Optimization of Enzymatic Production of Oligopeptides from Apricot Almonds Meal with Neutrase and N120P
Abstract
:1. Introduction
2. Results and Discussion
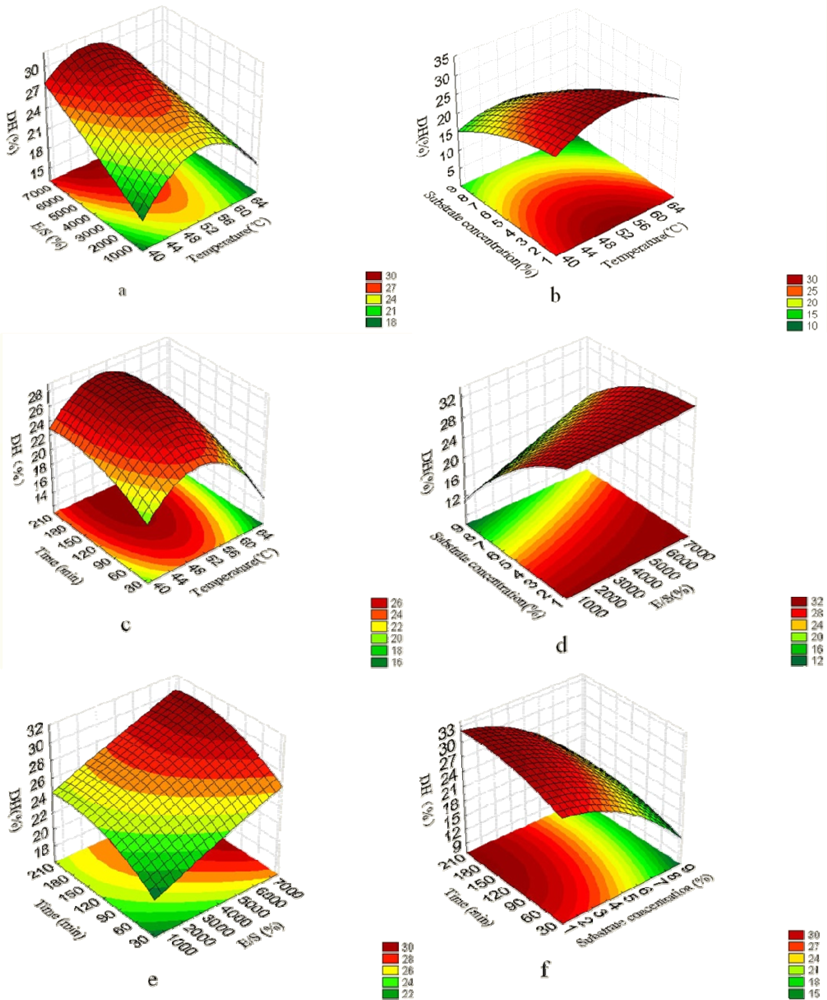
2.1. Analysis of Response Surface
2.2. Fitting the Model
2.3. Optimization of Hydrolysis Parameters and Validation of the Model
3. Experimental Section
3.1. Materials
3.2. Preparation of Apricot Kernel Oligopeptides
3.3. Experimental Design
4. Conclusions
Acknowledgments
References
- Ahrens, S; Venkatachalam, MN; Lapsley, K; Sathe, SK. Almond (Prunus dulcis L.) protein quality. Plant Foods Hum. Nutr 2005, 60, 123–128. [Google Scholar]
- Cherif, A; Sebei, K; Boukhchina, S; Kallel, H; Belkacemi, K; Arul, J. Kernel fatty acid and triacylglycerol composition for three almond cultivars during maturation. J. Am. Oil Chem. Soc 2004, 81, 901–905. [Google Scholar]
- Gezer, I; Haciseferoğulları, H; Demir, F. Some physical properties of Hacıhalilo_glu apricot pit and its kernel. J. Food Eng 2002, 56, 49–57. [Google Scholar]
- Özkal, SG; Yener, ME; Bayındırlı, L. Response surfaces of apricot kernel oil yield in supercritical carbon dioxide. Food Sci. Technol 2005, 38, 611–616. [Google Scholar]
- Abd E1-Aal, MH; Hamza, MA; Rahma, EH. In vitro digestibility, physicochemical and functional properties of apricot kernel proteins. Food Chem 1986, 19, 197–211. [Google Scholar]
- Sze-Tao, KWC; Sathe, SK. Functional properties and in vitro digestibility of almond (Prunus dulcis L.) protein isolate. Food Chem 2000, 69, 153–160. [Google Scholar]
- Tiwari, RS; Venkatachalam, M; Girdhari, MS; Su, M; Roux, KH; Sathe, SK. Effect of food matrix on amandin, almond (Prunus dulcis L.) major protein, immunorecognition and recovery. Food Sci. Technol 2010, 43, 675–683. [Google Scholar]
- Gauthier, SF; Pouliot, Y; Saint-Sauveur, D. Imunomodulatory peptides obtained by the enzymatic hydrolysis of whey proteins. Int. Dairy J 2006, 16, 197–211. [Google Scholar]
- Iván, L-E; Recio, I. Antibacterial activity of peptides and folding variants from milk proteins. Int. Dairy J 2006, 16, 1294–1305. [Google Scholar]
- Shimizu, M; Sawashita, N; Morimatsu, F; Ichikawa, J; Taguchi, Y; Ijiri, Y; Yamamoto, J. Antithrombotic papain-hydrolyzed peptides isolated from pork meat. Thrombosis Res 2009, 123, 753–757. [Google Scholar]
- Kuba, M; Tana, C; Tawata, S; Yasuda, M. Production of angiotensin I-converting enzyme inhibitory peptides from soybean protein with Monascus purpureus acid proteinase. Process Biochem 2005, 40, 2191–2196. [Google Scholar]
- Wang, YK; He, HL; Chen, XL; Sun, CY; Zhang, YZ; Zhou, BC. Production of novel angiotensin I-converting enzyme inhibitory peptides by fermentation of marine shrimp Acetes chinensis with Lactobacillus fermentum SM 605. Appl. Microbiol. Biotechnol 2008, 79, 785–791. [Google Scholar]
- van Beresteijn, ECH; Peeters, RA; Kaper, J; Meijer, RJGM; Robben, AJPM; Schmidt, DG. Molecular mass distribution, immunological properties and nutritive value of whey protein hydrolysates. J. Food Prot 1994, 57, 619–625. [Google Scholar]
- Ena, JM; van Beresteijn, ECH; Robben, AJPM; Schmid, DG. Whey protein antigenicity reduction by fungal proteinases and a pepsin/pancreatin combination. J. Food Sci 1995, 60, 104–116. [Google Scholar]
- Vázquez-Lara, L; Tello-Solis, SR; Gomez-Ruiz, L; Garcia-Garibay, M; Rodriguez-Serrano, GM. Degradation of α-Lactalbumin and B-Lactoglobulin by Actinidin. Food Biotechnol 2003, 17, 117–128. [Google Scholar]
- Zhang, Y; Wang, Q. Preparation of peanut functional oligopeptides by two-step-hydrolysis. Trans. Chin. Soc. Agric. Eng 2007, 23, 258–263. [Google Scholar]
- Wang, C; Feng, B; Dong, Y. Optimization of the enzymatic hydrolysis of oat bran by Protamex protease response surface method. Food Sci. Technol 2008, 5, 134–137. [Google Scholar]
- Gonzalez, P; Camacho, F; Jurado, E; Paez, MP; Guadix, EM. Enzymatic hydrolysis of whey proteins: I. Kinetic models. Biotechnol. Bioeng 1994, 44, 523–528. [Google Scholar]
- Zhang, Y; Wang, Q; Zhou, S. Peanut protein hydrolyzing by Alcalase to prepare peanut oligopeptides. Trans. Chin. Soc. Agric. Eng 2008, 24, 275–279. [Google Scholar]
- Ravikumar, K; Ramalingam, S; Krishnan, S; Balu, K. Optimization of process variables by the application of response surface methodology for dye removal using a novel adsorbent. Dyes Pigm 2006, 70, 18–26. [Google Scholar]
- Sheng, X; Wang, Z; Xu, S. Study on functional properties and structure of sugary almond protein. Sci. Technol. Food Ind 2008, 29, 133–136. [Google Scholar]
- Wang, S; Wen, Z; Li, H. Analysis of the Nutritional Components and Contents of Sweet Apricot kernel. Beijing Agric 2008, 9, 13–15. [Google Scholar]
- Petersen, D; Nielsen, PM; Cambmann, C. Determination of the Degree of Hydrolysis (DH) based on OPA Reaction, ED-9512723; Novo Nordisk A/S: Bagsværd, Denmark, 1995. [Google Scholar]
- Owueu-Apenten, RK. Food Protein Analysis: Quantitative Effecting on Processing, 1st ed; Marcel Dekker, Inc: New York, NY, USA, 2002; pp. 69–87. [Google Scholar]
- Jang, A; Lee, M. Purification and identification of angiotensin converting enzyme inhibitory peptides from beef hydrolysates. Meat Sci 2005, 69, 653–661. [Google Scholar]
| Trial No. | Independent Variables | DH (%) | Yield Rate (%) | |||||
|---|---|---|---|---|---|---|---|---|
| X1 | X2 | X3 | X4 | Actual Value | Predicted Value | Actual Value | Predicted Value | |
| 1 | 59.9 | 5852 | 6.8 | 174 | 21.29 | 21.04 | 52.54 | 52.07 |
| 2 | 59.9 | 5852 | 3.2 | 66 | 27.04 | 26.21 | 59.58 | 59.15 |
| 3 | 59.9 | 1898 | 6.8 | 66 | 15.07 | 15.10 | 41.47 | 40.08 |
| 4 | 59.9 | 1898 | 3.2 | 174 | 28.02 | 27.86 | 57.54 | 57.05 |
| 5 | 45.1 | 5852 | 6.8 | 66 | 24.21 | 24.24 | 46.68 | 47.02 |
| 6 | 45.1 | 5852 | 3.2 | 174 | 30.56 | 30.40 | 63.22 | 64.46 |
| 7 | 45.1 | 1898 | 6.8 | 174 | 19.5 | 20.20 | 49.32 | 49.60 |
| 8 | 45.1 | 1898 | 3.2 | 66 | 24.8 | 24.92 | 57.75 | 58.07 |
| 9 | 40 | 3875 | 5 | 120 | 23.84 | 23.36 | 53.89 | 52.514 |
| 10 | 65 | 3875 | 5 | 120 | 18.68 | 19.34 | 46.38 | 47.97 |
| 11 | 52.5 | 500 | 5 | 120 | 24.4 | 23.93 | 50.51 | 51.20 |
| 12 | 52.5 | 7200 | 5 | 120 | 29.08 | 29.74 | 59.21 | 58.73 |
| 13 | 52.5 | 3875 | 2 | 120 | 30.57 | 31.11 | 70.32 | 69.71 |
| 14 | 52.5 | 3875 | 8 | 120 | 19.37 | 19.00 | 48.2 | 48.71 |
| 15 | 52.5 | 3875 | 5 | 30 | 23.31 | 23.64 | 51.52 | 52.14 |
| 16 | 52.5 | 3875 | 5 | 210 | 27.58 | 27.44 | 60.47 | 60.06 |
| 17 | 52.5 | 3875 | 5 | 120 | 27.49 | 26.99 | 58.46 | 59.21 |
| 18 | 52.5 | 3875 | 5 | 120 | 27.69 | 26.98 | 56.4 | 59.21 |
| 19 | 52.5 | 3875 | 5 | 120 | 27.48 | 26.98 | 60 | 59.21 |
| 20 | 52.5 | 3875 | 5 | 120 | 26.97 | 26.98 | 61.03 | 59.21 |
| 21 | 52.5 | 3875 | 5 | 120 | 26.49 | 26.98 | 59.23 | 59.21 |
| 22 | 52.5 | 3875 | 5 | 120 | 26.49 | 26.98 | 60 | 59.21 |
| 23 | 52.5 | 3875 | 5 | 120 | 26.49 | 26.98 | 59.3 | 59.21 |
| Source | DF | SS | MS | F-value | Prob > F |
|---|---|---|---|---|---|
| DH (%) | |||||
| Residual | 11 | 343.122 | 31.193 | 69.655 | 0.0001** |
| Lack of fit | 11 | 4.926 | 0.448 | ||
| Pure error | 5 | 3.199 | 0.639 | 2.22405 | 0.125 |
| Cor total | 6 | 1.7264 | 0.288 | ||
| Regression | 22 | 348.048 | |||
| R2 = 0.9859 | R2Adj = 0.9858 | ||||
| Yield rate (%) | |||||
| Residual | 11 | 928.407 | 84.401 | 39.172 | 0.0001** |
| Lack of fit | 11 | 23.701 | 2.1546 | ||
| Pure error | 5 | 10.674 | 2.135 | 0.983 | 0.469 |
| Cor total | 6 | 13.027 | 2.171 | ||
| Regression | 22 | 952.108 | |||
| R2 = 0.9751 | R2Adj = 0.9502 |
| Variable | DH | Yield Rate | ||||||
|---|---|---|---|---|---|---|---|---|
| Regression coefficients | Standard error | t-value | p-value | Regression coefficients | Standar d error | t-value | p-value | |
| X1 | −0.8936 | 0.1810 | 6.6025 | 0.0002 | −0.7163 | 0.3971 | 3.4050 | 0.0143 |
| X2 | 0.9445 | 0.1678 | 9.5353 | 0.0001 | 0.8618 | 0.3682 | 5.6358 | 0.0005 |
| X3 | −0.9864 | 0.1810 | 19.8892 | 0.0001 | −0.9785 | 0.3971 | 15.7214 | 0.0001 |
| X4 | 0.8830 | 0.1678 | 6.2399 | 0.0002 | 0.8729 | 0.3682 | 5.9345 | 0.0004 |
| X1 X1 | −0.9631 | 0.1810 | 11.8620 | 0.0001 | −0.9332 | 0.3971 | 8.6124 | 0.0001 |
| X2 X2 | −0.0965 | 0.1678 | 0.3214 | 0.7891 | −0.7757 | 0.3682 | 4.0760 | 0.0052 |
| X3 X3 | −0.7736 | 0.1810 | 4.0490 | 0.0054 | 0.0447 | 0.3971 | 0.1484 | 0.9016 |
| X4 X4 | −0.6767 | 0.1678 | 3.0486 | 0.0247 | −0.6691 | 0.3682 | 2.9863 | 0.0272 |
| X1 X2 | −0.6401 | 0.2365 | 2.7632 | 0.0381 | 0.5982 | 0.5189 | 2.4761 | 0.0584 |
| X1 X3 | −0.7468 | 0.2365 | 3.7247 | 0.0088 | 0.1339 | 0.5189 | 0.4480 | 0.7097 |
| X1 X4 | 0.6998 | 0.2365 | 3.2492 | 0.0182 | 0.0667 | 0.5189 | 0.2216 | 0.8536 |
| Coded Level | Independent Variables | |||
|---|---|---|---|---|
| X1 (°C) | X2 (units g−1 protein) | X3 (%) | X4 (min) | |
| 1.682(+γ) | 65 | 7200 | 8 | 210 |
| 1 | 59.9 | 5852 | 6.8 | 174 |
| 0 | 52.5 | 3875 | 5 | 120 |
| −1 | 45.1 | 1898 | 3.2 | 66 |
| −1.682(−γ) | 40 | 550 | 2 | 30 |
© 2010 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, C.; Wang, Q.; Tian, J. Optimization of Enzymatic Production of Oligopeptides from Apricot Almonds Meal with Neutrase and N120P. Int. J. Mol. Sci. 2010, 11, 4952-4961. https://doi.org/10.3390/ijms11124952
Wang C, Wang Q, Tian J. Optimization of Enzymatic Production of Oligopeptides from Apricot Almonds Meal with Neutrase and N120P. International Journal of Molecular Sciences. 2010; 11(12):4952-4961. https://doi.org/10.3390/ijms11124952
Chicago/Turabian StyleWang, Chunyan, Qiang Wang, and Jinqiang Tian. 2010. "Optimization of Enzymatic Production of Oligopeptides from Apricot Almonds Meal with Neutrase and N120P" International Journal of Molecular Sciences 11, no. 12: 4952-4961. https://doi.org/10.3390/ijms11124952




