Chemical Fingerprint and Multicomponent Quantitative Analysis for the Quality Evaluation of Cyclocarya paliurus Leaves by HPLC–Q–TOF–MS
Abstract
:1. Introduction
2. Results and Discussion
2.1. Optimization of Sample Extraction and Chromatographic Conditions
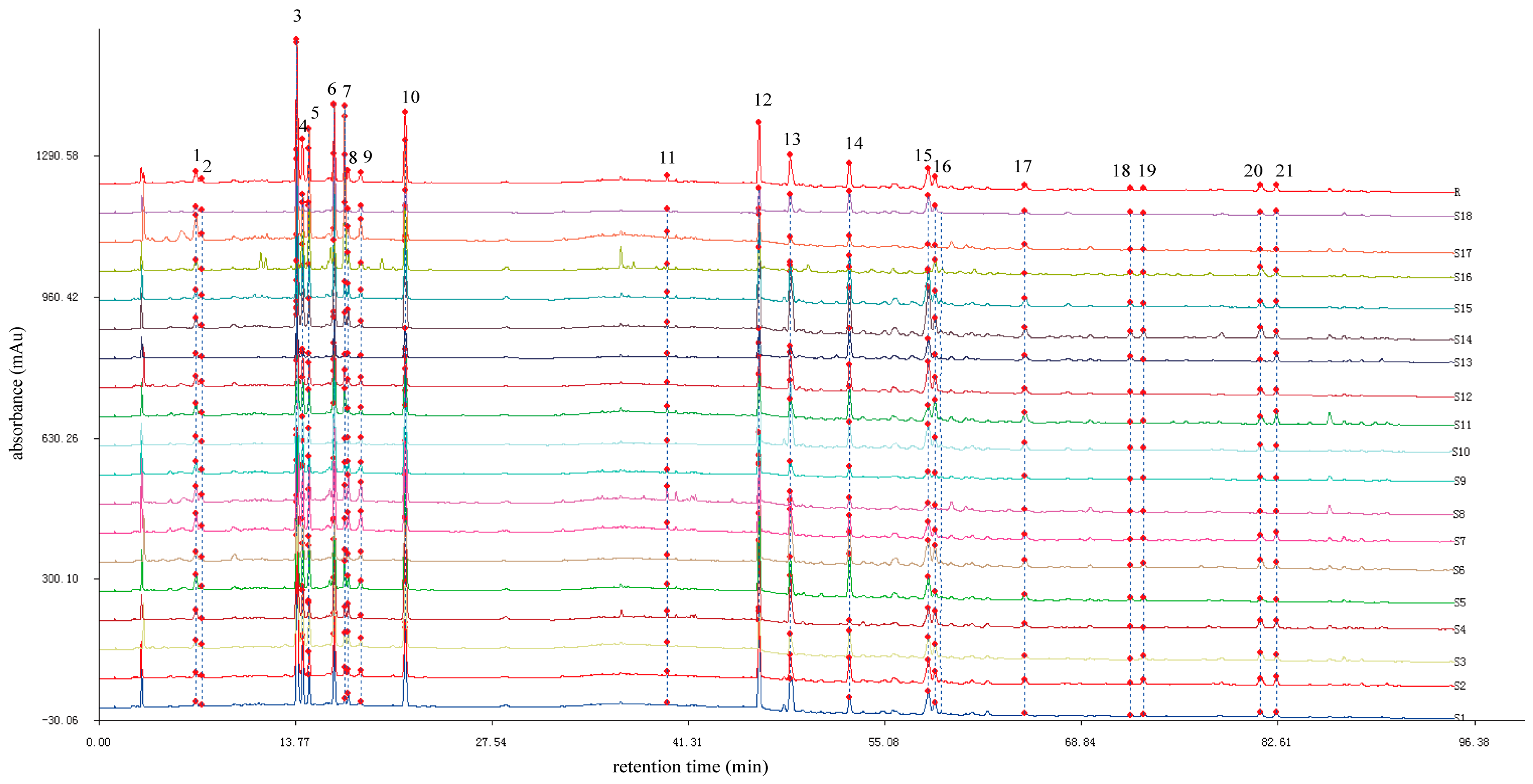
2.2. HPLC Fingerprint Establishment and Similarity Analysis
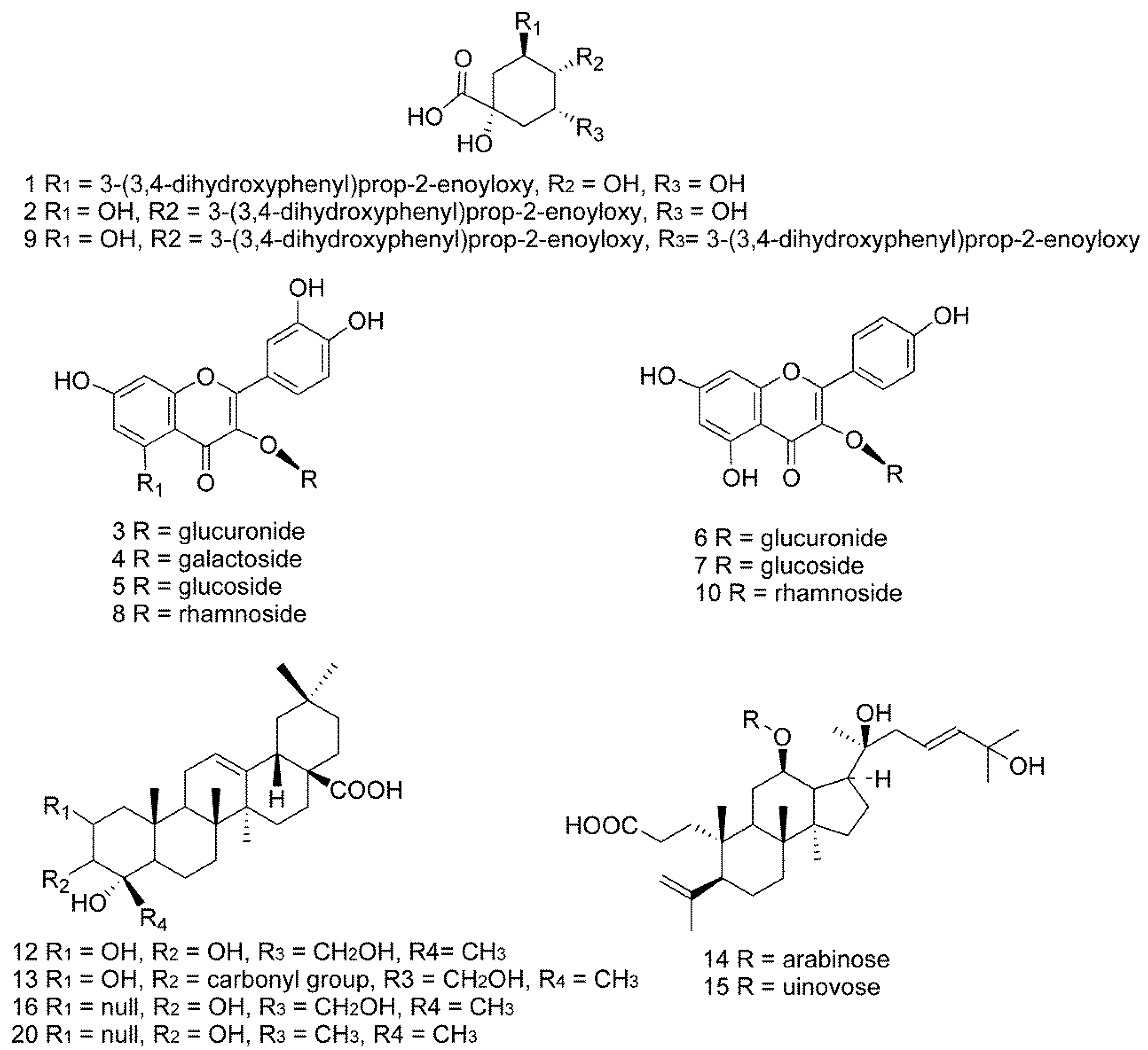
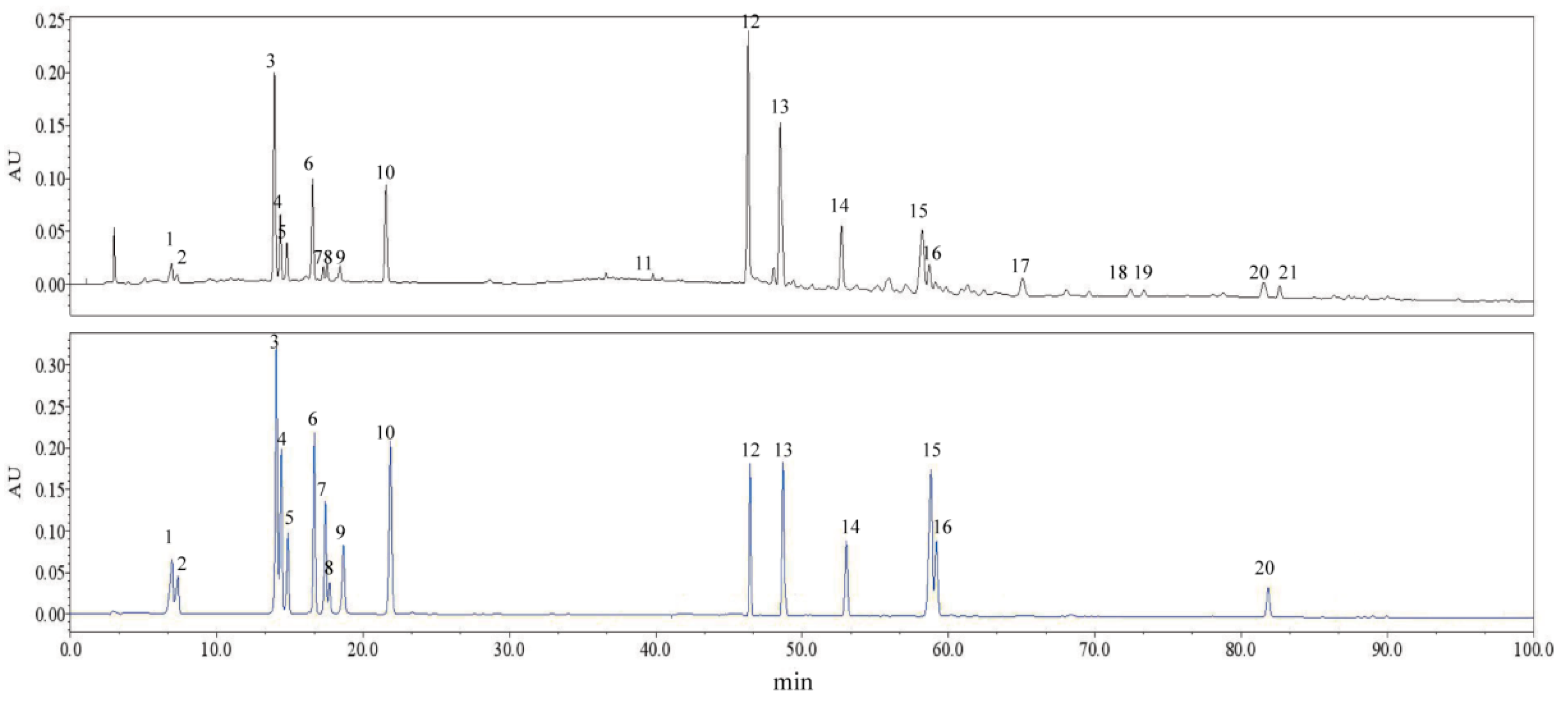
2.3. Identification of Characteristic Common Peaks in C. paliurus Leaves
2.3.1. Identification of Phenolic Acids
2.3.2. Identification of Flavonoids
2.3.3. Identification of Triterpenoids
2.4. Method Validation
2.5. Quantification of Chemical Components in C. paliurus Leaves
3. Materials and Methods
3.1. Plant Materials
3.2. Chemical Reagents and References
3.3. HPLC Instrument and Chromatographic Conditions
3.4. HPLC–Q–TOF–MS Confirmation Analysis
3.5. Sample and Standard Solution Preparation
3.6. Method Validation
3.7. Data Analysis
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Fang, S.Z.; Wang, J.Y.; Wei, Z.Y.; Zhu, Z.X. Methods to break seed dormancy in Cyclocarya paliurus (Batal.) Iljinskaja. Sci. Hortic. 2006, 110, 305–309. [Google Scholar] [CrossRef]
- Birari, R.B.; Bhutani, K.K. Pancreatic lipase inhibitors from natural source: Unexplored potential. Drug Discov. Today 2007, 12, 879–889. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.H.; Xie, M.Y.; Nie, S.P.; Shen, M.Y.; Wang, Y.X.; Li, C. Isolation, chemical composition and antioxidant activities of a watersoluble polysaccharide from Cyclocarya paliurus (Batal.) Iljinskaja. Food Chem. 2010, 119, 1626–1632. [Google Scholar] [CrossRef]
- Hou, X.L.; Liu, X.X.; Wang, S.; Zhou, X.L.; Wei, F.; Miao, J.H. Effect of the flavonoids from Cyclocarya paliurus on spontaneous hypertension rats. Pharm. Clin. Chin. Mater. Med. 2014, 30, 62–69. [Google Scholar]
- Kurihara, H.; Fukami, H.; Kusumoto, A.; Toyoda, Y.; Shibata, H.; Matsui, Y. Hypoglycemic action of Cyclocarya paliurus (Batal.) Iljinskaja in normal and diabetic mice. Biosci. Biotechnol. Biochem. 2003, 67, 877–880. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Shen, Q.; Lu, J.C.; Li, J.Y.; Liu, W.Y.; Yang, J.J.; Li, J.; Xiao, K. Phenolic compounds from the leaves of Cyclocarya paliurus (Batal.) Ijinskaja and their inhibitory activity against PTP1B. Food Chem. 2010, 119, 1491–1496. [Google Scholar] [CrossRef]
- Kurihara, H.; Asami, S.; Shibata, H.; Fukami, H.; Tanaka, T. Hypolipemic effect of Cyclocarya paliurus (Batal) Iljinskaja in lipid-loaded mice. Biol. Pharm. Bull. 2003, 26, 383–385. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.W.; Ouyang, K.H.; Zhao, J.; Chen, H.; Xiong, L.; Wang, W.J. Structural characterization and hypolipidemic effect of Cyclocarya paliurus polysaccharide in rat. Int. J. Biol. Macromol. 2016, 91, 1073–1080. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.F.; Gao, T.H.; Zhong, R.L.; Lin, Z.; Jiang, C.H.; Ouyang, S.; Zhao, M.; Che, C.T.; Zhang, J.; Yin, Z.Q. Antihyperlipidaemic effect of triterpenic acid-enriched fraction from Cyclocarya paliurus leaves in hyperlipidaemic rats. Pharm. Biol. 2017, 55, 712–721. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Xie, J.H.; Jia, S.; Huang, L.X.; Wang, Z.J.; Li, C.; Xie, M.Y. Immunomodulatory effects of an acetylated Cyclocarya paliurus polysaccharide on murine macrophages raw264.7. Int. J. Biol. Macromol. 2017, 98, 576–581. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.H.; Liu, X.; Shen, M.Y.; Nie, S.P.; Zhang, H.; Li, C.; Gong, D.M.; Xie, M.Y. Purification, physicochemical characterization and anticancer activity of a polysaccharide from Cyclocarya paliurus leaves. Food Chem. 2013, 136, 1453–1460. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Huang, N.; Lu, J.C.; Li, X.; Wang, Y.H.; Yang, L.M.; Zheng, Y.T.; Xiao, K. Water-soluble phenolic compounds and their anti-HIV-1 activities from the leaves of Cyclocarya paliurus. J. Food Drug Anal. 2010, 18, 398–404. [Google Scholar]
- Xie, J.H.; Shen, M.Y.; Xie, M.Y.; Nie, S.P.; Chen, Y.; Li, C.; Huang, D.F.; Wang, Y.X. Ultrasonic-assisted extraction, antimicrobial and antioxidant activities of Cyclocarya paliurus (Batal.) Iljinskaja polysaccharides. Carbohydr. Polym. 2012, 89, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Shu, R.G.; Xu, C.R.; Li, L.N.; Yu, Z.L. Cyclocariosides II and III: Two secodammarane triterpenoid saponins from Cyclocarya paliurus. Planta Med. 1995, 61, 551–553. [Google Scholar] [CrossRef] [PubMed]
- Shu, R.G.; Xu, C.R.; Li, L.N. Studies on the sweet principles from the leaves of Cyclocarya paliurus (Batal.) Iljinsk. Acta Pharm. Sin. 1995, 30, 757–761. [Google Scholar]
- Kennelly, E.J.; Cai, L.; Long, L.; Shamon, L.; Zaw, K.; Zhou, B.N.; Pezzuto, J.M.; Kinghorn, A.D. Novel highly sweet secodammarane glycosides from Pterocarya paliurus. J. Agric. Food Chem. 1995, 43, 2602–2607. [Google Scholar] [CrossRef]
- Wright, M.; Byrd, J.; Gao, Y.; Stubblefield, J.; Park, H.; Dunlap, N. Isolation and structural clarification of triterpenes from Cyclocarya paliurus: Cyclocaric acid A and B. Planta Med. 2014, 80, 139–143. [Google Scholar] [CrossRef]
- Li, J.K.; He, X.Y.; Li, M.Y.; Zhao, W.; Liu, L.; Kong, X.H. Chemical fingerprint and quantitative analysis for quality control of polyphenols extracted from pomegranate peel by HPLC. Food Chem. 2015, 176, 7–11. [Google Scholar] [CrossRef] [PubMed]
- Fang, S.Z.; Yang, W.X.; Chu, X.L.; Shang, X.L.; She, C.Q.; Fu, X.X. Provenance and temporal variations in selected flavonoids in leaves of Cyclocarya paliurus. Food Chem. 2011, 124, 1382–1386. [Google Scholar] [CrossRef]
- Deng, B.; Shang, X.L.; Fang, S.Z.; Li, Q.Q.; Fu, X.X.; Su, J. Integrated effects of light intensity and fertilization on growth and flavonoid accumulation in Cyclocarya paliurus. J. Agric. Food Chem. 2012, 60, 6286–6292. [Google Scholar] [CrossRef] [PubMed]
- Shang, X.L.; Wu, Z.F.; Yin, Z.Y.; Zhang, J.; Liu, Z.J.; Fang, S.Z. Simultaneous determination of flavonoids and triterpenoids in Cyclocarya paliurus leaves using high-performance liquid chromatography. Afr. J. Tradit. Complemt. Altern. Med. 2015, 12, 125–134. [Google Scholar] [CrossRef]
- Gu, M.; Ouyang, F.; Su, Z.G. Comparison of high-speed counter-current chromatography and high-performance liquid chromatography on fingerprinting of Chinese traditional medicine. J. Chromatogr. A 2004, 1022, 139–144. [Google Scholar] [CrossRef] [PubMed]
- Chou, G.X.; Xu, S.J.; Liu, D.; Koh, G.Y.; Zhang, J.; Liu, Z.J. Quantitative and fingerprint analyses of Chinese sweet tea plant (Rubus suavissimus S. Lee). J. Agric. Food Chem. 2009, 57, 1076–1083. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Sun, J.H.; Yu, L.L.; Chen, P. Chromatographic and mass spectrometric fingerprinting analyses of Angelica sinensis (Oliv.) Diels-derived dietary supplements. Anal. Bioanal. Chem. 2013, 405, 4477–4485. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.L.; Yin, L.; Wu, F.; Zhang, X. Investigation of a novel method for quality control of Chinese herbal compound prescription: HPLC fingerprint and multi-index components combining blending technology control for quality stability of Zhou’s prescription extract. Anal. Methods 2014, 6, 4158–4170. [Google Scholar] [CrossRef]
- Yudthavorasit, S.; Wongravee, K.; Leepipatpiboon, N. Characteristic fingerprint based on gingerol derivative analysis for discrimination of ginger (Zingiber officinale) according to geographical origin using HPLC-DAD combined with chemometrics. Food Chem. 2014, 158, 101–111. [Google Scholar] [CrossRef] [PubMed]
- Arceusz, A.; Wesolowski, M. Quality consistency evaluation of Melissa officinalis L. commercial herbs by HPLC fingerprint and quantitation of selected phenolic acids. J. Pharm. Biomed. Anal. 2013, 83, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Engström, M.T.; Pälijärvi, M.; Salminen, J.P. Rapid fingerprint analysis of plant extracts for ellagitannins, gallic acid, and quinic acid derivatives and quercetin-, kaempferol- and myricetin-based flavonol glycosides by UPLC-QqQ-MS/MS. J. Agric. Food Chem. 2015, 63, 4068–4079. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Cao, X.S.; Ferchaud, V.; Qi, Y.D.; Jiang, H.; Tang, F.; Yue, Y.D.; Chin, K.L. Variations in chemical fingerprints and major flavonoid contents from the leaves of thirty-one accessions of Hibiscus sabdariffa L. Biomed. Chromatogr. 2016, 30, 880–887. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wu, T.; Zhu, J.H.; Wan, L.L.; Yu, Q.; Li, X.X.; Cheng, Z.H.; Guo, C. Combinative method using HPLC fingerprint and quantitative analyses for quality consistency evaluation of an herbal medicinal preparation produced by different manufacturers. J. Pharm. Biomed. Anal. 2010, 52, 597–602. [Google Scholar] [CrossRef] [PubMed]
- Qin, K.M.; Wang, B.; Li, W.D.; Cai, H.; Chen, D.N.; Liu, X.; Yin, F.Z.; Cai, B.C. Quality assessment of raw and processed Arctium. lappa L. through multicomponent quantification, chromatographic fingerprint, and related chemometric analysis. J. Sep. Sci. 2015, 38, 1491–1498. [Google Scholar] [CrossRef] [PubMed]
- Viapiana, A.; Strucklewicka, W.; Konieczynski, P.; Wesolowski, M.; Kaliszan, R. An approach based on HPLC-fingerprint and chemometrics to quality consistency evaluation of Matricaria chamomillal L. commercial samples. Front. Plant Sci. 2016, 7, 1561–1571. [Google Scholar] [CrossRef] [PubMed]
- Ni, Y.N.; Song, R.M.; Kokot, S. Analysis of HPLC fingerprints: Discrimination of raw and processed Rhubarb samples with the aid of chemometrics. Anal. Methods 2012, 4, 171–176. [Google Scholar] [CrossRef]
- Kite, G.C.; Porter, E.A.; Denison, F.C.; Grayer, R.J.; Veitch, N.C.; Butler, I.; Simmods, M.S.J. Data-directed scan sequence for the general assignment of C-glycosylflavone O-glycosides in plant extracts by liquid chromatography-ion trap mass spectrometry. J. Chromatogr. A 2006, 1104, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Liu, Y.Q.; Han, L.F.; Liu, J.Z.; Liu, W.Y.; Feng, F.; Zhang, J.; Xie, N. Chemical constituents and quality control of two Dracocephalum species based on high-performance liquid chromatographic fingerprints coupled with tandem mass spectrometry and chemometrics. J. Sep. Sci. 2016, 39, 4071–4085. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.L.; Zhong, P.P.; Wang, Y.X. Antioxidant activities and identification of bioactive components of Cyclocarya paliurus leaves by UPLC-QTOF-MS/MS. Food Sci. 2017, 38, 122–128. [Google Scholar]
- Cui, B.S.; Li, S. New triterpenoid saponins from the leaves of Cyclocarya paliurus. Chin. Chem. Lett. 2015, 26, 585–589. [Google Scholar] [CrossRef]
- Jiang, C.H.; Wang, Q.Q.; Wei, Y.J.; Yao, N.; Wu, Z.F.; Ma, Y.L.; Lin, Z.; Zhao, M.; Che, C.T.; Yao, X.M.; et al. Cholesterol-lowering effects and potential mechanisms of different polar extracts from Cyclocarya paliurus leaves in hyperlipidemic mice. J. Ethnopharmacol. 2015, 176, 17–26. [Google Scholar] [CrossRef] [PubMed]
- André, C.M.; Oufir, M.; Hoffmann, L.; Hausman, J.F.; Rogez, H.; Larondelle, Y.; Evers, D. Influence of environment and genotype on polyphenol compounds and vitro antioxidant capacity of native Andean potatoes (Solanum tuberosum L.). J. Food Compos. Anal. 2009, 22, 517–524. [Google Scholar] [CrossRef]
- Liu, Y.; Qian, C.Y.; Ding, S.H.; Shang, X.L.; Yang, W.X.; Fang, S.Z. Effect of regime and provenance on leaf characteristics, growth and flavonoid accumulation in Cyclocarya paliurus (Batal) Iljinskaja coppices. Bot. Stud. 2016, 57, 28–40. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, S.; Zietz, M.; Schreiner, M.; Rohn, S.; Kroh, L.W.; Krumbein, A. Genotypic and climatic influence on the concentration and composition of flavonoids in kales (Brassica oleracea var. sabellica). Food Chem. 2010, 119, 1293–1299. [Google Scholar] [CrossRef]
- Lu, J.L.; Lu, Y.J.; Niu, Y.G.; Whent, M.; Ramadan, M.F.; Costa, J.; Yu, L.L. Effect of genotype, environment, and their interaction on phytochemical compositions and antioxidant properties of soft winter wheat flour. Food Chem. 2013, 138, 454–462. [Google Scholar]
- Deng, B.; Cao, Y.N.; Fang, S.Z.; Shang, X.L.; Yang, W.X.; Qian, C.Y. Variation and stability of growth and leaf flavonoid content in Cyclocarya paliurus across environments. Ind. Crops Prod. 2015, 76, 386–393. [Google Scholar] [CrossRef]
- Solar, A.; Colarič, M.; Usenik, V.; Stampar, F. Seasonal variations of selected flavonoids, phenolic acids and quinones in annual shoots of common walnut (Juglans regia L.). Plant Sci. 2006, 170, 453–461. [Google Scholar] [CrossRef]
- Fu, X.X.; Zhou, X.D.; Deng, B.; Shang, X.L.; Fang, S.Z. Seasonal and genotypic variation of water-soluble polysaccharide content in leaves of Cyclocarya paliurus. South. For. 2015, 77, 231–236. [Google Scholar]
Sample Availability: Samples of the compounds 3-O-caffeoylquinic acid, 4-O-caffeoylquinic acid, quercetin-3-O-glucuronide, quercetin-3-O-galactoside, isoquercitrin, kaempferol-3-O-glucuronide, kaempferol 3-O-glucoside, quercetin-3-O-rhamnoside, 4, 5-di-O-caffeoylquinic acid, kaempferol-3-O-rhamnoside, arjunolic acid, cyclocaric acid B, pterocaryoside B, pterocaryoside A, hederagenin and oleanolic acid are available from the authors. |

| Sample No. | Origins | Latitude (N) | Longitude (E) | Similarity |
|---|---|---|---|---|
| S1 | Qimen, Anhui | 30°1′11′′ | 117°31′44′′ | 0.956 |
| S2 | Mingxi, Fujian | 26°34′7′′ | 116°33′46′′ | 0.987 |
| S3 | Pucheng, Fujian | 27°55′43′′ | 118°45′46′′ | 0.978 |
| S4 | Longlin, Guangxi | 24°21′36′′ | 104°34′12′′ | 0.968 |
| S5 | Longsheng, Guangxi | 25°22′12′′ | 109°31′48′′ | 0.971 |
| S6 | Liping, Guizhou | 26°20′24′′ | 109°14′24′′ | 0.836 |
| S7 | Yinjiang, Guizhou | 27°44′24′′ | 108°30′36′′ | 0.975 |
| S8 | Hefeng, Hubei | 29°31′12′′ | 110°15′00′′ | 0.926 |
| S9 | Wufeng, Hubei | 30°11′26′′ | 110°53′52′′ | 0.957 |
| S10 | Jianghua, Hunan | 24°55′2′′ | 112°1′37′′ | 0.847 |
| S11 | Nanzhao, Henan | 33°28′35′′ | 112°00′05′′ | 0.987 |
| S12 | Shangcheng, Henan | 31°25′12′′ | 115°19′12′′ | 0.917 |
| S13 | Suining, Hunan | 26°22′24′′ | 110°7′47′′ | 0.876 |
| S14 | Fenyi, Jiangxi | 27°22′12′′ | 114°18′36′′ | 0.882 |
| S15 | Xiushui, Jiangxi | 28°9′7′′ | 114°31′8′′ | 0.960 |
| S16 | Qingchuan, Sichuan | 32°15′00′′ | 104°30′36′′ | 0.523 |
| S17 | Lueyang, Shanxi | 33°22′12′′ | 105°50′24′′ | 0.888 |
| S18 | Longquan, Zhejiang | 27°32′24′′ | 119°6′36′′ | 0.936 |
| Peak No. | tR (min) | [M ‒ H]− | MS/MS Fragment Ion (m/z) | Formula | Identification |
|---|---|---|---|---|---|
| 1 | 7.4 | 353.9874 | 191.0554 | C16H8O9 | 3-O-Caffeoylquinic acid |
| 2 | 7.8 | 353.0875 | 191.0552;179.0339;135.0444 | C16H8O9 | 4-O-Caffeoylquinic acid |
| 3 | 14.9 | 477.0673 | 301.0351;178.9979;151.003 | C21H18O13 | Quercetin-3-O-glucuronide |
| 4 | 15.4 | 463.0888 | 301.0343;271.0244;178.9979;151.0029 | C21H20O12 | Quercetin-3-O-galactoside |
| 5 | 15.8 | 463.0888 | 301.0338;271.0239;178.9981;151.0029 | C21H20O12 | Isoquercitrin |
| 6 | 17.6 | 461.9727 | 286.0431;285.0401;113.0235;85.0296 | C21H18O12 | Kaempferol-3-O-glucuronide |
| 7 | 18.5 | 447.0935 | 285.0388;284.0319;255.0295 | C21H20O11 | Kaempferol-3-O-glucoside |
| 8 | 18.8 | 447.0973 | 301.0337;285.0388;255.0294;217.0133 | C21H20O11 | Quercetin-3-O-rhamnoside |
| 9 | 19.8 | 515.1199 | 353.0864;191.0553;179.0342;135.0443 | C25H24O12 | 4,5-di-O-Caffeoylquinic acid |
| 10 | 23.4 | 431.0981 | 285.0394;255.0295;227.0342 | C21H20O10 | Kaempferol-3-O-rhamnoside |
| 11 | 40.8 | 577.136 | 397.2221;285.0394;163.0389;145.0289 | C30H26O12 | Kaempferol-3-(6′′-(Z)-cinnamylglucoside) |
| 12 | 47.1 | 487.3429 | 445.2942;401.3056;389.2698 | C30H48O5 | Arjunolic acid |
| 13 | 49.7 | 485.3275 | / | C30H46O5 | Cyclocaric acid B |
| 14 | 54.2 | 621.4001 | 521.3107;489.3571 | C35H58O9 | Pterocaryoside B |
| 15 | 60.0 | 635.4162 | 535.3265;489.3573 | C36H60O9 | Pterocaryoside A |
| 16 | 60.5 | 471.3481 | 145.0286 | C30H48O4 | Hederagenin |
| 17 | 67.2 | 621.4017 | 489.3578;469.3318 | C35H58O9 | Cyclocarioside J |
| 18 | 74.4 | 635.4177 | 489.3579 | C36H60O9 | Cyclocarioside III |
| 19 | 75.2 | 603.3894 | 521.3107;489.3573;471.347 | C35H56O8 | Cyclocarioside II |
| 20 | 83.5 | 455.3549 | / | C30H48O3 | Oleanolic acid |
| 21 | 84.3 | 277.2192 | 146.9664; 197.9641;248.9617 | C18H30O2 | Unknown |
| Compound | Regressive Equation a | Linear Range (ug/mL) | R2 | LOD (ng/mL) | LOQ (ng/mL) | Precision | Repeatability RSD% (n = 6) | Recovery | ||
|---|---|---|---|---|---|---|---|---|---|---|
| Intraday RSD% (n = 6) | Interday RSD% (n = 18) | Mean | RSD% (n = 6 ) | |||||||
| 3-O-caffeoylquinic acid | y = 4943.9x − 37,710 | 8–640 | 0.9998 | 64.25 | 214.21 | 0.88 | 2.14 | 0.83 | 98.72 | 1.89 |
| 4-O-caffeoylquinic acid | y = 6245.2x − 14,821 | 4–192 | 0.9998 | 57.93 | 197.58 | 0.71 | 1.94 | 1.92 | 96.53 | 1.24 |
| Quercetin-3-O-glucuronide | y = 23,136x − 119,324 | 2.5–600 | 0.9997 | 40.28 | 128.74 | 0.43 | 1.21 | 0.53 | 98.32 | 2.13 |
| Quercetin-3-O-galactoside | y = 20,402x − 28,339 | 2–480 | 0.9994 | 52.94 | 174.17 | 0.48 | 1.59 | 0.92 | 102.43 | 0.92 |
| Isoquercitrin | y = 21,231x + 2369.2 | 2–180 | 0.9997 | 58.42 | 192.52 | 0.98 | 2.35 | 1.89 | 97.89 | 2.02 |
| Kaempferol-3-O-glucuronide | y = 16,702x − 92,304 | 2–640 | 0.9995 | 43.98 | 153.14 | 0.62 | 1.93 | 2.13 | 100.93 | 2.61 |
| Kaempferol-3-O-glucoside | y = 18,066x − 87,126 | 2–480 | 0.9991 | 53.85 | 187.37 | 1.26 | 2.78 | 1.89 | 99.23 | 2.67 |
| Quercetin-3-O-rhamnoside | y = 15,638x − 21,262 | 2–240 | 0.9993 | 62.48 | 199.32 | 0.49 | 0.98 | 0.57 | 98.92 | 1.95 |
| 4,5-di-O-Caffeoylquinic acid | y = 9428.7x +52,403 | 2.5–140 | 0.9997 | 58.97 | 201.22 | 0.63 | 1.52 | 0.82 | 101.29 | 1.82 |
| Kaempferol-3-O-rhamnoside | y = 15,648x + 12,295 | 5–150 | 0.9997 | 64.13 | 211.81 | 0.58 | 1.68 | 1.52 | 97.69 | 1.91 |
| Arjunolic acid | y = 5465.1x + 33,575 | 5–1500 | 0.9994 | 78.13 | 260.17 | 0.48 | 1.19 | 1.04 | 102.38 | 2.48 |
| Cyclocaric acid B | y = 8579.6x − 91,948 | 7.5–1080 | 0.9993 | 90.16 | 296.36 | 0.92 | 2.83 | 2.05 | 102.9 | 2.52 |
| Pterocaryoside B | y = 3716.1x + 52,647 | 6.7–1608 | 0.9991 | 72.46 | 210.27 | 1.23 | 2.42 | 2.04 | 98.19 | 2.61 |
| Pterocaryoside A | y = 3746.1x + 56,466 | 10–1200 | 0.9991 | 86.39 | 268.62 | 0.84 | 1.96 | 1.29 | 102.85 | 1.85 |
| Hederagenin | y = 6054.7x + 25,521 | 8–640 | 0.9997 | 72.82 | 232.3 | 0.91 | 2.53 | 1.83 | 101.83 | 1.69 |
| Oleanolic acid | y = 7144.8x + 963.97 | 1–200 | 0.9994 | 58.41 | 185.72 | 1.23 | 2.59 | 2.43 | 98.28 | 2.93 |
| Sample No. | Content (mg g−1) | |||
|---|---|---|---|---|
| 3-O-Caffeoylquinic Acid | 4-O-Caffeoylquinic Acid | 4,5-di-O-Caffeoylquinic Acid | TPC b | |
| S1 | 0.50 ± 0.03j | 0.15 ± 0.00i | 0.11 ± 0.00l | 0.77 ± 0.04h |
| S2 | 0.52 ± 0.00j | 0.21 ± 0.01h | 0.17 ± 0.00hi | 0.91 ± 0.00g |
| S3 | 0.80 ± 0.04hi | 0.32 ± 0.01fg | 0.22 ± 0.01f | 1.34 ± 0.07f |
| S4 | 0.82 ± 0.00ghi | 0.11 ± 0.01j | 0.08 ± 0.00m | 1.01 ± 0.01g |
| S5 | 1.31 ± 0.00d | 0.29 ± 0.01g | 0.30 ± 0.01d | 1.90 ± 0.02c |
| S6 | 0.89 ± 0.02fg | 0.37 ± 0.01e | 0.25 ± 0.00e | 1.52 ± 0.02e |
| S7 | 1.54 ± 0.01b | 0.50 ± 0.02c | 0.62 ± 0.00b | 2.66 ± 0.03b |
| S8 | 1.42 ± 0.01c | 0.67 ± 0.00a | 0.63 ± 0.03b | 2.71 ± 0.02b |
| S9 | 1.00 ± 0.01e | 0.34 ± 0.02f | 0.32 ± 0.02c | 1.66 ± 0.04d |
| S10 | 0.78 ± 0.04i | 0.31 ± 0.02g | 0.18 ± 0.00hi | 1.27 ± 0.11f |
| S11 | 1.00 ± 0.05e | 0.22 ± 0.01h | 0.14 ± 0.01jk | 1.37 ± 0.07f |
| S12 | 0.94 ± 0.00ef | 0.44 ± 0.02d | 0.21 ± 0.00fg | 1.60 ± 0.01de |
| S13 | 0.46 ± 0.00j | 0.12 ± 0.00j | Trace c | 0.58 ± 0.01i |
| S14 | 0.92 ± 0.01ef | 0.22 ± 0.00h | 0.16 ± 0.01ij | 1.31 ± 0.06f |
| S15 | 0.81 ± 0.04ghi | 0.30 ± 0.01g | 0.19 ± 0.01gh | 1.31 ± 0.09f |
| S16 | 0.89 ± 0.03fgh | 0.13 ± 0.00ij | 0.22 ± 0.00f | 1.25 ± 0.03f |
| S17 | 2.34 ± 0.00a | 0.62 ± 0.03b | 0.66 ± 0.01a | 3.61 ± 0.01a |
| S18 | 0.54 ± 0.03j | 0.22 ± 0.01h | 0.12 ± 0.01kl | 0.88 ± 0.08gh |
| Sample No. | Content (mg g−1) | |||||||
|---|---|---|---|---|---|---|---|---|
| Quercetin-3-O-glucuronide | Quercetin-3-O-galactoside | Isoquercitrin | Kaempferol-3-O-glucuronide | Kaempferol-3-O-glucoside | Quercetin-3-O-rhamnoside | Kaempferol-3-O-rhamnoside | TFC b | |
| S1 | 2.15 ± 0.06c | 0.79 ± 0.02ab | 0.37 ± 0.02f | 1.16 ± 0.00gh | 0.19 ± 0.00jk | 0.21 ± 0.00e | 1.48 ± 0.02f | 6.35 ± 0.12de |
| S2 | 1.60 ± 0.05d | 0.52 ± 0.02d | 0.31 ± 0.01g | 1.13 ± 0.01h | 0.22 ± 0.01i | 0.14 ± 0.00gh | 1.04 ± 0.02h | 4.95 ± 0.12g |
| S3 | 2.09 ± 0.11c | 0.71 ± 0.04c | 0.55 ± 0.02c | 1.19 ± 0.06gh | 0.27 ± 0.01fg | 0.24 ± 0.01d | 1.26 ± 0.06g | 6.29 ± 0.38de |
| S4 | 2.14 ± 0.01c | 0.42 ± 0.01f | 0.20 ± 0.01i | 1.39 ± 0.03e | 0.24 ± 0.00h | 0.24 ± 0.00d | 2.00 ± 0.14c | 6.63 ± 0.15de |
| S5 | 1.67 ± 0.01d | 0.55 ± 0.00d | 0.51 ± 0.00d | 1.15 ± 0.00h | 0.43 ± 0.00c | 0.22 ±0.00e | 2.17 ± 0.08b | 6.70 ± 0.06d |
| S6 | 0.79 ± 0.01g | 0.53 ± 0.00d | 0.30 ± 0.00gh | 0.81 ± 0.01j | 0.20 ± 0.01ijk | 0.11 ± 0.01i | 0.81 ± 0.00i | 3.55 ± 0.06h |
| S7 | 2.56 ± 0.01b | 0.82 ± 0.01a | 0.51 ± 0.01d | 1.30 ± 0.01ef | 0.25 ± 0.01gh | 0.31 ± 0.00b | 1.73 ± 0.01d | 7.48 ± 0.02c |
| S8 | 3.98 ± 0.09a | 0.46 ± 0.00e | 0.53 ± 0.01cd | 2.30 ± 0.08a | 0.30 ± 0.02de | 0.43 ± 0.02a | 2.63 ± 0.1a | 10.63 ± 0.33a |
| S9 | 2.11 ± 0.09c | 0.40 ± 0.01fg | 0.19 ± 0.01i | 1.64 ± 0.01d | 0.21 ± 0.01ij | 0.19 ± 0.01f | 1.57 ± 0.05ef | 6.32 ± 0.10de |
| S10 | 1.09 ± 0.05f | 0.38 ± 0.02g | 0.21 ± 0.01i | 0.83 ± 0.04ij | 0.16 ± 0.01l | 0.15 ± 0.01gh | 0.82 ± 0.041i | 3.65 ± 0.18h |
| S11 | 1.22 ± 0.06e | 0.44 ± 0.02ef | 0.29 ± 0.01gh | 1.17 ± 0.06gh | 0.42 ± 0.02c | 0.12 ± 0.01hi | 1.16 ± 0.16gh | 4.81 ± 0.24g |
| S12 | 0.86 ± 0.01g | 0.41 ± 0.01fg | 0.28 ± 0.01h | 1.25 ± 0.01fg | 0.30 ± 0.00de | 0.13 ± 0.00h | 0.57 ± 0.01j | 3.80 ± 0.02h |
| S13 | 0.50 ± 0.00h | 0.11 ± 0.00h | 0.05 ± 0.00k | 0.67 ± 0.01k | 0.11 ± 0.01m | 0.08 ± 0.00j | 0.55 ± 0.01j | 2.07 ± 0.02i |
| S14 | 1.07 ± 0.03f | 0.78 ± 0.01b | 0.37 ± 0.01f | 1.68 ± 0.06d | 0.28 ± 0.01ef | 0.30 ± 0.00b | 1.71 ± 0.01de | 6.20 ± 0.06e |
| S15 | 1.64 ± 0.07d | 0.69 ± 0.03c | 0.44 ± 0.02e | 1.95 ± 0.10c | 0.31 ± 0.02d | 0.25 ± 0.01cd | 1.26 ± 0.10gh | 6.56 ± 0.48de |
| S16 | 0.16 ± 0.00i | 0.80 ± 0.00ab | 0.74 ± 0.00b | 0.57 ± 0.00l | 2.04 ± 0.00a | 0.26 ± 0.00c | 1.23 ± 0.00g | 5.80 ± 0.00f |
| S17 | 2.15 ± 0.01c | 0.56 ± 0.01d | 1.27 ± 0.01a | 2.21 ± 0.01b | 1.81 ± 0.01b | 0.22 ± 0.00e | 1.94 ± 0.00c | 10.15 ± 0.05b |
| S18 | 0.58 ± 0.03h | 0.13 ± 0.01h | 0.10 ± 0.01j | 0.91 ± 0.05i | 0.18 ± 0.01kl | 0.06 ± 0.00j | 0.38 ± 0.05k | 2.36 ± 0.11i |
| Sample No. | Content (mg g−1) | ||||||
|---|---|---|---|---|---|---|---|
| Arjunolic Acid | Cyclocaric Acid B | Pterocaryoside B | Pterocaryoside A | Hederagenin | Oleanolic Acid | TTC b | |
| S1 | 5.52 ± 0.23c | 1.00 ± 0.05f | 1.61 ± 0.081i | 3.78 ± 0.09d | 1.02 ± 0.00hi | 0.26 ± 0.01h | 13.18 ± 0.49e |
| S2 | 3.13 ± 0.16f | 1.06 ± 0.02f | 2.62 ± 0.13g | 3.82 ± 0.06d | 0.96 ± 0.01i | 0.40 ± 0.00e | 11.99 ± 0.53fg |
| S3 | 2.22 ± 0.11g | 0.89 ± 0.04g | 2.62 ± 0.13g | 3.18 ± 0.16f | 1.14 ± 0.06fg | 0.45 ± 0.02d | 10.50 ± 0.20h |
| S4 | 4.29 ± 0.15d | 1.82 ± 0.02c | 0.46 ± 0.03k | 2.89 ± 0.14fg | 1.32 ± 0.03cd | 0.46 ± 0.00d | 11.24 ± 0.20gh |
| S5 | 4.29 ± 0.15d | 1.51 ± 0.01e | 4.70 ± 0.12c | 3.49 ± 0.09e | 0.65 ± 0.03k | 0.17 ± 0.00i | 14.79 ± 0.45d |
| S6 | 5.77 ± 0.29bc | 2.17 ± 0.00b | 3.46 ± 0.00d | 4.88 ± 0.04c | 1.75 ± 0.01a | 0.54 ± 0.00c | 18.57 ± 0.54c |
| S7 | 3.92 ± 0.04e | 1.45 ± 0.02e | 2.20 ± 0.11h | 2.95 ± 0.07f | 0.74 ± 0.04j | 0.37 ± 0.00f | 11.62 ± 0.34fgh |
| S8 | 2.11 ± 0.11g | 0.70 ± 0.02h | 1.10 ± 0.02j | 1.39 ± 0.06i | 0.45 ± 0.01m | 0.10 ± 0.00jk | 5.85 ± 0.21k |
| S9 | 2.13 ± 0.11g | 0.68 ± 0.00hi | 0.95 ± 0.00j | 1.05 ± 0.05j | 0.41 ± 0.02m | 0.16 ± 0.00i | 5.38 ± 0.41kl |
| S10 | 5.98 ± 0.02b | 2.86 ± 0.14a | 2.99 ± 0.15f | 4.91 ± 0.25c | 1.10 ± 0.06gh | 0.41 ± 0.02e | 18.25 ± 1.17c |
| S11 | 2.02 ± 0.10g | 0.85 ± 0.04g | 3.10 ± 0.16ef | 2.65 ± 0.13g | 1.70 ± 0.09a | 0.40 ± 0.02e | 10.72 ± 0.57h |
| S12 | 1.66 ± 0.08h | 1.63 ± 0.00d | 2.89 ± 0.11f | 4.81 ± 0.04c | 1.40 ± 0.03c | 0.29 ± 0.00g | 12.67 ± 0.26ef |
| S13 | 1.62 ± 0.01h | 0.59 ± 0.01i | 3.31 ± 0.13de | 3.17 ± 0.14f | 0.40 ± 0.00m | 0.11 ± 0.00j | 9.19 ± 0.26i |
| S14 | 6.69 ± 0.26a | 2.80 ± 0.06a | 5.16 ± 0.14b | 8.38 ± 0.16a | 1.52 ± 0.08b | 0.67 ± 0.00a | 25.22 ± 0.30a |
| S15 | 5.47 ± 0.27c | 1.90 ± 0.10c | 5.61 ± 0.28a | 6.06 ± 0.30b | 1.26 ± 0.02de | 0.36 ± 0.01f | 20.67 ± 1.03b |
| S16 | 1.39 ± 0.07h | 0.39 ± 0.00j | 0.64 ± 0.03k | 0.67 ± 0.03k | 1.20 ± 0.05ef | 0.59 ± 0.02b | 4.89 ± 0.15kl |
| S17 | 1.67 ± 0.08h | 0.43 ± 0.01j | 1.17 ± 0.03j | 0.73 ± 0.00k | 0.28 ± 0.00n | 0.08 ± 0.00k | 4.36 ± 0.22l |
| S18 | 1.35 ± 0.07h | 0.68 ± 0.03hi | 2.06 ± 0.10h | 2.27 ± 0.11h | 0.54 ± 0.03l | 0.17 ± 0.00i | 7.08 ± 0.35j |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cao, Y.; Fang, S.; Yin, Z.; Fu, X.; Shang, X.; Yang, W.; Yang, H. Chemical Fingerprint and Multicomponent Quantitative Analysis for the Quality Evaluation of Cyclocarya paliurus Leaves by HPLC–Q–TOF–MS. Molecules 2017, 22, 1927. https://doi.org/10.3390/molecules22111927
Cao Y, Fang S, Yin Z, Fu X, Shang X, Yang W, Yang H. Chemical Fingerprint and Multicomponent Quantitative Analysis for the Quality Evaluation of Cyclocarya paliurus Leaves by HPLC–Q–TOF–MS. Molecules. 2017; 22(11):1927. https://doi.org/10.3390/molecules22111927
Chicago/Turabian StyleCao, Yanni, Shengzuo Fang, Zhiqi Yin, Xiangxiang Fu, Xulan Shang, Wanxia Yang, and Huimin Yang. 2017. "Chemical Fingerprint and Multicomponent Quantitative Analysis for the Quality Evaluation of Cyclocarya paliurus Leaves by HPLC–Q–TOF–MS" Molecules 22, no. 11: 1927. https://doi.org/10.3390/molecules22111927
APA StyleCao, Y., Fang, S., Yin, Z., Fu, X., Shang, X., Yang, W., & Yang, H. (2017). Chemical Fingerprint and Multicomponent Quantitative Analysis for the Quality Evaluation of Cyclocarya paliurus Leaves by HPLC–Q–TOF–MS. Molecules, 22(11), 1927. https://doi.org/10.3390/molecules22111927





