Identification of Candidate Polymorphisms on Stress Oxidative and DNA Damage Repair Genes Related with Clinical Outcome in Breast Cancer Patients
Abstract
:1. Introduction
2. Results and Discussion
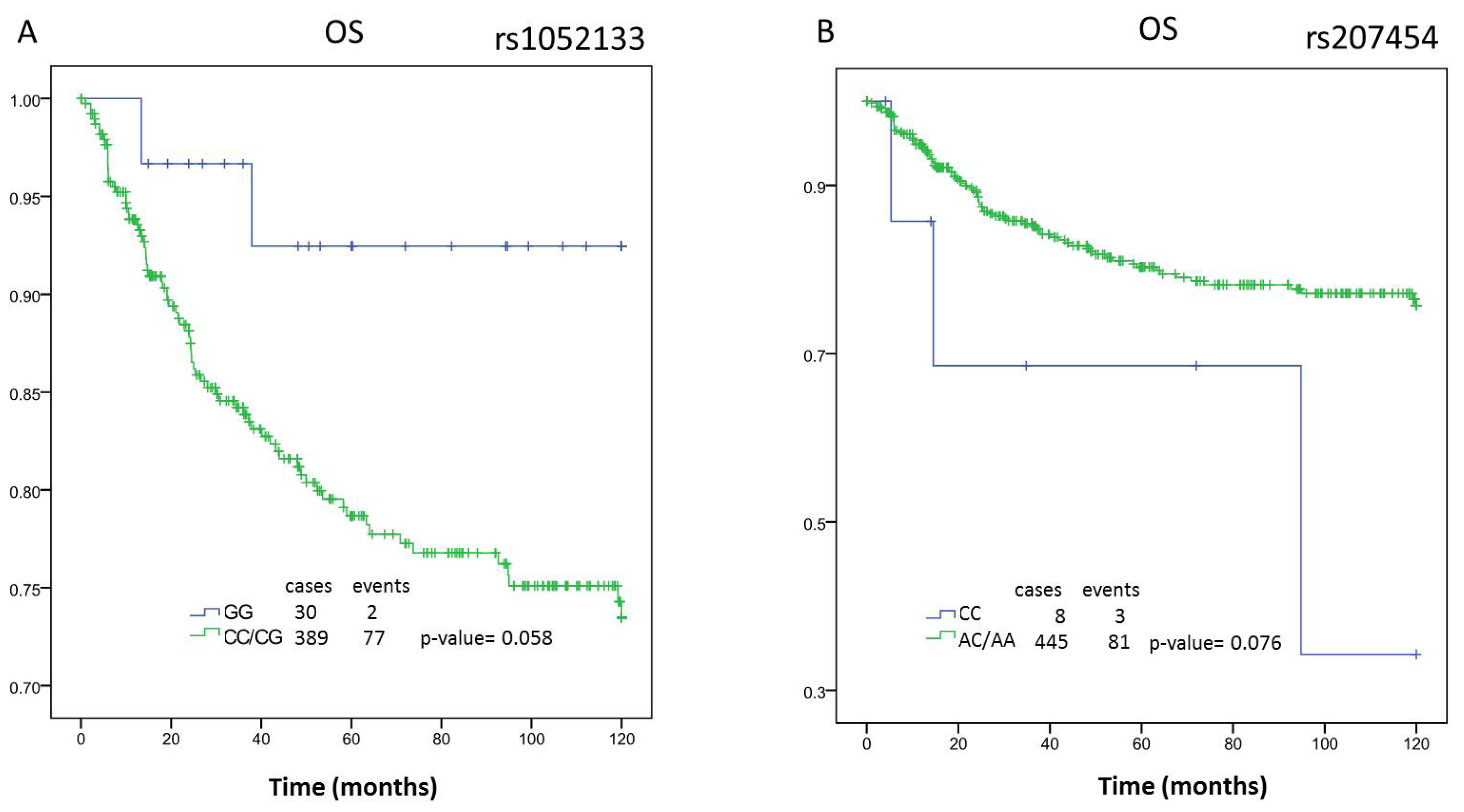
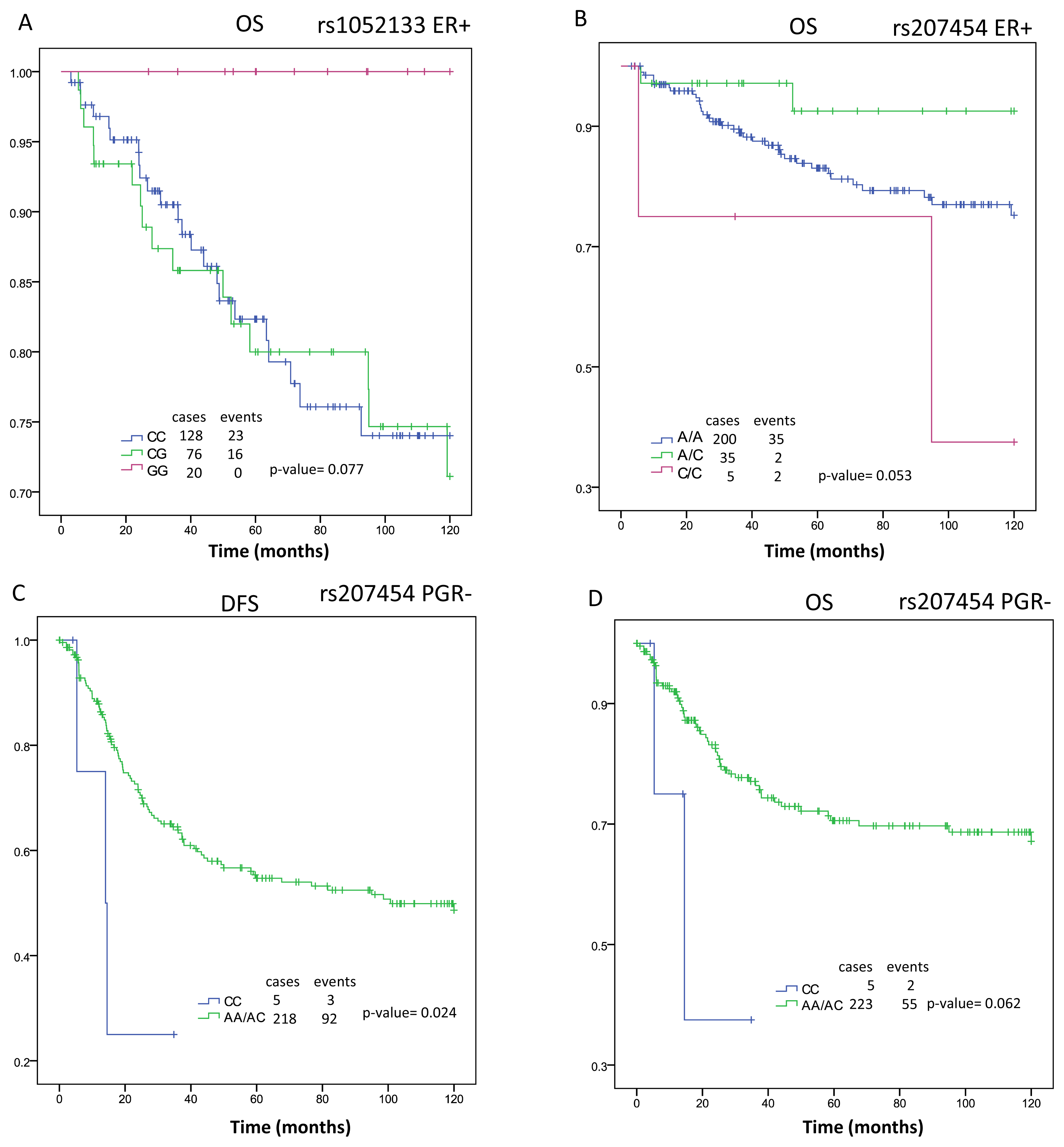
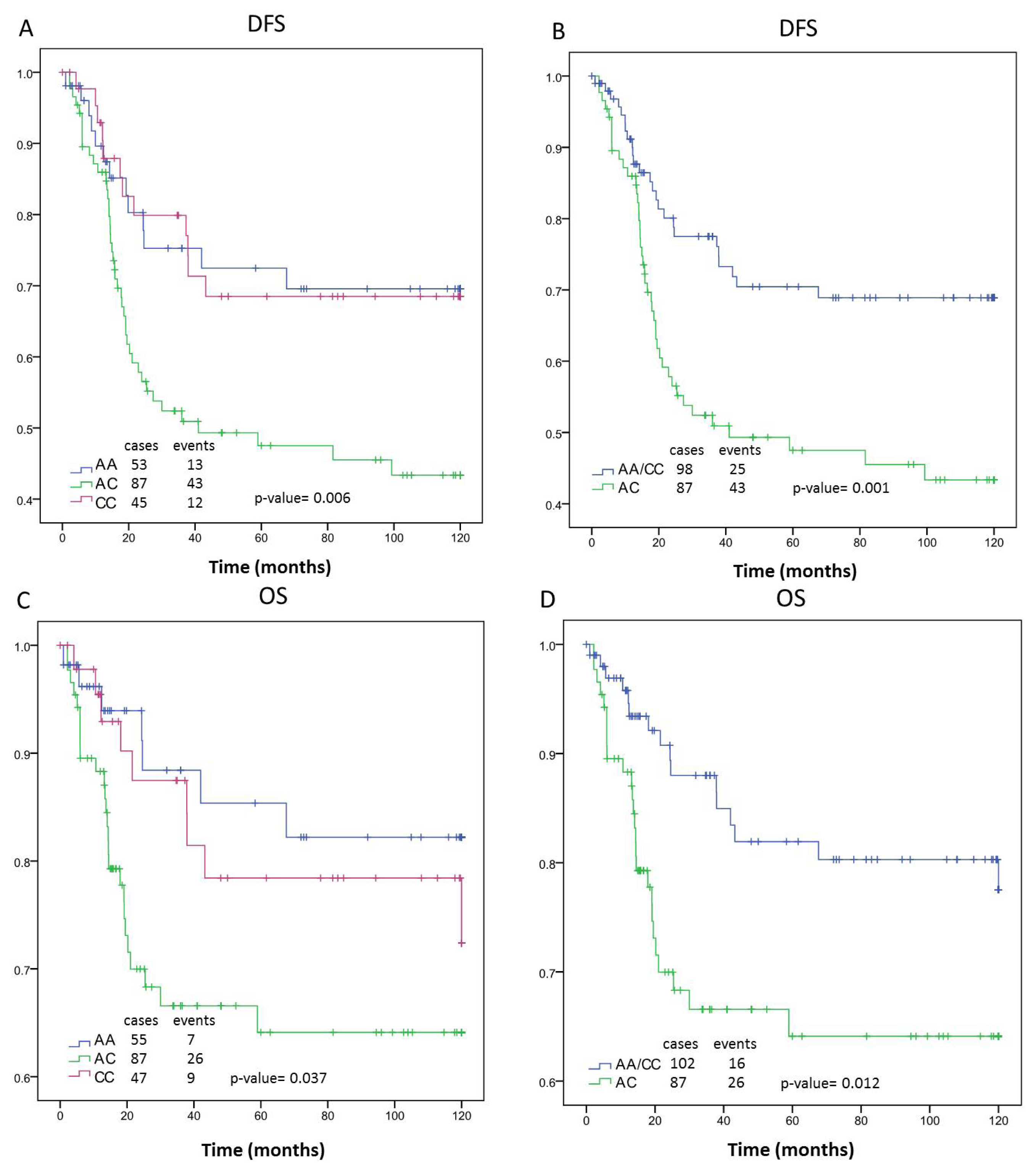
2.1. Survival Analysis
2.2. Discussion
3. Experimental Section
3.1. Study Population
3.2. DNA Extraction and Genotyping
3.3. Statistical Analyses
4. Conclusions
Acknowledgments
- Conflict of InterestThe authors declare no conflict of interest.
References
- Arteaga, C.L.; Sliwkowski, M.X.; Osborne, C.K.; Perez, E.A.; Puglisi, F.; Gianni, L. Treatment of HER2-positive breast cancer: current status and future perspectives. Nat. Rev. Clin. Oncol 2012, 9, 16–32. [Google Scholar]
- Dumitrescu, R.G.; Cotarla, I. Understanding breast cancer risk—Where do we stand in 2005? J. Cell. Mol. Med 2005, 9, 208–221. [Google Scholar]
- Fasching, P.A.; Pharoah, P.D.; Cox, A.; Nevanlinna, H.; Bojesen, S.E.; Karn, T.; Broeks, A.; van Leeuwen, F.E.; van’t Veer, L.J.; Udo, R.; et al. The role of genetic breast cancer susceptibility variants as prognostic factors. Hum. Mol. Genet 2012, 21, 3926–3939. [Google Scholar]
- Ferraldeschi, R.; Arnedos, M.; Hadfield, K.D.; A’Hern, R.; Drury, S.; Wardley, A.; Howell, A.; Evans, D.G.; Roberts, S.A.; Smith, I.; et al. Polymorphisms of CYP19A1 and response to aromatase inhibitors in metastatic breast cancer patients. Breast Cancer Res. Treat 2012, 133, 1191–1198. [Google Scholar]
- Johnson, N.; Walker, K.; Gibson, L.J.; Orr, N.; Folkerd, E.; Haynes, B.; Palles, C.; Coupland, B.; Schoemaker, M.; Jones, M.; et al. CYP3A Variation, Premenopausal Estrone Levels, and Breast Cancer Risk. J. Natl. Cancer Inst 2012, 104, 657–669. [Google Scholar]
- Ma, X.; Beeghly-Fadiel, A.; Lu, W.; Shi, J.; Xiang, Y.B.; Cai, Q.; Shen, H.; Shen, C.Y.; Ren, Z.; Keitaro, M.; et al. Pathway analyses identify TGFBR2 as potential breast cancer susceptibility gene: Results from a consortium study among Asians. Cancer Epidemiol. Biomarkers Prev 2012, 21, 1176–1184. [Google Scholar]
- Pabalan, N.; Jarjanazi, H.; Sung, L.; Li, H.; Ozcelik, H. Menopausal status modifies breast cancer risk associated with the myeloperoxidase (MPO) G463A polymorphism in Caucasian women: A meta-analysis. PLoS One 2012, 7, e32389. [Google Scholar]
- Rodrigues, P.; Furriol, J.; Tormo, E.; Ballester, S.; Lluch, A.; Eroles, P. The single-nucleotide polymorphisms +936 C/T VEGF and −710 C/T VEGFR1 are associated with breast cancer protection in a Spanish population. Breast Cancer Res. Treat 2012, 133, 769–778. [Google Scholar]
- Wang, J.; Wang, B.; Bi, J.; Li, K.; Di, J. MDR1 gene C3435T polymorphism and cancer risk: A meta-analysis of 34 case-control studies. J. Cancer Res. Clin. Oncol 2012, 138, 979–989. [Google Scholar]
- Zhou, X.; Wu, C. Association of p73 G4C14-A4T14 polymorphisms with genetic susceptibilities to breast cancer: A case-control study. Med. Oncol 2012, 29, 3216–3221. [Google Scholar]
- Goode, E.L.; Ulrich, C.M.; Potter, J.D. Polymorphisms in DNA repair genes and associations with cancer risk. Cancer Epidemiol. Biomarkers Prev 2002, 11, 1513–1530. [Google Scholar]
- Wu, H.; Kang, H.; Liu, Y.; Tong, W.; Liu, D.; Yang, X.; Lian, M.; Yao, W.; Zhao, H.; Huang, D.; et al. Roles of ABCB1 gene polymorphisms and haplotype in susceptibility to breast carcinoma risk and clinical outcomes. J. Cancer Res. Clin. Oncol 2012, 138, 1449–1462. [Google Scholar]
- Yang, X.; Liu, D.; Wu, H.; Kang, H.; Pang, H.; Huang, D.; Sha, X.; Wang, E.; Wang, Z.; Wei, M. Association of XPC polymorphisms with susceptibility and clinical outcome to chemotherapy in breast cancer patients. Cancer Sci. 2012. [Google Scholar] [CrossRef]
- Ambrosone, C.B.; Tian, C.; Ahn, J.; Kropp, S.; Helmbold, I.; von Fournier, D.; Haase, W.; Sautter-Bihl, M.L.; Wenz, F.; Chang-Claude, J. Genetic predictors of acute toxicities related to radiation therapy following lumpectomy for breast cancer: A case-series study. Breast Cancer Res 2006, 8, R40. [Google Scholar]
- Choi, J.Y.; Barlow, W.E.; Albain, K.S.; Hong, C.C.; Blanco, J.G.; Livingston, R.B.; Davis, W.; Rae, J.M.; Yeh, I.T.; Hutchins, L.F.; et al. Nitric oxide synthase variants and disease-free survival among treated and untreated breast cancer patients in a Southwest Oncology Group clinical trial. Clin. Cancer Res 2009, 15, 5258–5266. [Google Scholar]
- Fagerholm, R.; Hofstetter, B.; Tommiska, J.; Aaltonen, K.; Vrtel, R.; Syrjakoski, K.; Kallioniemi, A.; Kilpivaara, O.; Mannermaa, A.; Kosma, V.M.; et al. NAD(P)H: quinone oxidoreductase 1 NQO1*2 genotype (P187S) is a strong prognostic and predictive factor in breast cancer. Nat. Genet 2008, 40, 844–853. [Google Scholar]
- Hubackova, M.; Vaclavikova, R.; Ehrlichova, M.; Mrhalova, M.; Kodet, R.; Kubackova, K.; Vrana, D.; Gut, I.; Soucek, P. Association of superoxide dismutases and NAD(P)H quinone oxidoreductases with prognosis of patients with breast carcinomas. Int. J. Cancer 2012, 130, 338–348. [Google Scholar]
- Martin, R.C.; Ahn, J.; Nowell, S.A.; Hein, D.W.; Doll, M.A.; Martini, B.D.; Ambrosone, C.B. Association between manganese superoxide dismutase promoter gene polymorphism and breast cancer survival. Breast Cancer Res 2006, 8, R45. [Google Scholar]
- Udler, M.; Maia, A.T.; Cebrian, A.; Brown, C.; Greenberg, D.; Shah, M.; Caldas, C.; Dunning, A.; Easton, D.; Ponder, B.; et al. Common germline genetic variation in antioxidant defense genes and survival after diagnosis of breast cancer. J. Clin. Oncol 2007, 25, 3015–3023. [Google Scholar]
- Yao, S.; Barlow, W.E.; Albain, K.S.; Choi, J.Y.; Zhao, H.; Livingston, R.B.; Davis, W.; Rae, J.M.; Yeh, I.T.; Hutchins, L.F.; et al. Manganese superoxide dismutase polymorphism, treatment-related toxicity and disease-free survival in SWOG 8897 clinical trial for breast cancer. Breast Cancer Res. Treat 2010, 124, 433–439. [Google Scholar]
- Acharya, A.; Das, I.; Chandhok, D.; Saha, T. Redox regulation in cancer: A double-edged sword with therapeutic potential. Oxid. Med. Cell. Longev 2010, 3, 23–34. [Google Scholar]
- Mignotte, B.; Vayssiere, J.L. Mitochondria and apoptosis. Eur. J. Biochem 1998, 252, 1–15. [Google Scholar]
- Modica-Napolitano, J.S.; Singh, K.K. Mitochondria as targets for detection and treatment of cancer. Expert Rev. Mol. Med 2002, 4, 1–19. [Google Scholar]
- Costantini, P.; Jacotot, E.; Decaudin, D.; Kroemer, G. Mitochondrion as a novel target of anticancer chemotherapy. J. Natl. Cancer Inst 2000, 92, 1042–1053. [Google Scholar]
- Hussain, S.P.; Hofseth, L.J.; Harris, C.C. Radical causes of cancer. Nat. Rev. Cancer 2003, 3, 276–285. [Google Scholar]
- Hirano, T. Repair system of 7, 8-dihydro-8-oxoguanine as a defense line against carcinogenesis. J. Radiat. Res 2008, 49, 329–340. [Google Scholar]
- Klungland, A.; Bjelland, S. Oxidative damage to purines in DNA: role of mammalian Ogg1. DNA Repair 2007, 6, 481–488. [Google Scholar]
- Paz-Elizur, T.; Sevilya, Z.; Leitner-Dagan, Y.; Elinger, D.; Roisman, L.C.; Livneh, Z. DNA repair of oxidative DNA damage in human carcinogenesis: Potential application for cancer risk assessment and prevention. Cancer Lett 2008, 266, 60–72. [Google Scholar]
- Yuan, W.; Xu, L.; Feng, Y.; Yang, Y.; Chen, W.; Wang, J.; Pang, D.; Li, D. The hOGG1 Ser326Cys polymorphism and breast cancer risk: A meta-analysis. Breast Cancer Res. Treat 2010, 122, 835–842. [Google Scholar]
- Castro, G.D.; Delgado de Layno, A.M.; Costantini, M.H.; Castro, J.A. Cytosolic xanthine oxidoreductase mediated bioactivation of ethanol to acetaldehyde and free radicals in rat breast tissue. Its potential role in alcohol-promoted mammary cancer. Toxicology 2001, 160, 11–18. [Google Scholar]
- Soini, Y.; Karihtala, P.; Mantyniemi, A.; Turunen, N.; Paakko, P.; Kinnula, V. Glutamate-L-cysteine ligase in breast carcinomas. Histopathology 2004, 44, 129–135. [Google Scholar]
- Pritsos, C.A.; Gustafson, D.L. Xanthine dehydrogenase and its role in cancer chemotherapy. Oncol. Res 1994, 6, 477–481. [Google Scholar]
- Comings, D.E.; MacMurray, J.P. Molecular heterosis: A review. Mol. Genet. Metab 2000, 71, 19–31. [Google Scholar]
- Montano, M.M.; Deng, H.; Liu, M.; Sun, X.; Singal, R. Transcriptional regulation by the estrogen receptor of antioxidative stress enzymes and its functional implications. Oncogene 2004, 23, 2442–2453. [Google Scholar]
- Urata, Y.; Ihara, Y.; Murata, H.; Goto, S.; Koji, T.; Yodoi, J.; Inoue, S.; Kondo, T. 17 Beta-estradiol protects against oxidative stress-induced cell death through the glutathione/glutaredoxin-dependent redox regulation of Akt in myocardiac H9c2 cells. J. Biol. Chem 2006, 281, 13092–13102. [Google Scholar]
- Galaris, D.; Skiada, V.; Barbouti, A. Redox signaling and cancer: the role of “labile” iron. Cancer Lett 2008, 266, 21–29. [Google Scholar]
- Halliwell, B. Oxidative stress and cancer: have we moved forward? Biochem. J 2007, 401, 1–11. [Google Scholar]
| n = 470 | |||
|---|---|---|---|
| Median | Range | ||
| Age (years) | 53 | (20–86) | |
| Follow up * (months) | 52.44 | (0–120) | |
| cases | % | ||
| Menopause status | Premenopausal | 77 | 28.00 |
| Postmenopausal | 187 | 68.00 | |
| Perimenopausal | 11 | 4.00 | |
| Histologic Type | Ductal infiltrate | 317 | 69.06 |
| Inflammatory | 69 | 15.03 | |
| CDI intraductal | 46 | 10.02 | |
| Others | 27 | 5.88 | |
| Tumor size | T0 | 7 | 1.65 |
| T1 | 109 | 25.65 | |
| T2 | 205 | 48.24 | |
| T3 | 37 | 8.71 | |
| T4 | 67 | 15.9 | |
| Node stage | N0 | 153 | 34.77 |
| N1 | 275 | 62.5 | |
| N2 | 9 | 2.05 | |
| N3-N9 | 3 | 0.7 | |
| Metastasis diagnosis | M0 | 425 | 96.59 |
| M1 | 15 | 3.41 | |
| Metastasis relapse | Primary | 136 | 28.4 |
| Secondary | 64 | 13.6 | |
| Stage | I | 53 | 21.9 |
| II | 136 | 56.2 | |
| III | 53 | 21.9 | |
| ER status | − | 219 | 46.79 |
| + | 249 | 53.21 | |
| PGR status | − | 239 | 50.96 |
| + | 230 | 49.04 | |
| HER2 overexpression | − | 195 | 84.78 |
| + | 35 | 15.22 | |
| Drug treatment | Anthracyclines | 159 | 69.74 |
| Tamoxifen | 45 | 19.74 | |
| Others | 24 | 10.53 | |
| Administration Treatment | Adjuvant | 326 | 72.4 |
| Neoadjuvant | 99 | 22.00 | |
| No treatment | 25 | 5.56 | |
| Radiotherapy | No | 294 | 62.55 |
| Yes | 153 | 32.55 | |
| NA | 23 | 4.90 | |
| Genotype Overall | ER+ | ER− | PGR+ | PGR− | ER− and PGR− | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| months | (95% CI) | p | months | (95% CI) | p | months | (95% CI) | p | months | (95% CI) | p | months | (95% CI) | p | months | (95% CI) | p | ||||||||
| DFS | AA | n = 402 | 90.74 | (82.74–98.74) | 0.015 | n = 215 | 91.29 | (81.57–101.02) | 0.887 | n = 185 | 90.1 | (76.26–103.94) | 0.006 | n = 196 | 97.96 | (88.65–107.27) | 0.111 | n = 205 | 81.51 | (68.10–94.93) | 0.095 | n = 150 | 87.64 | (71.58–103.7) | 0.018 |
| AC | 78.62 | (71.07–86.17) | 92.03 | (83.09–100.97) | 65.01 | (53.59–76.43) | 94.26 | (85.06–103.47) | 64.82 | (53.97–75.67) | 60.49 | (47.96–73.02) | |||||||||||||
| CC | 93.24 | (83.98–102.49) | 95.11 | (82.86–107.36) | 89.36 | (74.83–103.88) | 108.65 | (99.69–117.61) | 78.42 | (63.78–93.07) | 85.4 | (67.92–102.89) | |||||||||||||
| AA/CC | 91.77 | (85.71–97.82) | 0.004 | 92.82 | (85.23–100.41) | 0.64 | 89.63 | (79.59–99.68) | 0.001 | 102.05 | (95.30–108.80) | 0.095 | 80.00 | (70.08–89.92) | 0.032 | 86.4 | (74.51–98.30) | 0.005 | |||||||
| AC | 78.62 | (71.07–86.17) | 92.03 | (83.09–100.97) | 65.01 | (53.59–76.43) | 94.26 | (85.06–103.47) | 64.82 | (53.97–75.67) | 60.49 | (47.96–73.02) | |||||||||||||
| AA/AC | 83.92 | (78.37–89.47) | 0.114 | 91.79 | (85.19–98.38) | 0.742 | 73.88 | (64.77–83.00) | 0.091 | 96.13 | (89.60–102.67) | 0.06 | 71.1 | (62.53–79.67) | 0.421 | 69.54 | (59.28–79.80) | 0.133 | |||||||
| CC | 93.24 | (83.98–102.49) | 95.11 | (82.86–107.36) | 89.36 | (74.83–103.88) | 108.65 | (99.69–117.61) | 78.42 | (63.78–93.07) | 85.4 | (67.92–102.89) | |||||||||||||
| CC/AC | 83.81 | (77.88–89.75) | 0.126 | 93.09 | (85.86–100.32) | 0.855 | 73.02 | (63.75–82.29) | 0.068 | 99.59 | (92.78–106.39) | 0.996 | 69.51 | (60.74–78.29) | 0.128 | 68.14 | (57.67–78.61) | 0.093 | |||||||
| AA | 90.74 | (82.74–98.74) | 91.29 | (81.57–101.02) | 90.1 | (76.26–103.94) | 97.96 | (88.65–107.27) | 81.51 | (68.10–94.93) | 87.64 | (71.58–103.7) | |||||||||||||
| OS | AA | n = 409 | 100.73 | (93.74–107.72) | 0.12 | n = 218 | 99.00 | (90.07–107.94) | 0.381 | n = 189 | 103.82 | (92.73–114.92) | 0.037 | n = 199 | 106.84 | (99.15–114.53) | 0.309 | n = 209 | 92.75 | (80.48–105.02) | 0.238 | n = 154 | 102.62 | (89.83–115.41) | 0.052 |
| AC | 93.02 | (85.77–100.26) | 102.85 | (94.67–111.03) | 82.99 | (71.32–94.66) | 104.93 | (96.69–113.18) | 81.91 | (70.69–93.12) | 80.18 | (67.04–93.31) | |||||||||||||
| CC | 104.15 | (96.24–112.06) | 107.54 | (98.17–116.90) | 99.42 | (85.9–112.95) | 113.35 | (106.05–120.65) | 94.47 | (80.80–108.13) | 98.81 | (82.48–115.15) | |||||||||||||
| AA/CC | 102.12 | (96.87–107.37) | 0.05 | 102.85 | (94.67–111.03) | 0.918 | 101.63 | (92.90–110.36) | 0.012 | 109.39 | (103.89–114.90) | 0.457 | 93.49 | (84.35–102.63) | 0.091 | 100.69 | (90.38–111.00) | 0.017 | |||||||
| AC | 93.02 | (85.77–100.26) | 82.99 | (71.32–94.66) | 104.93 | (96.69–113.18) | 81.91 | (70.69–93.12) | 80.18 | (67.04–93.31) | |||||||||||||||
| AA/AC | 96.37 | (91.25–101.48) | 0.136 | 100.94 | (94.89–106.99) | 0.23 | 90.55 | (81.93–99.18) | 0.398 | 105.83 | (100.18–111.48) | 0.126 | 86.03 | (77.61–94.45) | 0.361 | 87.96 | (78.09–97.84) | 0.357 | |||||||
| CC | 104.15 | (96.24–112.06) | 107.54 | (98.17–116.90) | 99.42 | (85.90–112.95) | 113.35 | (106.05–120.65) | 94.47 | (80.80–108.13) | 98.81 | (82.48–115.15) | |||||||||||||
| CC/AC | 97.23 | (91.79–102.67) | 0.527 | 104.62 | (98.39–110.84) | 0.246 | 88.55 | (79.48–97.62) | 0.06 | 108.25 | (102.44–114.07) | 0.527 | 86.53 | (77.80–95.26) | 0.357 | 86.22 | (75.77–96.67) | 0.087 | |||||||
| AA | 100.73 | (93.74–107.72) | 99.00 | (90.07–107.94) | 103.82 | (92.73–114.92) | 106.84 | (99.15–114.53) | 92.75 | (80.48–105.02) | 102.62 | (89.83–115.41) | |||||||||||||
| rs1052133 | rs207454 | rs3736729 | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Variables | AA | AC | CC | AA | AC | CC | AA | AC | CC |
| Age | 0.147 | 0.227 | 0.593 | ||||||
| Estrogen receptor | 0.592 | 0.201 | 0.287 | ||||||
| Progesterone receptor | 0.199 | 0.538 | 0.499 | ||||||
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Rodrigues, P.; Furriol, J.; Bermejo, B.; Chaves, F.J.; Lluch, A.; Eroles, P. Identification of Candidate Polymorphisms on Stress Oxidative and DNA Damage Repair Genes Related with Clinical Outcome in Breast Cancer Patients. Int. J. Mol. Sci. 2012, 13, 16500-16513. https://doi.org/10.3390/ijms131216500
Rodrigues P, Furriol J, Bermejo B, Chaves FJ, Lluch A, Eroles P. Identification of Candidate Polymorphisms on Stress Oxidative and DNA Damage Repair Genes Related with Clinical Outcome in Breast Cancer Patients. International Journal of Molecular Sciences. 2012; 13(12):16500-16513. https://doi.org/10.3390/ijms131216500
Chicago/Turabian StyleRodrigues, Patricia, Jessica Furriol, Begoña Bermejo, Felipe Javier Chaves, Ana Lluch, and Pilar Eroles. 2012. "Identification of Candidate Polymorphisms on Stress Oxidative and DNA Damage Repair Genes Related with Clinical Outcome in Breast Cancer Patients" International Journal of Molecular Sciences 13, no. 12: 16500-16513. https://doi.org/10.3390/ijms131216500
APA StyleRodrigues, P., Furriol, J., Bermejo, B., Chaves, F. J., Lluch, A., & Eroles, P. (2012). Identification of Candidate Polymorphisms on Stress Oxidative and DNA Damage Repair Genes Related with Clinical Outcome in Breast Cancer Patients. International Journal of Molecular Sciences, 13(12), 16500-16513. https://doi.org/10.3390/ijms131216500




