Protein GB1 Folding and Assembly from Structural Elements
Abstract
:1. Introduction
2. Results and Discussion
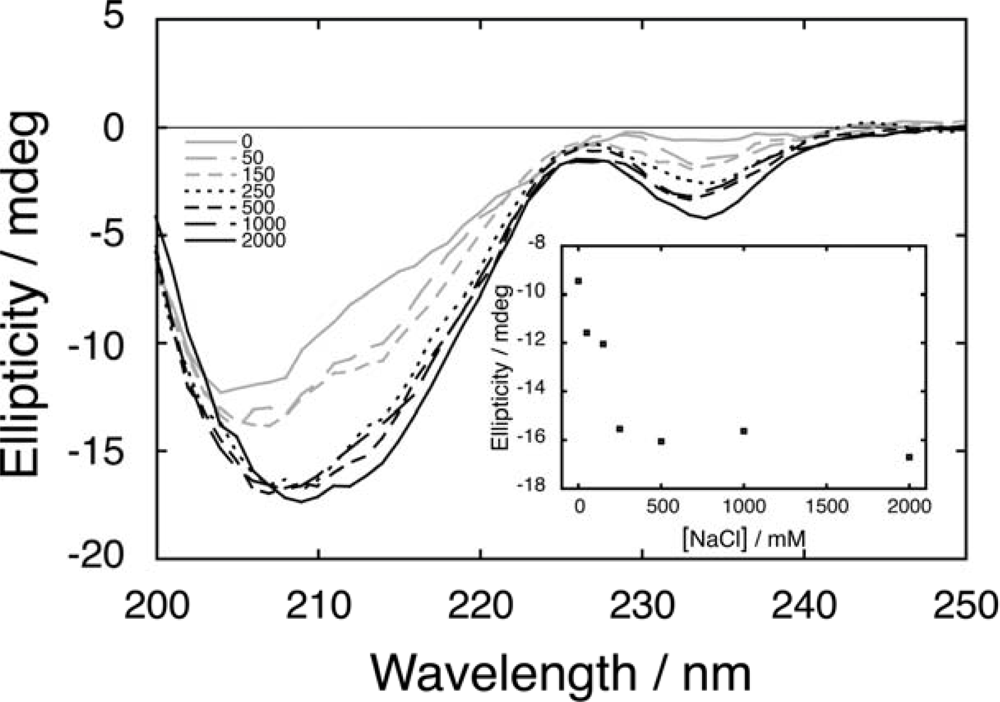
2.1. Salt dependence of CD spectra for fragment mixtures
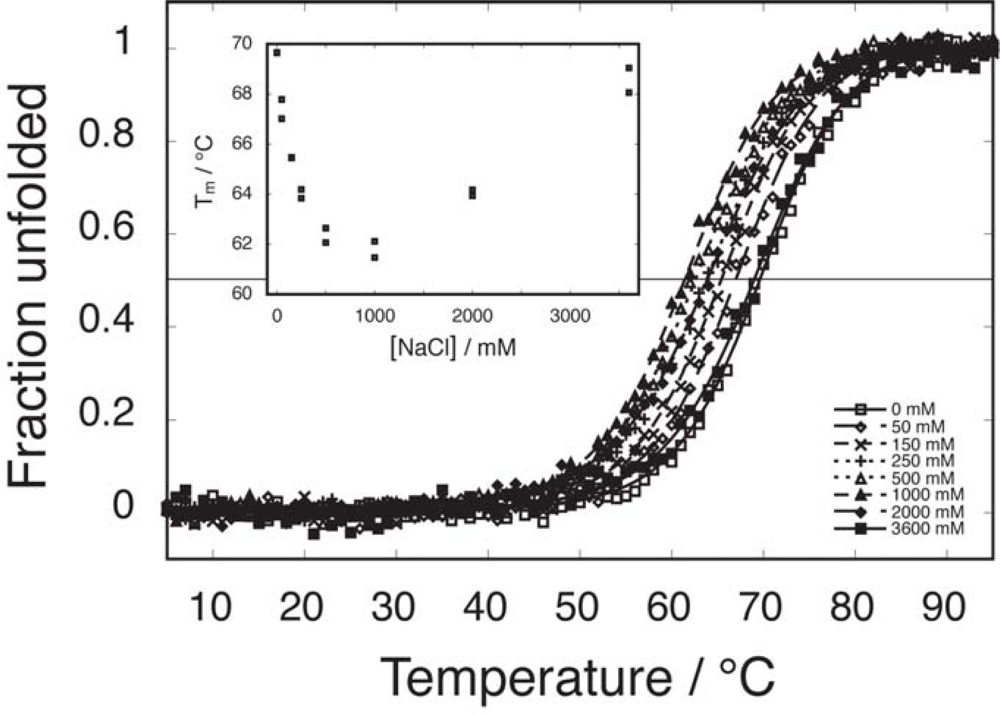
2.2. Salt-dependent residual structure in C16
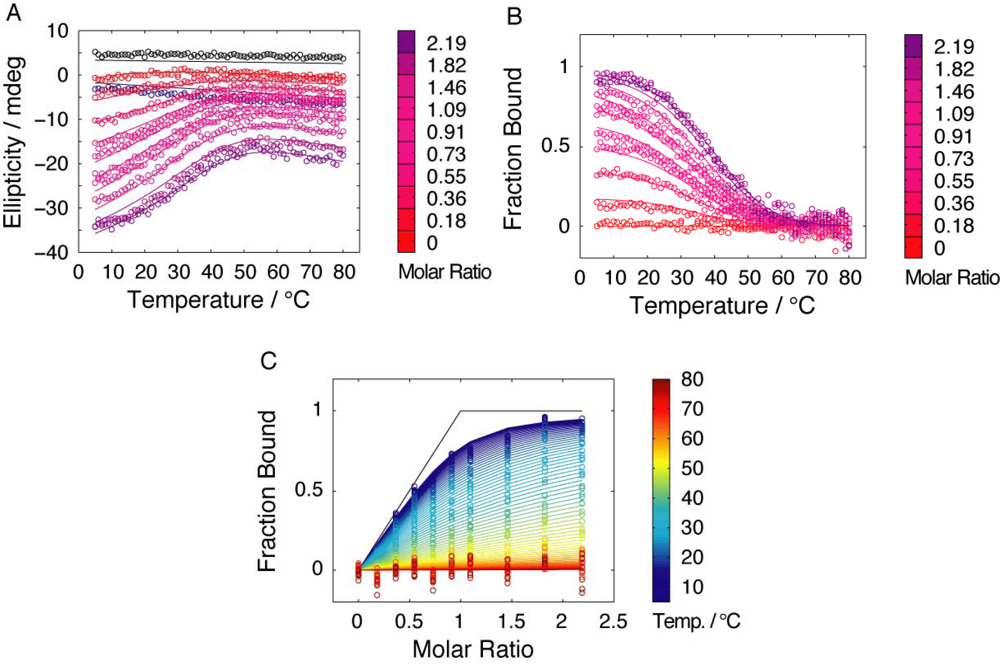
2.3. Temperature dependence of CD spectra for fragment mixtures
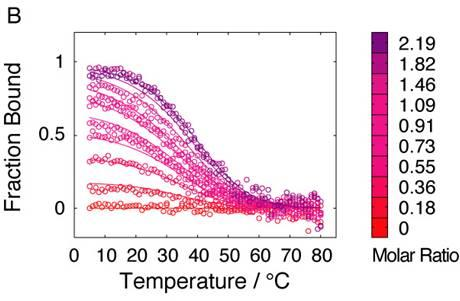
2.4. Affinity between N41-QDD and C16
2.5. Prefolding of C16 favors complex formation
2.6. Comparable salt effects on assembly and folding equilibria
3. Experimental Section
3.1. Cloning, expression and purification of PGB1 fragments
3.2. Expression and purification of intact PGB1-QDD
3.3. Circular dichroism spectroscopy
3.4. 1H-NMR spectroscopy
3.5. Binding model
3.6. Data Analysis
4. Conclusions
Acknowledgments
References and Notes
- Kalman, SM; Liderstrom-Lang, K; Ottesen, M; Richards, FM. Degradation of Ribonuclease by Subtilisin. Biochim. Biophys. Acta 1955, 16, 297–299. [Google Scholar]
- Richards, FM. On the enzymatic activity of subtilisin-modified ribonuclease. Proc. Natl. Acad. Sci. USA 1958, 44, 162–166. [Google Scholar]
- Taniuchi, H; Parr, GR; Juillerat, MA. Complementation in folding and fragment exchange. Methods Enzymol 1986, 131, 185–217. [Google Scholar]
- Hakansson, M; Linse, S. Protein reconstitution and 3D domain swapping. Curr. Protein Pept. Sci 2002, 3, 629–642. [Google Scholar]
- Carey, J; Lindman, S; Bauer, M; Linse, S. Protein reconstitution and three-dimensional domain swapping. Limits and benefits of covalency. Protein Sci 2007, 16, 2317–2333. [Google Scholar]
- Berggård, T; Julenius, K; Ogard, A; Drakenberg, T; Linse, S. Fragment Complementation Studies of Protein Stabilization by Hydrophobic Core Residues. Biochemistry 2001, 40, 1257–1264. [Google Scholar]
- Xue, W-F; Szczepankiewicz, O; Bauer, MC; Thulin, E; Linse, S. Intra- vs. intermolecular interactions – Contribution of surface charges to protein assembly. J. Mol. Biol 2006, 358, 1244–1255. [Google Scholar]
- Dell’Orco, D; Xue, W-F; Thulin, E; Linse, S. Electrostatic contributions to the kinetics and thermodynamics of protein assembly. Biophys. J 2005, 88, 1991–2002. [Google Scholar]
- Linse, S; Voorhies, M; Norström, E; Schultz, DA. An EF-hand phage display study of calmodulin subdomain pairing. J. Mol. Biol 2000, 296, 473–486. [Google Scholar]
- Ruiz-Sanz, J; Gay, GD; Otzen, DE; Fersht, AR. Protein-fragments as models for events in protein-folding pathways - Protein engineering analysis of the association of 2 complementary fragments of the barley chymotrypsin inhibitor-2 (Ci-2). Biochemistry 1995, 34, 1695–1701. [Google Scholar]
- Ptitsyn, OB. Protein folding – hypotheses and experiments. J. Protein Chem 1987, 6, 273–293. [Google Scholar]
- Gianni, S; Guydosh, NR; Khan, F; Caldas, TD; Mayor, U; White, GW; DeMarco, ML; Daggett, V; Fersht, AR. Unifying features in protein-folding mechanisms. Proc. Natl. Acad. Sci. USA 2003, 100, 13286–13291. [Google Scholar]
- Haspel, N; Tsai, CJ; Wolfson, H; Nussinov, R. Hierarchical protein folding pathways: a computational study of protein fragments. Proteins 2003, 51, 203–215. [Google Scholar]
- Linse, S; Linse, B. Protein folding through kinetic discrimination. J. Am. Chem. Soc 2007, 129, 8481–8486. [Google Scholar]
- Holmgren, A. Thioredoxin-C’ - reconstitution of an active form of escherichia-coli thioredoxin from 2 noncovalently linked cyanogen bromide peptide fragments. FEBS Lett 1972, 24, 351–354. [Google Scholar]
- Hantgan, RR; Taniuchi, H. Formation of a Biologically-Active, Ordered complex from 2 overlapping fragments of cytochrome-C. J. Biol. Chem 1977, 252, 1367–1374. [Google Scholar]
- Smith, VF; Matthews, CR. Testing the role of chain connectivity on the stability and structure of dihydrofolate reductase from E. coli: Fragment complementation and circular permutation reveal stable, alternatively folded forms. Protein Sci 2001, 10, 116–128. [Google Scholar]
- Gutte, B; Lin, MC; Caldi, DG; Merrifie, RB. Reactivation of des(119–124, 120–124, or 121–124) ribonuclease-a by mixture with synthetic cooh-terminal peptides of varying lengths. J. Biol. Chem 1972, 247, 4763–4767. [Google Scholar]
- Taniuchi, H; Anfinsen, CB. An experimental approach to study of folding of staphylococcal nuclease. J. Biol. Chem 1969, 244, 3864–3875. [Google Scholar]
- Shaw, GS; Hodges, RS; Kay, CM; Sykes, BD. Relative stabilities of synthetic peptide homodimeric and heterodimeric troponin-C domains. Protein Sci 1994, 3, 1010–1019. [Google Scholar]
- Shuman, CF; Jiji, R; Akerfeldt, KS; Linse, S. Reconstitution of calmodulin from domains and subdomains: Influence of target peptide. J. Mol. Biol 2006, 358, 870–881. [Google Scholar]
- Finn, BE; Kördel, J; Thulin, E; Sellers, P; Forsén, S. Dissection of calbindin D9k into two Ca(2+) - binding subdomains by a combination of mutagenesis and chemical cleavage. FEBS Lett 1992, 298, 211–214. [Google Scholar]
- Linse, S; Thulin, E; Gifford, LK; Radzewsky, D; Hagan, J; Wilk, RR; Åkerfeldt, KS. Domain organization of calbindin D(28k) as determined from the association of six synthetic EFhand fragments. Protein Sci 1997, 6, 2385–2396. [Google Scholar]
- Palczewska, M; Groves, P; Batta, G; Heise, B; Kuźnicki, J. Calretinin and calbindin D28k have different domain organizations. Protein Sci 2003, 12, 180–184. [Google Scholar]
- Kobayashi, N; Honda, S; Yoshii, H; Uedaira, H; Munekata, E. Complement assembly of 2 fragments of the streptococcal protein-G B1 domain in aqueous-solution. FEBS Lett 1995, 366, 99–103. [Google Scholar]
- de Prat Gay, G; Ruiz-Sanz, J; Davis, B; Fersht, AR. The structure of the transition state for the association of two fragments of the barley chymotrypsin inhibitor 2 to generate native-like protein: Implications for mechanisms of protein folding. Proc. Natl. Acad. Sci. USA 1994, 91, 10943–10946. [Google Scholar]
- Jourdan, M; Searle, MS. Cooperative assembly of a nativelike ubiquitin structure through peptide fragment complexation: energetics of peptide association and folding. Biochemistry 2000, 39, 12355–12364. [Google Scholar]
- Xue, W-F; Carey, J; Linse, S. Multi-method global analysis of thermodynamics and kinetics in reconstitution of monellin. Proteins 2004, 57, 586–595. [Google Scholar]
- Gronenborn, AM; Filpula, DR; Essig, NZ; Achari, A; Whitlow, M; Wingfield, PT; Clore, GM. A novel, highly stable fold of the immunoglobulin binding domain of streptococcal protein G. Science 1991, 253, 657–661. [Google Scholar]
- Gallagher, T; Alexander, P; Bryan, P; Gilliland, GL. Two crystal structures of the B1 immunoglobulin-binding domain of streptococcal protein G and comparison with NMR. Biochemistry 1994, 33, 4721–4729. [Google Scholar]
- Blanco, FJ; Rivas, G; Serrano, L. A short linear peptide that folds into a native stable betahairpin in aqueous solution. Nat. Struct. Biol 1994, 1, 584–590. [Google Scholar]
- Honda, S; Kobayashi, N; Munekata, E; Uedaira, H. Fragment reconstitution of a small protein: folding energetics of the reconstituted immunoglobulin binding domain B1 of streptococcal protein G. Biochemistry 1999, 38, 1203–1213. [Google Scholar]
- Olsen, K; Fesinmeyer, RM; Stewart, JM; Andersen, NH. Hairpin folding rates reflect mutations within and remote from the turn region. Proc Natl Acad Sci USA 2005, 102, 15483–15487. [Google Scholar]
- Muñoz, V; Thompson, PA; Hofrichter, J; Eaton, WA. Folding dynamics and mechanism of β-hairpin formation. Nature 1997, 390, 196–199. [Google Scholar]
- Koradi, R; Billeter, M; Wuthrich, K. MOLMOL: a program for display and analysis of macromolecular structures. J Mol Graph 1996, 14, 29–32. [Google Scholar]
- Lindman, S; Xue, W-F; Szczepankiewicz, O; Bauer, MC; Nilsson, H; Linse, S. Salting the charged surface – pH and salt dependence of protein G B1 stability. Biophys. J 2006, 90, 2911–2921. [Google Scholar]
- Kobayashi, N; Honda, S; Munekata, E. Fragment reconstitution of a small protein: disulfide mutant of a short C-terminal fragment derived from streptococcal protein G. Biochemistry 1999, 38, 3228–3234. [Google Scholar]
- Lindman, S; Linse, S; Mulder, FA; André, I. Electrostatic contributions to residue-specific protination equilibria and proton binding capacitance for a small protein. Biochemistry 2006, 45, 13993–14002. [Google Scholar]
- Lindman, S; Linse, S; Mulder, FA; André, I. pK(a) values for side-chain carboxyl groups of a PGB1 variant explain salt and pH-dependent stability. Biophys. J 2007, 92, 257–266. [Google Scholar]
- Efron, B; Gong, G. A leisurely look at the bootstrap, the jackknife and cross-validation. Am. Stat 1983, 37, 36–48. [Google Scholar]

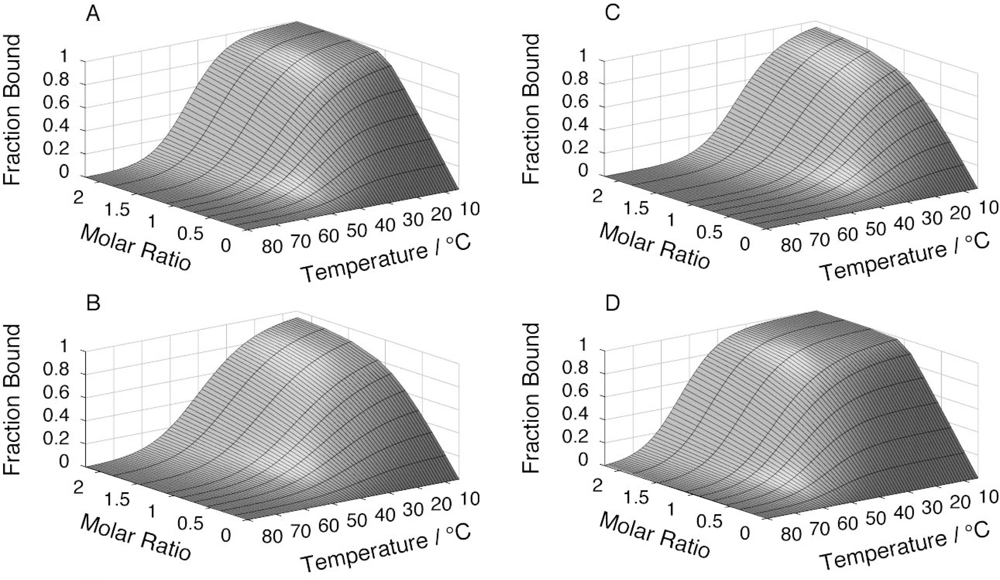
| [NaCl] | logKA at 5 °C | logKA at 25 °C | logKA.Eff at 5 °C | logKA.Eff at 25 °C |
|---|---|---|---|---|
| no added | 5.4 ± 0.4 | 4.3 ± 0.2 | 5.4 ± 0.3 | 4.2 ± 0.2 |
| 150 mM | 4.3 ± 0.1 | 3.7 ± 0.1 | 4.3 ± 0.1 | 3.6 ± 0.1 |
| 500 mM | 4.4 ± 0.2 | 3.4 ± 0.2 | 4.3 ± 0.2 | 3.4 ± 0.1 |
| 2000 mM | 6.0 ± 0.8 | 4.9 ± 0.4 | 5.7 ± 0.6 | 4.8 ± 0.4 |
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Bauer, M.C.; Xue, W.-F.; Linse, S. Protein GB1 Folding and Assembly from Structural Elements. Int. J. Mol. Sci. 2009, 10, 1552-1566. https://doi.org/10.3390/ijms10041552
Bauer MC, Xue W-F, Linse S. Protein GB1 Folding and Assembly from Structural Elements. International Journal of Molecular Sciences. 2009; 10(4):1552-1566. https://doi.org/10.3390/ijms10041552
Chicago/Turabian StyleBauer, Mikael C., Wei-Feng Xue, and Sara Linse. 2009. "Protein GB1 Folding and Assembly from Structural Elements" International Journal of Molecular Sciences 10, no. 4: 1552-1566. https://doi.org/10.3390/ijms10041552