Structures and Ribosomal Interaction of Ribosome-Inactivating Proteins
Abstract
:1. Introduction
2. Distribution and Classification of RIPs
3. Crystal Structure and Structural Comparison of RIPs
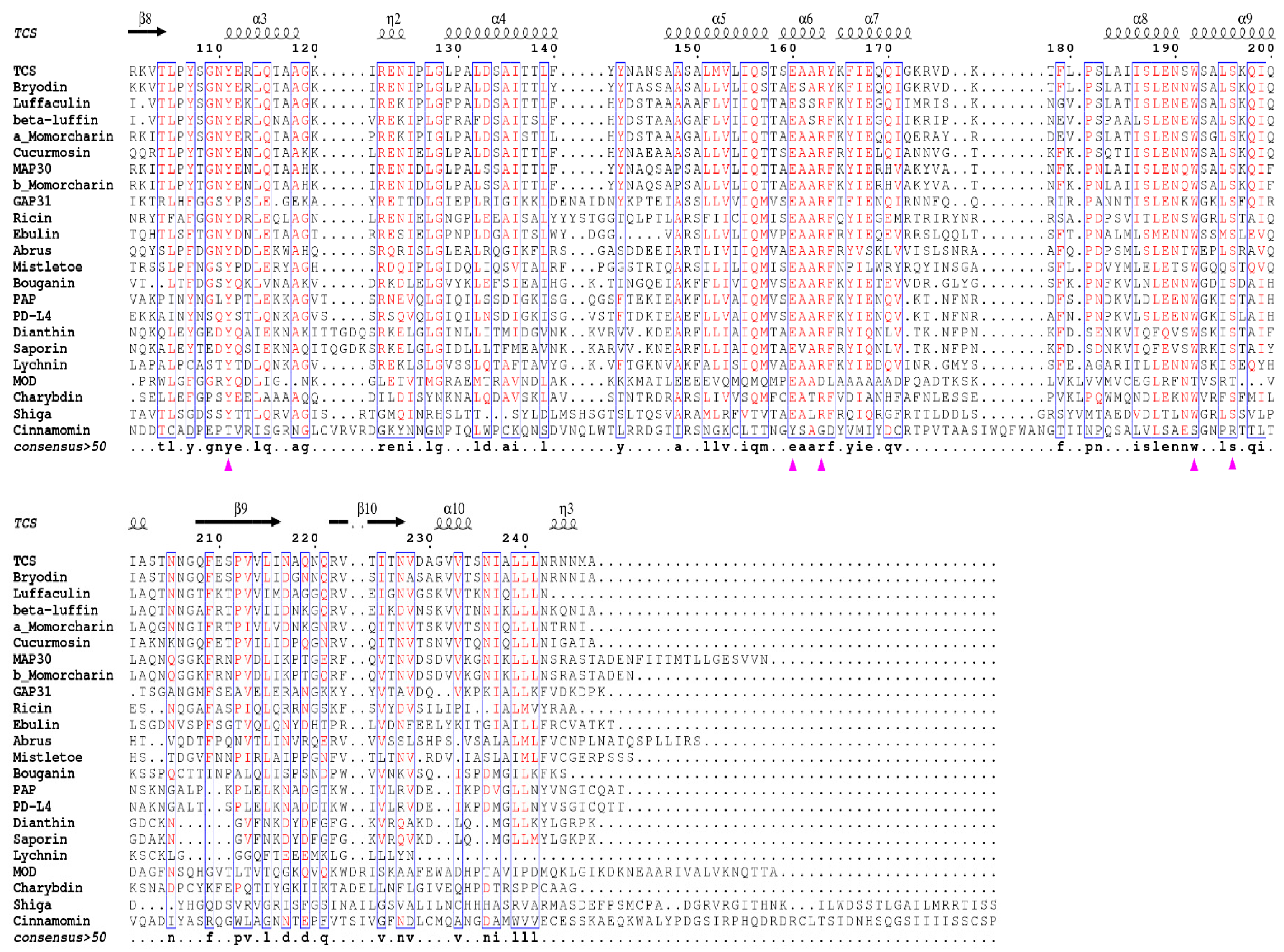
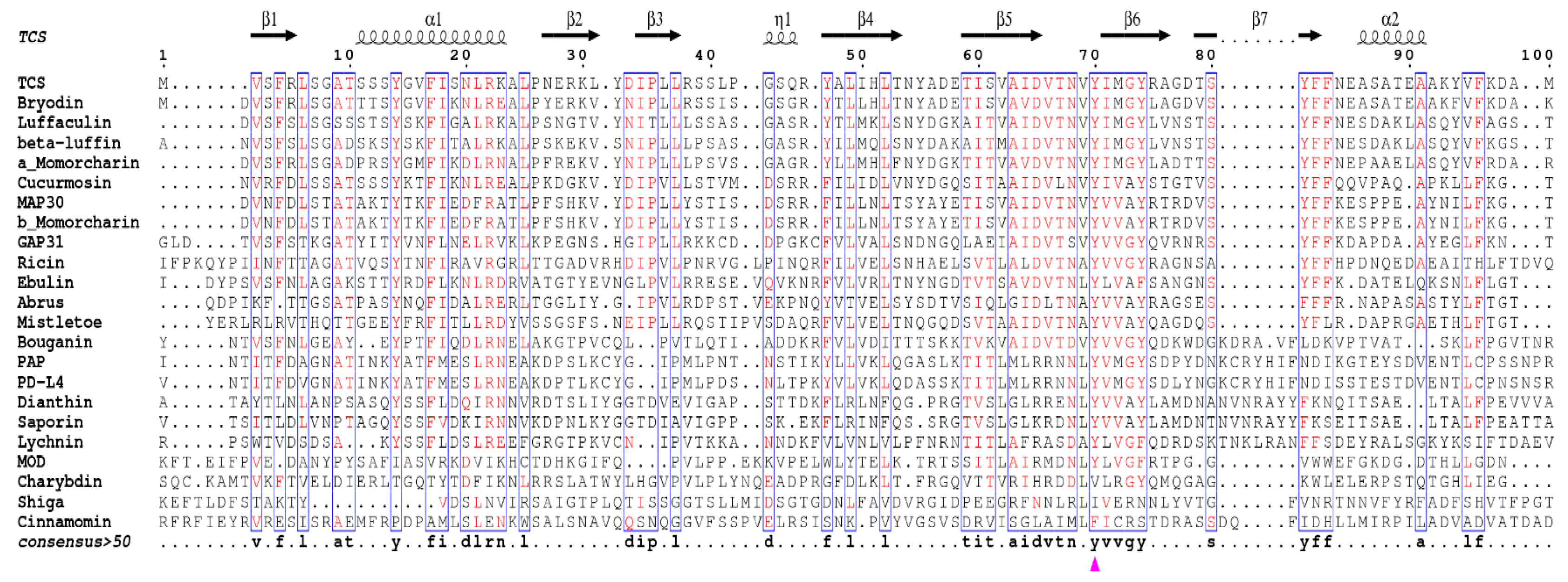
4. Invariant Residues in the Catalytic Subunit
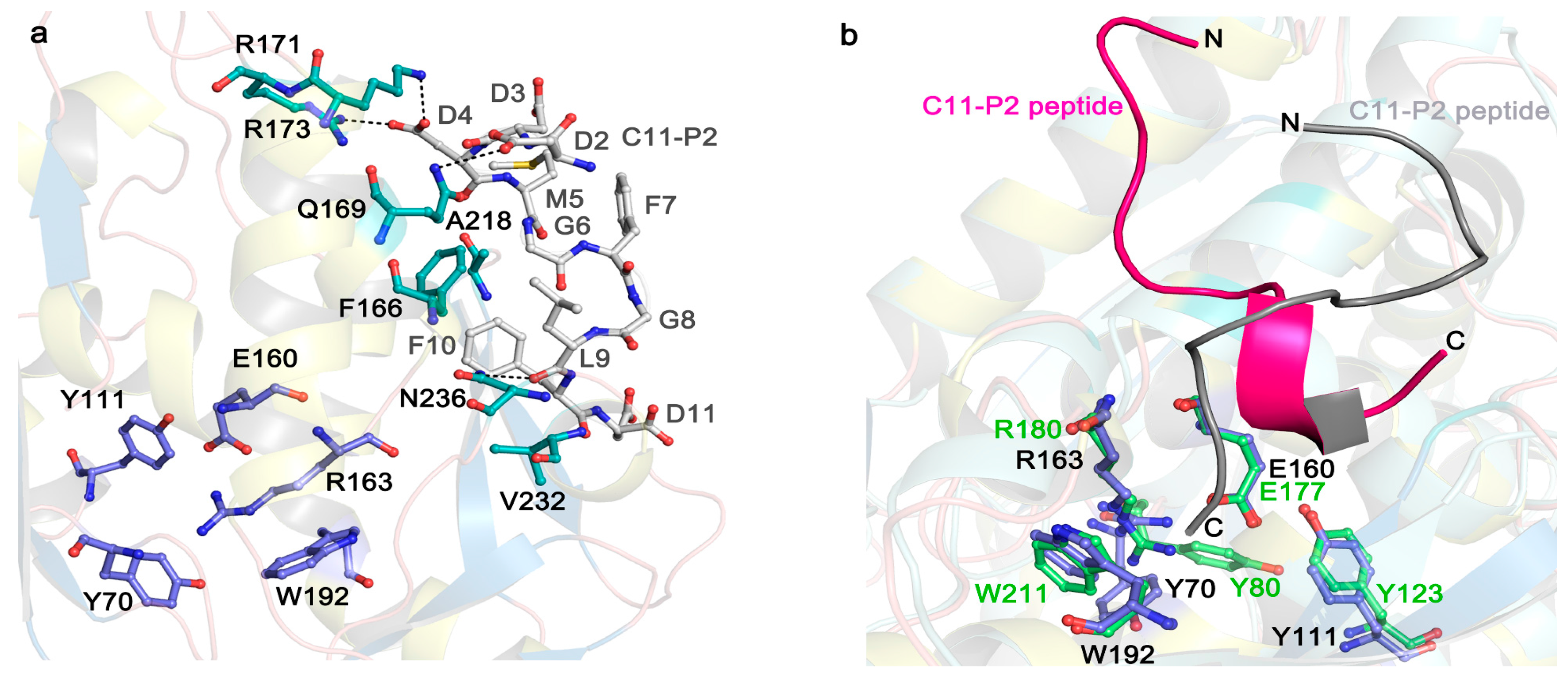
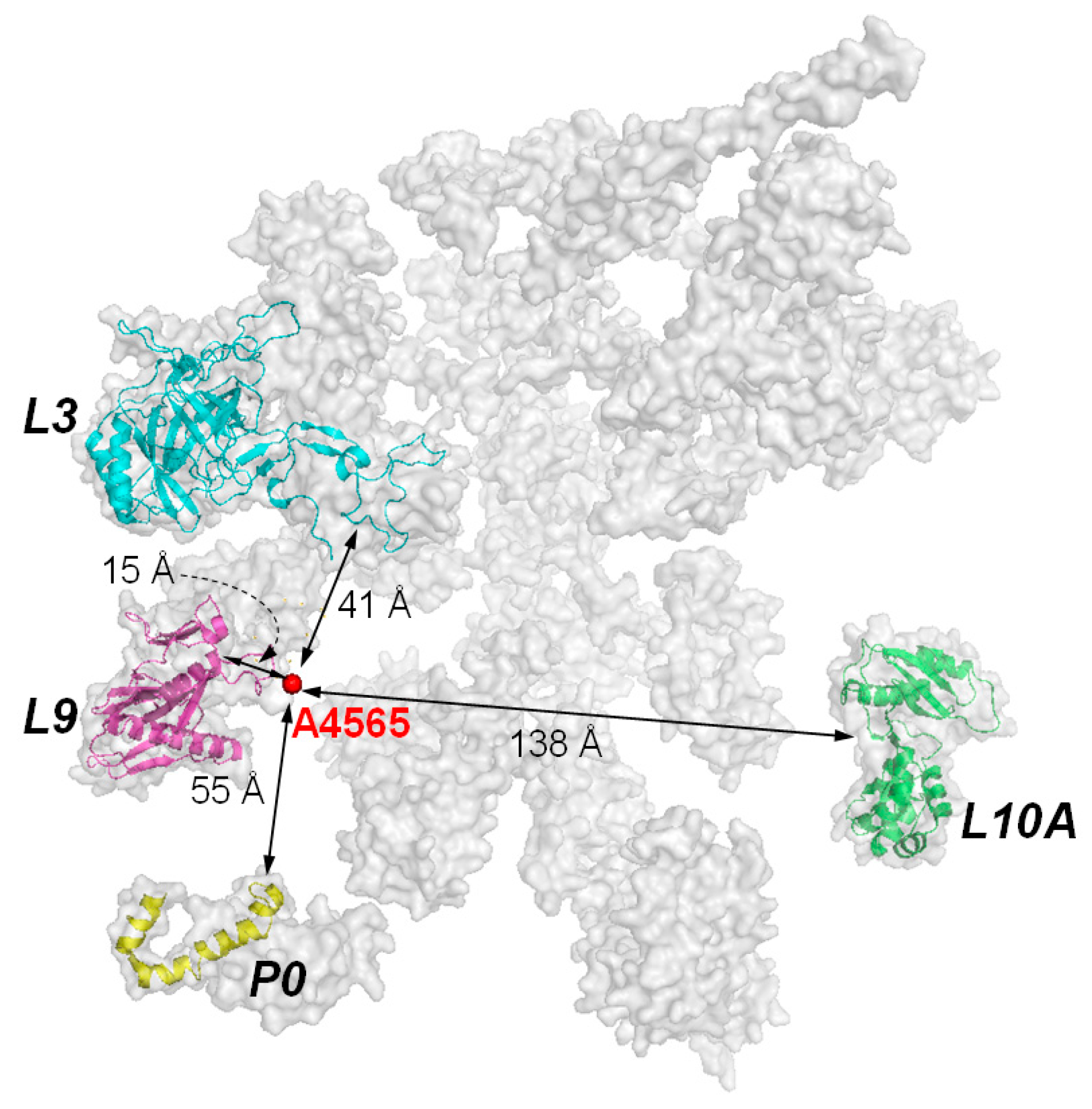
5. The Interaction of RIPs with Ribosomes
6. Kingdom Specificity of RIPs to Ribosome
7. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Stillmark, H. Uber ricin, eines gifiges ferment aus den samen von Ricinus communis L. Und anderen euphorbiacen. Ph.D. Thesis, University of Dorpat, Dorpat, Estonia, 1888. [Google Scholar]
- Poisoning And Drug Overdose: Medicine, Toxicology, 5th ed.; Olson, K.R.; Anderson, I.B. (Eds.) McGraw-Hill Companies: New York, NY, USA, 2006.
- Endo, Y.; Tsurugi, K. RNA N-glycosidase activity of ricin A-chain. Mechanism of action of the toxic lectin ricin on eukaryotic ribosomes. J. Biol. Chem. 1987, 262, 8128–8130. [Google Scholar] [PubMed]
- Barbieri, L.; Battelli, M.G.; Stirpe, F. Ribosome-inactivating proteins from plants. Biochim. Biophys. Acta 1993, 1154, 237–282. [Google Scholar] [CrossRef]
- Correll, C.C.; Munishkin, A.; Chan, Y.L.; Ren, Z.; Wool, I.G.; Steitz, T.A. Crystal structure of the ribosomal RNA domain essential for binding elongation factors. Proc. Natl. Acad. Sci. USA 1998, 95, 13436–13441. [Google Scholar] [CrossRef] [PubMed]
- Lancaster, L.; Lambert, N.J.; Maklan, E.J.; Horan, L.H.; Noller, H.F. The sarcin-ricin loop of 23S rRNA is essential for assembly of the functional core of the 50S ribosomal subunit. RNA 2008, 14, 1999–2012. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Tumer, N.E. Pokeweed antiviral protein cleaves double-stranded supercoiled DNA using the same active site required to depurinate rRNA. Nucleic Acids Res. 1999, 27, 1900–1905. [Google Scholar] [CrossRef] [PubMed]
- Lacadena, J.; Alvarez-Garcia, E.; Carreras-Sangra, N.; Herrero-Galan, E.; Alegre-Cebollada, J.; Garcia-Ortega, L.; Onaderra, M.; Gavilanes, J.G.; del Martinez Pozo, A. Fungal ribotoxins: Molecular dissection of a family of natural killers. FEMS Microbiol. Rev. 2007, 31, 212–237. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Collier, R.J. Understanding the mode of action of diphtheria toxin: A perspective on progress during the 20th century. Toxicon 2001, 39, 1793–1803. [Google Scholar] [CrossRef]
- Endo, Y.; Tsurugi, K. The RNA N-glycosidase activity of ricin A-chain. The characteristics of the enzymatic activity of ricin A-chain with ribosomes and with rRNA. J. Biol. Chem. 1988, 263, 8735–8739. [Google Scholar] [PubMed]
- Gasperi-Campani, A.; Brognara, I.; Baiocchi, D.; Roncuzzi, L. Mitochondrial DNA D-loop as a new target of saporin 6 nuclease activity. Toxicon 2005, 45, 475–480. [Google Scholar] [CrossRef] [PubMed]
- Li, X.D.; Chen, W.F.; Liu, W.Y.; Wang, G.H. Large-scale preparation of two new ribosome-inactivating proteins—Cinnamomin and Camphorin from the seeds of Cinnamomum camphora. Protein Expr. Purif. 1997, 10, 27–31. [Google Scholar] [CrossRef] [PubMed]
- Helmy, M.; Lombard, S.; Pieroni, G. Ricin RCA60: Evidence of its phospholipase activity. Biochem. Biophys. Res. Commun. 1999, 258, 252–255. [Google Scholar] [CrossRef] [PubMed]
- Lee-Huang, S.; Kung, H.F.; Huang, P.L.; Bourinbaiar, A.S.; Morell, J.L.; Brown, J.H.; Tsai, W.P.; Chen, A.Y.; Huang, H.I. Human immunodeficiency virus type 1 (HIV-1) inhibition, DNA-binding, RNA-binding, and ribosome inactivation activities in the N-terminal segments of the plant anti-HIV protein GAP31. Proc. Natl. Acad. Sci. USA 1994, 91, 12208–12212. [Google Scholar] [CrossRef] [PubMed]
- Day, P.J.; Lord, J.M.; Roberts, L.M. The deoxyribonuclease activity attributed to ribosome-inactivating proteins is due to contamination. Eur. J. Biochem. 1998, 258, 540–545. [Google Scholar] [CrossRef] [PubMed]
- Motto, M.; Lupotto, E. The genetics and properties of cereal ribosome-inactivating proteins. Mini Rev. Med. Chem. 2004, 4, 493–503. [Google Scholar] [CrossRef] [PubMed]
- Toxic Plant Proteins, Plant Cell Monographs; Lord, J.M.; Hartley, M.R. (Eds.) Springer: Berlin/Heidelberg, Germany, 2010.
- Girbes, T.; Ferreras, J.M.; Arias, F.J.; Stirpe, F. Description, distribution, activity and phylogenetic relationship of ribosome-inactivating proteins in plants, fungi and bacteria. Mini Rev. Med. Chem. 2004, 4, 461–476. [Google Scholar] [CrossRef] [PubMed]
- Ribosome-Inactivating Proteins. Ricin and Related Proteins; Stirpe, F.; Lappi, D.A. (Eds.) Wiley Blackwell: Oxford, UK, 2014.
- Reinbothe, S.; Reinbothe, C.; Lehmann, J.; Becker, W.; Apel, K.; Parthier, B. JIP60, a methyl jasmonate-induced ribosome-inactivating protein involved in plant stress reactions. Proc. Natl. Acad. Sci. USA 1994, 91, 7012–7016. [Google Scholar] [CrossRef] [PubMed]
- Rippmann, J.F.; Michalowski, C.B.; Nelson, D.E.; Bohnert, H.J. Induction of a ribosome-inactivating protein upon environmental stress. Plant Mol. Biol. 1997, 35, 701–709. [Google Scholar] [CrossRef] [PubMed]
- Dowd, P.F.; Holmes, R.A.; Pinkerton, T.S.; Johnson, E.T.; Lagrimini, L.M.; Boston, R.S. Relative activity of a tobacco hybrid expressing high levels of a tobacco anionic peroxidase and maize ribosome-inactivating protein against Helicoverpa zea and Lasioderma serricorne. J. Agric. Food Chem. 2006, 54, 2629–2634. [Google Scholar] [CrossRef] [PubMed]
- Iglesias, R.; Citores, L.; Ragucci, S.; Russo, R.; Di Maro, A.; Ferreras, J.M. Biological and antipathogenic activities of ribosome-inactivating proteins from Phytolacca dioica L. Biochim. Biophys. Acta 2016, 1860, 1256–1264. [Google Scholar] [CrossRef] [PubMed]
- Pascal, J.M.; Day, P.J.; Monzingo, A.F.; Ernst, S.R.; Robertus, J.D.; Iglesias, R.; Perez, Y.; Ferreras, J.M.; Citores, L.; Girbes, T. 2.8-Å crystal structure of a nontoxic type-II ribosome-inactivating protein, ebulin l. Proteins 2001, 43, 319–326. [Google Scholar] [CrossRef] [PubMed]
- Paton, J.C.; Paton, A.W. Pathogenesis and diagnosis of Shiga toxin-producing Escherichia coli infections. Clin. Microbiol. Rev. 1998, 11, 450–479. [Google Scholar] [PubMed]
- Walsh, T.A.; Morgan, A.E.; Hey, T.D. Characterization and Molecular Cloning of a Proenzyme Form of a Ribosome-Inactivating Protein From Maize. Novel Mechanism of Proenzyme Activation by Proteolytic Removal of a 2.8-kiloDalton Internal Peptide Segment. J. Biol. Chem. 1991, 266, 23422–23427. [Google Scholar] [PubMed]
- Hey, T.D.; Hartley, M.; Walsh, T.A. Maize ribosome-inactivating protein (b-32). Homologs in related species, effects on maize ribosomes, and modulation of activity by pro-peptide deletions. Plant Physiol. 1995, 107, 1323–1332. [Google Scholar] [CrossRef] [PubMed]
- Chaudhry, B.; Muller-Uri, F.; Cameron-Mills, V.; Gough, S.; Simpson, D.; Skriver, K.; Mundy, J. The barley 60 kDa jasmonate-induced protein (JIP60) is a novel ribosome-inactivating protein. Plant J. 1994, 6, 815–824. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef] [PubMed]
- Chan, S.H.; Shaw, P.C.; Mulot, S.F.; Xu, L.H.; Chan, W.L.; Tam, S.C.; Wong, K.B. Engineering of a mini-trichosanthin that has lower antigenicity by deleting its C-terminal amino acid residues. Biochem. Biophys. Res. Commun. 2000, 270, 279–285. [Google Scholar] [CrossRef] [PubMed]
- Chan, D.S.; Chu, L.O.; Lee, K.M.; Too, P.H.; Ma, K.W.; Sze, K.H.; Zhu, G.; Shaw, P.C.; Wong, K.B. Interaction between trichosanthin, a ribosome-inactivating protein, and the ribosomal stalk protein P2 by chemical shift perturbation and mutagenesis analyses. Nucleic Acids Res. 2007, 35, 1660–1672. [Google Scholar] [CrossRef] [PubMed]
- DeLano, W. The Pymol Molecular Graphics System; DeLano Scientific: San Carlos, CA, USA, 2002. [Google Scholar]
- Hartley, M.R.; Lord, J.M. Cytotoxic ribosome-inactivating lectins from plants. Biochim. Biophys. Acta 2004, 1701, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Hatakeyama, T.; Yamasaki, N.; Funatsu, G. Identification of the tryptophan residue located at the low-affinity saccharide binding site of ricin D. J. Biochem. 1986, 100, 781–788. [Google Scholar] [PubMed]
- Shaw, P.C.; Mulot, S.; Ma, S.K.; Xu, Q.F.; Yao, H.B.; Wu, S.; Lu, X.H.; Dong, Y.C. Structure/function relationship study of Tyr14 and Arg22 in trichosanthin, a ribosome-inactivating protein. Eur. J. Biochem. 1997, 245, 423–427. [Google Scholar] [CrossRef] [PubMed]
- Shaw, P.C.; Lee, K.M.; Wong, K.B. Recent advances in trichosanthin, a ribosome-inactivating protein with multiple pharmacological properties. Toxicon 2005, 45, 683–689. [Google Scholar] [CrossRef] [PubMed]
- Wong, K.B.; Ke, Y.B.; Dong, Y.C.; Li, X.B.; Guo, Y.W.; Yeung, H.W.; Shaw, P.C. Structure/function relationship study of Gln156, Glu160 and Glu189 in the active site of trichosanthin. Eur. J. Biochem. 1994, 221, 787–791. [Google Scholar] [CrossRef] [PubMed]
- Robertus, J.D.; Monzingo, A.F. The structure of ribosome inactivating proteins. Mini Rev. Med. Chem. 2004, 4, 477–486. [Google Scholar] [CrossRef] [PubMed]
- Mak, A.N.; Wong, Y.T.; An, Y.J.; Cha, S.S.; Sze, K.H.; Au, S.W.; Wong, K.B.; Shaw, P.C. Structure-function study of maize ribosome-inactivating protein: Implications for the internal inactivation region and the sole glutamate in the active site. Nucleic Acids Res. 2007, 35, 6259–6267. [Google Scholar] [CrossRef] [PubMed]
- Robert, X.; Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 2014, 42, W320–W324. [Google Scholar] [CrossRef] [PubMed]
- Rajamohan, F.; Ozer, Z.; Mao, C.; Uckun, F.M. Active center cleft residues of pokeweed antiviral protein mediate its high-affinity binding to the ribosomal protein L3. Biochemistry 2001, 40, 9104–9114. [Google Scholar] [CrossRef] [PubMed]
- Chan, S.H.; Hung, F.S.; Chan, D.S.; Shaw, P.C. Trichosanthin interacts with acidic ribosomal proteins P0 and P1 and mitotic checkpoint protein MAD2b. Eur. J. Biochem. 2001, 268, 2107–2112. [Google Scholar] [CrossRef] [PubMed]
- Vater, C.A.; Bartle, L.M.; Leszyk, J.D.; Lambert, J.M.; Goldmacher, V.S. Ricin A chain can be chemically cross-linked to the mammalian ribosomal proteins L9 and L10e. J. Biol. Chem. 1995, 270, 12933–12940. [Google Scholar] [PubMed]
- Hudak, K.A.; Dinman, J.D.; Tumer, N.E. Pokeweed antiviral protein accesses ribosomes by binding to L3. J. Biol. Chem. 1999, 274, 3859–3864. [Google Scholar] [CrossRef] [PubMed]
- Di, R.; Tumer, N.E. Expression of a truncated form of ribosomal protein L3 confers resistance to pokeweed antiviral protein and the fusarium mycotoxin deoxynivalenol. Mol. Plant Microbe Interact. 2005, 18, 762–770. [Google Scholar] [CrossRef] [PubMed]
- McCluskey, A.J.; Poon, G.M.; Bolewska-Pedyczak, E.; Srikumar, T.; Jeram, S.M.; Raught, B.; Gariepy, J. The catalytic subunit of Shiga-like toxin 1 interacts with ribosomal stalk proteins and is inhibited by their conserved C-terminal domain. J. Mol. Biol. 2008, 378, 375–386. [Google Scholar] [CrossRef] [PubMed]
- Chiou, J.C.; Li, X.P.; Remacha, M.; Ballesta, J.P.; Tumer, N.E. The ribosomal stalk is required for ribosome binding, depurination of the rRNA and cytotoxicity of ricin A chain in Saccharomyces cerevisiae. Mol. Microbiol. 2008, 70, 1441–1452. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Mayoral, F.; Garcia-Ortega, L.; Alvarez-Garcia, E.; Bruix, M.; Gavilanes, J.G.; del Pozo, A.M. Modeling the highly specific ribotoxin recognition of ribosomes. FEBS Lett. 2005, 579, 6859–6864. [Google Scholar] [CrossRef] [PubMed]
- Korennykh, A.V.; Piccirilli, J.A.; Correll, C.C. The electrostatic character of the ribosomal surface enables extraordinarily rapid target location by ribotoxins. Nat. Struct. Mol. Biol. 2006, 13, 436–443. [Google Scholar] [CrossRef] [PubMed]
- Korennykh, A.V.; Correll, C.C.; Piccirilli, J.A. Evidence for the importance of electrostatics in the function of two distinct families of ribosome inactivating toxins. RNA 2007, 13, 1391–1396. [Google Scholar] [CrossRef] [PubMed]
- May, K.L.; Yan, Q.; Tumer, N.E. Targeting ricin to the ribosome. Toxicon 2013, 69, 143–151. [Google Scholar] [CrossRef] [PubMed]
- Li, X.P.; Chiou, J.C.; Remacha, M.; Ballesta, J.P.; Tumer, N.E. A two-step binding model proposed for the electrostatic interactions of ricin A chain with ribosomes. Biochemistry 2009, 48, 3853–3863. [Google Scholar] [CrossRef] [PubMed]
- Li, X.P.; Grela, P.; Krokowski, D.; Tchorzewski, M.; Tumer, N.E. Pentameric organization of the ribosomal stalk accelerates recruitment of ricin A chain to the ribosome for depurination. J. Biol. Chem. 2010, 285, 41463–41471. [Google Scholar] [CrossRef] [PubMed]
- Too, P.H.; Ma, M.K.; Mak, A.N.; Wong, Y.T.; Tung, C.K.; Zhu, G.; Au, S.W.; Wong, K.B.; Shaw, P.C. The C-terminal fragment of the ribosomal P protein complexed to trichosanthin reveals the interaction between the ribosome-inactivating protein and the ribosome. Nucleic Acids Res. 2009, 37, 602–610. [Google Scholar] [CrossRef] [PubMed]
- Savino, C.; Federici, L.; Ippoliti, R.; Lendaro, E.; Tsernoglou, D. The crystal structure of saporin SO6 from Saponaria officinalis and its interaction with the ribosome. FEBS Lett. 2000, 470, 239–243. [Google Scholar] [CrossRef]
- Shi, W.W.; Tang, Y.S.; Sze, S.Y.; Zhu, Z.N.; Wong, K.B.; Shaw, P.C. Crystal structure of ribosome-inactivating protein ricin A chain in complex with the C-terminal peptide of the ribosomal stalk protein P2. Toxins 2016, 8, 296. [Google Scholar] [CrossRef] [PubMed]
- Choi, A.K.; Wong, E.C.; Lee, K.M.; Wong, K.B. Structures of eukaryotic ribosomal stalk proteins and its complex with trichosanthin, and their implications in recruiting ribosome-inactivating proteins to the ribosomes. Toxins 2015, 7, 638–647. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.M.; Yusa, K.; Chu, L.O.; Yu, C.W.; Oono, M.; Miyoshi, T.; Ito, K.; Shaw, P.C.; Wong, K.B.; Uchiumi, T. Solution structure of human P1*P2 heterodimer provides insights into the role of eukaryotic stalk in recruiting the ribosome-inactivating protein trichosanthin to the ribosome. Nucleic Acids Res. 2013, 41, 8776–8787. [Google Scholar] [CrossRef] [PubMed]
- Diaconu, M.; Kothe, U.; Schlunzen, F.; Fischer, N.; Harms, J.M.; Tonevitsky, A.G.; Stark, H.; Rodnina, M.V.; Wahl, M.C. Structural basis for the function of the ribosomal L7/12 stalk in factor binding and GTPase activation. Cell 2005, 121, 991–1004. [Google Scholar] [CrossRef] [PubMed]
- Naganuma, T.; Nomura, N.; Yao, M.; Mochizuki, M.; Uchiumi, T.; Tanaka, I. Structural basis for translation factor recruitment to the eukaryotic/archaeal ribosomes. J. Biol. Chem. 2010, 285, 4747–4756. [Google Scholar] [CrossRef] [PubMed]
- Nolan, R.D.; Grasmuk, H.; Drews, J. The binding of tritiated elongation-factors 1 and 2 to ribosomes from Krebs II mouse ascites-tumore cells. The influence of various antibiotics and toxins. Eur. J. Biochem. 1976, 64, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Bargis-Surgey, P.; Lavergne, J.P.; Gonzalo, P.; Vard, C.; Filhol-Cochet, O.; Reboud, J.P. Interaction of elongation factor eEF-2 with ribosomal P proteins. Eur. J. Biochem. 1999, 262, 606–611. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Puentes, C.; Benson, S.; Olsnes, S.; Pihl, A. Protective effect of elongation factor 2 on the inactivation of ribosomes by the toxic lectins abrin and ricin. Eur. J. Biochem. 1976, 64, 437–443. [Google Scholar] [CrossRef] [PubMed]
- Cawley, D.B.; Hedblom, M.L.; Houston, L.L. Protection and rescue of ribosomes from the action of ricin A chain. Biochemistry 1979, 18, 2648–2654. [Google Scholar] [CrossRef] [PubMed]
- Ayub, M.J.; Smulski, C.R.; Ma, K.W.; Levin, M.J.; Shaw, P.C.; Wong, K.B. The C-terminal end of P proteins mediates ribosome inactivation by trichosanthin but does not affect the pokeweed antiviral protein activity. Biochem. Biophys. Res. Commun. 2008, 369, 314–319. [Google Scholar] [CrossRef] [PubMed]
- Wong, Y.T.; Ng, Y.M.; Mak, A.N.; Sze, K.H.; Wong, K.B.; Shaw, P.C. Maize ribosome-inactivating protein uses Lys158-Lys161 to interact with ribosomal protein P2 and the strength of interaction is correlated to the biological activities. PLoS ONE 2012, 7, e49608. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Mak, A.N.; Shaw, P.C.; Sze, K.H. Solution structure of an active mutant of maize ribosome-inactivating protein (MOD) and its interaction with the ribosomal stalk protein P2. J. Mol. Biol. 2010, 395, 897–907. [Google Scholar] [CrossRef] [PubMed]
- Hartley, M.R.; Legname, G.; Osborn, R.; Chen, Z.; Lord, J.M. Single-chain ribosome inactivating proteins from plants depurinate Escherichia coli 23S ribosomal RNA. FEBS Lett. 1991, 290, 65–68. [Google Scholar] [CrossRef]
- Suh, J.K.; Hovde, C.J.; Robertus, J.D. Shiga toxin attacks bacterial ribosomes as effectively as eucaryotic ribosomes. Biochemistry 1998, 37, 9394–9398. [Google Scholar] [CrossRef] [PubMed]
- Pizzo, E.; Di Maro, A. A new age for biomedical applications of ribosome inactivating proteins (RIPs): From bioconjugate to nanoconstructs. J. Biomed. Sci. 2016, 23, 54. [Google Scholar] [CrossRef] [PubMed]
- Puri, M.; Kaur, I.; Perugini, M.A.; Gupta, R.C. Ribosome-inactivating proteins: Current status and biomedical applications. Drug Discov. Today 2012, 17, 774–783. [Google Scholar] [CrossRef] [PubMed]
- Stirpe, F. Ribosome-inactivating proteins: From toxins to useful proteins. Toxicon 2013, 67, 12–16. [Google Scholar] [CrossRef] [PubMed]
- Bolognesi, A.; Polito, L. Immunotoxins and other conjugates: Pre-clinical studies. Mini Rev. Med. Chem. 2004, 4, 563–583. [Google Scholar] [CrossRef] [PubMed]
- Gilabert-Oriol, R.; Weng, A.; Mallinckrodt, B.; Melzig, M.F.; Fuchs, H.; Thakur, M. Immunotoxins constructed with ribosome-inactivating proteins and their enhancers: A lethal cocktail with tumor specific efficacy. Curr. Pharm. Des. 2014, 20, 6584–6643. [Google Scholar] [CrossRef] [PubMed]
- Hess, P.R.; Barnes, C.; Woolard, M.D.; Johnson, M.D.; Cullen, J.M.; Collins, E.J.; Frelinger, J.A. Selective deletion of antigen-specific CD8+ T cells by MHC class I tetramers coupled to the type I ribosome-inactivating protein saporin. Blood 2007, 109, 3300–3307. [Google Scholar] [CrossRef] [PubMed]
- Alexeev, L.P.; Dolbin, A.G.; Toptygin, A.Y.; Moysuk, Y.G.; Prokofiev, S.A.; Tonevitsky, A.G.; Sechkin, A.V.; Shumakov, V.A.; Khaitov, M.R. New method of kidney allograft pretreatment (six-year follow-up). Transplant. Proc. 2001, 33, 1240–1241. [Google Scholar] [CrossRef]
- LaCasse, E.C.; Bray, M.R.; Patterson, B.; Lim, W.M.; Perampalam, S.; Radvanyi, L.G.; Keating, A.; Stewart, A.K.; Buckstein, R.; Sandhu, J.S.; et al. Shiga-Like Toxin-1 Receptor on Human Breast Cancer, Lymphoma, and Myeloma and Absence From CD34+ Hematopoietic Stem Cells: Implications for Ex Vivo Tumor Purging and Autologous Stem Cell Transplantation. Blood 1999, 94, 2901–2910. [Google Scholar] [PubMed]
- Parikh, B.A.; Tumer, N.E. Antiviral activity of ribosome inactivating proteins in medicine. Mini Rev. Med. Chem. 2004, 4, 523–543. [Google Scholar] [CrossRef] [PubMed]
- Tumer, N.E.; Hudak, K.; Di, R.; Coetzer, C.; Wang, P.; Zoubenko, O. Pokeweed antiviral protein and its applications. Curr. Top. Microbiol. Immunol. 1999, 240, 139–158. [Google Scholar] [PubMed]
- Nielsen, K.; Payne, G.A.; Boston, R.S. Maize ribosome-inactivating protein inhibits normal development of Aspergillus nidulans and Aspergillus flavus. Mol. Plant Microbe Interact. 2001, 14, 164–172. [Google Scholar] [CrossRef] [PubMed]
- Lorenzetti, I.; Meneguzzi, A.; Fracasso, G.; Potrich, C.; Costantini, L.; Chiesa, E.; Legname, G.; Menestrina, G.; Tridente, G.; Colombatti, M. Genetic grafting of membrane-acting peptides to the cytotoxin dianthin augments its ability to de-stabilize lipid bilayers and enhances its cytotoxic potential as the component of transferrin-toxin conjugates. Int. J. Cancer 2000, 86, 582–589. [Google Scholar] [CrossRef]
- Von Mallinckrodt, B.; Thakur, M.; Weng, A.; Gilabert-Oriol, R.; Durkop, H.; Brenner, W.; Lukas, M.; Beindorff, N.; Melzig, M.F.; Fuchs, H. Dianthin-EGF is an effective tumor targeted toxin in combination with saponins in a xenograft model for colon carcinoma. Future Oncol. 2014, 10, 2161–2175. [Google Scholar] [CrossRef] [PubMed]
- Weyergang, A.; Selbo, P.K.; Berg, K. Photochemically stimulated drug delivery increases the cytotoxicity and specificity of EGF-saporin. J. Control. Release 2006, 111, 165–173. [Google Scholar] [CrossRef] [PubMed]
- Bostad, M.; Kausberg, M.; Weyergang, A.; Olsen, C.E.; Berg, K.; Hogset, A.; Selbo, P.K. Light-triggered, efficient cytosolic release of IM7-saporin targeting the putative cancer stem cell marker CD44 by photochemical internalization. Mol. Pharm. 2014, 11, 2764–2776. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.R.; Au, K.Y.; Zheng, H.Y.; Gao, L.M.; Zhang, X.; Luo, R.H.; Law, S.K.; Mak, A.N.; Wong, K.B.; Zhang, M.X.; et al. The recombinant maize ribosome-inactivating protein transiently reduces viral load in SHIV89.6 infected Chinese rhesus macaques. Toxins 2015, 7, 156–169. [Google Scholar] [CrossRef] [PubMed]
- Law, S.K.; Wang, R.R.; Mak, A.N.; Wong, K.B.; Zheng, Y.T.; Shaw, P.C. A switch-on mechanism to activate maize ribosome-inactivating protein for targeting HIV-infected cells. Nucleic Acids Res. 2010, 38, 6803–6812. [Google Scholar] [CrossRef] [PubMed]



| RIP | Source | Ligand | PDB | Resolution (Å) | Cα RMSD |
|---|---|---|---|---|---|
| Type 1 | |||||
| Bouganin | Bougainvillea spectabilis | - | 3CTK | 1.8 | 1.1 |
| Bryodin 1 | Bryonia dioica | - | 1BRY | 2.1 | 0.4 |
| Charybdin | Charybdis maritima | 2-N-morpholino-ethanesulfonic acid | 2B7U | 1.6 | 1.9 |
| Cucurmosin | Cucurbita moschata | - | 3BWH | 1.0 | 0.4 |
| Dianthin | Dianthus caryophyllus | Adenine | 1LPD | 1.7 | 2.6 |
| Luffaculin 1 | Luffa acutangula | N-Acetyl d-glucosamine | 2OQA | 1.4 | 0.5 |
| beta-luffin | Luffa cylindrica | N-Acetyl-d-glucosamine | 1NIO | 2 | 0.5 |
| GAP31 | Gelonium multiforum | 3KTZ | 1.6 | 0.745 | |
| Lychnin | Lychnis chalcedonica | - | 2G5X | 1.7 | 1.9 |
| MAP30 | Momordica charantia | - | 1D8V | NMR | 1.6 |
| α-Momorcharin | Momordica charantia | Adenine | 1AHA | 2.2 | 0.5 |
| β-Momorcharin | Momordica charantia | Modified hexasaccharide | 1CF5 | 2.6 | 0.6 |
| PAP | Phytolacca americana | 2-(acetylamino)-2-deoxy-A-d-glucopyranose | 1GIK | 1.8 | 1.6 |
| PD-L4 | Phytolacca dioica | 2Z4U | 1.1 | 1.7 | |
| TCS | Trichosanthes kirilowii | Adenine | 2JDL | 1.8 | - |
| Saporin | Saponaria officinalis | - | 1QI7 | 2 | 2.0 |
| Type 2 | |||||
| Abrus agglutinin 1 | Abrus precatorius | N-Acetyl-d-glucosamine | 2Q3N | 3.5 | 1.0 |
| Cinnamomin | Cinnamonum camphora | - | 2VLC | 2.95 | 0.752 |
| Ebulin | Sambucus ebulus | Beta-d-galactose | 1HWM | 2.8 | 0.9 |
| Mistletoe lectin 1 | Viscum album | N-Acetyl-d-glucosamine | 1ONK | 2.1 | 1.0 |
| Ricin | Ricinus communis | Adenine | 1IFS | 2 | 1.1 |
| Shiga toxin | Bacteriophage 933W | Adenine | 2GA4 | 1.8 | 2.9 |
| Atypical RIP | |||||
| MOD | Zea mays | - | 2PQI | 2.5 | 3.4 |
| Maize RIP | Zea mays | 2PQG | 2.4 | 3.8 | |
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shi, W.-W.; Mak, A.N.-S.; Wong, K.-B.; Shaw, P.-C. Structures and Ribosomal Interaction of Ribosome-Inactivating Proteins. Molecules 2016, 21, 1588. https://doi.org/10.3390/molecules21111588
Shi W-W, Mak AN-S, Wong K-B, Shaw P-C. Structures and Ribosomal Interaction of Ribosome-Inactivating Proteins. Molecules. 2016; 21(11):1588. https://doi.org/10.3390/molecules21111588
Chicago/Turabian StyleShi, Wei-Wei, Amanda Nga-Sze Mak, Kam-Bo Wong, and Pang-Chui Shaw. 2016. "Structures and Ribosomal Interaction of Ribosome-Inactivating Proteins" Molecules 21, no. 11: 1588. https://doi.org/10.3390/molecules21111588
APA StyleShi, W.-W., Mak, A. N.-S., Wong, K.-B., & Shaw, P.-C. (2016). Structures and Ribosomal Interaction of Ribosome-Inactivating Proteins. Molecules, 21(11), 1588. https://doi.org/10.3390/molecules21111588






