DNA Barcodes of Asian Houbara Bustard (Chlamydotis undulata macqueenii)
Abstract
:1. Introduction
2. Results
3. Discussion
4. Experimental Section
5. Conclusions
Acknowledgments
References
- Cramp, S.; Simmons, K.E.L. Handbook of the Birds of Europe, the Middle East and North Africa; Oxford University Press: Oxford, UK, 1980; Volume II, pp. 636–668. [Google Scholar]
- Jennings, M.C. The Birds of Saudia Arabia: Past, Present and Future. Proceedings of the 1st Symposium Wildlife Conservation and Development in Saudi Arabia, Riyadh, Saudi Arabia, February 1989, Abuzinada, A., Gorriup, P., Nader, I., Eds.; Saudi Wildlife Commission: Riyadh, Saudi Arabia; pp. 255–262.
- Seddon, P.J.; Jaime, M.S.; van Heezik, V.; Paillat, P.; Gaucher, P.; Combreau, O. Restoration of houbara bustard populations in Saudi Arabia: Developments and future directions. Oryx 1995, 29, 136–142. [Google Scholar]
- Combreau, O.; Saint-Jame, M.; Seddon, P.; Rambaud, F.; van Heezic, Y.; Paillat, P.; Gaucher, P.; Smith, T. A Program for Houbara Bustard Restoration in Saudi Arabia. In Integrating People and Wildlife for a Sustainable Future; Proceedings of the 1st International Wildlife Management Congress, Bethesda, MD, USA, 1995, Bissonette, J.A., Krausman, P.R., Eds.; Wildlife Society: Bethesda, MD, USA; pp. 520–524.
- Wernery, U.; Liu, C.; Baskar, V.; Guerineche, Z.; Khazanehdari, K.A.; Saleem, S.; Kinne, J.; Wernery, R.; Griffin, D.K.; Chang, I. Primordial germ cell-mediated chimera technology produces viable pure-line Houbara bustard offspring: Potential for repopulating an endangered species. PLoS One 2010, 5. [Google Scholar] [CrossRef] [Green Version]
- Ivy, J.A.; Miller, A.; Lacy, R.C.; Dewoody, J.A. Methods and prospects for using molecular data in captive breeding programs: An empirical example using parma wallabies (Macropus parma). J. Hered 2009, 100, 441–454. [Google Scholar]
- Blonk, R.J.; Komen, H.; Kamstra, A.; van Arendonk, J.A. Estimating breeding values with molecular relatedness and reconstructed pedigrees in natural mating populations of common sole, Solea solea. Genetics 2010, 184, 213–219. [Google Scholar]
- Russello, M.A.; Amato, G. On the horns of a dilemma: Molecular approaches refine ex situ conservation in crisis. Mol. Ecol 2007, 16, 2405–2406. [Google Scholar]
- Khan, H.A.; Arif, I.A.; Shobrak, M.; Homaidan, A.A.; Farhan, A.H.; Sadoon, M.A. Application of mitochondrial genes sequences for measuring the genetic diversity of Arabian oryx. Genes Genet. Syst 2011, 86, 67–72. [Google Scholar]
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L.; deWaard, J.R. Biological identifications through DNA barcodes. Proc. R. Soc. Lond. B 2003, 270, 313–321. [Google Scholar]
- Hebert, P.D.N.; Ratnasingham, S.; deWaard, J.R. Barcoding animal life: Cytochrome c oxidase subunit 1 divergences among closely related species. Proc. R. Soc. Lond. B 2003, 270, S596–S599. [Google Scholar]
- Kerr, K.C.; Stoeckle, M.Y.; Dove, C.J.; Weigt, L.A.; Francis, C.M.; Hebert, P.D. Comprehensive DNA barcode coverage of North American birds. Mol. Ecol. Notes 2007, 7, 535–543. [Google Scholar]
- Tavares, E.S.; Baker, A.J. Single mitochondrial gene barcodes reliably identify sister-species in diverse clades of birds. BMC Evol. Biol 2008, 8. [Google Scholar] [CrossRef]
- Moritz, C.; Cicero, C. DNA barcoding: Promise and pitfalls. PLoS Biol 2004, 2. [Google Scholar] [CrossRef]
- Hebert, P.D.; Stoeckle, M.Y.; Zemlak, T.S.; Francis, C.M. Identification of birds through DNA barcodes. PLoS Biol 2004, 2. [Google Scholar] [CrossRef]
- Kerr, K.C.; Birks, S.M.; Kalyakin, M.V.; Red’kin, Y.A.; Koblik, E.A.; Hebert, P.D. Filling the gap-COI barcode resolution in eastern Palearctic birds. Front. Zool 2009, 6. [Google Scholar] [CrossRef]
- Yoo, H.S.; Eah, J.Y.; Kim, J.S.; Kim, Y.J.; Min, M.S.; Paek, W.K.; Lee, H.; Kim, C.B. DNA barcoding Korean birds. Mol. Cell 2006, 22, 323–327. [Google Scholar]
- Vilaça, S.T.; Lacerda, D.R.; Sari, H.E.R.; Santos, F.R. DNA-based identification applied to Thamnophilidae (Passeriformes) species: The first barcodes of Neotropical birds. Rev. Bras. Ornitol 2006, 14, 7–13. [Google Scholar]
- Chaves, A.V.; Clozato, C.L.; Lacerda, D.R.; Sari, E.H.R.; Santos, F.R. Molecular taxonomy of brazilian tyrant-flycatchers (Passeriformes: Tyrannidae). Mol. Ecol. Resour 2008, 8, 1169–1177. [Google Scholar]
- Khan, H.A.; Arif, I.A.; Shobrak, M. DNA barcodes of arabian partridge and philby’s rock partridge: Implications for phylogeny and species identification. Evol. Bioinform 2010, 6, 151–158. [Google Scholar]
- Arif, I.A.; Khan, H.A.; Shobrak, M.; Williams, J. Cytochrome c oxidase subunit I barcoding of the green bee-eater (Merops orientalis). Genet. Mol. Res 2011, 10. in press. [Google Scholar]
- Howlett, J.C.; Hölzer, W.; Bailey, T.A.; Wernery, U.; Samour, J.H.; Naldo, J.L. Serum bile acids in captive bustards. Zentr. Vet. B 1999, 46, 701–705. [Google Scholar]
- Bailey, T.A.; Wernery, U.; Howlett, J.; Naldo, J.; Samour, J.H. Age-related plasma chemistry changes in houbara and kori bustards in the United Arab Emirates. J. Wildl. Dis 1999, 35, 31–37. [Google Scholar]
- Gelinaud, G.; Combreau, O.; Seddon, P.J. First breeding by captive-bred houbara bustards introduced in central Saudi Arabia. J. Arid Environ 1997, 35, 527–534. [Google Scholar]
- Greth, A.; Andral, B.; Gerbermann, H.; Vassart, M.; Gerlach, H.; Launay, F. Chlamydiosis in a captive group of Houbara bustards (Chlamydotis undulata). Avian Dis 1993, 37, 1117–1120. [Google Scholar]
- Jalme, M.S.; Gaucher, P.; Paillat, P. Artificial insemination in Houbara bustards (Chlamydotis undulata): Influence of the number of spermatozoa and insemination frequency on fertility and ability to hatch. J. Reprod. Fertil 1994, 100, 93–103. [Google Scholar]
- D’aloia, M.A.; Samour, J.H.; Howlett, J.C.; Bailey, T.A.; Naldo, J. Normal blood chemistry of the Houbara bustard (Chlamydotis undulata). Avian Pathol 1996, 25, 167–173. [Google Scholar]
- Ostrowski, S.; Ancrenaz, M.; Saint-Jalme, M.; Greth, A. Concurrent avian pox and Newcastle disease infection in a Houbara bustard (Chlamydotts undulatd). Avian Pathol 1995, 24, 573–577. [Google Scholar]
- Khan, O.A.; Shuaib, M.A.; Rhman, S.S.; Ismail, M.M.; Hammad, Y.A.; Baky, M.H.; Fusaro, A.; Salviato, A.; Cattoli, G. Isolation and identification of highly pathogenic avian influenza H5N1 virus from Houbara bustards (Chlamydotis undulata macqueenii) and contact falcons. Avian Pathol 2009, 38, 35–39. [Google Scholar]
- Facon, C.; Guerin, J.L.; Lacroix, F. Assessment of newcastle disease vaccination of houbara bustard breeders (Chlamydotis undulata undulata). J. Wildl. Dis 2005, 41, 768–774. [Google Scholar]
- Idaghdour, Y.; Broderick, D.; Korrida, A.; Chbel, F. Mitochondrial control region diversity of the houbara bustard Chlamydotis undulata complex and genetic structure along the Atlantic seaboard of North Africa. Mol. Ecol 2004, 13, 43–54. [Google Scholar]
- Khan, H.A.; Arif, I.A.; Bahkali, A.H.; Al Farhan, A.H.; Al Homaidan, A.A. Bayesian, maximum parsimony and UPGMA models for inferring the phylogenies of antelopes using mitochondrial markers. Evol. Bioinform 2008, 4, 263–270. [Google Scholar]
- Tateno, Y.; Takezaki, N.; Nei, M. Relative efficiencies of the maximum-likelihood, neighbor-joining, and maximum-parsimony methods when substitution rate varies with site. Mol. Biol. Evol 1994, 11, 261–277. [Google Scholar]
- Arif, I.A.; Khan, H.A. Molecular markers for biodiversity analysis of wildlife animals: A brief review. Anim. Biodivers. Conserv 2009, 32, 9–17. [Google Scholar]
- Arif, I.A.; Khan, H.A.; Bahkali, A.H.; Al Homaidan, A.A.; Al Farhan, A.H.; Al Sadoon, M.; Shobrak, M. DNA marker technology for wildlife conservation. Saudi J. Biol. Sci 2011, 18, 219–225. [Google Scholar]
- Rach, J.; Desalle, R.; Sarkar, I.N.; Schierwater, B.; Hadrys, H. Character-based DNA barcoding allows discrimination of genera, species and populations in Odonata. Proc. Biol. Sci 2008, 275, 237–247. [Google Scholar]
- Fleischer, R.C.; Kirchman, J.J.; Dumbacher, J.P.; Bevier, L.; Dove, C.; Rotzel, N.C.; Edwards, S.V.; Lammertink, M.; Miglia, K.J.; Moore, W.S. Mid-Pleistocene divergence of Cuban and North American ivory-billed woodpeckers. Biol. Lett 2006, 2, 466–469. [Google Scholar]
- Tavares, E.S.; de Kroon, G.H.; Baker, A.J. Phylogenetic and coalescent analysis of three loci suggest that the Water Rail is divisible into two species, Rallus aquaticus and R. indicus. BMC Evol. Biol 2010, 10. [Google Scholar] [CrossRef]
- Yang, R.; Wu, X.; Yan, P.; Li, X. Using DNA barcodes to identify a bird involved in a birdstrike at a Chinese airport. Mol. Biol. Rep 2010, 37, 3517–3523. [Google Scholar]
- Liu, L.; Pearl, D.K.; Brumfield, R.T.; Edwards, S.V. Estimating species trees using multiple-allele DNA sequence data. Evolution 2008, 62, 2080–2091. [Google Scholar]
- Arif, I.A.; Bakir, M.A.; Khan, H.A. Inferring the phylogeny of Bovidae using mitochondrial DNA sequences: Resolving power of individual genes relative to complete genome. Evol. Bioinform 2012, 8, 139–150. [Google Scholar]
- Cummings, M.P.; Otto, S.P.; Wakeley, J. Sampling properties of DNA sequence data in phylogenetic analysis. Mol. Biol. Evol 1995, 12, 814–822. [Google Scholar]
- Barcode of lift data system. Available online: http://www.boldsystems.org Accessed on 5 January 2012.
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. ClustalW and ClustalX version 2. Bioinformatics 2007, 23, 2947–2948. [Google Scholar]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol 1987, 4, 406–425. [Google Scholar]
- Tamura, K.; Nei, M.; Kumar, S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc. Natl. Acad. Sci. USA 2004, 101, 11030–11035. [Google Scholar]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol 2007, 24, 1596–1599. [Google Scholar]
- Huelsenbeck, J.P.; Ronquist, F.R. MrBayes: Bayesian inference of phylogenetic trees. Bioinformatics 2001, 17, 754–755. [Google Scholar]
- Page, R.D.M. Treeview: An application to display phylogenetic trees on personal computers. Comput. Appl. Biosci 1996, 12, 357–358. [Google Scholar]

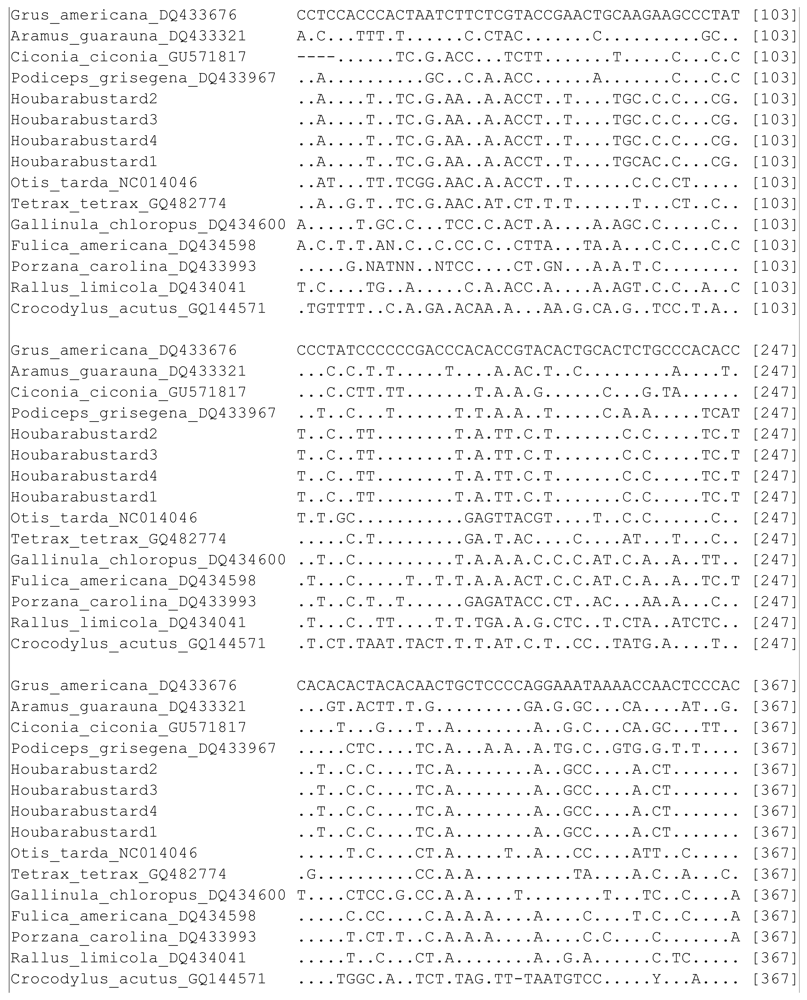

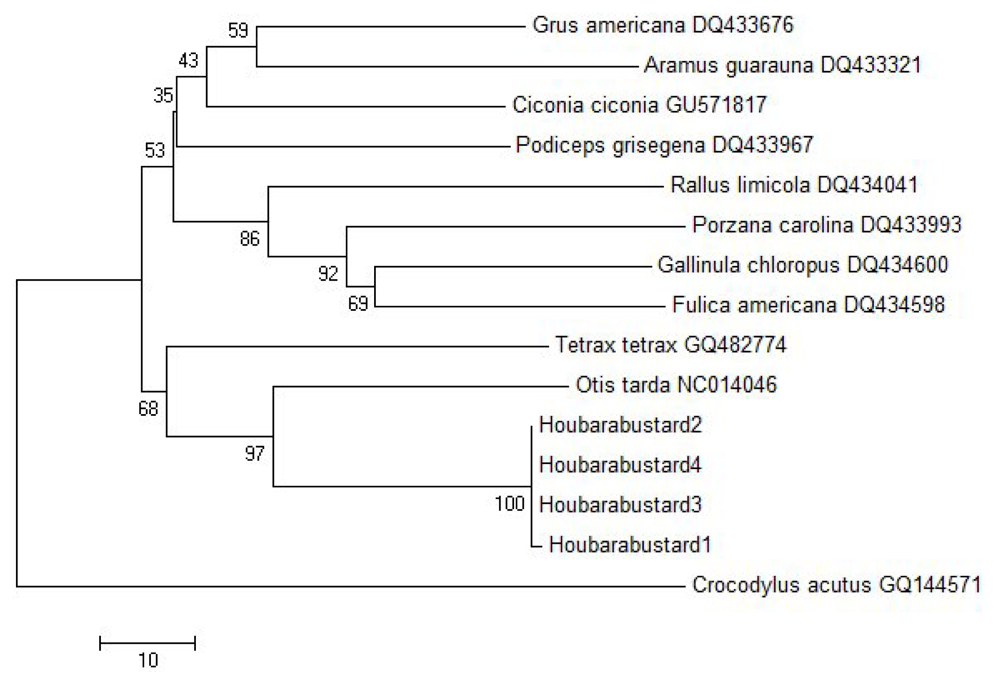
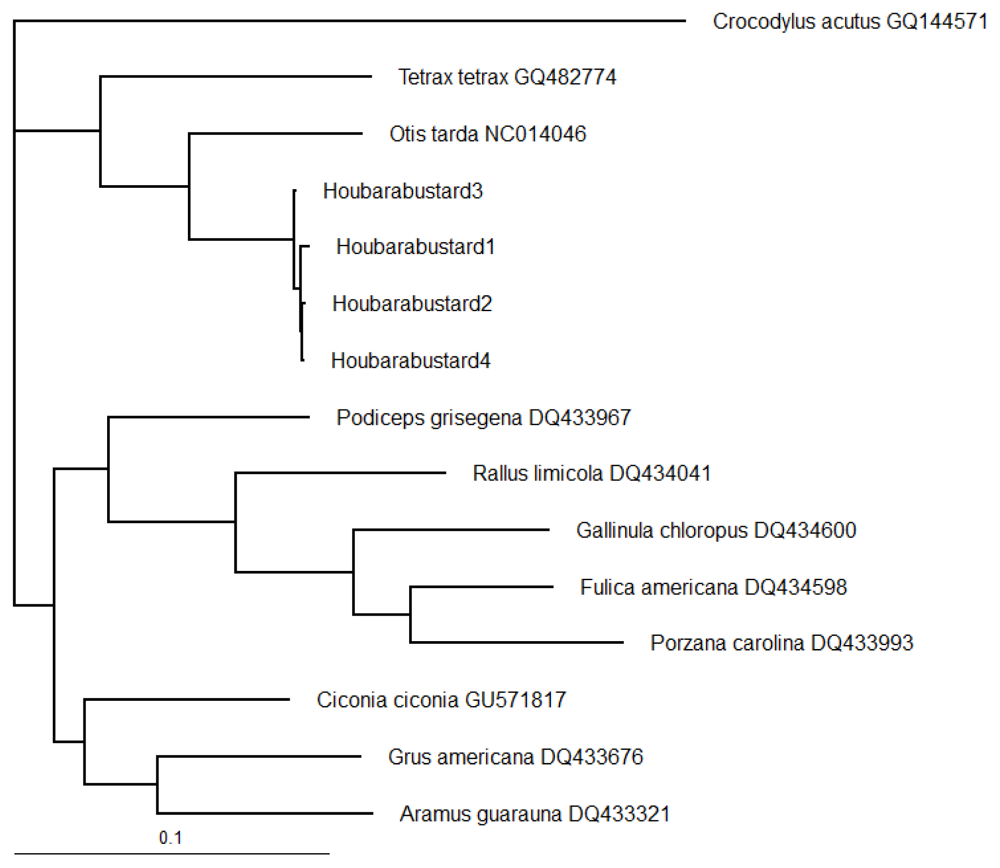
| Sp. | GA | AG | CC | PG | HB1 | HB2 | HB3 | HB4 | OT | TT | GC | FA | PC | RL | CA |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GA | 68 | 68 | 76 | 81 | 81 | 81 | 82 | 83 | 78 | 87 | 91 | 82 | 90 | 124 | |
| AG | 0.131 | 73 | 77 | 97 | 97 | 97 | 98 | 100 | 93 | 97 | 99 | 100 | 104 | 124 | |
| CC | 0.133 | 0.145 | 70 | 74 | 74 | 74 | 75 | 89 | 84 | 90 | 86 | 82 | 87 | 129 | |
| PG | 0.154 | 0.155 | 0.139 | 74 | 74 | 74 | 75 | 86 | 94 | 89 | 83 | 94 | 82 | 124 | |
| HB1 | 0.167 | 0.210 | 0.150 | 0.150 | 0 | 0 | 1 | 58 | 79 | 93 | 98 | 100 | 97 | 123 | |
| HB2 | 0.167 | 0.210 | 0.150 | 0.150 | 0.000 | 0 | 1 | 58 | 79 | 93 | 98 | 100 | 97 | 123 | |
| HB3 | 0.167 | 0.210 | 0.150 | 0.150 | 0.000 | 0.000 | 1 | 58 | 79 | 93 | 98 | 100 | 97 | 123 | |
| HB4 | 0.169 | 0.213 | 0.152 | 0.152 | 0.002 | 0.002 | 0.002 | 59 | 80 | 94 | 99 | 101 | 98 | 124 | |
| OT | 0.170 | 0.218 | 0.189 | 0.180 | 0.106 | 0.106 | 0.106 | 0.108 | 78 | 95 | 96 | 98 | 98 | 131 | |
| TT | 0.159 | 0.200 | 0.175 | 0.202 | 0.156 | 0.156 | 0.156 | 0.158 | 0.153 | 97 | 97 | 90 | 96 | 124 | |
| GC | 0.187 | 0.216 | 0.193 | 0.192 | 0.203 | 0.203 | 0.203 | 0.205 | 0.206 | 0.216 | 59 | 68 | 76 | 138 | |
| FA | 0.199 | 0.223 | 0.182 | 0.175 | 0.219 | 0.219 | 0.219 | 0.222 | 0.212 | 0.217 | 0.111 | 68 | 82 | 134 | |
| PC | 0.172 | 0.223 | 0.171 | 0.205 | 0.224 | 0.224 | 0.224 | 0.227 | 0.216 | 0.197 | 0.130 | 0.130 | 91 | 147 | |
| RL | 0.193 | 0.233 | 0.184 | 0.170 | 0.213 | 0.213 | 0.213 | 0.216 | 0.214 | 0.212 | 0.153 | 0.168 | 0.192 | 130 | |
| CA | 0.312 | 0.315 | 0.340 | 0.316 | 0.310 | 0.310 | 0.310 | 0.313 | 0.336 | 0.309 | 0.383 | 0.368 | 0.431 | 0.342 |
| Class | Order | Family | Genus | Species | Similarity(%) |
|---|---|---|---|---|---|
| Aves | Gruiformes | Otididae | Otis | tarda | 90.43 |
| Aves | Sphenisciformes | Spheniscidae | Pygoscelis | adeliae | 88.39 |
| Aves | Sphenisciformes | Spheniscidae | Pygoscelis | adeliae | 88.39 |
| Aves | Sphenisciformes | Spheniscidae | Pygoscelis | adeliae | 88.39 |
| Aves | Gruiformes | Otididae | Ardeotis | kori | 88.27 |
| Aves | Gruiformes | Otididae | Ardeotis | kori | 88.12 |
| Aves | Gruiformes | Otididae | Ardeotis | kori | 88.12 |
| Aves | Ciconiiformes | Ciconiidae | Ciconia | ciconia | 88.12 |
| Aves | Ciconiiformes | Ciconiidae | Ciconia | ciconia | 88.01 |
| Aves | Ciconiiformes | Ciconiidae | Ciconia | ciconia | 88.01 |
| Aves | Procellariiformes | Hydrobatidae | Oceanites | nereis | 88.01 |
| Aves | Procellariiformes | Hydrobatidae | Oceanites | nereis | 88.01 |
| Aves | Procellariiformes | Hydrobatidae | Oceanites | nereis | 88.01 |
| Aves | Procellariiformes | Hydrobatidae | Oceanites | nereis | 88.01 |
| Aves | Procellariiformes | Hydrobatidae | Oceanites | nereis | 88.01 |
| Aves | Ciconiiformes | Ciconiidae | Ciconia | ciconia | 87.83 |
| Aves | Podicipediformes | Podicipedidae | Podiceps | grisegena | 87.69 |
| Aves | Podicipediformes | Podicipedidae | Podiceps | grisegena | 87.69 |
| Aves | Podicipediformes | Podicipedidae | Podiceps | grisegena | 87.68 |
| Aves | Podicipediformes | Podicipedidae | Podiceps | grisegena | 87.66 |
| GenBank Accession | Order | Family | Genus | Species | No. of Base Pairs |
|---|---|---|---|---|---|
| DQ434041 | Gruiformes | Rallidae | Rallus | limicola | 697 |
| DQ434600 | Gruiformes | Rallidae | Gallinula | chloropus | 696 |
| DQ434598 | Gruiformes | Rallidae | Fulica | americana | 697 |
| DQ433993 | Gruiformes | Rallidae | Porzana | carolina | 695 |
| DQ433676 | Gruiformes | Gruidae | Grus | americana | 681 |
| DQ433321 | Gruiformes | Aramidae | Aramus | guarauna | 697 |
| GQ482774 | Gruiformes | Otididae | Tetrax | tetrax | 694 |
| NC014046 | Gruiformes | Otididae | Otis | tarda | 694 |
| GU571817 | Ciconiiformes | Ciconiidae | Ciconia | ciconia | 648 |
| DQ433967 | Podicipediformes | Podicipedidae | Podiceps | grisegena | 672 |
| GQ144571 * | Crocodylia | Crocodylidae | Crocodylus | acutus | 645 |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Arif, I.A.; Khan, H.A.; Williams, J.B.; Shobrak, M.; Arif, W.I. DNA Barcodes of Asian Houbara Bustard (Chlamydotis undulata macqueenii). Int. J. Mol. Sci. 2012, 13, 2425-2438. https://doi.org/10.3390/ijms13022425
Arif IA, Khan HA, Williams JB, Shobrak M, Arif WI. DNA Barcodes of Asian Houbara Bustard (Chlamydotis undulata macqueenii). International Journal of Molecular Sciences. 2012; 13(2):2425-2438. https://doi.org/10.3390/ijms13022425
Chicago/Turabian StyleArif, Ibrahim A., Haseeb A. Khan, Joseph B. Williams, Mohammad Shobrak, and Waad I. Arif. 2012. "DNA Barcodes of Asian Houbara Bustard (Chlamydotis undulata macqueenii)" International Journal of Molecular Sciences 13, no. 2: 2425-2438. https://doi.org/10.3390/ijms13022425





