Cloning, Soluble Expression and Purification of High Yield Recombinant hGMCSF in Escherichia coli
Abstract
:1. Introduction
2. Results and Discussion
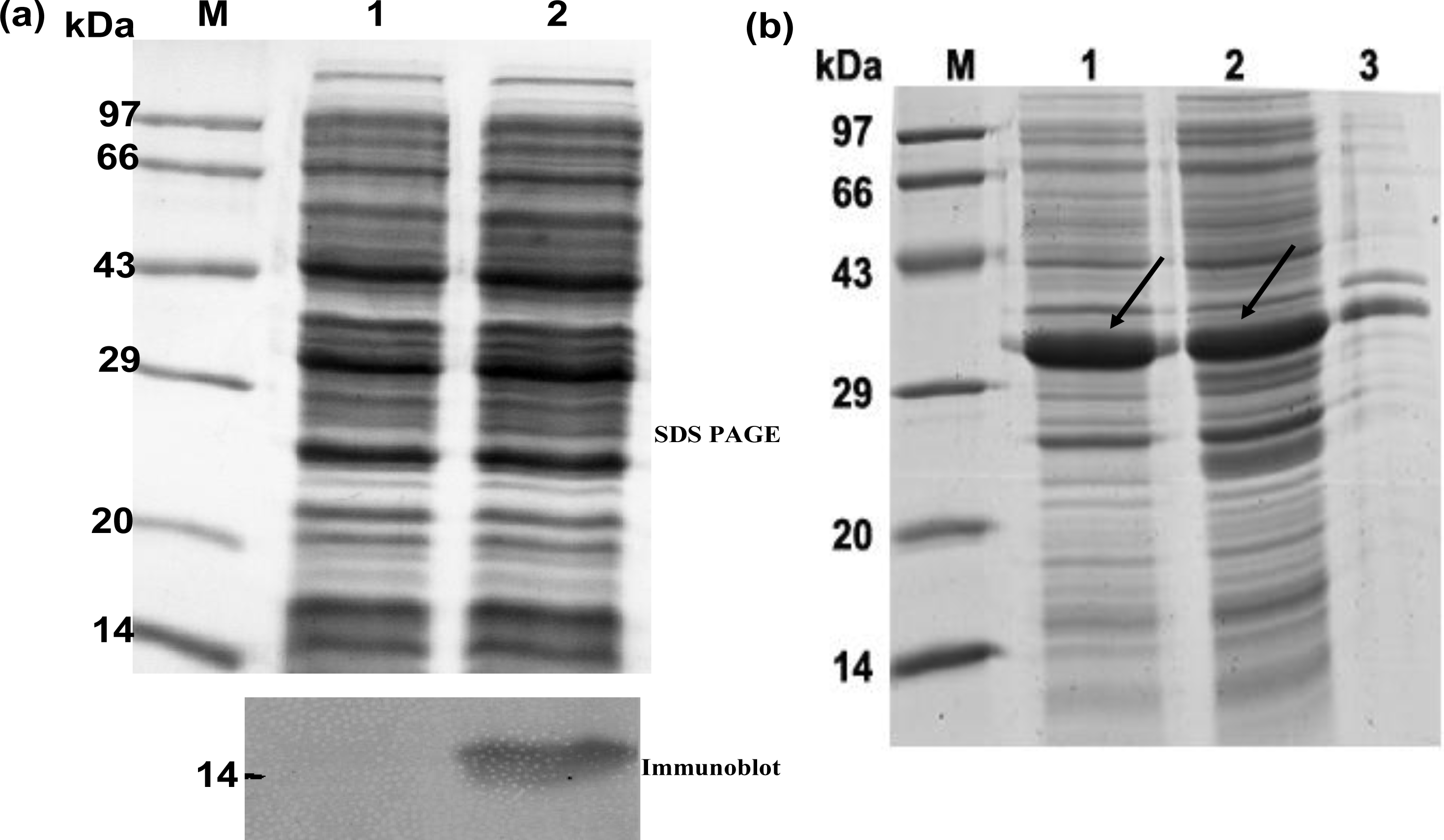
2.1. Cloning and Expression of hGMCSF
2.2. Purification of TRX-hGMCSF Followed by Separation of hGMCSF from the Fusion Tag
3. Experimental Section
3.1. Cloning of hGMCSF in pET21a Vector and in pET32a as a TRX Fusion Tag
3.2. Expression of pET21a-rhGMCSF and pET32a-rhGMCSF in Shake Flask
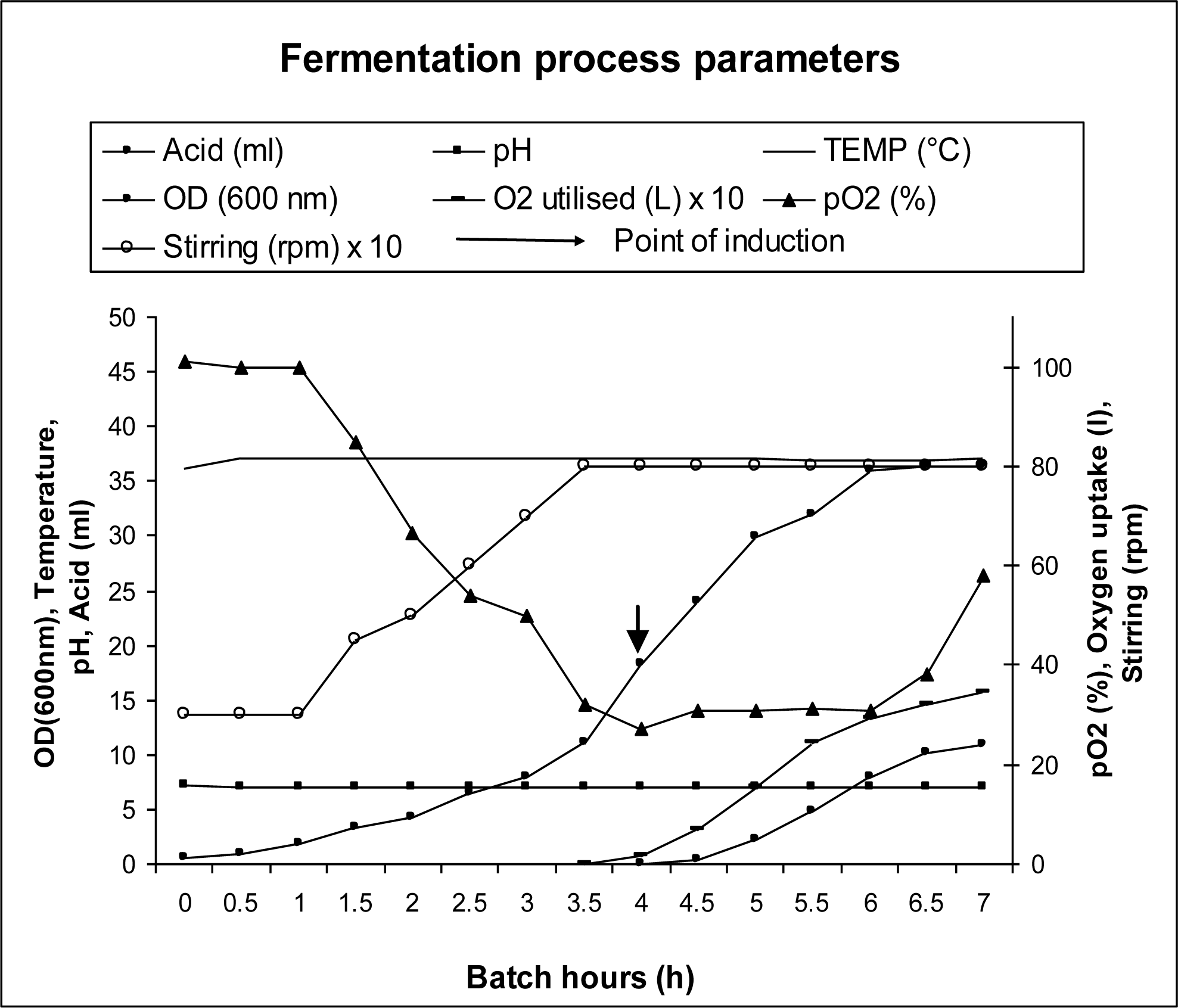
3.3. Bioreactor Studies
3.4. Purification hGMCSF Using Ni2+-NTA Column
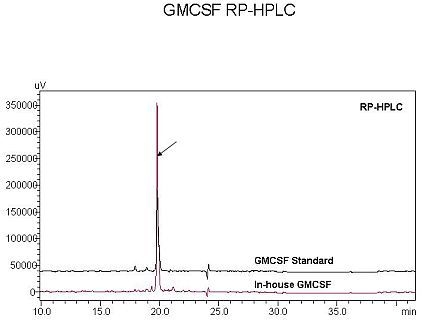
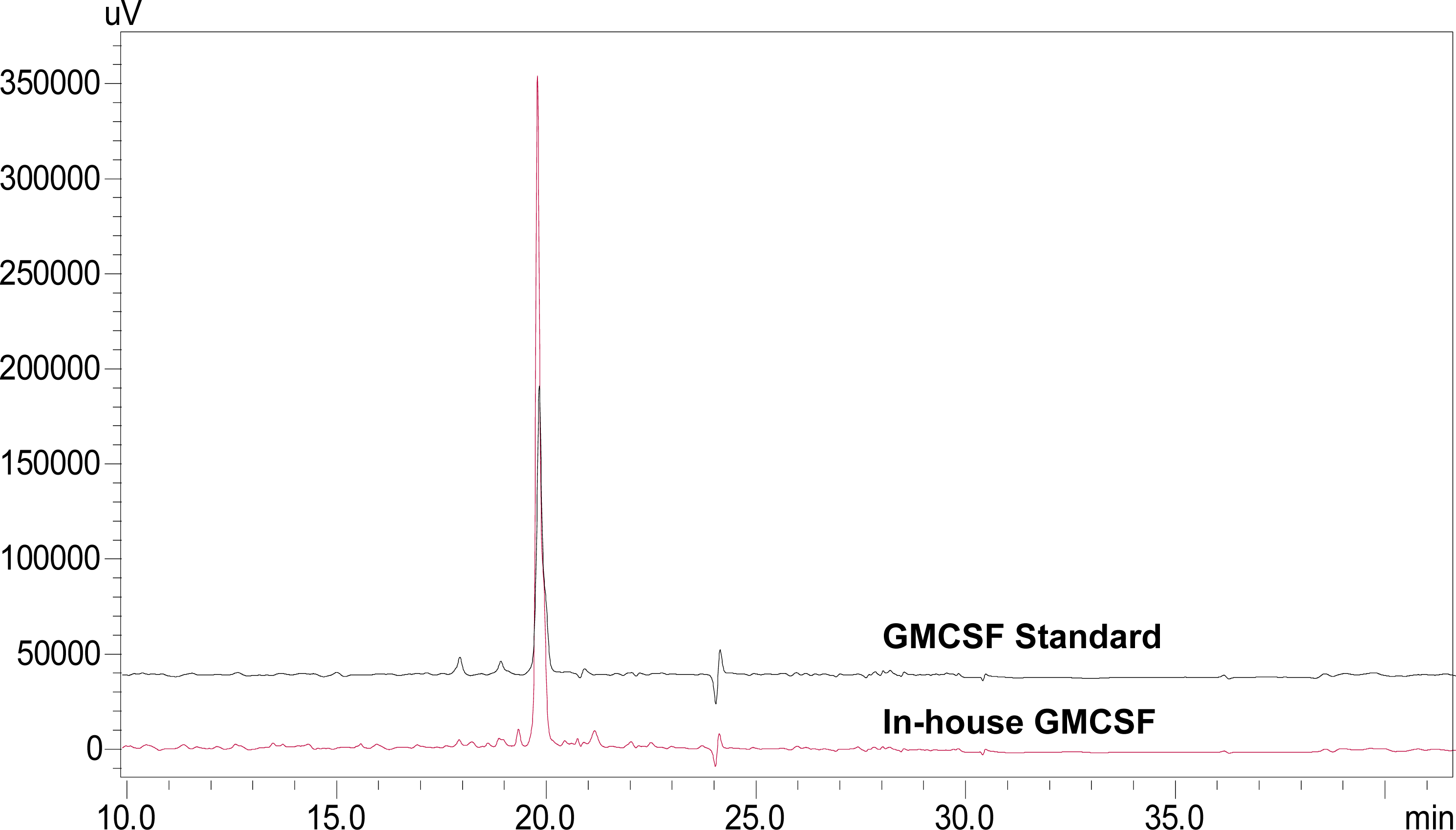
3.5. Characterization of rhGMCSF by RP-HPLC
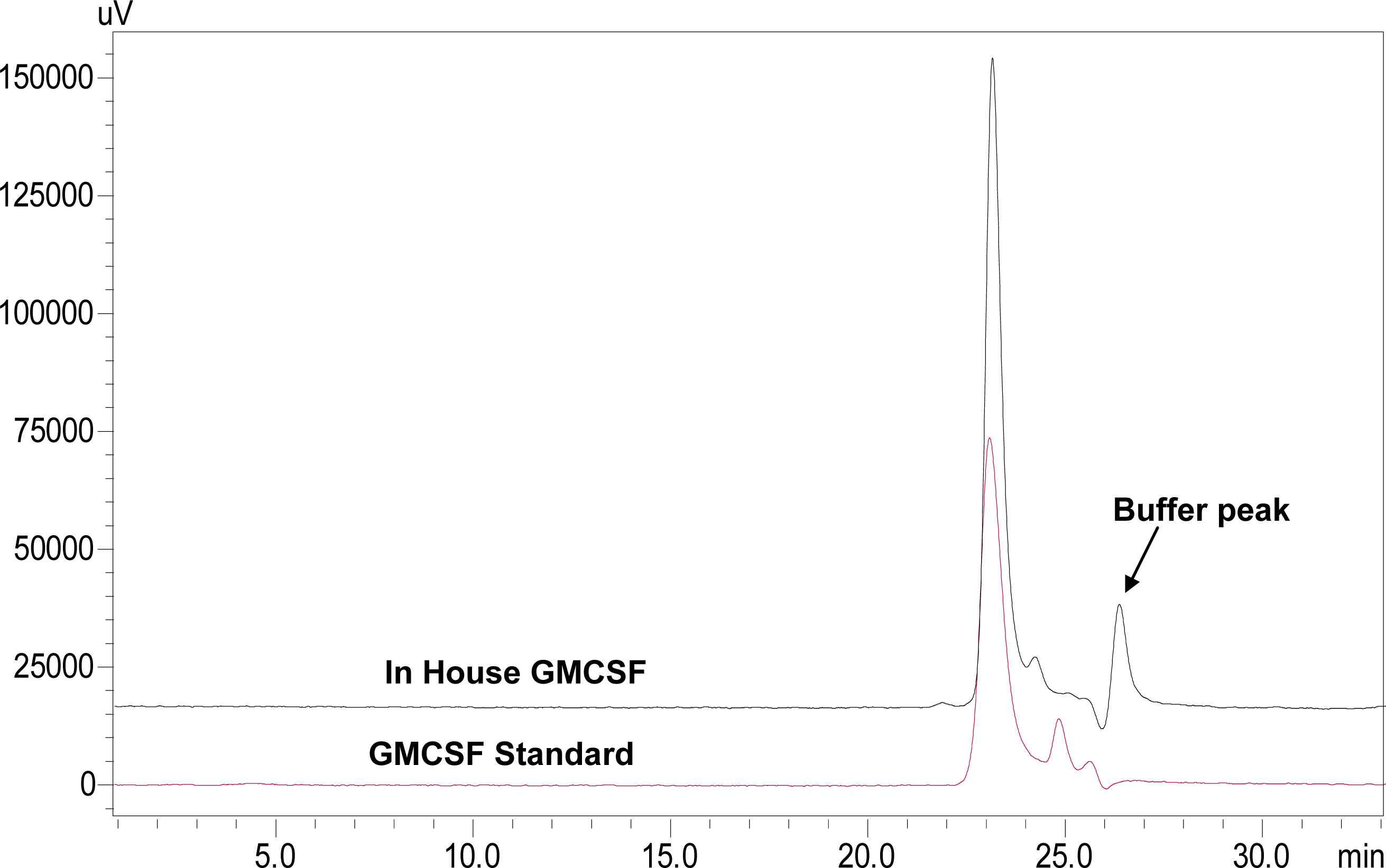
3.6. Characterization of rhGMCSF by SE-HPLC
3.7. Western Blot Analysis
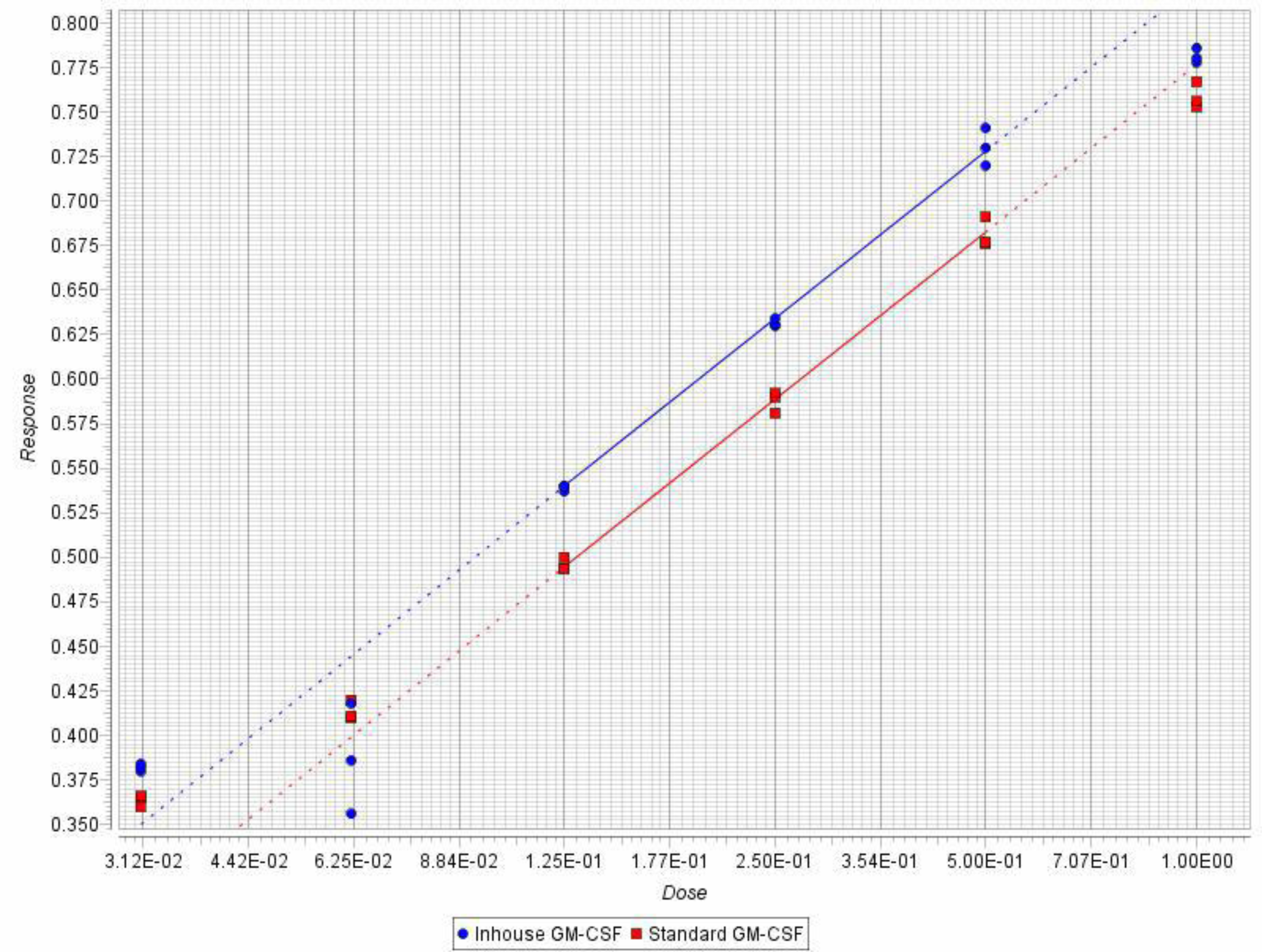
3.8. Bioassay for hGMCSF
4. Conclusions
Acknowledgments
References
- Schwanke, RC; Renard, G; Chies, JM; Campos, MM; Batista, EL, Jr; Santos, DS; Basso, LA. Molecular cloning, expression in Escherichia coli and production of bioactive homogeneous recombinant human granulocyte and macrophage colony stimulating factor. Int. J. Biol. Macromol 2009, 45, 97–102. [Google Scholar]
- Armitage, JO. Emerging applications of recombinant human granulocyte-macrophage colony-stimulating factor. EMBO J 1985, 4, 2575–2581. [Google Scholar]
- DeLamarter, JF; Mermod, JJ; Liang, CM; Eliason, JF; Thatcher, DR. Recombinant murine GM-CSF from E. coli has biological activity and is neutralized by a specific antiserum. EMBO J 1985, 4, 2575–2581. [Google Scholar]
- Bhopale, GM; Nanda, RK. Recombinant DNA expression products for human therapeutic use. Curr. Sci 2005, 89, 614–622. [Google Scholar]
- Berrow, NS; Bussow, K; Coutard, B; Diprosw, J; Ekberg, M. Recombinant protein expression and solubility screening in E. coli: A comparative study. Acta Crystallogr 2006, 67, 1218–1226. [Google Scholar]
- Sorenson, HP; Mortensen, KK. Advanced genetic strategies for recombinant protein expression in E. coli. Biotechnology 2005, 115, 13–28. [Google Scholar]
- Sahdev, S; Khattar, SK; Saini, KS. Production of active eukaryotic proteins through bacterial expression system: A review of the existing biotechnology strategies. Mol. Cell Biochem 2008, 307, 249–264. [Google Scholar]
- Yin, J; Li, G; Ren, X; Herrler, GA. Comparative evaluation of the advantages and limitations of frequently used expression systems for foreign genes. J. Biotechnol 2007, 127, 335–347. [Google Scholar]
- Valeria, DM; Gunter, S; Stephanie, B; Ario, DM. The solubility and stability of recombinant proteins are increased by their fusion to NusA. Biochem. Biophys. Res. Commun 2004, 322, 766–771. [Google Scholar]
- Cabrita, LD; Dai, W; Bottomley, SP. A family of E. coli expression vectors for laboratory scale and high throughput soluble protein production. BMC Biotechnol 2006, 6, 12–19. [Google Scholar]
- Srinivasa, BK; Muthukumaran, T; Antony, A; Samuel, SDPS; Balamurali, M; Murugan, V; Meenakshisundaram, S. Single step intein-mediated purification of hGMCSF expressed in salt-inducible E. coli. Biotechnol. Lett 2009, 31, 659–664. [Google Scholar]
- Lee, AY; Chung, HK; Kyong Bae, E; Hwang, JS; Sung, BW; Cho, CW; Kim, JK; Lee, K; Han, JY; Lee, CT; Youn, HJ. A recombinant human G-CSF/GM-CSF fusion protein from E. coli showing colony stimulating activity on human bone marrow cells. Biotechnol. Lett 2003, 25, 205–211. [Google Scholar]
- Smith, HE. Transcriptional response of E. coli to recombinant protein insolubility. J. Struct. Funct. Genomics 2007, 8, 27–35. [Google Scholar]
- Banerjee, S; Apte-Deshpande, A; Mandi, N; Padmanabhan, S. A novel cytokine derived fusion tag for over-expression of heterologous proteins in E. coli. Int. J Biol. Life Sci 2009, 1, 139–143. [Google Scholar]
- Salunkhe, S; Prasad, B; Sabnis-Prasad, K; Apte-Deshpande, A; Padmanabhan, S. Expression and purification of SAK-fused human interferon alpha in Escherichia coli. J. Microbial. Biochem. Technol 2009, 1, 5–10. [Google Scholar]
- Harrison, RG. Expression of soluble heterologous proteins via fusion with NusA protein. Innovations 2000, 11, 4–7. [Google Scholar]
- Ackland, CE; Berndt, WG; Frezza, JE; Landgraf, BE; Pritchard, KW; Ciardelli, TL. Monitoring of protein conformation by high-performance size-exclusion liquid chromatography and scanning diode array second-derivative UV absorption spectroscopy. J. Chromatogr. A 1991, 540, 187–198. [Google Scholar]
- Kaushansky, K; O’Hara, PJ; Berkner, K; Segal, GM; Hagen, S; Adamson, JW. Genomic cloning, characterization, and multilineage growth-promoting activity of human granulocyte-macrophage colony-stimulating factor. Proc. Natl. Acad. Sci. USA 1986, 83, 3101–3105. [Google Scholar]
- Moonen, P. Increased biological activity of deglycosylated recombinant human granulocyte/macrophage colony-stimulating factor produced by yeast or animal cells. Proc. Natl. Acad. Sci. USA 1987, 84, 4428–4431. [Google Scholar]
- Wong, GG; Witek, JS; Temple, PA; Wilkens, KM; Leary, AC; Luxenberg, DP; Jones, SS; Brown, EL; Kay, RM; Orr, EC. Human GM-CSF: Molecular cloning of the complementary DNA and purification of the natural and recombinant proteins. Science 1985, 228, 810–815. [Google Scholar]
- Miyajima, A; Otsn, K; Schreurs, J; Bond, MW; Abrams, JS; Arai, K. Expression of murine and human granulocyte-macrophage colony-stimulating factors in S. cerevisiae: Mutagenesis of the potential glycosylation sites. EMBO J 1986, 5, 1193–1197. [Google Scholar]
- Burgess, AW; Begley, CG; Johnson, GR; Lopez, AF; Williamson, DJ; Mermod, JJ; Simpson, RJ; Schmitz, A; de Lamarter, JF. Purification and properties of bacterially synthesized human granulocyte-macrophage colony stimulating factor. Blood 1987, 69, 43–51. [Google Scholar]
- Belew, M; Zhou, Y; Wang, S; Nystrom, LE; Janson, JC. Purification of recombinant human granulocyte-macrophage colony-stimulating factor from the inclusion bodies produced by transformed Escherichia coli cells. J. Chromatogr 1994, 679, 67–83. [Google Scholar]
- Srinivasa, BK; Antony, A; Muthukumaran, T; Meenakshisundaram, S. Construction of intein-mediated hGMCSF expression vector and its purification in Pichia pastoris. Protein Expr. Purif 2008, 57, 201–205. [Google Scholar]
- Cowgill, C; Ozturk, AG; John, R. Protein Refolding and Scale Up; Shukla, AA, Etzel, MR, Gadam, S, Eds.; Taylor & Francis: New York, NY, USA, 2007; pp. 124–158. [Google Scholar]
- Sletta, H; Tondervik, A; Hakvag, S; Aune, TEV; Nedal, A; Aune, R; Evensen, G; Valla, S; Ellingsen, TE; Brautaset, T. The presence of N-terminal Secretion signal sequences leads to strong stimulation of the total expression levels of three tested medically important proteins during high-cell-density cultivations of Escherichia coli. Appl. Environ. Microbiol 2007, 73, 906–912. [Google Scholar]
- Mandi, N; Soorapaneni, S; Rewanwar, S; Kotwal, P; Prasad, BJ; Mandal, G; Padmanabhan, S. High-yielding recombinant staphylokinase in bacterial expression system-cloning, expression, purification and activity studies. Protein Expr. Purif 2009, 64, 69–75. [Google Scholar]

| Fraction | Volume (mL) | Protein (mg/mL) | Total Protein (mg) | Total Activity (U) | Specific Activity (U/mg) | Fold Purification | %Yield |
|---|---|---|---|---|---|---|---|
| Crude cell lysate | 10 | 2.39 | 23.9 | 2.19 × 108 | 9.16 × 106 | 1.0 | 100 |
| Purified TRX-GMCSF fusion | 20 | 0.43 | 8.6 | 11.24 × 107 | 1.3 × 107 | 1.42 | 51 |
| Purified GMCSF | 20 | 0.22 | 4.4 | 10.20 × 107 | 2.3 × 107 | 2.5 | 46 |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Das, K.M.P.; Banerjee, S.; Shekhar, N.; Damodaran, K.; Nair, R.; Somani, S.; Raiker, V.P.; Jain, S.; Padmanabhan, S. Cloning, Soluble Expression and Purification of High Yield Recombinant hGMCSF in Escherichia coli. Int. J. Mol. Sci. 2011, 12, 2064-2076. https://doi.org/10.3390/ijms12032064
Das KMP, Banerjee S, Shekhar N, Damodaran K, Nair R, Somani S, Raiker VP, Jain S, Padmanabhan S. Cloning, Soluble Expression and Purification of High Yield Recombinant hGMCSF in Escherichia coli. International Journal of Molecular Sciences. 2011; 12(3):2064-2076. https://doi.org/10.3390/ijms12032064
Chicago/Turabian StyleDas, Krishna M.P., Sampali Banerjee, Nivedita Shekhar, Karpagavalli Damodaran, Rahul Nair, Sandeep Somani, Veena P. Raiker, Shweta Jain, and Sriram Padmanabhan. 2011. "Cloning, Soluble Expression and Purification of High Yield Recombinant hGMCSF in Escherichia coli" International Journal of Molecular Sciences 12, no. 3: 2064-2076. https://doi.org/10.3390/ijms12032064