Studying the Structural Significance of Galectin Design by Playing a Modular Puzzle: Homodimer Generation from Human Tandem-Repeat-Type (Heterodimeric) Galectin-8 by Domain Shuffling
Abstract
:1. Introduction
2. Results
2.1. Protein Production and Characterization
2.1.1. Engineering and Yield
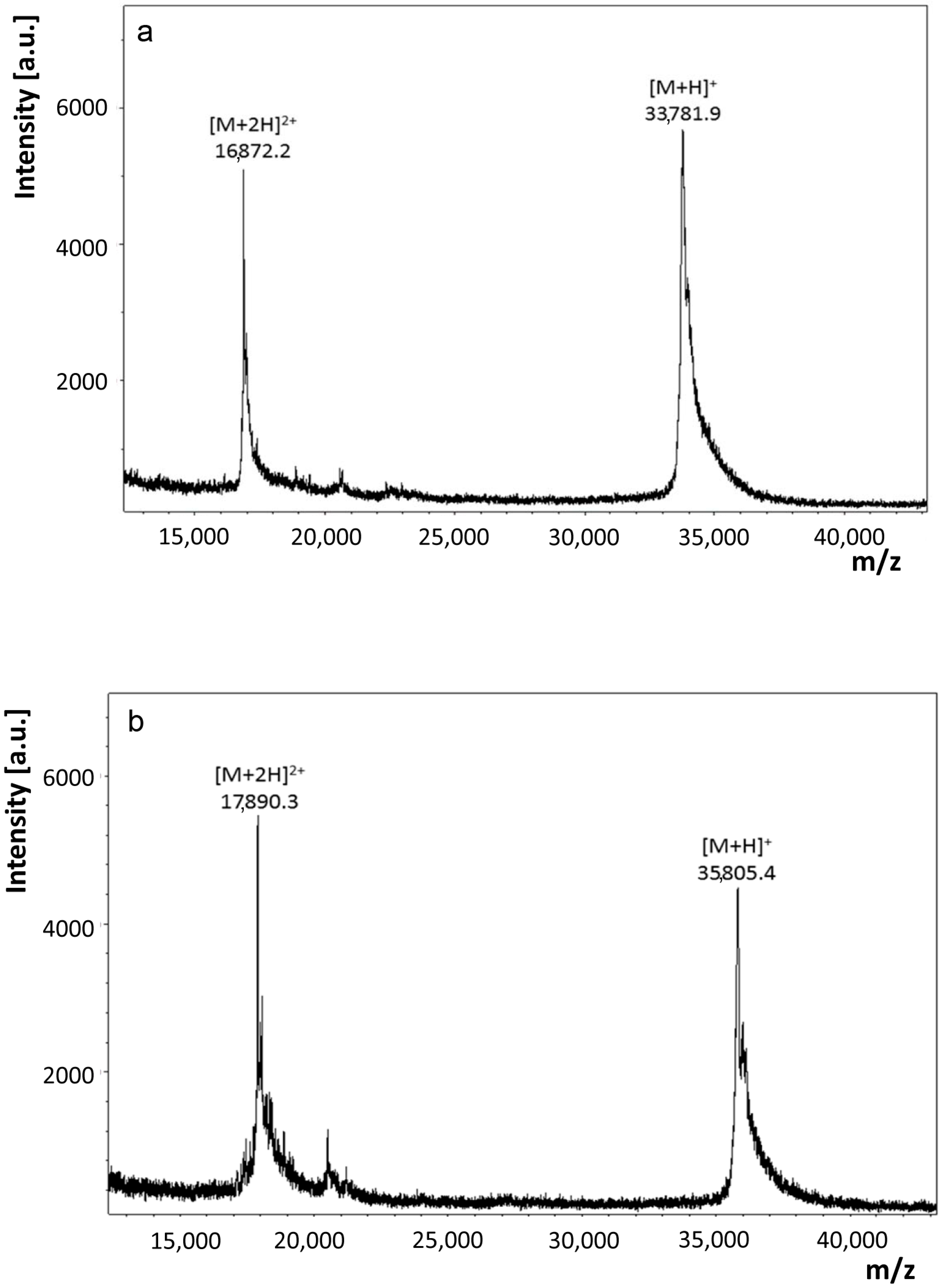
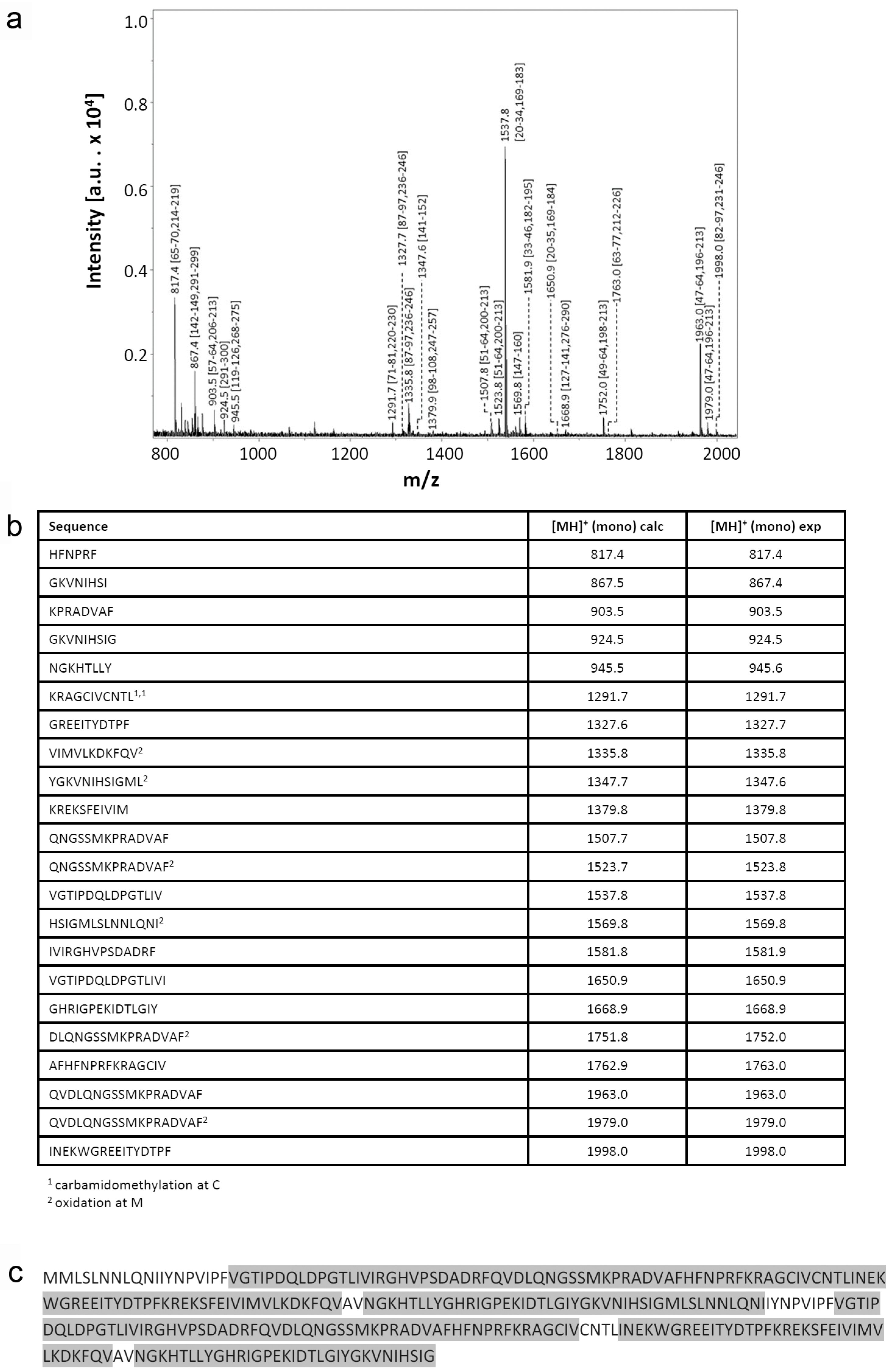
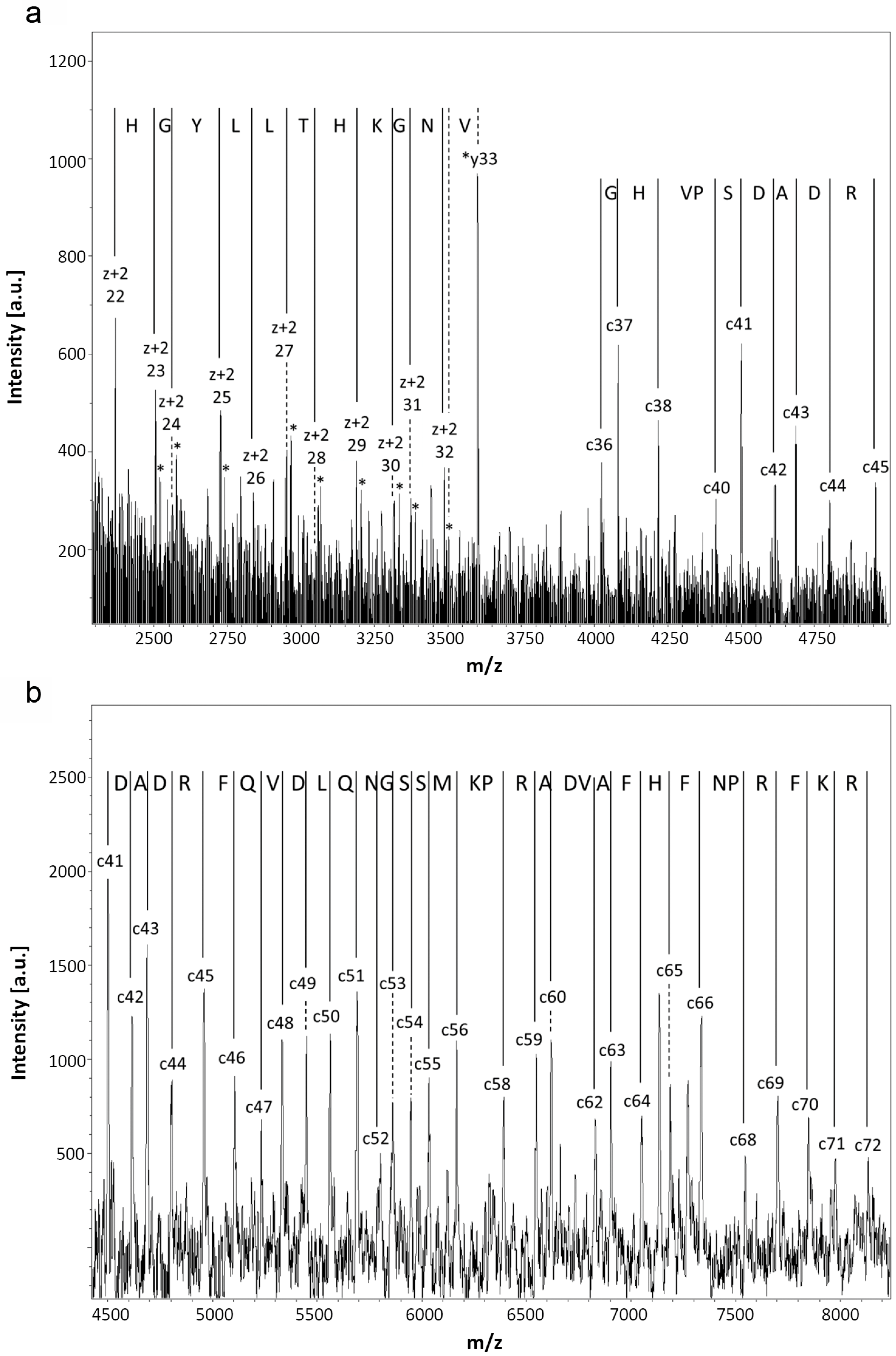
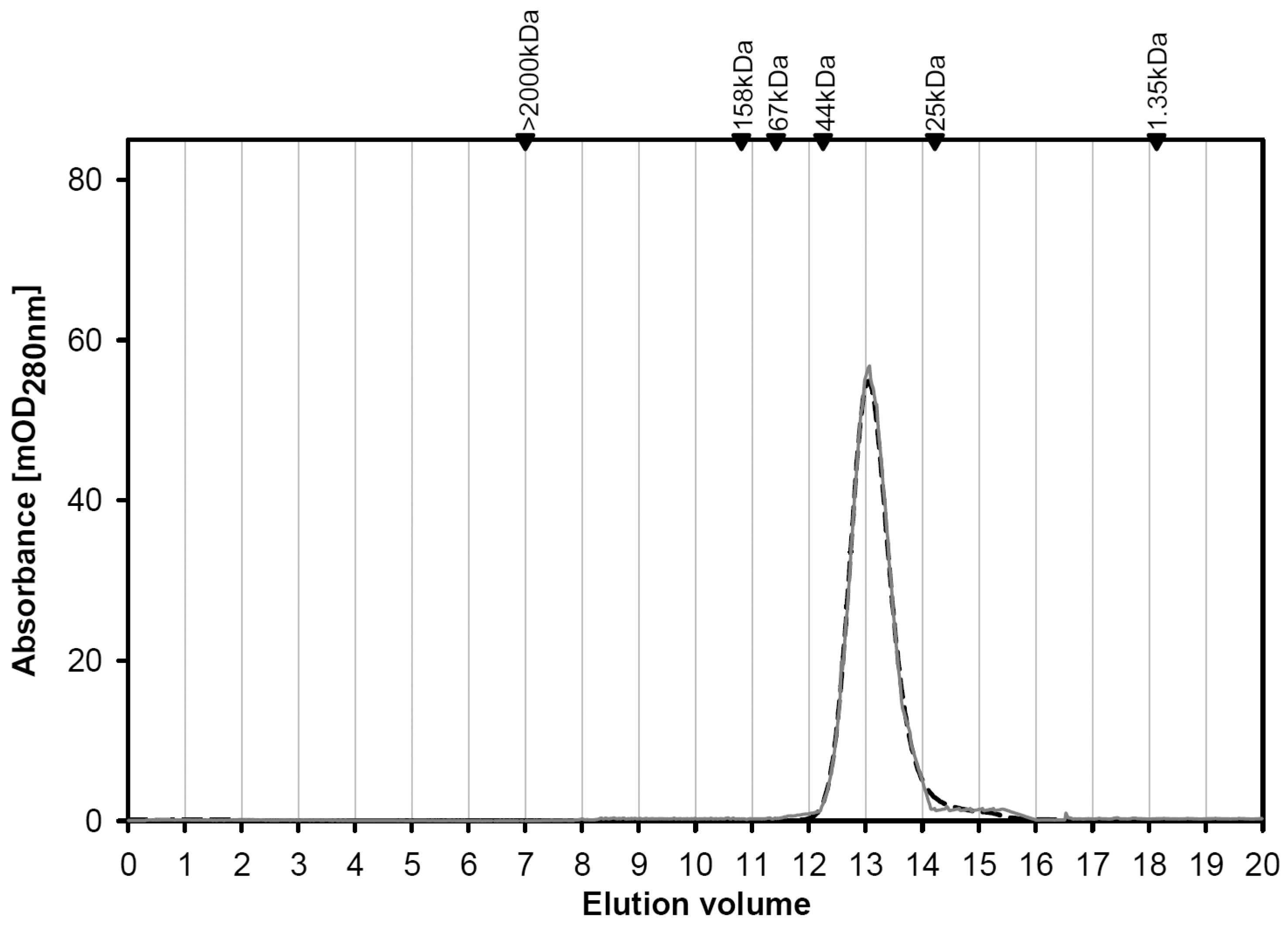
2.1.2. Structural Characterization
2.1.3. Glycan Specificity Profile
2.1.4. Cell Binding and Aggregation
3. Discussion
4. Materials and Methods
4.1. cDNA Engineering and Protein Production
4.2. Analytical Procedures
4.3. Glycan Array
4.4. Solid-Phase and Cell Assays
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kopitz, J. Glycolipids. In The Sugar Code: Fundamentals of Glycosciences; Gabius, H.-J., Ed.; Wiley-VCH: Weinheim, Germany, 2009; pp. 177–198. [Google Scholar]
- Zuber, C.; Roth, J. N-Glycosylation. In The Sugar Code: Fundamentals of Glycosciences; Gabius, H.-J., Ed.; Wiley-VCH: Weinheim, Germany, 2009; pp. 87–110. [Google Scholar]
- Corfield, A.P.; Berry, M. Glycan variation and evolution in the eukaryotes. Trends Biochem. Sci. 2015, 40, 351–359. [Google Scholar] [CrossRef] [PubMed]
- Kudelka, M.R.; Ju, T.; Heimburg-Molinaro, J.; Cummings, R.D. Simple sugars to complex disease: Mucin-type O-glycans in cancer. Adv. Cancer Res. 2015, 126, 53–135. [Google Scholar] [PubMed]
- Schengrund, C.-L. Gangliosides: Glycosphingolipids essential for normal neural development and function. Trends Biochem. Sci. 2015, 40, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Corfield, A. Eukaryotic protein glycosylation: A primer for histochemists and cell biologists. Histochem. Cell Biol. 2017, 147, 119–147. [Google Scholar] [CrossRef] [PubMed]
- Kopitz, J. Lipid glycosylation: A primer for histochemists and cell biologists. Histochem. Cell Biol. 2017, 147, 175–198. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, H. Analytical aspects: Analysis of protein-bound glycans. In The Sugar Code: Fundamentals of Glycosciences; Gabius, H.-J., Ed.; Wiley-VCH: Weinheim, Germany, 2009; pp. 71–83. [Google Scholar]
- Nishimura, S.-I. Toward automated glycan analysis. Adv. Carbohydr. Chem. Biochem. 2011, 65, 219–271. [Google Scholar] [PubMed]
- Lazar, I.M.; Lee, W.; Lazar, A.C. Glycoproteomics on the rise: Established methods, advanced techniques, sophisticated biological applications. Electrophoresis 2013, 34, 113–125. [Google Scholar] [CrossRef] [PubMed]
- Novotny, M.V.; Alley, W.R.; Mann, B.F. Analytical glycobiology at high sensitivity: Current approaches and directions. Glycoconj. J. 2013, 30, 89–117. [Google Scholar] [CrossRef] [PubMed]
- Gabius, H.-J.; Roth, J. An introduction to the sugar code. Histochem. Cell Biol. 2017, 147, 111–117. [Google Scholar] [CrossRef] [PubMed]
- Roth, J. The lectins: Molecular probes in cell biology and membrane research. Exp. Pathol. 1978, Suppl., 11–29. [Google Scholar]
- Caselitz, J. Lectins and blood group substances as “tumor markers”. Curr. Top. Pathol. 1987, 77, 245–277. [Google Scholar] [PubMed]
- Danguy, A.; Akif, F.; Pajak, B.; Gabius, H.-J. Contribution of carbohydrate histochemistry to glycobiology. Histol. Histopathol. 1994, 9, 155–171. [Google Scholar] [PubMed]
- Drake, P.M.; Cho, W.; Li, B.; Prakobphol, A.; Johansen, E.; Anderson, N.L.; Regnier, F.E.; Gibson, B.W.; Fisher, S.J. Sweetening the pot: Adding glycosylation to the biomarker discovery equation. Clin. Chem. 2010, 56, 223–236. [Google Scholar] [CrossRef] [PubMed]
- André, S.; Kaltner, H.; Manning, J.C.; Murphy, P.V.; Gabius, H.-J. Lectins: Getting familiar with translators of the sugar code. Molecules 2015, 20, 1788–1823. [Google Scholar] [CrossRef] [PubMed]
- Solís, D.; Bovin, N.V.; Davis, A.P.; Jiménez-Barbero, J.; Romero, A.; Roy, R.; Smetana, K.; Gabius, H.-J. A guide into glycosciences: How chemistry, biochemistry and biology cooperate to crack the sugar code. Biochim. Biophys. Acta 2015, 1850, 186–235. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Hsueh, P.; Kletter, D.; Bern, M.; Haab, B. The detection and discovery of glycan motifs in biological samples using lectins and antibodies: New methods and opportunities. Adv. Cancer Res. 2015, 126, 167–202. [Google Scholar] [PubMed]
- Manning, J.C.; Romero, A.; Habermann, F.A.; García Caballero, G.; Kaltner, H.; Gabius, H.-J. Lectins: A primer for histochemists and cell biologists. Histochem. Cell Biol. 2017, 147, 199–222. [Google Scholar] [CrossRef] [PubMed]
- Kilpatrick, D.C. Animal lectins: A historical introduction and overview. Biochim. Biophys. Acta 2002, 1572, 187–197. [Google Scholar] [CrossRef]
- Gabius, H.-J.; Kaltner, H.; Kopitz, J.; André, S. The glycobiology of the CD system: A dictionary for translating marker designations into glycan/lectin structure and function. Trends Biochem. Sci. 2015, 40, 360–376. [Google Scholar] [CrossRef] [PubMed]
- Gabius, H.-J.; Manning, J.C.; Kopitz, J.; André, S.; Kaltner, H. Sweet complementarity: The functional pairing of glycans with lectins. Cell. Mol. Life Sci. 2016, 73, 1989–2016. [Google Scholar] [CrossRef] [PubMed]
- Von der Lieth, C.-W.; Siebert, H.-C.; Kozár, T.; Burchert, M.; Frank, M.; Gilleron, M.; Kaltner, H.; Kayser, G.; Tajkhorshid, E.; Bovin, N.V.; et al. Lectin ligands: New insights into their conformations and their dynamic behavior and the discovery of conformer selection by lectins. Acta Anat. 1998, 161, 91–109. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gabius, H.-J.; André, S.; Jiménez-Barbero, J.; Romero, A.; Solís, D. From lectin structure to functional glycomics: Principles of the sugar code. Trends Biochem. Sci. 2011, 36, 298–313. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.C.; Lee, R.T.; Rice, K.G.; Ichikawa, Y.; Wong, T.C. Topography of binding sites of animal lectins: Ligands’ view. Pure Appl. Chem. 1991, 63, 499–506. [Google Scholar] [CrossRef]
- Chabre, Y.M.; Roy, R. The chemist’s way to prepare multivalency. In The Sugar Code: Fundamentals of Glycosciences; Gabius, H.-J., Ed.; Wiley-VCH: Weinheim, Germany, 2009; pp. 53–70. [Google Scholar]
- Murphy, P.V.; André, S.; Gabius, H.-J. The third dimension of reading the sugar code by lectins: Design of glycoclusters with cyclic scaffolds as tools with the aim to define correlations between spatial presentation and activity. Molecules 2013, 18, 4026–4053. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cecioni, S.; Imberty, A.; Vidal, S. Glycomimetics versus multivalent glycoconjugates for the design of high affinity lectin ligands. Chem. Rev. 2015, 115, 525–561. [Google Scholar] [CrossRef] [PubMed]
- Roy, R.; Murphy, P.V.; Gabius, H.-J. Multivalent carbohydrate-lectin interactions: How synthetic chemistry enables insights into nanometric recognition. Molecules 2016, 21, 629. [Google Scholar] [CrossRef] [PubMed]
- Hirabayashi, J. Recent topics on galectins. Trends Glycosci. Glycotechnol. 1997, 9, 1–180. [Google Scholar]
- Cooper, D.N.W. Galectinomics: Finding themes in complexity. Biochim. Biophys. Acta 2002, 1572, 209–231. [Google Scholar] [CrossRef]
- Kaltner, H.; Toegel, S.; García Caballero, G.; Manning, J.C.; Ledeen, R.W.; Gabius, H.-J. Galectins: Their network and roles in immunity/tumor growth control. Histochem. Cell Biol. 2017, 147, 239–256. [Google Scholar] [CrossRef] [PubMed]
- Zick, Y.; Eisenstein, M.; Goren, R.A.; Hadari, Y.R.; Levy, Y.; Ronen, D. Role of galectin-8 as a modulator of cell adhesion and cell growth. Glycoconj. J. 2004, 19, 517–526. [Google Scholar] [CrossRef] [PubMed]
- Cludts, S.; Decaestecker, C.; Mahillon, V.; Chevalier, D.; Kaltner, H.; André, S.; Remmelink, M.; Leroy, X.; Gabius, H.-J.; Saussez, S. Galectin-8 up-regulation during hypopharyngeal and laryngeal tumor progression and comparison with galectins-1, -3 and -7. Anticancer Res. 2009, 29, 4933–4940. [Google Scholar] [PubMed]
- Cattaneo, V.; Tribulatti, M.V.; Campetella, O. Galectin-8 tandem-repeat structure is essential for T-cell proliferation but not for co-stimulation. Biochem. J. 2011, 434, 153–160. [Google Scholar] [CrossRef] [PubMed]
- Cattaneo, V.; Tribulatti, M.V.; Carabelli, J.; Carestia, A.; Schattner, M.; Campetella, O. Galectin-8 elicits pro-inflammatory activities in the endothelium. Glycobiology 2014, 24, 966–973. [Google Scholar] [CrossRef] [PubMed]
- Vinik, Y.; Shatz-Azoulay, H.; Vivanti, A.; Hever, N.; Levy, Y.; Karmona, R.; Brumfeld, V.; Baraghithy, S.; Attar-Lamdar, M.; Boura-Halfon, S.; et al. The mammalian lectin galectin-8 induces RANKL expression, osteoclastogenesis, and bone mass reduction in mice. eLife 2015, 4, e05914. [Google Scholar] [CrossRef] [PubMed]
- Houzelstein, D.; Gonçalves, I.R.; Fadden, A.J.; Sidhu, S.S.; Cooper, D.N.W.; Drickamer, K.; Leffler, H.; Poirier, F. Phylogenetic analysis of the vertebrate galectin family. Mol. Biol. Evol. 2004, 21, 1177–1187. [Google Scholar] [CrossRef] [PubMed]
- Hirabayashi, J.; Hashidate, T.; Arata, Y.; Nishi, N.; Nakamura, T.; Hirashima, M.; Urashima, T.; Oka, T.; Futai, M.; Müller, W.E.G.; et al. Oligosaccharide specificity of galectins: A search by frontal affinity chromatography. Biochim. Biophys. Acta 2002, 1572, 232–254. [Google Scholar] [CrossRef]
- Ideo, H.; Seko, A.; Ishizuka, I.; Yamashita, K. The N-terminal carbohydrate recognition domain of galectin-8 recognizes specific glycosphingolipids with high affinity. Glycobiology 2003, 13, 713–723. [Google Scholar] [CrossRef] [PubMed]
- Ideo, H.; Matsuzaka, T.; Nonaka, T.; Seko, A.; Yamashita, K. Galectin-8-N-domain recognition mechanism for sialylated and sulfated glycans. J. Biol. Chem. 2011, 286, 11346–11355. [Google Scholar] [CrossRef] [PubMed]
- Carlsson, S.; Oberg, C.T.; Carlsson, M.C.; Sundin, A.; Nilsson, U.J.; Smith, D.; Cummings, R.D.; Almkvist, J.; Karlsson, A.; Leffler, H. Affinity of galectin-8 and its carbohydrate recognition domains for ligands in solution and at the cell surface. Glycobiology 2007, 17, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Stowell, S.R.; Arthur, C.M.; Slanina, K.A.; Horton, J.R.; Smith, D.F.; Cummings, R.D. Dimeric Galectin-8 induces phosphatidylserine exposure in leukocytes through polylactosamine recognition by the C-terminal domain. J. Biol. Chem. 2008, 283, 20547–20559. [Google Scholar] [CrossRef] [PubMed]
- Vokhmyanina, O.A.; Rapoport, E.M.; André, S.; Severov, V.V.; Ryzhov, I.; Pazynina, G.V.; Korchagina, E.; Gabius, H.-J.; Bovin, N.V. Comparative study of the glycan specificities of cell-bound human tandem-repeat-type galectins-4, -8 and -9. Glycobiology 2012, 22, 1207–1217. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, H.; Yamashita, S.; Teraoka, M.; Itoh, A.; Nakakita, S.; Nishi, N.; Kamitori, S. X-ray structure of a protease-resistant mutant form of human galectin-8 with two carbohydrate recognition domains. FEBS J. 2012, 279, 3937–3951. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, F.M.; Scholz, B.A.; Buzamet, E.; Kopitz, J.; André, S.; Menéndez, M.; Romero, A.; Solís, D.; Gabius, H.-J. Natural single amino acid polymorphism (F19Y) in human galectin-8: Detection of structural alterations and increased growth-regulatory activity on tumor cells. FEBS J. 2014, 281, 1446–1464. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, F.M.; Gilles, U.; Lindner, I.; André, S.; Romero, A.; Reusch, D.; Gabius, H.-J. Combining crystallography and hydrogen-deuterium exchange to study galectin-ligand complexes. Chem. Eur. J. 2015, 21, 13558–13568. [Google Scholar] [CrossRef] [PubMed]
- Romaniuk, M.A.; Tribulatti, M.V.; Cattaneo, V.; Lapponi, M.J.; Molinas, F.C.; Campetella, O.; Schattner, M. Human platelets express and are activated by galectin-8. Biochem. J. 2010, 432, 535–547. [Google Scholar] [CrossRef] [PubMed]
- Joziasse, D.H.; Schiphorst, W.E.C.M.; van den Eijnden, D.H.; van Kuik, J.A.; van Halbeek, H.; Vliegenthart, J.F.G. Branch specificity of bovine colostrum CMP-sialic acid: Galβ1→4GlcNAc-R α2→6-sialyltransferase. Sialylation of bi-, tri-, and tetraantennary oligosaccharides and glycopeptides of the N-acetyllactosamine type. J. Biol. Chem. 1987, 262, 2025–2033. [Google Scholar] [PubMed]
- Amano, M.; Eriksson, H.; Manning, J.C.; Detjen, K.M.; André, S.; Nishimura, S.-I.; Lehtiö, J.; Gabius, H.-J. Tumour suppressor p16INK4a: Anoikis-favouring decrease in N/O-glycan/cell surface sialylation by down-regulation of enzymes in sialic acid biosynthesis in tandem in a pancreatic carcinoma model. FEBS J. 2012, 279, 4062–4080. [Google Scholar] [CrossRef] [PubMed]
- Gunther, G.R.; Wang, J.L.; Yahara, I.; Cunningham, B.A.; Edelman, G.M. Concanavalin A derivatives with altered biological activities. Proc. Natl. Acad. Sci. USA 1973, 70, 1012–1016. [Google Scholar] [CrossRef] [PubMed]
- Fraser, A.R.; Hemperly, J.J.; Wang, J.L.; Edelman, G.M. Monovalent derivatives of concanavalin A. Proc. Natl. Acad. Sci. USA 1976, 73, 790–794. [Google Scholar] [CrossRef] [PubMed]
- Beppu, M.; Terao, T.; Osawa, T. Covalently cross-linked monovalent, divalent, and tetravalent derivatives of concanavalin A. J. Biochem. 1979, 85, 1275–1287. [Google Scholar] [PubMed]
- Kurokawa, T.; Tsuda, M.; Sugino, Y. Purification and characterization of a lectin from Wistaria floribunda seeds. J. Biol. Chem. 1976, 251, 5686–5693. [Google Scholar] [PubMed]
- Kaku, H.; Shibuya, N. Preparation of a stable subunit of Japanese elderberry (Sambucus sieboldiana) bark lectin and its application for the study of cell surface carbohydrates by flow cytometry. FEBS Lett. 1992, 306, 176–180. [Google Scholar] [CrossRef]
- Swanson, M.D.; Boudreaux, D.M.; Salmon, L.; Chugh, J.; Winter, H.C.; Meagher, J.L.; André, S.; Murphy, P.V.; Oscarson, S.; Roy, R.; et al. Engineering a therapeutic lectin by uncoupling mitogenicity from antiviral activity. Cell 2015, 163, 746–758. [Google Scholar] [CrossRef] [PubMed]
- Bättig, P.; Saudan, P.; Gunde, T.; Bachmann, M.F. Enhanced apoptotic activity of a structurally optimized form of galectin-1. Mol. Immunol. 2004, 41, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Bi, S.; Earl, L.A.; Jacobs, L.; Baum, L.G. Structural features of galectin-9 and galectin-1 that determine distinct T cell death pathways. J. Biol. Chem. 2008, 283, 12248–12258. [Google Scholar] [CrossRef] [PubMed]
- Earl, L.A.; Bi, S.; Baum, L.G. Galectin multimerization and lattice formation are regulated by linker region structure. Glycobiology 2011, 21, 6–12. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Moussodia, R.-O.; Murzeau, C.; Sun, H.J.; Klein, M.L.; Vértesy, S.; André, S.; Roy, R.; Gabius, H.-J.; Percec, V. Dissecting molecular aspects of cell interactions using glycodendrimersomes with programmable glycan presentation and engineered human lectins. Angew. Chem. Int. Ed. 2015, 54, 4036–4040. [Google Scholar] [CrossRef] [PubMed]
- Vértesy, S.; Michalak, M.; Miller, M.C.; Schnölzer, M.; André, S.; Kopitz, J.; Mayo, K.H.; Gabius, H.-J. Structural significance of galectin design: Impairment of homodimer stability by linker insertion and partial reversion by ligand presence. Protein Eng. Des. Sel. 2015, 28, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.C.; Honjo, Y.; Nangia-Makker, P.; Hogan, V.; Mazurak, N.; Bresalier, R.S.; Raz, A. The NH2 terminus of galectin-3 governs cellular compartmentalization and functions in cancer cells. Cancer Res. 1999, 59, 6239–6245. [Google Scholar] [PubMed]
- Menon, R.P.; Hughes, R.C. Determinants in the N-terminal domains of galectin-3 for secretion by a novel pathway circumventing the endoplasmic reticulum-Golgi complex. Eur. J. Biochem. 1999, 264, 569–576. [Google Scholar] [CrossRef] [PubMed]
- Kopitz, J.; Vértesy, S.; André, S.; Fiedler, S.; Schnölzer, M.; Gabius, H.-J. Human chimera-type galectin-3: Defining the critical tail length for high-affinity glycoprotein/cell surface binding and functional competition with galectin-1 in neuroblastoma cell growth regulation. Biochimie 2014, 104, 90–99. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, A.K.; Vértesy, S.; Michalak, M.; Manning, J.C.; André, S.; Kübler, D.; Kopitz, J.; Kaltner, H.; Gabius, H.-J. Playing modular puzzle with adhesion/growth-regulatory galectins: Design and testing of a hybrid to unravel structure-activity relationships. Protein Pept. Lett. 2016, 23, 1003–1012. [Google Scholar] [CrossRef] [PubMed]
- Sato, M.; Nishi, N.; Shoji, H.; Seki, M.; Hashidate, T.; Hirabayashi, J.; Kasai, K.-I.; Hata, Y.; Suzuki, S.; Hirashima, M.; et al. Functional analysis of the carbohydrate recognition domains and a linker peptide of galectin-9 as to eosinophil chemoattractant activity. Glycobiology 2002, 12, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.H.; Nakagawa, R.; Kashio, Y.; Ito, A.; Shoji, H.; Nishi, N.; Hirashima, M.; Yamauchi, A.; Nakamura, T. Characterization of galectin-9-induced death of Jurkat T cells. J. Biochem. 2007, 141, 157–172. [Google Scholar] [CrossRef] [PubMed]
- Göhler, A.; André, S.; Kaltner, H.; Sauer, M.; Gabius, H.-J.; Doose, S. Hydrodynamic properties of human adhesion/growth-regulatory galectins studied by fluorescence correlation spectroscopy. Biophys. J. 2010, 98, 3044–3053. [Google Scholar] [CrossRef] [PubMed]
- Kopitz, J.; Ballikaya, S.; André, S.; Gabius, H.-J. Ganglioside GM1/galectin-dependent growth regulation in human neuroblastoma cells: Special properties of bivalent galectin-4 and significance of linker length for ligand selection. Neurochem. Res. 2012, 37, 1267–1276. [Google Scholar] [CrossRef] [PubMed]
- André, S.; Wang, G.N.; Gabius, H.-J.; Murphy, P.V. Combining glycocluster synthesis with protein engineering: An approach to probe into the significance of linker length in a tandem-repeat-type lectin (galectin-4). Carbohydr. Res. 2014, 389, 25–38. [Google Scholar] [CrossRef] [PubMed]
- Jovanović, M.; Peter-Katalinić, J. Preliminary mass spectrometry characterization studies of galectin-3 samples, prior to carbohydrate-binding studies using affinity mass spectrometry. Rapid Commun. Mass Spectrom. 2017, 31, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Tu, Z.; Hsieh, H.W.; Tsai, C.M.; Hsu, C.W.; Wang, S.G.; Wu, K.J.; Lin, K.I.; Lin, C.H. Synthesis and characterization of sulfated Gal-β1,3/4-GlcNAc disaccharides through consecutive protection/glycosylation steps. Chem. Asian J. 2013, 8, 1536–1550. [Google Scholar] [CrossRef] [PubMed]
- Carlsson, S.; Carlsson, M.C.; Leffler, H. Intracellular sorting of galectin-8 based on carbohydrate fine specificity. Glycobiology 2007, 17, 906–912. [Google Scholar] [CrossRef] [PubMed]
- Levy, Y.; Arbel-Goren, R.; Hadari, Y.R.; Eshhar, S.; Ronen, D.; Elhanany, E.; Geiger, B.; Zick, Y. Galectin-8 functions as a matricellular modulator of cell adhesion. J. Biol. Chem. 2001, 276, 31285–31295. [Google Scholar] [CrossRef] [PubMed]
- Friedel, M.; André, S.; Goldschmidt, H.; Gabius, H.-J.; Schwartz-Albiez, R. Galectin-8 enhances adhesion of multiple myeloma cells to vascular endothelium and is an adverse prognostic factor. Glycobiology 2016, 26, 1048–1058. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.; Huang, H.; Tateno, H.; Nakakita, S.; Sato, T.; Narimatsu, H.; Yao, X.; Hirabayashi, J. Engineering of a 3’-sulpho-Galβ1-4GlcNAc-specific probe by a single amino acid substitution of a fungal galectin. J. Biochem. 2015, 157, 197–200. [Google Scholar] [CrossRef] [PubMed]
- Bhide, G.P.; Colley, K.J. Sialylation of N-glycans: Mechanism, cellular compartmentalization and function. Histochem. Cell Biol. 2017, 147, 149–174. [Google Scholar] [CrossRef] [PubMed]
- Roy, R.; Cao, Y.; Kaltner, H.; Kottari, N.; Shiao, T.C.; Belkhadem, K.; André, S.; Manning, J.C.; Murphy, P.V.; Gabius, H.-J. Teaming up synthetic chemistry and histochemistry for activity screening in galectin-directed inhibitor design. Histochem. Cell Biol. 2017, 147, 285–301. [Google Scholar] [CrossRef] [PubMed]
- Yabe, R.; Itakura, Y.; Nakamura-Tsuruta, S.; Iwaki, J.; Kuno, A.; Hirabayashi, J. Engineering a versatile tandem repeat-type α2-6sialic acid-binding lectin. Biochem. Biophys. Res. Commun. 2009, 384, 204–209. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.; Tateno, H.; Hirabayashi, J. Lectin engineering, a molecular evolutionary approach to expanding the lectin utilities. Molecules 2015, 20, 7637–7656. [Google Scholar] [CrossRef] [PubMed]
- Kaltner, H.; Solís, D.; André, S.; Lensch, M.; Manning, J.C.; Mürnseer, M.; Sáiz, J.L.; Gabius, H.-J. Unique chicken tandem-repeat-type galectin: Implications of alternative splicing and a distinct expression profile compared to those of the three proto-type proteins. Biochemistry 2009, 48, 4403–4416. [Google Scholar] [CrossRef] [PubMed]
- Gabius, H.-J.; Wosgien, B.; Hendrys, M.; Bardosi, A. Lectin localization in human nerve by biochemically defined lectin-binding glycoproteins, neoglycoprotein and lectin-specific antibody. Histochemistry 1991, 95, 269–277. [Google Scholar] [CrossRef] [PubMed]
- Kaltner, H.; García Caballero, G.; Sinowatz, F.; Schmidt, S.; Manning, J.C.; André, S.; Gabius, H.-J. Galectin-related protein: An integral member of the network of chicken galectins. 2. From expression profiling to its immunocyto- and histochemical localization and application as tool for ligand detection. Biochim. Biophys. Acta 2016, 1860, 2298–2312. [Google Scholar] [CrossRef] [PubMed]
- Manning, J.C.; García Caballero, G.; Knospe, C.; Kaltner, H.; Gabius, H.-J. Network analysis of adhesion/growth-regulatory galectins and their binding sites in adult chicken retina and choroid. J. Anat. 2017, 231, 23–37. [Google Scholar] [CrossRef] [PubMed]
- García Caballero, G.; Flores-Ibarra, A.; Michalak, M.; Khasbiullina, N.; Bovin, N.V.; André, S.; Manning, J.C.; Vértesy, S.; Ruiz, F.M.; Kaltner, H.; et al. Galectin-related protein: An integral member of the network of chicken galectins. 1. From strong sequence conservation of the gene confined to vertebrates to biochemical characteristics of the chicken protein and its crystal structure. Biochim. Biophys. Acta 2016, 1860, 2285–2297. [Google Scholar] [CrossRef] [PubMed]
- García Caballero, G.; Kaltner, H.; Michalak, M.; Shilova, N.; Yegres, M.; André, S.; Ludwig, A.K.; Manning, J.C.; Schmidt, S.; Schnölzer, M.; et al. Chicken GRIFIN: A homodimeric member of the galectin network with canonical properties and a unique expression profile. Biochimie 2016, 128–129, 34–47. [Google Scholar] [CrossRef] [PubMed]
- André, S.; Kaltner, H.; Lensch, M.; Russwurm, R.; Siebert, H.-C.; Fallsehr, C.; Tajkhorshid, E.; Heck, A.J.R.; von Knebel-Döberitz, M.; Gabius, H.-J.; et al. Determination of structural and functional overlap/divergence of five proto-type galectins by analysis of the growth-regulatory interaction with ganglioside GM1 in silico and in vitro on human neuroblastoma cells. Int. J. Cancer 2005, 114, 46–57. [Google Scholar] [CrossRef] [PubMed]
- André, S.; Sansone, F.; Kaltner, H.; Casnati, A.; Kopitz, J.; Gabius, H.-J.; Ungaro, R. Calix[n]arene-based glycoclusters: Bioactivity of thiourea-linked galactose/lactose moieties as inhibitors of binding of medically relevant lectins to a glycoprotein and cell-surface glycoconjugates and selectivity among human adhesion/growth-regulatory galectins. ChemBioChem 2008, 9, 1649–1661. [Google Scholar] [PubMed]
- Gabius, H.-J.; Engelhardt, R.; Rehm, S.; Cramer, F. Biochemical characterization of endogenous carbohydrate-binding proteins from spontaneous murine rhabdomyosarcoma, mammary adenocarcinoma, and ovarian teratoma. J. Natl. Cancer Inst. 1984, 73, 1349–1357. [Google Scholar] [PubMed]
Sample Availability: Not available. |



| Structure | Median | MAD # |
|---|---|---|
| Neu5Acα2,3Galβ1,3(6-O-Su)GlcNAc | 59,796 | 1931 |
| Neu5Acα2,3Galβ1,4Glc-Gly-Phe * | 43,636 | 4717 |
| Neu5Acα2,8Neu5Acα2,3Galβl,4Glc | 31,204 | 1141 |
| Neu5Aα2,3Galβ1,3GlcNAcβ1,3Galβ1,4Glc | 18,278 | 6705 |
| Neu5Acα2,3Galβ1,3GlcNAc | 17,215 | 1619 |
| Neu5Acα2,3Galβ1,4Glc | 8888 | 294 |
| 3-O-Su-Galβ1,4(6-O-Su)Glc | 5279 | 146 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ludwig, A.-K.; Michalak, M.; Shilova, N.; André, S.; Kaltner, H.; Bovin, N.V.; Kopitz, J.; Gabius, H.-J. Studying the Structural Significance of Galectin Design by Playing a Modular Puzzle: Homodimer Generation from Human Tandem-Repeat-Type (Heterodimeric) Galectin-8 by Domain Shuffling. Molecules 2017, 22, 1572. https://doi.org/10.3390/molecules22091572
Ludwig A-K, Michalak M, Shilova N, André S, Kaltner H, Bovin NV, Kopitz J, Gabius H-J. Studying the Structural Significance of Galectin Design by Playing a Modular Puzzle: Homodimer Generation from Human Tandem-Repeat-Type (Heterodimeric) Galectin-8 by Domain Shuffling. Molecules. 2017; 22(9):1572. https://doi.org/10.3390/molecules22091572
Chicago/Turabian StyleLudwig, Anna-Kristin, Malwina Michalak, Nadya Shilova, Sabine André, Herbert Kaltner, Nicolai V. Bovin, Jürgen Kopitz, and Hans-Joachim Gabius. 2017. "Studying the Structural Significance of Galectin Design by Playing a Modular Puzzle: Homodimer Generation from Human Tandem-Repeat-Type (Heterodimeric) Galectin-8 by Domain Shuffling" Molecules 22, no. 9: 1572. https://doi.org/10.3390/molecules22091572





