Non Coding RNAs and Viruses in the Framework of the Phylogeny of the Genes, Epigenesis and Heredity
Abstract
:1. Introduction
2. Results and Discussion
2.1. The Discovery and the Origin of Non-Coding RNAs
2.2. Functions of Non Coding RNAs
2.3. The Non-Coding RNAs and Their Relationship with Epigenesis and Genomic Imprinting
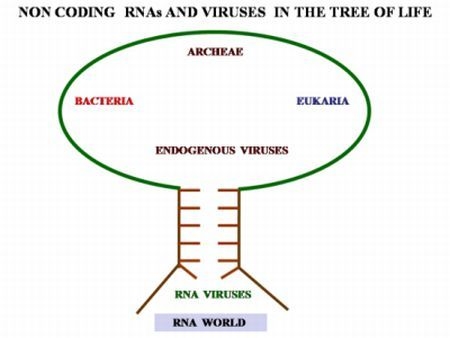
2.4. The Non-Coding RNAs and Their Relationships with Viruses
2.5. Sexuality Between Viruses and Related Genetic Elements
3. Conclusions
Acknowledgments
- Conflict of InterestThe author declares no conflict of interest.
References
- Crick, F. Central dogma of molecular biology. Nature 1970, 227, 561–563. [Google Scholar]
- Benzer, S. Fine structure of a genetic region in bacteriophage. Science 1955, 41, 344–354. [Google Scholar]
- Benzer, S. On the topology of the genetic fine structure. Proc. Natl. Acad. Sci. USA 1959, 45, 1607–1620. [Google Scholar]
- Yanofsky, C. Gene structure and protein structure. Sci. Am 1967, 216, 80–94. [Google Scholar]
- Taft, R.J.; Pheasant, M.; Mattick, J.S. The relationship between non-protein-coding DNA and eukaryotic complexity. Bioessays 2007, 29, 288–299. [Google Scholar]
- Mattick, J.S. Non-coding RNAs: The architects of eukaryotic complexity. EMBO Rep 2001, 2, 986–991. [Google Scholar]
- Frías, L.D. La hetrocromatina y su rol Functional: ¿Qué es un gen? : Desde el Dogma Central de la Biología Molecular hasta la Secuenciación del Genoma Humano (in Spanish); RIL Editores: Santiago, Chile, 2004; pp. 1–148. [Google Scholar]
- Frías, D.L. The history of the mendelian gene. Riv. Biol 2007, 100, 69–92. [Google Scholar]
- Frías, L.D. The phylogeny of the gene, from “Lower” naked structural genes to “Higher” non-transcriptional genes. Riv. Biol 2007, 100, 221–246. [Google Scholar]
- Frías, L.D. Omissions in the synthetic theory of evolution. Biol. Res 2010, 43, 299–306. [Google Scholar]
- Pearson, H. What is a gene? Nature 2006, 441, 399–401. [Google Scholar]
- Parris, G. A Hypothetical master development program for multi-cellular organisms: Ontogeny and phylogeny. Bioscience Hypotheses 2009, 2, 3–12. [Google Scholar]
- Gilbert, W. The RNA world. Nature 1986, 319, 618–618. [Google Scholar]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet 2009, 10, 155–159. [Google Scholar]
- Eddy, S.R. Non-coding RNA genes and the modern RNA world. Nat. Rev. Genet 2001, 12, 919–929. [Google Scholar]
- Mattick, J.S.; Makunin, I.V. Non-coding RNA. Hum. Mol. Genet 2006, 1, 17–29. [Google Scholar]
- Berget, S.M.; Moore, C.; Sharp, P.A. Spliced segments at the 5′ termine of adenoviruses 2 late mRNA. Proc. Natl. Acad. Sci. USA 1977, 74, 3171–3175. [Google Scholar]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The Caenorhabditis elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in Caenorhabditis elegans. Cell 1993, 75, 855–862. [Google Scholar]
- Teixeira, A.; Tahiri-Alaoui, A.; West, S.; Thomas, B.; Ramadass, A.; Martianov, I.; Dye, M.; James, W.; Proudfoot, N.J.; Akoulitchev, A. Autocatalytic RNAcleavage in the human beta-globin pre-mRNA promotes transcriptiontermination. Nature 2004, 432, 526–530. [Google Scholar]
- Salehi-Ashtiani, K.; Lupták, A.; Litovchick, A.; Szostak, J.W. A genomewide search for ribozymes reveals an HDV-like sequence in the human CPEB3 gene. Science 2006, 313, 1788–1792. [Google Scholar]
- Martick, M.; Horan, L.H.; Noller, H.F.; Scott, W.G. A discontinuous hammerhead ribozyme embedded in a mammalian messenger RNA. Nature 2008, 454, 899–902. [Google Scholar]
- de la Peña, M.; García-Robles, I. Intronic hammerhead ribozymes are ultraconserved in the human genome. EMBO Rep 2010, 11, 711–716. [Google Scholar]
- Prody, G.A.; Bakos, J.T.; Buzayan, J.M.; Schneider, I.R.; Bruening, G. Autolytic processing of dimeric plant virus satellite RNA. Science 1986, 231, 1577–1580. [Google Scholar]
- Palmer, J.D.; Logsdon, J.M., Jr. The recent origins of introns. Curr. Opin. Genet. Dev 1991, 4, 470–477. [Google Scholar]
- Cavalier-Smith, T. Intron phylogeny: A new hypothesis. Trends Genet. 1991, 7, 145–148. [Google Scholar]
- Filée, J.; Forterre, P.; Lauren, J. The role played by viruses in the evolution of their hosts: A view based on informational protein phylogenies. Res. Microbiol 2003, 154, 237–243. [Google Scholar]
- Zheng, Z.M. Viral oncogenes, noncoding RNAs, and RNA splicing in human tumor viruses. Int. J. Biol. Sci 2010, 6, 730–755. [Google Scholar]
- Zheng, Z.M.; Baker, C.C. Papillomavirus genome structure, expression, and post-transcriptional regulation. Front. Biosci 2006, 11, 2286–2302. [Google Scholar]
- Xiong, Y.; Eickbush, T.H. Similarity of reverse transcriptase-like sequences of viruses, transposable elements, and mitochondrial introns. Mol. Biol. Evol 1988, 5, 675–690. [Google Scholar]
- Fischer, M.G.; Suttle, C.A. A virophage at the origin of large DNA transposons. Science 2011, 332, 231–234. [Google Scholar]
- la Scola, B.; Desnues, Ch.; Pagnier, I.; Robert, C.; Barrassi, L.; Fourmous, G.; Merchant, M.; Susan-Monti, M.; Forterre, P.; Koonin, E.; et al. The virophage as a unique parasite of the giant mimivirus. Nature 2008, 455, 100–104. [Google Scholar]
- Diener, T.O. Potato spindle tuber virus. IV. A replicating, low molecular weight RNA. Virology 1971, 45, 411–428. [Google Scholar]
- Navarro, B.; Flores, R. Chrysanthemum chlorotic mottle viroid: Unusual structural properties of a subgroup of self-cleaving viroids with hammerhead ribozymes. Proc. Natl. Acad. Sci. USA 1997, 94, 11262–11267. [Google Scholar]
- Diener, T.O. Circular RNAs: Relics of precellular evolution. Proc. Natl. Acad. Sci. USA 1989, 86, 9370–9374. [Google Scholar]
- Smalheiser, N.R.; Torvik, V.I. Alu elements within human mRNAs are probable micro RNA targets. Trends Genet 2006, 22, 532–536. [Google Scholar]
- Smalheiser, N.R.; Torvik, V.I. Mammalian microRNAs derived from genomic repeats. Trends Genet 2005, 21, 322–326. [Google Scholar]
- Berger, A.; Strub, K. Multiple roles of Alu-related non-coding RNAs. Prog. Mol. Subcell. Biol 2011, 51, 119–146. [Google Scholar]
- Stort, G. An expanding universe of noncoding RNAs. Science 2002, 296, 1260–1263. [Google Scholar]
- Ryan, J.; Taft, R.J.; Pang, K.C.; Mercer, T.R.; Dinger, M.; Mattick, J.S. Non-coding RNAs: Regulators of disease. J. Pathol 2002, 220, 126–139. [Google Scholar]
- Sánchez, L. Sex determining mechanism in insects. Int. J. Dev. Biol 2008, 52, 837–856. [Google Scholar]
- Taft, R.J.; Pang, K.C.; Mercer, T.R.; Dinger, M.; Mattick, J.S. Non-coding RNAs: Regulators of disease. J. Pathol 2010, 220, 126–139. [Google Scholar]
- Munker, R.; Calin, G.A. MicroRNA profiling in cancer. Clin. Sci 2011, 121, 141–158. [Google Scholar]
- Backofen, R.; Bernhart, S.H.; Flamm, C.; Fried, C.; Fritzsch, G.; Hackermuller, J.; Hertel, J.; Hofacker, I.L.; Missal, K.; Mosig, A.; et al. RNAs everywhere: Genome-wide annotation of structured RNAs. J. Exp. Zool. B 2007, 308, 1–25. [Google Scholar]
- Smalheiser, N.R.; Lugli, G. MicroRNA regulation of synaptic plasticity. Neuromol. Med 2009, 11, 133–140. [Google Scholar]
- Bredy, T.W.; Lin, Q.; Wei, W.; Baker-Andresen, D.; Mattick, J. MicroRNA regulation of neural plasticity and memory. Neurobiol. Learn. Mem 2011, 96, 89–94. [Google Scholar]
- Smalheiser, N.R.; Lugli, G.; Thimmapuram, J.; Cook, E.H.; Larson, J. Endogenous siRNAs and noncoding RNA-derived small RNAs are expressed in adult mouse hippocampus and are up-regulated in olfactory discrimination training. RNA 2011, 17, 166–181. [Google Scholar]
- Cazalla, D.; Yario, T.; Steitz, J.A. Down-regulation of a host microRNA by a Herpesvirus saimiri noncoding RNA. Science 2010, 328, 1563–1566. [Google Scholar]
- Pijlman, G.P.; Funk, A.; Kondratieva, N.; Leung, J.; Torres, S.; van der Aa, L.; Liu, W.J.; Palmenberg, A.C.; Shi, P.Y.; Hall, R.A.; et al. A highly structured, nuclease-resistant, noncoding RNA produced by flaviviruses is required for pathogenicity. Cell Host Microbe 2008, 4, 579–591. [Google Scholar]
- Koerner, M.V.; Pauler, F.M.; Huang, R.; Barlow, D.P. The function of non-coding RNAs in genomic imprinting. Development 2009, 136, 1771–1783. [Google Scholar]
- Satterlee, J.S.; Barbee, S.; Jin, P.; Krichevsky, A.; Salama, S.; Schratt, G.; Wu, D.Y. Noncoding RNAs in the brain. J. Neurosci 2007, 27, 11856–11859. [Google Scholar]
- Tian, D.; Sun, S.; Lee, J.T. The long noncoding RNA, Jpx, is a molecular switch for X chromosome inactivation. Cell 2010, 143, 390–403. [Google Scholar]
- Dinger, M.E.; Amaral, P.P.; Mercer, T.R.; Pang, K.C.; Bruce, S.J.; Gardiner, B.B.; Askarian-Amiri, M.E.; Ru, K.; Soldà, G.; Simons, C.; et al. Long noncoding RNAs in mouse embryonic stem cell pluripotency and differentiation. Genome Res 2008, 18, 1433–1445. [Google Scholar]
- Thakur, N.; Tiwari, V.K.; Thomassin, H.; Pandey, R.R.; Kanduri, M.; Gondor, A.; Grange, T.; Ohlsson, R.; Kanduri, C. An antisense RNA regulates the bidirectional silencing property of the Kcnq1 imprinting control region. Mol. Cell. Biol 2004, 24, 7855–7862. [Google Scholar]
- Kurakawa, R. Long noncoding RNAs as a regulator for transcription. Prog. Mol. Subcell. Biol 2011, 51, 29–41. [Google Scholar]
- Raman, R.P.; Kanduri, C. Transcriptional and post transcriptional programming by long noncoding RNAs. Prog. Mol. Subcell. Biol 2011, 51, 1–27. [Google Scholar]
- Azzalin, C.M.; Reichenbach, P.; Khoriauli, L.; Giulotto, E.; Lingner, J. Telomeric repeat containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 2007, 318, 798–801. [Google Scholar]
- Schoeftner, S.; Blasco, M.A. Developmentally regulated transcription of mammalian telomeres by DNA dependent RNA polymerase II. Nat. Cell Biol 2007, 10, 228–236. [Google Scholar]
- Arora, R.; Brun, C.M.C.; Azzalin, C.M. TERRA: Long noncoding RNAs at eukaryotic telomeres. Prog. Mol. Subcell. Biol 2011, 51, 65–94. [Google Scholar]
- Orom, U.A.; Shiekhattar, R. Long non-coding RNAs and enhancers. Curr. Opin Genet. Dev 2011, 21, 194–198. [Google Scholar]
- Ahmad, A.; Zhang, Y.; Cao, X.F. Decoding the epigenetic language of plant development. Mol. Plant 2010, 3, 719–728. [Google Scholar]
- Chuang, J.C.; Jones, P.A. Epigenetics and microRNAs. Pediatr. Res 2007, 61, 24R–29R. [Google Scholar]
- Autran, D.; Huanca-Mamani, W.; Calzada, J.P.V. Genomic imprinting in plants: The epigenetic version of an Oedipus complex. Curr. Opin. Plant Biol 2005, 8, 19–25. [Google Scholar]
- Anaka, M.; Lynn, A.; McGinn, P.; Lloyd, V.K. Genomic imprinting in Drosophila has properties of both mammalian and insect imprinting. Dev. Genes Evol 2009, 219, 59–66. [Google Scholar]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebert, B.E.; Mak, R.H.; Ferrando, A.A.; et al. MicroRNA expression profiles classify human cancers. Nature 2005, 435, 834–838. [Google Scholar]
- Benne, R.; van den Burg, J.; Brakenhoff, J.P.J.; Sloof, P.; van Boom, J.H.; Tromp, M.C. Major transcript of the frameshifted coxII gene from Trypanosome mitochondria contains four nucleotides that are not incoded in the DNA. Cell 1986, 46, 819–826. [Google Scholar]
- Benne, R. RNA editing in trypanosomes. Mol. Biol. Rep 1992, 16, 217–227. [Google Scholar]
- Lonergan, K.M.; Gray, M.W. Editing of transfer RNAs in Acanthamoeba castellanii mitochondria. Science 1993, 259, 812–816. [Google Scholar]
- Rubio, M.A.T.; Pastar, I.; Gaston, K.W.; Ragone, F.L.; Janzen, C.J.; Cross, G.A.M.; Papavasiliou, F.N.; Alfonzo, J.D. An adenosine-to inosine tRNA-editing enzyme that can perform C-to-U deamination of DNA. Proc. Natl. Acad. Sci. USA 2007, 104, 7821–7826. [Google Scholar]
- Luciano, D.J.; Mirsky, H.; Vendetti, N.J.; Maas, S. RNA editing of a miRNA precursor. RNA 2004, 10, 1174–1177. [Google Scholar]
- Liang, H.; Landweber, L.F. Hypothesis: RNA editing of microRNA target sites in human? RNA 2007, 13, 463–467. [Google Scholar]
- Gott, J.M.; Emerson, R.B. Functions and mechanisms of RNA editing. Annu. Rev. Genet 2000, 34, 499–531. [Google Scholar]
- Blanc, V.; Davidson, N.O. C-to-U RNA editing: Mechanisms leading to genetic diversity. J. Biol. Chem 2003, 278, 1395–1398. [Google Scholar]
- Jobson, R.W.; Qiu, Y.-L. Did RNA editing in plant organellar genomes originate under natural selection or through genetic drift. Biol. Direct 2008, 3. [Google Scholar] [CrossRef]
- Mehler, M.F.; Mattick, J.S. Noncoding RNAs and RNA editing in brain development, functional diversification, and neurological disease. Physiol. Rev 2007, 87, 799–823. [Google Scholar]
- Reik, W.; Walter, J. Genomic imprinting: Parental influence in the genome. Nat. Rev. Genet 2001, 2, 21–32. [Google Scholar]
- Úbeda, F.; Gardner, A. A model for genomic imprinting in the social brain: Juvenile. Evolution 2010, 64, 2587–2600. [Google Scholar]
- Li, E.; Beard, C.; Jaenisch, R. Role for DNA methylation in genomic imprinting. Nature 1993, 366, 362–365. [Google Scholar]
- Filipowicz, W. RNAi: The nuts and bolts of the RISC machine. Cell 2005, 122, 17–20. [Google Scholar]
- Tang, G. siRNA and miRNA: An insight into RISCs. Trends Biochem. Sci 2005, 30, 106–114. [Google Scholar]
- Royo, H.; Cavaillé, J. Non-coding RNAs in imprinted gene clusters. Biol. Cell 2008, 100, 149–166. [Google Scholar]
- Zhang, Y.; Qu, L. Non-coding RNAs and the acquisition of genomic imprinting in mammals. Sci. China C Life Sci 2009, 52, 195–204. [Google Scholar]
- María, L.V.S.; Currotto, B.L.; Cortés, F.M.; Rojas, C.B.; Alliende, M.A.R. Methylation, cytogenetic and FISH tests in the molecular diagnosis of Prader-Willi and Angelman síndromes. Rev. Med. Chile 2001, 129, 1–9. [Google Scholar]
- Skuse, D.H.; James, R.S.; Bishop, D.V.M.; Coppin, B.; Dalton, P.; Aamodt-Leeper, G.; Bacarese-Hamilton, M.; Creswell, C.; McGurk, R.; Jacob, P.A. Evidence from Turner’s Syndrome of an imprinted X-linked locus affecting cognitive function. Nature 1997, 387, 705–708. [Google Scholar]
- Wilkins, J.F.; Haig, D. Inbreeding, maternal care and genomic imprinting. J. Theor. Biol 2003, 221, 559–564. [Google Scholar]
- Dinter-Gottlieb, G. Viroids and virusoids are related to group I introns. Proc. Natl. Acad. Sci. USA 1986, 83, 6250–6254. [Google Scholar]
- Moreira, D.; Lópes-García, P. Ten reasons to excluded viruses from the tree of life. Nat. Rev. Microbiol 2009, 7, 306–311. [Google Scholar]
- Villarreal, L.P. How viruses shape the tree of life. Future Virol 2006, 1, 587–595. [Google Scholar]
- Villarreal, L.P.; Witzany, G. Viruses are essential agents within the roots and stem of the tree of life. J. Theor Biol 2010, 262, 698–710. [Google Scholar]
- Mindell, D.P.; Rest, J.S.; Villarreal, L. Cracraft, J., Donoghue, M.J., Eds.; Viruses the Tree of Life. In Assembling the Tree of Life; Oxford University Press: New York, NY, USA, 2004; pp. 107–118. [Google Scholar]
- Combelas, N.; Holmblat, B.; Joffret, M.L.; Colbére-Garapin, F.; Delpeyroux, F. Recombination between poliovirus and cocxackie A viruses of species C: A model of viral genetic plasticity and emergence. Viruses 2011, 3, 1460–1484. [Google Scholar]
- Worobey, M.; Holmes, E.C. Evolutionary aspects of recombination in RNA viruses. J. Gen. Virol 1999, 80, 2535–2543. [Google Scholar]
- Zinder, N.D.; Lederberg, J. Genetic exchange Salmonella. J. Bacteriol 1952, 64, 679–699. [Google Scholar]
- Levine, L. Bases físicas da Herença. In Biologia do Gene (in Portuguese); Levine, L., Ed.; Edgard Blucher: São Paulo, Brazil, 1977; pp. 41–49. [Google Scholar]
- Mieczkowski, P.; Francene, A.; Lemoine, J.; Thomas, D.; Pete, T.D. Recombination between retrotransposons as a source of chromosome rearrangements in the yeast Saccharomyces cerevisiae. DNA Repair 2006, 5, 1010–1020. [Google Scholar]
- Geuking, M.B.; Weber, J.; Dewannieux, M.; Gorelik, E.; Heidmann, T.; Hengartner, H.; Zinkernagel, R.M.; Hangartner, L. Recombination of retrotransposon and exogenous RNA virus results in nonretroviral cDNA integration. Science 2009, 323, 393–396. [Google Scholar]
- Liu, H.; Fu, Y.; Jiang, D.; Li, G.; Xie, J.; Cheng, J.; Peng, Y.; Ghabrial, S.A.; Yi, X. Widespread horizontal gene transfer from double-stranded RNA viruses to eukaryotic nuclear genomes. J. Virol 2010, 84, 11876–11887. [Google Scholar]
- Villarreal, L.P. Domingo, E., Webster, R., Hollabd, J., Eds.; DNA Virus Contribution to Host Evolution. In Origin and Evolution of Viruses; Academic Press: New York, NY, USA, 1999; pp. 391–420. [Google Scholar]
- Villarreal, L.P. Viruses and the Evolution of Life; American Society for Microbiology Press: Washington, DC, USA, 2005; p. 472. [Google Scholar]
- Petronis, A. Epigenetics as a unifying principle in the aetiology of complex traits and diseases. Nature 2010, 465, 721–727. [Google Scholar]
- van Blerkom, L.M. Role of viruses in human evolution. Yearb. Phys. Anthropol 2003, 46, 14–46. [Google Scholar]
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Frías-Lasserre, D. Non Coding RNAs and Viruses in the Framework of the Phylogeny of the Genes, Epigenesis and Heredity. Int. J. Mol. Sci. 2012, 13, 477-490. https://doi.org/10.3390/ijms13010477
Frías-Lasserre D. Non Coding RNAs and Viruses in the Framework of the Phylogeny of the Genes, Epigenesis and Heredity. International Journal of Molecular Sciences. 2012; 13(1):477-490. https://doi.org/10.3390/ijms13010477
Chicago/Turabian StyleFrías-Lasserre, Daniel. 2012. "Non Coding RNAs and Viruses in the Framework of the Phylogeny of the Genes, Epigenesis and Heredity" International Journal of Molecular Sciences 13, no. 1: 477-490. https://doi.org/10.3390/ijms13010477
APA StyleFrías-Lasserre, D. (2012). Non Coding RNAs and Viruses in the Framework of the Phylogeny of the Genes, Epigenesis and Heredity. International Journal of Molecular Sciences, 13(1), 477-490. https://doi.org/10.3390/ijms13010477