The Complete Chloroplast Genomes of Two Lancea Species with Comparative Analysis
Abstract
1. Introduction
2. Materials and Methods
2.1. DNA Extraction and Sequencing
2.2. Chloroplast Genome Assembly and Annotation
2.3. Repeat Structure Analysis
2.4. Genome Comparison
2.5. Phylogenetic Analysis
3. Results
3.1. Characteristics of the Chloroplast Genomes
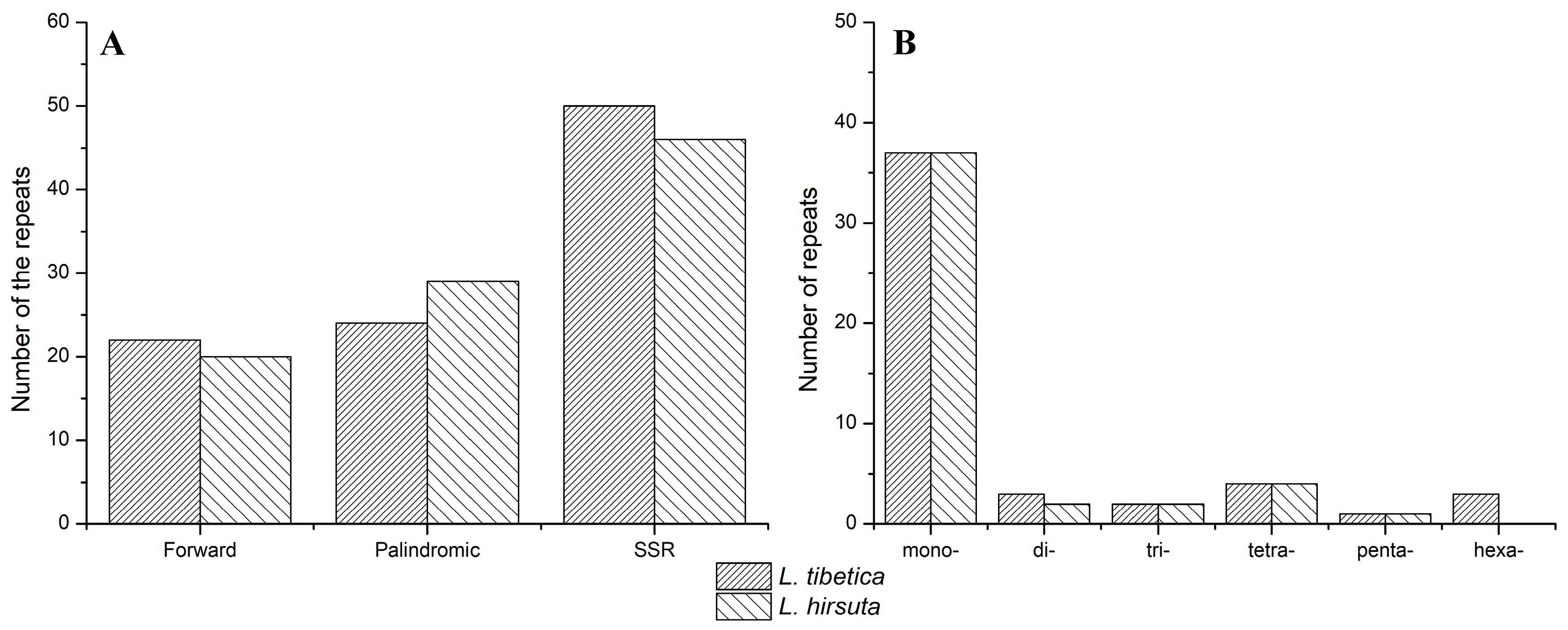
3.2. Repeat and SSR Analysis
3.3. IR Contraction and Expansion
3.4. Sequence Divergence and Divergence Hotspot
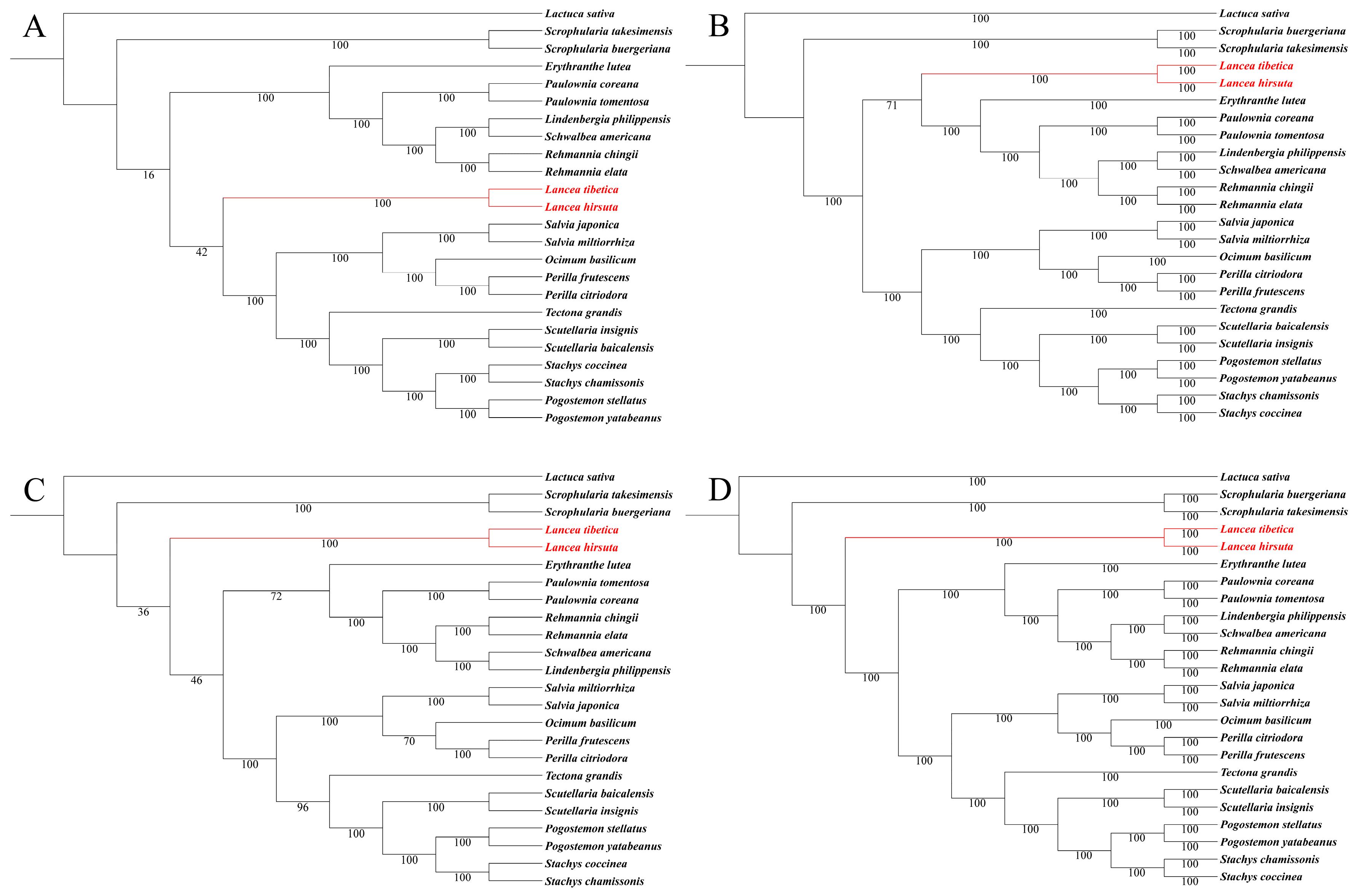
3.5. Phylogenomic Analysis
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Neuhaus, H.E.; Emes, M.J. Nonphotosynthetic metabolism in plastids. Annu. Rev. Plant Biol. 2000, 51, 111–140. [Google Scholar] [CrossRef] [PubMed]
- Sugiura, M. The chloroplast genome. Plant Mol. Biol. 1992, 19, 149–168. [Google Scholar] [CrossRef] [PubMed]
- Daniell, H.; Lin, C.; Yu, M.; Chang, W. Chloroplast genomes: Diversity, evolution, and applications in genetic engineering. Genome Biol. 2016, 17, 134. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Jin, J.; Chen, S.; Chase, M.W.; Soltis, D.E.; Li, H.; Yang, J.; Li, D.; Yi, T. Diversification of Rosaceae since the Late Cretaceous based on plastid phylogenomics. New Phytol. 2017, 214, 1355–1367. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Raven, P.H. Flora of China: Vol. 18 Scrophulariaceae through Gesneriaceae. Scrophulariaceae through Gesneriaceae; Missouri Botanical Garden: St. Louis, MO, USA, 1998. [Google Scholar]
- Chi, X.; Zhang, F.; Tian, Z.; Chen, S. New records of Lancea hirsuta in Qinghai-Tibetan Plateau. Acta Bot. Boreali-Occident. Sin. 2017, 37, 1447–1449. [Google Scholar]
- Song, Z.; Qian, Z.; Rumalla, C.S.; Smillie, T.J.; Khan, I.A. Identification of 11 marker compounds simultaneously in herb Lancea tibetica by using high-performance thin-layer chromatography. JPC 2011, 24, 312–315. [Google Scholar]
- Song, Z.; Wang, Y.; Qian, Z.; Smillie, T.J.; Khan, I.A. Quantitative determination of 10 phenylpropanoid and lignan compounds in Lancea tibetica by high-performance liquid chromatography with uv detection. Planta Medica 2011, 77, 1562–1566. [Google Scholar] [CrossRef] [PubMed]
- Su, B.; Zhu, Q.; Gao, K.; Yuan, C.; Jia, Z. Lignan and phenylpropanoid glycosides from Lancea tibetica and their antitumor activity. Planta Medica 1999, 65, 558–561. [Google Scholar] [CrossRef] [PubMed]
- Wettstein, R. Nolanaceae, Solanaceae, Scrophulariaceae; Engelmann: Leipzig, Germany, 1891. [Google Scholar]
- Olmstead, R.G.; Reeves, P.A. Evidence for the polyphyly of the Scrophulariaceae based on chloroplast rbcL and ndhF sequences. Ann. Mo. Bot. Gard. 1995, 82, 176–193. [Google Scholar] [CrossRef]
- Beardsley, P.M.; Olmstead, R.G. Redefining Phrymaceae: The placement of Mimulus, tribe Mimuleae, and Phryma. Am. J. Bot. 2002, 89, 1093–1102. [Google Scholar] [CrossRef] [PubMed]
- Refulio-Rodriguez, N.F.; Olmstead, R.G. Phylogeny of Lamiidae. Am. J. Bot. 2014, 101, 287–299. [Google Scholar] [CrossRef] [PubMed]
- The Angiosperm Phylogeny Group. An update of the angiosperm phylogeny group classification for the orders and families of flowering plants: APG IV. Bot. J. Linn. Soc. 2016, 181, 1–20. [Google Scholar]
- Oxelman, B.; Kornhall, P.; Olmstead, R.G.; Bremer, B. Further disintegration of Scrophulariaceae. Taxon 2005, 54, 411–425. [Google Scholar] [CrossRef]
- Tank, D.C.; Beardsley, P.M.; Kelchner, S.A.; Olmstead, R.G. Review of the systematics of Scrophulariaceae s.l. and their current disposition. Aust. Syst. Bot. 2006, 19, 289–307. [Google Scholar] [CrossRef]
- Xia, Z.; Wang, Y.; Smith, J.F. Familial placement and relations of Rehmannia and Triaenophora (Scrophulariaceae s.l.) inferred from five gene regions. Am. J. Bot. 2009, 96, 519–530. [Google Scholar] [CrossRef] [PubMed]
- Doyle, J.J. A rapid DNA isolation procedure for small amounts of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- Luo, R.; Liu, B.; Xie, Y.; Li, Z.; Huang, W.; Yuan, J.; He, G.; Chen, Y.; Qi, P.; Liu, Y. SOAPdenovo2: An empirically improved memory-efficient short-read de novo assembler. Gigascience 2012, 1, 18. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Shi, L.; Zhu, Y.; Chen, H.; Zhang, J.; Lin, X.; Guan, X. CpGAVAS, an integrated web server for the annotation, visualization, analysis, and genbank submission of completely sequenced chloroplast genome sequences. BMC Genom. 2012, 13, 715. [Google Scholar] [CrossRef] [PubMed]
- Lohse, M.; Drechsel, O.; Bock, R. Organellargenomedraw (OGDRAW): A tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr. Genet. 2007, 52, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Kurtz, S.; Choudhuri, J.V.; Ohlebusch, E.; Schleiermacher, C.; Stoye, J.; Giegerich, R. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 2001, 29, 4633–4642. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Li, Y.; Zhang, X.; Yue, B. MSDB: A user-friendly program for reporting distribution and building databases of microsatellites from genome sequences. J. Hered. 2012, 104, 154–157. [Google Scholar] [CrossRef] [PubMed]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 32, W273–W279. [Google Scholar] [CrossRef] [PubMed]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef] [PubMed]
- Kazutaka, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v3: An online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016, 44, W242–W245. [Google Scholar] [CrossRef] [PubMed]
- Yi, D.K.; Kim, K.J. Two complete chloroplast genome sequences of genus Paulownia (Paulowniaceae): Paulownia coreana and P. tomentosa. Mitochondrial DNA Part B 2016, 1, 627–629. [Google Scholar] [CrossRef][Green Version]
- Park, I.; Kim, W.J.; Yeo, S.M.; Choi, G.; Kang, Y.M.; Piao, R.; Moon, B.C. The complete chloroplast genome sequences of Fritillaria ussuriensis maxim. and Fritillaria cirrhosa D. Don, and comparative analysis with other Fritillaria species. Molecules 2017, 22, e982. [Google Scholar] [CrossRef] [PubMed]
- Shen, X.; Wu, M.; Liao, B.; Liu, Z.; Bai, R.; Xiao, S.; Li, X.; Zhang, B.; Xu, J.; Chen, S. Complete chloroplast genome sequence and phylogenetic analysis of the medicinal plant Artemisia annua. Molecules 2017, 22, e1330. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Li, Q.; Hu, Z.; Li, X.; Chen, S. The complete Amomum kravanh chloroplast genome sequence and phylogenetic analysis of the commelinids. Molecules 2017, 22, 1875. [Google Scholar] [CrossRef] [PubMed]
- Zeng, S.; Zhou, T.; Han, K.; Yang, Y.; Zhao, J.; Liu, Z.L. The complete chloroplast genome sequences of six Rehmannia species. Genes 2017, 8, 103. [Google Scholar] [CrossRef] [PubMed]
- Zhong, B.; Deusch, O.; Goremykin, V.V.; Penny, D.; Biggs, P.J.; Atherton, R.A.; Nikiforova, S.V.; Lockhart, P.J. Systematic error in seed plant phylogenomics. Genome Biol. Evol. 2011, 3, 1340–1348. [Google Scholar] [CrossRef] [PubMed]
- Som, A. Causes, consequences and solutions of phylogenetic incongruence. Brief. Bioinform. 2015, 16, 536–548. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Sequence data of two Lancea species are available from the authors. |
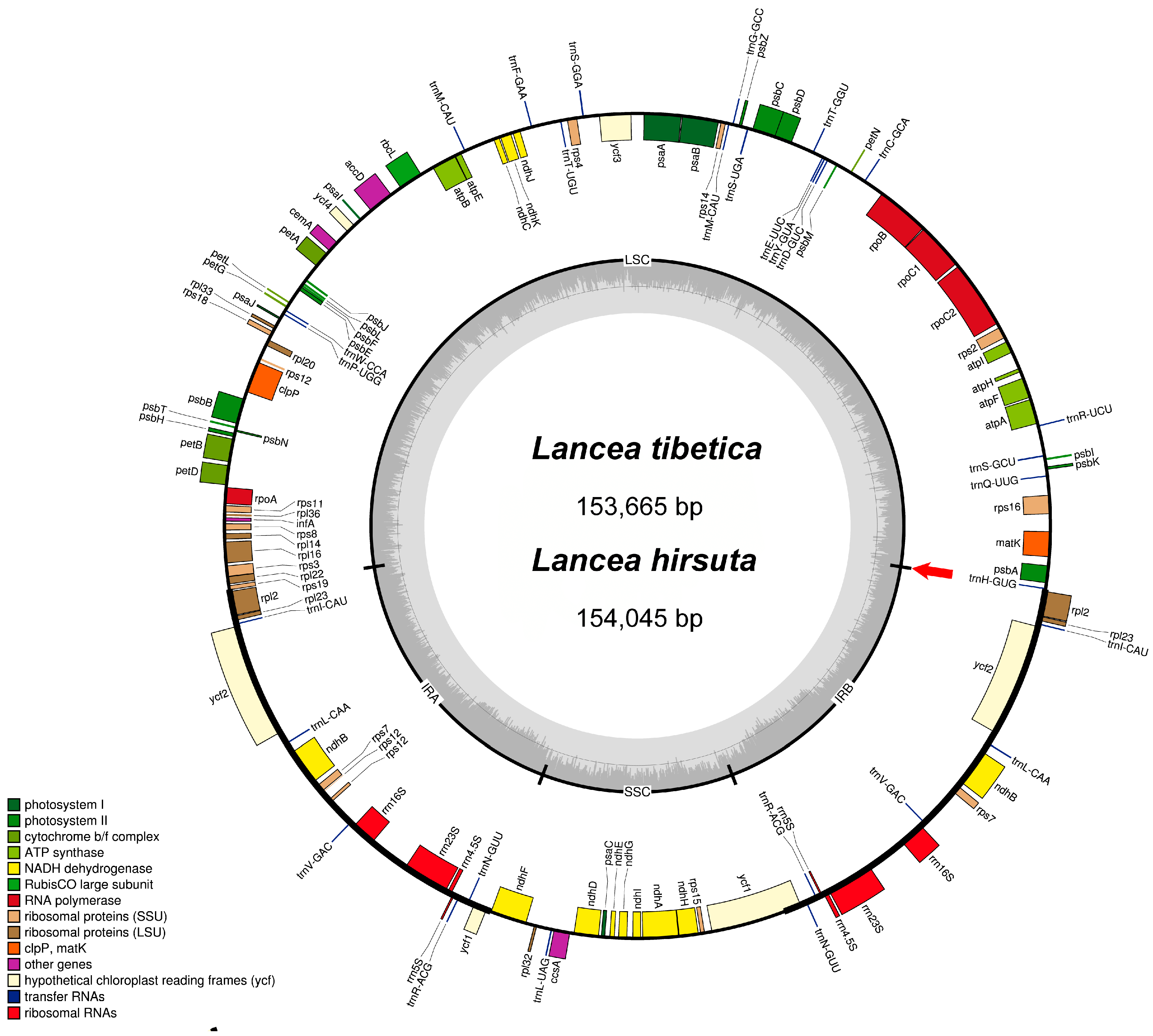



| Characteristics | L. tibetica | L. hirsuta |
|---|---|---|
| Total cpDNA size (bp) | 153,664 | 154,045 |
| Length of large single copy (LSC) region | 84,401 | 84,254 |
| Length of inverted repeat (IR) region | 25,624 | 25,838 |
| Length of small single copy (SSC) region | 18,016 | 17,781 |
| Total GC content (%) | 37.9 | 37.9 |
| LSC | 35.9 | 35.8 |
| IR | 43.3 | 43.2 |
| SSC | 30.0 | 32.0 |
| Total number of genes | 106 | 105 |
| Protein-coding genes | 79 | 79 |
| rRNAs genes | 4 | 4 |
| tRNAs genes | 23 | 22 |
| Category | Name |
|---|---|
| Rubisco | rbcL |
| Photosystem I | psaA, B, C, I, J |
| Photosystem II | psbA, B, C, D, E, F, H, I, J, K, L, M, N, T, Z |
| ATP synthase | atpA, B, E, * F, H, I |
| Cytochrome b/f complex | petA, * B, * D, G, L, N |
| Cytochrome c synthesis | ccsA |
| NADPH dehydrogenase | * ndhA, *,a B, C, D, E, F, G, H, I, J, K |
| Transcription | rpoA, B, * C1, C2 |
| Small subunit ribosomal proteins | rps2, 3, 4, a 7, 8, 11, ** 12, 14, 15, * 16, 18, 19, |
| Large subunit ribosomal proteins | *,a rpl2, 14, * 16, 20, 22, a 23, 32 33, 36 |
| Translation initiation factor | infA |
| Ribosomal RNA | a rrn4.5, a 5, a 16, a 23 |
| RNA processing | matK |
| Carbon metabolism | cemA |
| Fatty acid synthesis | accD |
| Proteolysis | ** c1pP |
| Unknown function protein-coding gene | ycf1, 2, ** 3, 4 |
| Transfer RNA | trnC-GCA, trnD-GUC, trnE-UUC, trnF-GAA, trnG-GCC, trnH-GUG, a trnI-CAU, a trnL-CAA, trnL-UAG, a trnM-CAU, a trnN-GUU, trnP-UGG, trnQ-UUG, a trnR-ACG, trnR-UCU, trnS-GCU, trnS-GGA, trnS-UGA, trnT-GGU, b trnT-UGU, a trnV-GAC, trnW-CCA, trnY-GUA |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chi, X.; Wang, J.; Gao, Q.; Zhang, F.; Chen, S. The Complete Chloroplast Genomes of Two Lancea Species with Comparative Analysis. Molecules 2018, 23, 602. https://doi.org/10.3390/molecules23030602
Chi X, Wang J, Gao Q, Zhang F, Chen S. The Complete Chloroplast Genomes of Two Lancea Species with Comparative Analysis. Molecules. 2018; 23(3):602. https://doi.org/10.3390/molecules23030602
Chicago/Turabian StyleChi, Xiaofeng, Jiuli Wang, Qingbo Gao, Faqi Zhang, and Shilong Chen. 2018. "The Complete Chloroplast Genomes of Two Lancea Species with Comparative Analysis" Molecules 23, no. 3: 602. https://doi.org/10.3390/molecules23030602
APA StyleChi, X., Wang, J., Gao, Q., Zhang, F., & Chen, S. (2018). The Complete Chloroplast Genomes of Two Lancea Species with Comparative Analysis. Molecules, 23(3), 602. https://doi.org/10.3390/molecules23030602






