethnobotany to Molecular Ecology; Romeo, J.T., Ed.; Pergamon: Amsterdam, The Netherlands, 2003; pp. 101–125. [Google Scholar]
- Agerbirk, N.; Olsen, C.E. Glucosinolate structures in evolution. Phytochemistry 2012, 77, 16–45. [Google Scholar]
- Wittstock, U.; Halkier, B.A. Glucosinolate research in the Arabidopsis era. Trends Plant Sci 2002, 7, 263–270. [Google Scholar]
- Bak, S.; Feyereisen, R. The involvement of two P450 enzymes, CYP83B1 and CYP83A1, in auxin homeostasis and glucosinolate biosynthesis. Plant Physiol 2001, 127, 108–118. [Google Scholar]
- Barlier, I.; Kowalczyk, M.; Marchant, A.; Ljung, K.; Bhalerao, R.; Bennett, M.; Sandberg, G.; Bellini, C. The SUR2 gene of Arabidopsis thaliana encodes the cytochrome P450 CYP83B1, a modulator of auxin homeostasis. Proc. Natl. Acad. Sci. USA 2000, 97, 14819–14824. [Google Scholar]
- Delarue, M.; Prinsen, E.; van Onckelen, H.; Caboche, M.; Bellini, C. Sur2 mutations of Arabidopsis thaliana define a new locus involved in the control of auxin homeostasis. Plant J 1998, 14, 603–611. [Google Scholar]
- Grubb, C.D.; Abel, S. Glucosinolate metabolism and its control. Trends Plant Sci 2006, 11, 89–100. [Google Scholar]
- Halkier, B.A.; Gershenzon, J. Biology and biochemistry of glucosinolates. Ann. Rev. Plant Biol 2006, 57, 303–333. [Google Scholar]
- Hansen, C.H.; Du, L.C.; Naur, P.; Olsen, C.E.; Axelsen, K.B.; Hick, A.J.; Pickett, J.A.; Halkier, B.A. CYP83B1 is the oxime-metabolizing enzyme in the glucosinolate pathway in Arabidopsis. J. Biol. Chem 2001, 276, 24790–24796. [Google Scholar]
- Piotrowski, M.; Schemenewitz, A.; Lopukhinat, A.; Mueller, A.; Janowitz, T.; Weiler, E.W.; Oecking, C. Desulfoglucosinolate sulfotransferases from Arabidopsis thaliana catalyze the final step in the biosynthesis of the glucosinolate core structure. J. Biol. Chem 2004, 279, 50717–50725. [Google Scholar]
- Smolen, G.; Bender, J. Arabidopsis cytochrome p450 cyp83B1 mutations activate the tryptophan biosynthetic pathway. Genetics 2002, 160, 323–332. [Google Scholar]
- Bak, S.; Tax, F.E.; Feldmann, K.A.; Galbraith, D.W.; Feyereisen, R. CYP83B1, a cytochrome P450 at the metabolic branch point in auxin and indole glucosinolate biosynthesis in Arabidopsis. Plant Cell 2001, 13, 101–111. [Google Scholar]
- Naur, P.; Petersen, B.L.; Mikkelsen, M.D.; Bak, S.; Rasmussen, H.; Olsen, C.E.; Halkier, B.A. CYP83A1 and CYP83B1, two nonredundant cytochrome P450 enzymes metabolizing oximes in the biosynthesis of glucosinolates in Arabidopsis. Plant Physiol 2003, 133, 67–72. [Google Scholar]
- Zang, Y.X.; Kim, H.U.; Kim, J.A.; Lim, M.H.; Jin, M.; Lee, S.C.; Kwon, S.J.; Lee, S.I.; Hong, J.K.; Park, T.H.; et al. Genome-wide identification of glucosinolate synthesis genes in Brassica rapa. FEBS J 2009, 276, 3559–3574. [Google Scholar]
- Bowers, J.E.; Chapman, B.A.; Rong, J.K.; Paterson, A.H. Unravelling angiosperm genome evolution by phylogenetic analysis of chromosomal duplication events. Nature 2003, 422, 433–438. [Google Scholar]
- Lysak, M.A.; Koch, M.; Pecinka, A.; Schubert, I. Chromosome triplication found across the tribe Brassiceae. Genome Res 2005, 15, 516–525. [Google Scholar]
- Yang, Y.W.; Lai, K.N.; Tai, P.Y.; Li, W.H. Rates of nucleotide substitution in angiosperm mitochondrial DNA sequences and dates of divergence between Brassica and other angiosperm lineages. J. Mol. Evol 1999, 48, 597–604. [Google Scholar]
- Mizutani, M.; Ward, E.; Ohta, E. Cytochrome p450 superfamily in Arabidopsis thaliana: Isolation of cDNAs, differential expression, and RFLP mapping of multiple cytochromes P450. Plant Mol. Biol 1998, 37, 39–52. [Google Scholar]
- Xu, W.; Bak, S.; Decker, A.; Paquette, S.M.; Feyereisen, R.; Galbraith, D.W. Microarray-based analysis of gene expression in very large gene families: The cytochrome P450 gene superfamily of Arabidopsis thaliana. Gene 2001, 272, 61–74. [Google Scholar]
- Brown, P.D.; Tokuhisa, J.G.; Reichelt, M.; Gershenzon, J. Variation of glucosinolate accumulation among different organs and developmental stages of Arabidopsis thaliana. Phytochemistry 2003, 62, 471–481. [Google Scholar]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server. In The Proteomics Protocols Handbook; Walker, J.M., Ed.; Humana Press Inc: Totowa, NJ, USA, 2005; pp. 571–607. [Google Scholar]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 1994, 22, 4673–4680. [Google Scholar]
- Rozen, S.; Skaletsky, H.J. Primer3 on the WWW for General Users and for Biologist Programmers. In Bioinformatics Methods and Protocols: Methods in Molecular Biology; Krawetz, S., Misener, S., Eds.; Humana Press Inc: Totowa, NJ, USA, 2000; pp. 365–386. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 2001, 25, 402–408. [Google Scholar]
- Yin, X.R.; Chen, K.S.; Allan, A.C.; Wu, R.M.; Zhang, B.; Lallu, N.; Ferguson, I.B. Ethylene-induced modulation of genes associated with the ethylene signaling pathway in ripening kiwifruit. J. Exp. Bot 2008, 59, 2097–2108. [Google Scholar]
- Krumbein, A.; Schonhof, I.; Schreiner, M. Composition and contents of phytochemicals (glucosinolates, carotenoids and chlorophylls) and ascorbic acid in selected Brassica species (B. juncea, B. rapa subsp. nipposinica var. chinoleifera, B. rapa subsp. chinensis and B. rapa subsp. rapa). J. Appl. Bot. Food Qual 2005, 79, 168–174. [Google Scholar]




| ZYYDE | HZYDE | NJZGB | CGB | SHQ | SYM | AJH | Leaf | Root | Petiole | |
|---|---|---|---|---|---|---|---|---|---|---|
| Aliphatic GS | 0.43 ± 0.07 | 0.94 ± 0.07 | 0.49 ± 0.07 | 0.54 ± 0.08 | 0.98 ± 0.04 | 2.90 ± 0.34 | 0.87 ± 0.10 | 1.89 ± 0.29 | 0.12 ± 0.02 | 0.48 ± 0.05 |
| Indole GS | 0.68 ± 0.03 | 0.76 ± 0.03 | 0.30 ± 0.01 | 0.45 ± 0.02 | 0.52 ± 0.02 | 0.59 ± 0.03 | 0.75 ± 0.10 | 0.25 ± 0.06 | 0.94 ± 0.10 | 0.24 ± 0.04 |
| Aromatic GS | 0.38 ± 0.02 | 0.51 ± 0.02 | 0.31 ± 0.03 | 0.59 ± 0.07 | 0.63 ± 0.04 | 0.46 ± 0.07 | 0.61 ± 0.05 | 0.28 ± 0.06 | 0.78 ± 0.07 | 0.25 ± 0.03 |
| Total GS | 1.49 ± 0.07 | 2.21 ± 0.09 | 1.10 ± 0.09 | 1.58 ± 0.11 | 2.13 ± 0.05 | 3.95 ± 0.41 | 2.23 ± 0.04 | 2.43 ± 0.24 | 1.84 ± 0.17 | 0.97 ± 0.05 |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhu, B.; Wang, Z.; Yang, J.; Zhu, Z.; Wang, H. Isolation and Expression of Glucosinolate Synthesis Genes CYP83A1 and CYP83B1 in Pak Choi (Brassica rapa L. ssp. chinensis var. communis (N. Tsen & S.H. Lee) Hanelt). Int. J. Mol. Sci. 2012, 13, 5832-5843. https://doi.org/10.3390/ijms13055832
Zhu B, Wang Z, Yang J, Zhu Z, Wang H. Isolation and Expression of Glucosinolate Synthesis Genes CYP83A1 and CYP83B1 in Pak Choi (Brassica rapa L. ssp. chinensis var. communis (N. Tsen & S.H. Lee) Hanelt). International Journal of Molecular Sciences. 2012; 13(5):5832-5843. https://doi.org/10.3390/ijms13055832
Chicago/Turabian StyleZhu, Biao, Zhizhou Wang, Jing Yang, Zhujun Zhu, and Huasen Wang. 2012. "Isolation and Expression of Glucosinolate Synthesis Genes CYP83A1 and CYP83B1 in Pak Choi (Brassica rapa L. ssp. chinensis var. communis (N. Tsen & S.H. Lee) Hanelt)" International Journal of Molecular Sciences 13, no. 5: 5832-5843. https://doi.org/10.3390/ijms13055832





