Introduction
Red-ox biotransformations catalyzed by microorganisms have been in the focus of organic chemists working in Biotransformations because the use of isolated enzymes is limited by their requirements for cofactors. The use of whole cells presents the advantage that the cofactors are produced and recycled by the cell’s metabolic pathways [
1]. Nevertheless, traditional biocatalysts such as the well known baker’s yeast give poor enantiomeric excesses, low yields or large biomass production, especially when large or medium ketones or alcohols are used as the substrates.
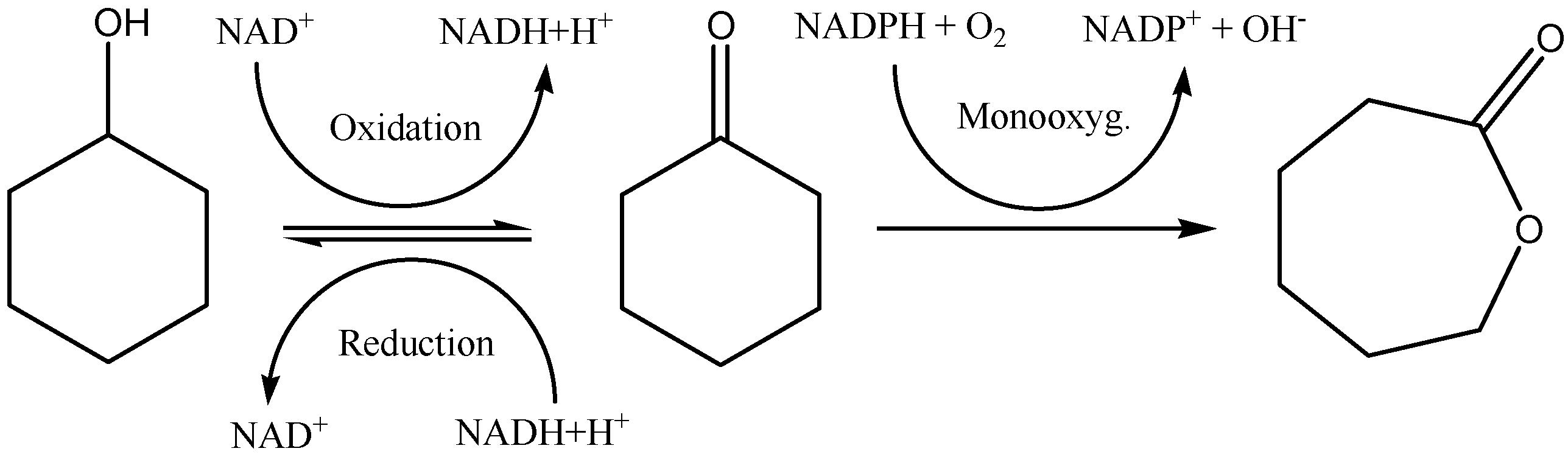
The oxidation of alcohols leading to aldehyde formation is an interesting reaction in Organic Chemistry. It has been previously developed using a
Gluconobacter oxidans strain as a whole cell biocatalyst [
2,
3,
4,
5]. The use of biocatalysts is very interesting for the stereoselective reduction of ketones and the stereospecific oxidation of secondary alcohols, reactions that sometimes cannot be performed using conventional chemistry procedures [
6]. Consecutive red-ox biotransformations can also be used to achieve desymmetrization and asymmetryzation processes, in order to increase the low enantiomeric excesses obtained using certain biocatalysts [
6].
The oxidation of alcohols and the reduction of ketones are commonly carried out by alcohol dehydrogenases (ADHs). These intracellular enzymes are ubiquitous, being present in microorganisms, plants and animals. ADHs are NAD+ or NADP+ dependent enzymes [
7,
8]. The formal enzymatic mechanism for the oxidation of alcohols is well established since 1951 [
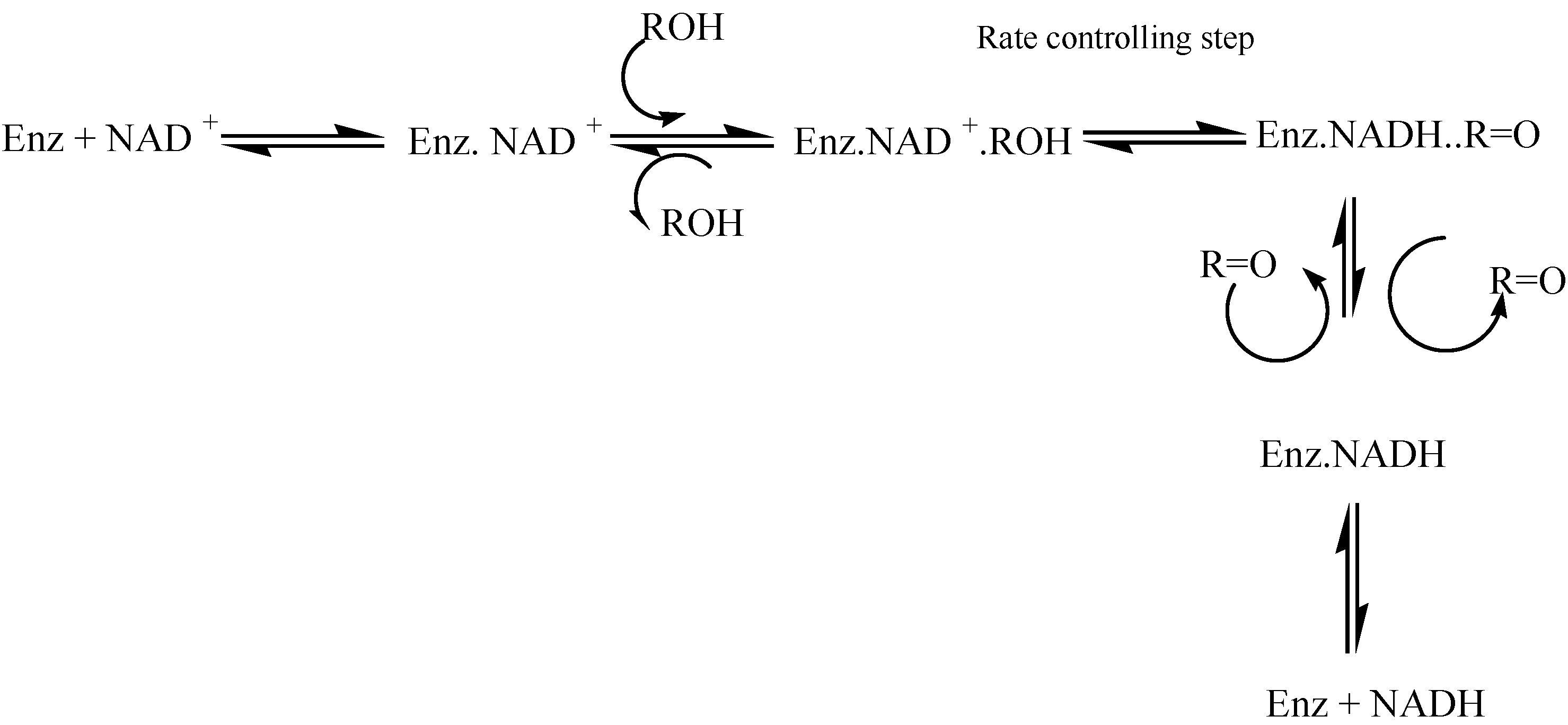
9]. The process consist of a series of equilibria where the rate controlling step is the hydride transfer from the alcohol to NAD(P)+ in the ternary “enzyme + NAD+ + alcohol” complex.
Scheme 1.
Alcohol dehydrogenase mechanism.
Scheme 1.
Alcohol dehydrogenase mechanism.
As a result of all this the discovery of new biocatalysts to increase the screening libraries is always of interest in Organic Chemistry [
10]. Nowadays, this is possible by the application of the newest screening and selection technologies that allow the rapid identification of enzyme activities from different sources.
Comparative Molecular field analysis (CoMFA) is a method for three dimensional (3D) quantitative structure-activity relationships (3D-QSAR) developed at Tripos
1. Although the concept of the approach has been known as DYLOMMS (dynamic lattice-oriented molecular modelling system) [
11] for over a decade, it was not until the 90s that the method became widely used after it was reborn as CoMFA in 1988 [
12,
13]. The methodology has been patented and the program is available as a QSAR package in SYBYL
1.
Comparative Molecular Field Analysis (CoMFA) applied to enzyme catalyzed biotransformations, as a QSAR-3D methodology, may produce a semiquantitative active site structure picture. CoMFA has proven its ability to predict biological properties of systems not amenable to direct analysis. The fundaments of this methodology are well described in the literature [
12,
13,
14,
15,
16,
17,
18,
19,
20] but CoMFA has only been applied a few times in Biocatalysis, to the best of our knowledge [
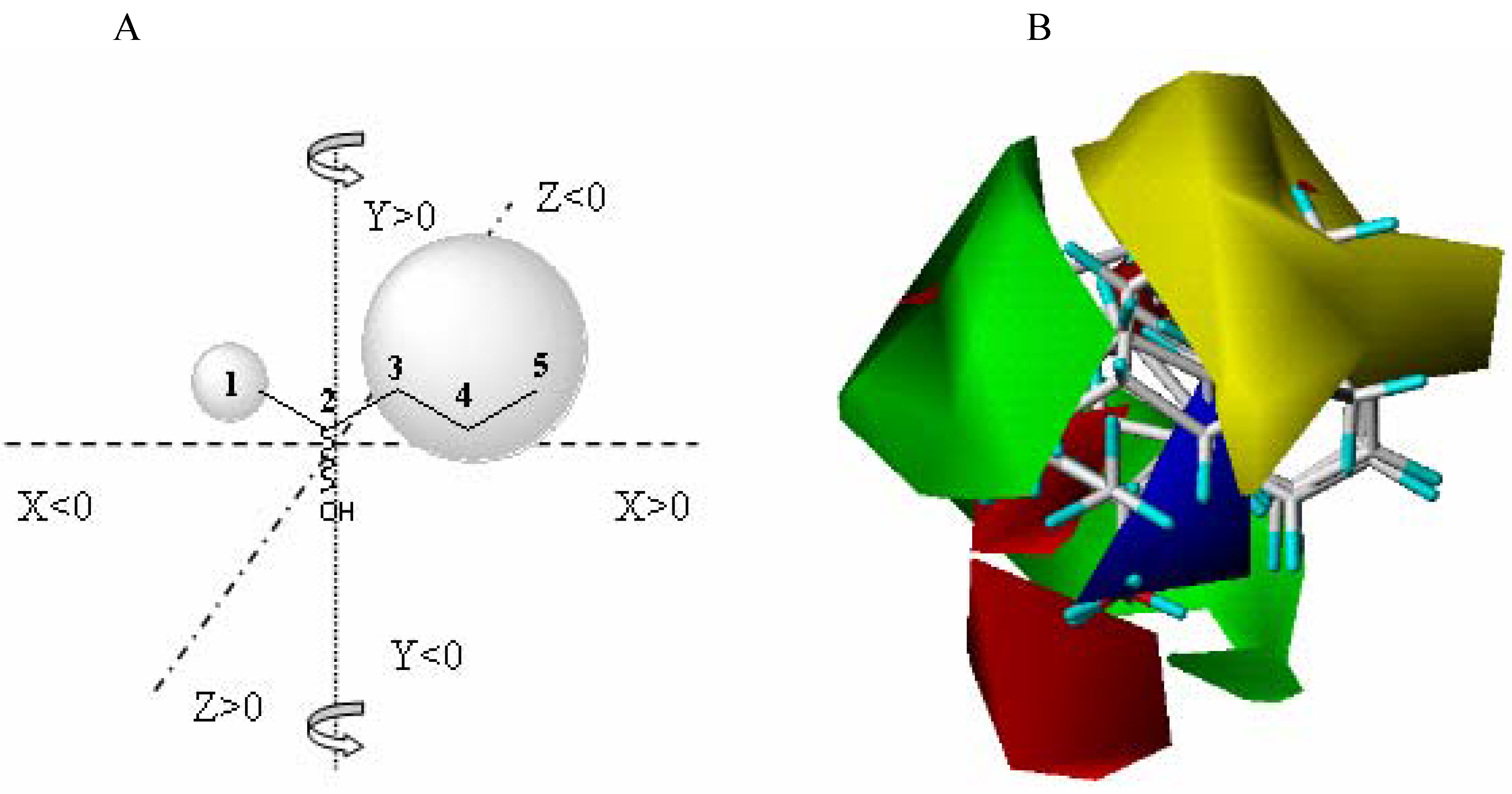
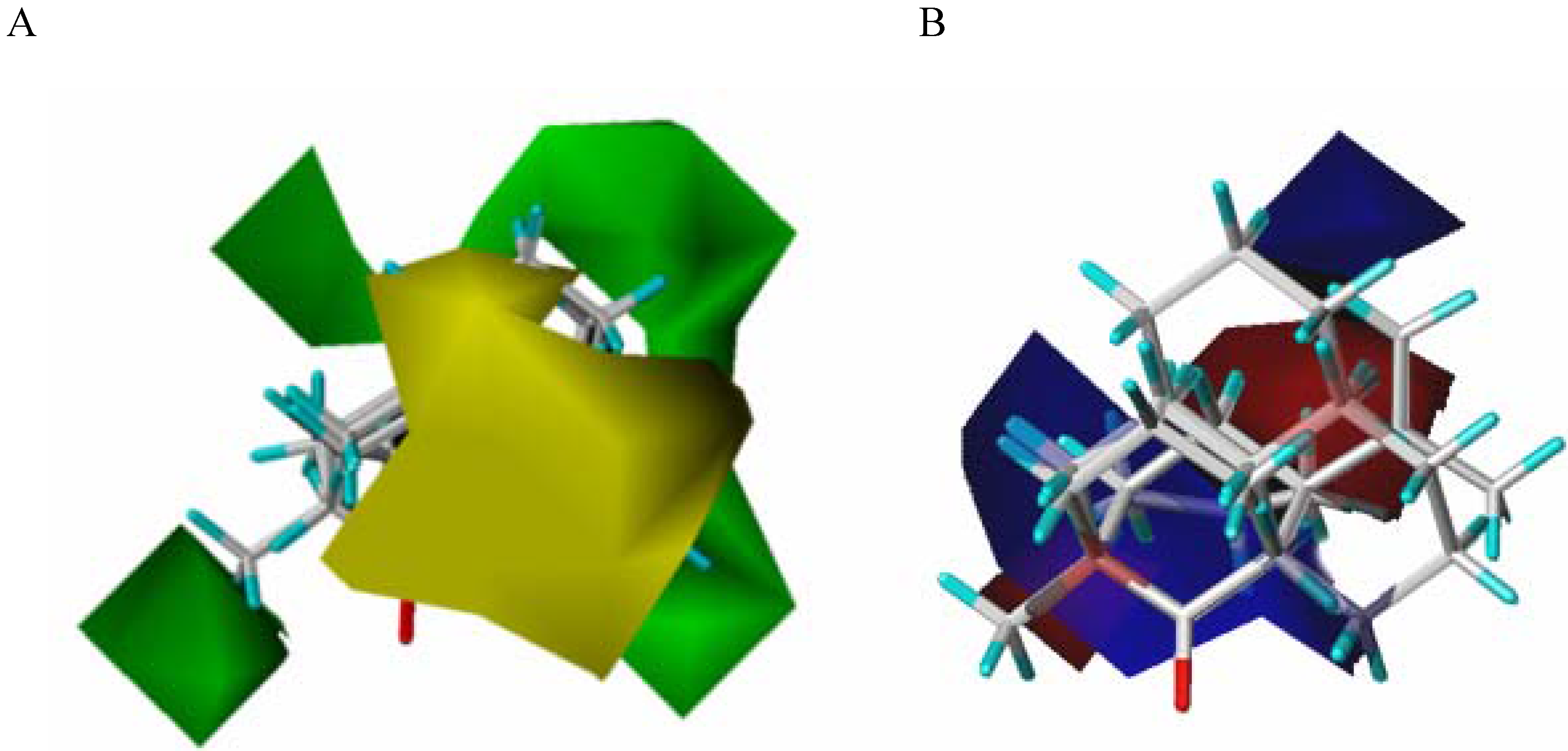
21]. The CoMFA model is a 3D representation of the steric and electrostatic zones of the active-site, directly related to amino acid composition. In Biocatalysis, the model is built starting from the experimental data obtained from the enzyme-substrate interaction (represented as a percentage conversion).
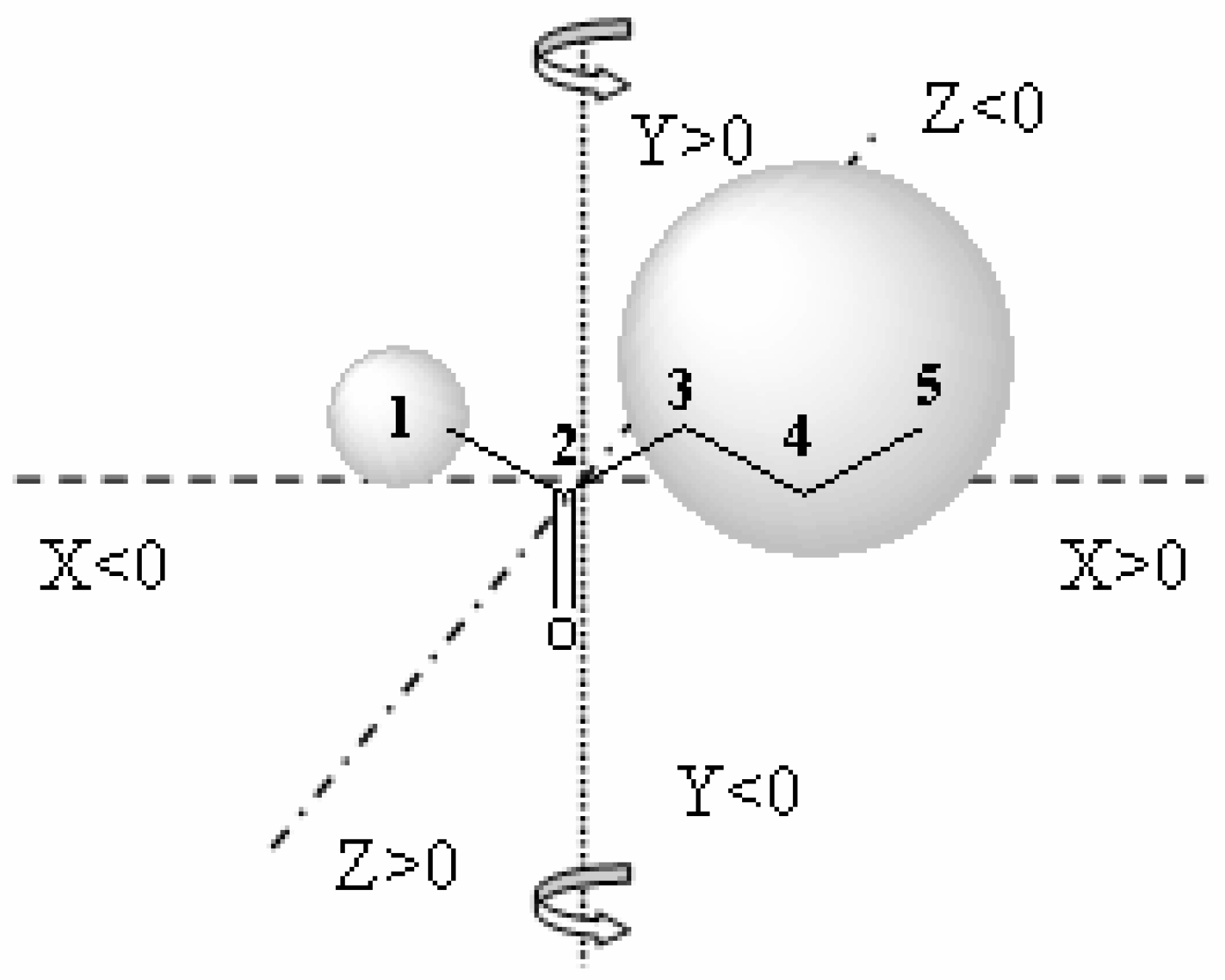
The CoMFA model is obtained starting from the fitting model of the substrates aligned according to a rational established criteria, and is complementary to the fitting model of the substrates (bioactive conformers) taking into account the yield (percentage conversion - experimental data) obtained against each substrate characterized by its steric and electrostatic properties.
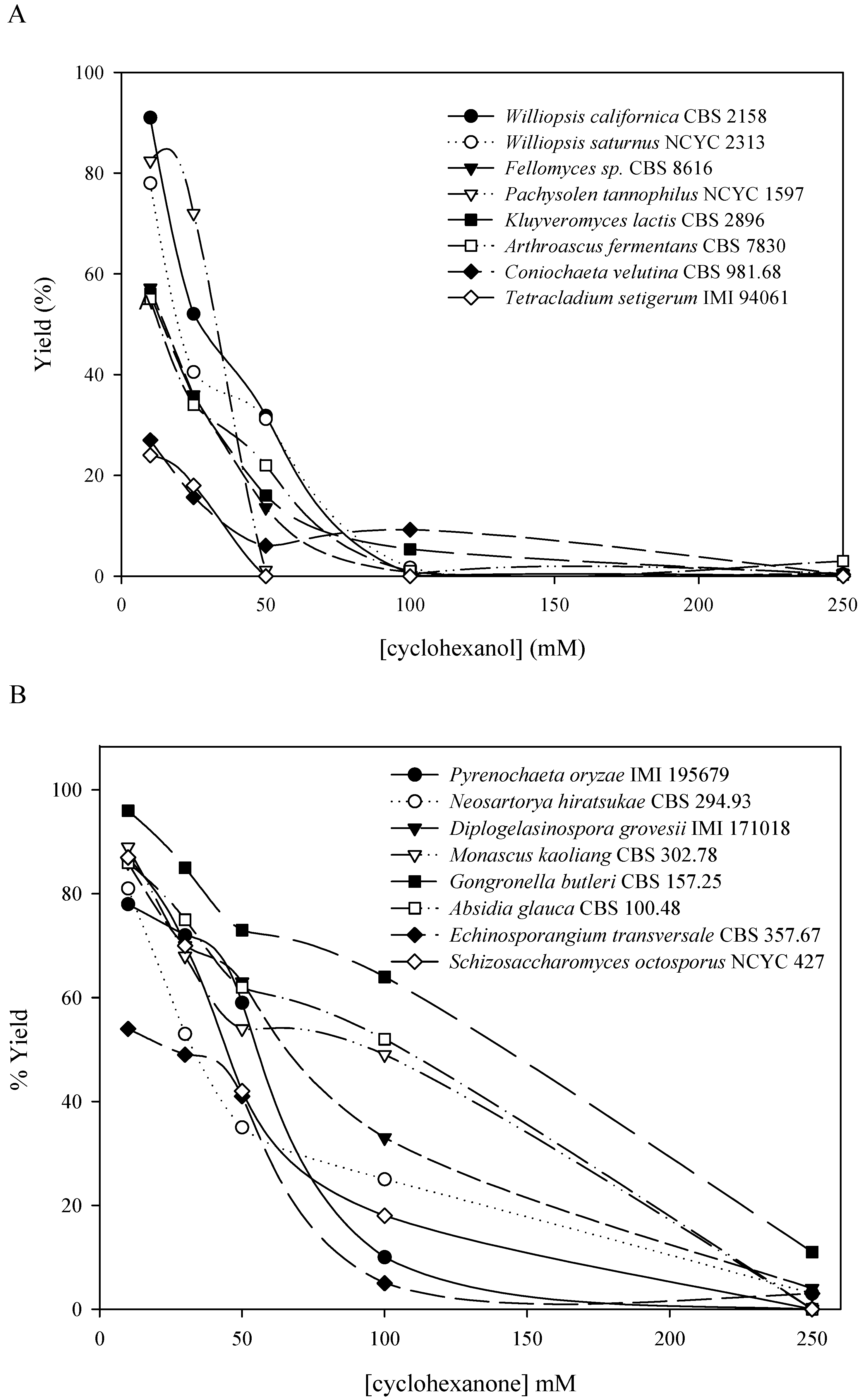
QSAR-3D/CoMFA methodology could be a good way to obtain a predictive model of the activity and substrate specificity of a biocatalyst. Using these models we can select a priori the appropriate biocatalyst for each substrate, saving time and reagents. In this paper, a hierarchical screening process looking for new alcohol oxidation and ketone reduction biocatalysts has been performed starting from a collection of 416 microorganisms from public libraries, selected trying to acheive the maximum biodiversity. We describe, comparatively, the results of both ketone-reduction and alcohol-oxidation microbial screenings. Afterwards the combinatorial screening of the alcohol oxidation biocatalysts and the design steps of the CoMFA model are also explained.
Experimental
Chemicals
All the substrates for the oxidation reactions were obtained from Sigma-Aldrich. All the culture media components were from Difco Laboratories and from Merck.
Microbial collection
The microorganisms used in this study were obtained from different public collections: American Type Culture Collection, U.S. (ATCC); Fungal Biodiversity Center, Utrecht, The Netherlands (CBS); International Mycological Institute, U.K. (IMI); German collection of cell cultures, Germany (DSMZ) and National Collection of Yeast Cultures, U.K. (NCYC).
Conservation of microorganisms
Bacteria, actinomycetes, yeasts, filamentous and marine fungi, were conserved as suspensions in a 30% glycerol solution. All these strains were stored at -80 ºC in Nunc cryotubes. Basidiomycetes were also conserved as cell suspensions in 30% glycerol and stored at -150 ºC in a liquid nitrogen tank.
Screening method: growing cells
Bacteria and Yeasts
Conical 100 mL flasks containing 20 mL of the selected culture media were inoculated with 50 μL of the microbial cells suspension in glycerol (previously thawed). Cultures were performed in an orbital shaker at 28 ºC and 250 rpm [
29,
30]. After 48 h of incubation, the substrate was added to the flasks at a 10 mM final concentration (in the case of the final screening reactions, the substrate concentration was set to 2.5 mM) [
29,
31]. The reaction time was 72 h. When the reaction was finished, the content of the conical flask was transferred into a Falcon tube and 5 mL of ethyl acetate (containing 1 mg/mL of hexadecane as internal standard) were added. After vortexing for 10 s, the organic phase was transferred to a 2 mL Hewlett-Packard vial. All the reactions were repeated three times and the results displayed are the arithmetic mean values.
Other microorganisms
For the other microbial groups the screening process was essentially the same as described above. Culture and reaction times were 72 h for actinomycetes and filamentous fungi. For basidiomycetes and marine fungi the culture time was set to 120 h and the reaction time was 72 h.
Analysis: Gas chromatography
The analysis of the samples from the reactions was performed with a Hewlett Packard 5890 Series II gas chromatograph provided with an automated sampler (Agilent Technologies) and an electrolytic hydrogen generator UHP-601 (Domnick Hunter). The capillary column was a Carbowax SGL-1000 (60m, 0.25mm, 0.25 μm) from Sugelabor (Spain). We defined the following analytical parameters. Ti 155 ºC; ti: = 1 min; Tf = 175 ºC; Tf = 10 min; Heating rate = 4 ºC/min ; Carrier flow (He) = 40 psi; Split ratio = 100 mL/min. Injector and Detector Temperatures = 250 ºC.
The analysis of the oxidation of chiral alcohols and the reduction of prochiral ketones were performed using a Varian 3400cx gas chromatograph equipped with an automated sampler. A CP 7502 Carbowax capillary column (25 m, 0.39 mm) from Sugelabor (Spain) was employed under the following conditions: Ti = 90 ºC; ti = 5 min; heating rate = 5 ºC/min; Tf: = 175 ºC; tf: = 7 min. Carrier flow (He) = 25 psi, and split ratio = 100 mL/min. Injector and detector temperatures = 250 ºC.
The yield calculation was performed by dividing the peak area of the product by that corresponding to the standard extraction value of the substrate in the same conditions. The standard extraction value was established using the data from 10 conical flasks with the same volume of culture medium, stirring speed, temperature and reaction time. The substrate was extracted using ethyl acetate and analyzed by gas chromatography. No significant variation in the value of the area was detected between flasks. The arithmetic mean of these areas was used for the yield determination. The standard extraction value is a reference value that serves to avoid potential problems owing to the physico-chemical properties of the organic substrates tested.
CoMFA analysis
The SYBYL/CoMFA® analysis was performed on a Silicon Graphics Octane-2 workstation at the Computer Service of the Universidad Autónoma of Barcelona. The optimization of the geometry of the database molecules in the postulated active conformation was performed using a 6-31G* basis set. The charges of each atom were calculated from the molecular electrostatic potential [
13] using a 6-31G* basis set. The solvating energies of the substrates were calculated with a continuous polarized model [
12,
13] using a 6-31G* basis set. The QSAR table for CoMFA included the steric and electronic field values, the stabilization energy of enzyme substrate complex, the solvating energy and the yield in alcohol obtained for each carbonylic group. The electrostatic and steric fields were calculated at each lattice intersection grid of 2 Å.
Partial Least Squares (PLS) analysis [
18,
19] was used to obtain straight lines from the matrix. The Leave One Out (LOO) cross validation method was used to select the number of main components of QSAR-3D analysis and to calculate the statistical parameters (q
2). The CoMFA model was generated using a no-cross-validation method and the number of components indicated by the LOO validation. QSAR 3D/CoMFA analysis was performed with the QSAR module of the SYBYL 6.5 program and the calculus was performed using GAUSSIAN-98 [
20].