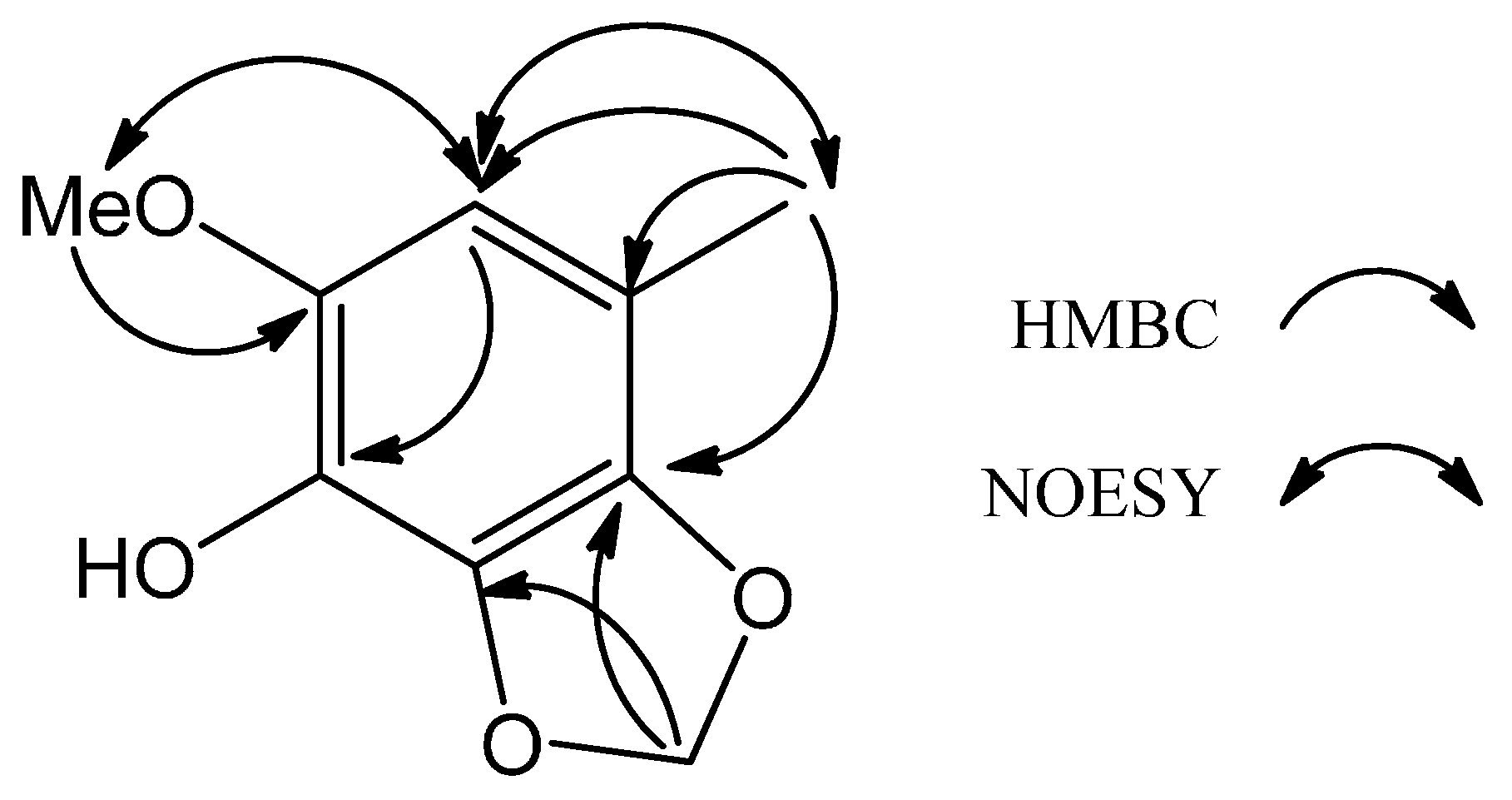1. Introduction
Antrodia camphorata Wu, Ryvarden & Chang (Polyporaceae), a fungus indigenous to Taiwan, was used by the aborigine as a hepatinica and anti-alcoholic agent initially and has been gradually used as a folk remedy for the treatment of liver cancer and various cardiovascular diseases in the past few decades [
1]. The fermentation and development of this fungus have already become one of the major components of the biotechnology industry in Taiwan recently, and some of the chemical entities isolated from the fermented products, e.g., antroquinonol [
2] and ergosta-7,9,22
E-trien-3β-ol [
3], were selected as leads for new drug development. Although over one hundred compounds have been identified from this fungus so far [
4,
5,
6], new chemical entities are still reported continually by virtue of varied culturing conditions for this fungus. In continuing our investigations on the chemical constituents of the mycelium of
Antrodia camphorata, one new phenylmethanoid, 6-methoxy-4-methyl-2,3-(methylenedioxy)phenol (
1), and one new stilbene, 4,4'-(ethane-1,2-diyl)bis(2,3,6-trimethoxyphenol)(
2), together with 2,3,6-trimethoxy-4-methylphenol (
3), 1(10→6)
abeo-ergosta-5,7,9,22-tetraen-3α-ol (
4), citreoanthrasteroid B (
5) and dankasterones A (
6) and B (
7) (
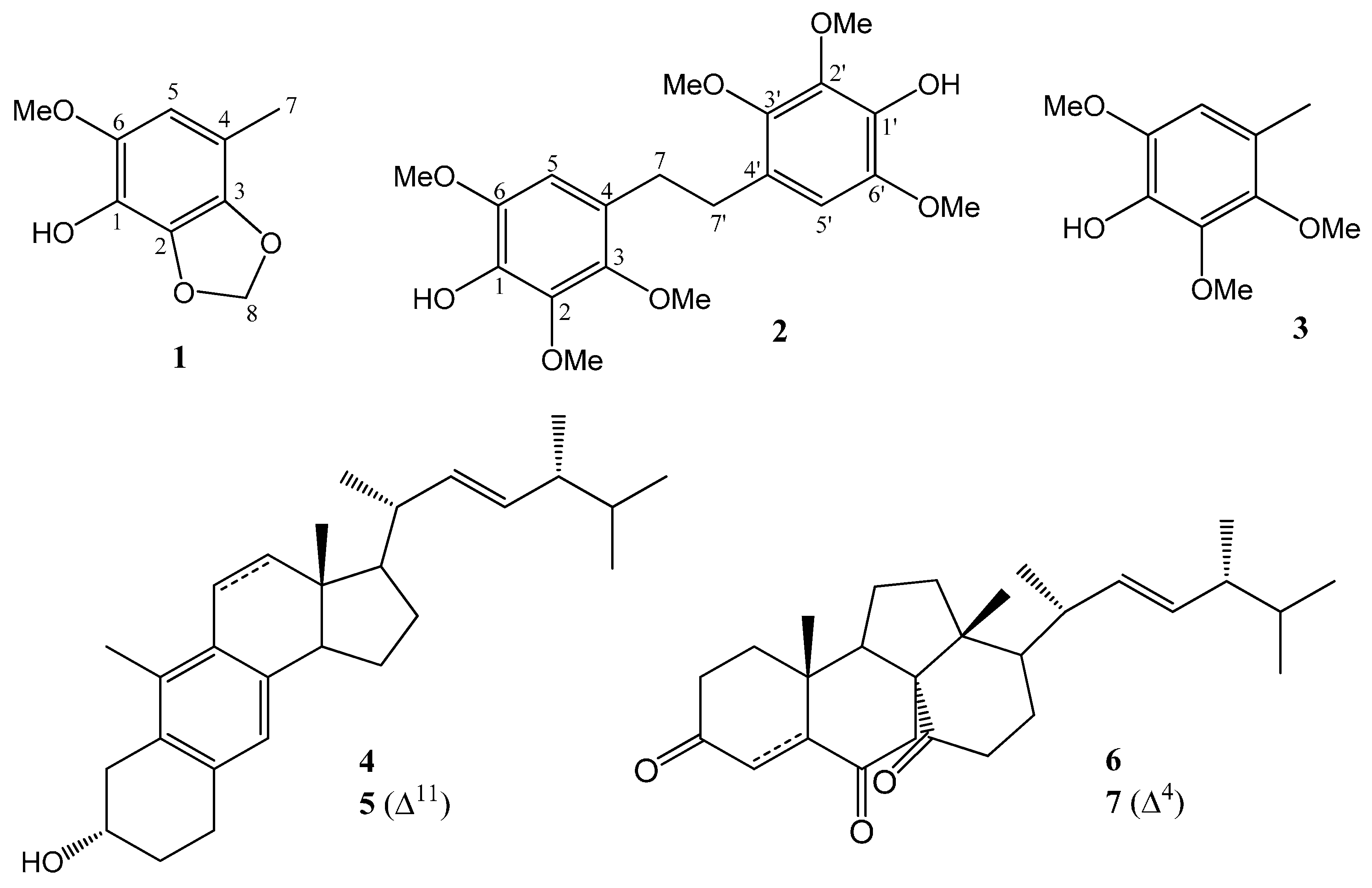
Figure 1) were isolated and characterized on the basis of the spectral analysis. The paper describes the isolation and identification of compounds
1–
7 from
A. camphorata along with their anticancer effects in murine colorectal cancer CT26 cells and human leukemia K562 cells.
Figure 1.
Chemical structures of 1–7 isolated in this study.
Figure 1.
Chemical structures of 1–7 isolated in this study.
2. Results and Discussion
From the methanolic extracts of the solid-state cultured, fermented dried powder of
A. camphorata, seven major compounds, including one new phenylmethanoid,
1, and one new stilbene,
2, along with a known phenylmethanoid,
3, and four steroids,
4–
7, were isolated by sequential separation on Si-gravity column and normal-phase HPLC. Compound
3 was obtained as an amorphous white solid whose
1H,
13C-NMR, IR, optical rotation and MS were consistent with those of synthetic 2,3,6-trimethoxy-4-methylphenol [
7], and this is the first time Compound
3 has been isolated from a natural resource. Compounds
4 and
5, two rare 1(10→6)
abeo-ergostane-type steroids, were identified as 1(10→6)
abeo-ergosta-5,7,9,22-tetraen-3α-ol (
4), obtained previously from the stromata of
Epichloe typhina [
8], and citreoanthrasteroid B (
5), isolated from a hybrid bacterial strain, KO 0231 [
9]. Compounds
6 and
7, two uncommon 13(14→8)
abeo-ergostane-type steroids, were characterized as respective dankasterones A and B, which were isolated previously from a
Halichondria sponge-derived fungus,
Gymnascella dankaliensis [
10].
Compound
1 was afforded as a colorless amorphous solid with the molecular formula of C
9H
10O
4 as established through the analysis of its
13C-NMR and HRESIMS data. The IR absorption peaks of
1 at 3277, 1626 and 1527 cm
−1 indicated that
1 contained a benzene moiety bearing a hydroxy functionality as reflected in its
1H- and
13C-NMR data. The
1H-NMR of
1 exhibited a phenyl proton at δ
H 6.27 (s, 1H), a benzene-borne methyl at δ
H 2.16 (s, 3H), a benzene-borne methoxyl at δ
H 3.82 (s, 3H) and a methylenedioxy group at δ
H 5.91 (s, 2H), which were further confirmed by their corresponding six phenyl resonances at δ
C 109.0 (d), 118.9 (s), 131.9 (s), 133.6 (s), 135.0 (s) and 136.8 (s), one methyl resonance at δ
C 15.8 (q), one methoxyl resonance at δ
C 57.2 (q) and one dioxygenated methylene resonance at δ
C 101.5 (t) in the
13C-NMR of
1. Further analysis of the 2D NMR data of
1, key cross-peaks of δ
H 2.16 (H
3-7)/δ
C 109.0 (C-5), 118.9 (C-4) and 131.9 (C-3), δ
H 3.82 (OMe-6)/δ
C 136.8 (C-6), δ
H 5.91 (H
2-8)/δ
C 131.9 (C-3) and 135.0 (C-2), δ
H 6.27 (H-5)/δ
C 133.6 (C-1) in the HMBC spectrum and mutual-correlated cross peaks of δ
H 6.27 (H-5)/δ
H 3.82 (OMe-6) and δ
H 2.16 (H
3-7) in the NOESY spectrum (
Figure 2) established the locations of all of the functional groups attached to the benzene ring. Thus,
1 was deduced as the shown phenylmethanoid and was named 6-methoxy-4-methyl-2,3-(methylenedioxy)phenol.
Figure 2.
Key HMBC and NOESY of 1.
Figure 2.
Key HMBC and NOESY of 1.
Compound
2, obtained as a colorless amorphous solid, was a symmetrical chemical entity as judged from its molecular formula, C
20H
26O
8, deduced from HRESIMS and only ten resonances in the
13C-NMR spectrum. The IR spectrum of
2 confirmed the presence of a hydroxyl group (3410 cm
−1) and a benzene ring (1610 and 1503 cm
−1). The
13C-NMR along with the DEPT of
2 displayed only ten signals (indeed, twenty signals due to symmetry), including one methylene carbon at δ
C 31.4, three benzene-borne methoxyl carbons at δ
C 56.4, 60.7 and 61.0 and six phenyl carbons at δ
C 107.2, 125.2, 137.3, 140.4, 143.2 and 145.2. The
1H-NMR spectrum showed signals for the methylene group at δ
H 2.78 (s, 2H), three methoxyl groups at δ
H 3.79 (s, 3H), 3.80 (s, 3H) and 3.91 (s, 3H), one hydroxyl group at δ
H 5.45 (brs, 1H) and one phenyl proton at δ
H 6.38 (s, 1H). The above assignments indicated that
2 had a stilbene skeleton with three methoxyl and one hydroxyl groups on each benzene ring. The locations of three methoxy, one hydroxy and one phenyl proton were further corroborated by key HMBC interpretations, including δ
H 2.78 (H-7, -7')/δ
C 107.2 (C-5, -5'), 125.2 (C-4, -4') and 145.2 (C-3, -3'), δ
H 3.79 (OMe-3, -3')/δ
C 145.2 (C-3, -3'), δ
H 3.80 (OMe-6, -6')/δ
C 143.2 (C-6, -6'), δ
H 3.91 (OMe-2, -2')/δ
C 140.4 (C-2, -2'), δ
H 5.45 (OH-1, -1')/δ
C 137.3 (C-1, -1'), 140.3 (C-2, -2'), 143.2 (C-6, -6') and δ
H 6.38 (H-5, -5')/δ
C 137.3 (C-1, -1') and 145.2 (C-3, -3'), as well as key NOESY correlations, including δ
H 6.38 (H-5, -5')/δ
H 2.78 (H-7, -7') and 3.80 (OMe-6, -6') and δ
H 3.79 (OMe-3, -3')/δ
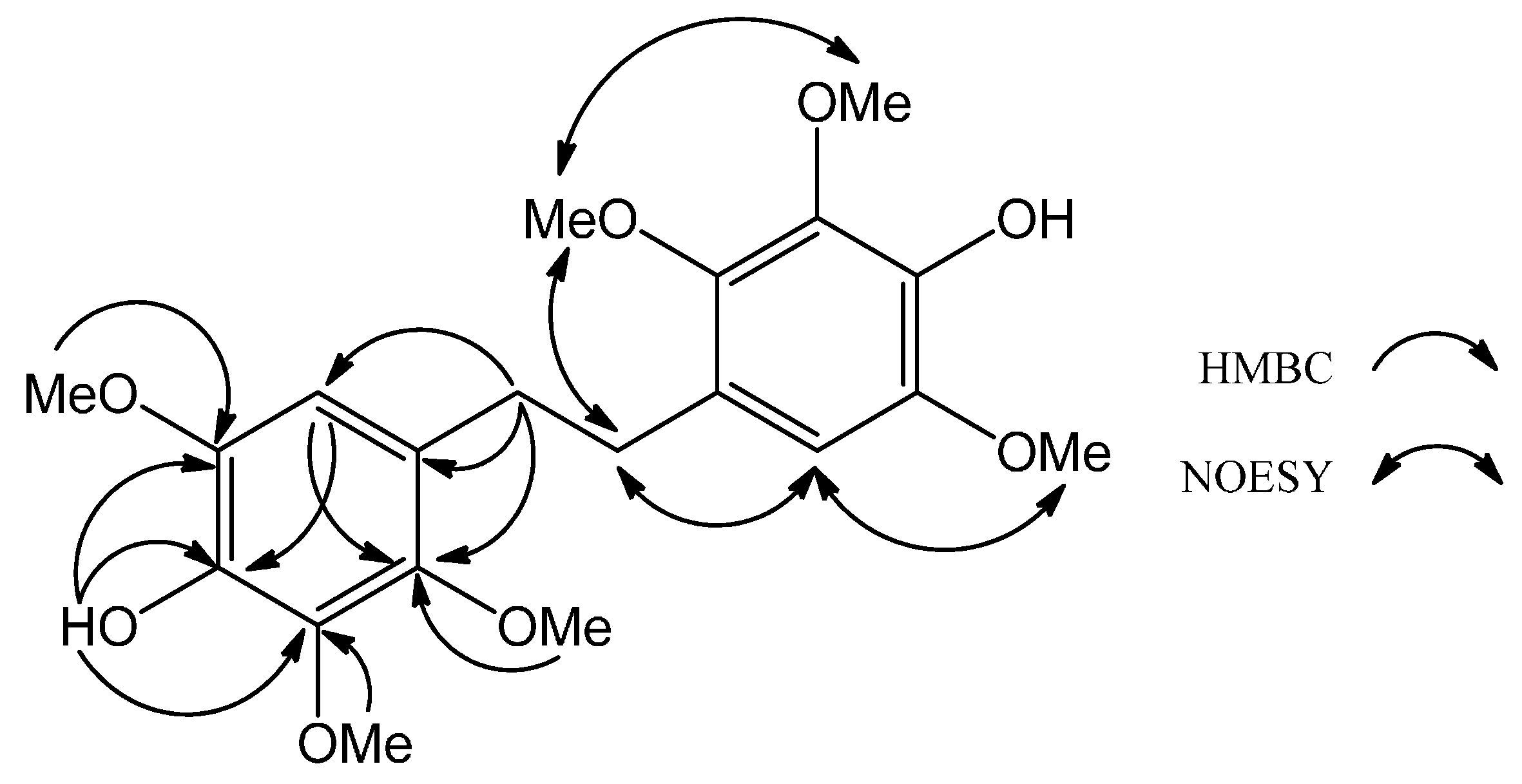
H 2.78 (H-7, -7') and 3.91 (OMe-2, -2') (
Figure 3). Accordingly,
1 was determined as the shown dihydrostilbene and was named 4,4′-(ethane-1,2-diyl)bis(2,3,6-trimethoxyphenol). Compound
2 was speculated from the oxidative coupling between two molecules of Compound
3.
For the anticancer activity,
Table 1 shows the IC
50 values of compounds
1‒
7 against murine colorectal cancer CT26 cells and human leukemia K562 cells. Compounds
1‒
3 exhibited no obvious effect toward CT26 and K562 cells with their IC
50 values higher than 20 μM, and
4–
7 showed significant cytotoxicity toward murine colorectal CT26 and human leukemia K562 cancer cell lines with IC
50 values ranging from 6.7 to 18.2 μM and from 12.5 to 23.1 μM, respectively. At the same condition, the IC
50 value of staurosporine against K562 leukemia cells was 16.7 nM.
Figure 3.
Key HMBC and NOESY of 2.
Figure 3.
Key HMBC and NOESY of 2.
Table 1.
IC50 values of compounds 1–7 against colorectal cancer CT26 and leukemia K562 cells.
Table 1.
IC50 values of compounds 1–7 against colorectal cancer CT26 and leukemia K562 cells.
| Compounds | IC50 (μM) |
|---|
| CT26 | K562 |
|---|
| 1 | >20 | >20 |
| 2 | >20 | >20 |
| 3 | >20 | >20 |
| 4 | 15.3 | 19.9 |
| 5 | 18.2 | 12.5 |
| 6 | 6.7 | >20 |
| 7 | 8.4 | 23.1 |
3. Experimental
3.1. General
Optical rotations were measured on a JASCO DIP-1000 polarimeter (Tokyo, Japan). 1H and 13C-NMR were acquired on a Bruker DMX-500 (Ettlingen, Germany). Low resolution and high resolution mass spectra were obtained using an API4000 triple quadrupole mass spectrometer (Applied Biosystems, Foster City, CA, USA) and a Synapt High Definition Mass Spectrometry system with an ESI interface and a TOF analyzer (Waters Corp., Manchester, UK), respectively. IR spectra were recorded on a JASCO FT/IR 4100 spectrometer (Tokyo, Japan). TLC was performed using silica gel 60 F254 plates (200 µm, Merck, Taipei, Taiwan).
3.2. Fungal Material
Freeze-dried powder of Antrodia camphorata was provided by Kang Jian Biotech Corp. Ltd, Nantan, Taiwan R.O.C.
3.3. Extraction and Isolation
The dried powder (2.83 kg) of solid-state cultured Antrodia camphorata was extracted with 12 L methanol for three times, and the concentrated residues (323.8 g) were suspended in H2O and further partitioned three times with equal volumes of ethyl acetate, then concentrated in vacuum to dryness (131.5 g). The ethyl acetate extract was applied onto an open column with silica gel. The column was eluted with mixtures of n-hexane, ethyl acetate and methanol, and each 1 L was collected as one fraction. Fractions 16–33 were combined and evaporated to dryness (1.4 g), which were further purified by HPLC on a semi-preparative Phenomenex Luna Si column (5 µm, 10 × 250 mm) with n-hexane–ethyl acetate (90:10, v/v) as the eluent, 2 mL/min, obtaining 1 (4.6 mg), 2 (6.4 mg) and 3 (8.9 mg). Fractions 92–124 were combined and evaporated to dryness (33.1 g), which were further purified by HPLC on the same column with n-hexane–ethyl acetate (50:50, v/v) as the eluent, 2 mL/min, obtaining 4 (5.6 mg), 5 (7.4 mg), 6 (10.3) and 7 (9.2 mg).
6-Methoxy-4-methyl-2,3-(methylenedioxy)phenol (1): Colorless amorphous solid; IR (neat): νmax 3277, 3015, 1626, 1527, 1440, 1341, 1208, 1129, 1043, 925 cm−1. 1H-NMR (CDCl3, 400 MHz): δH 2.16 (3H, s, H-7), 3.82 (3H, s, OMe-6), 5.91 (2H, s, H-8), 6.27 (1H, s, H-6). 13C-NMR (CDCl3, 100 MHz): δC 15.8 (C-7), 57.2 (Ome-6), 101.5 (C-8), 109.0 (C-5), 118.9 (C-4), 131.9 (C-3), 133.6 (C-1), 135.0 (C-2), 136.8 (C-6). ESIMS: m/z = 183 [M+H]+. HRESIMS: m/z = 183.0653 [M+H]+ (calcd. for C9H11O4, 183.0657).
4,4'-(Ethane-1,2-diyl)bis(2,3,6-trimethoxyphenol) (2): Colorless amorphous solid; IR (neat): νmax 3410, 3012, 2956, 2860, 1610, 1503, 1420, 1325, 1248, 1198, 1125, 1076, 970, 880. 1H-NMR (CDCl3, 400 MHz): δH 2.78 (4H, s, H-7, -7'), 3.79 (6H, s, OMe-3, -3'), 3.80 (6H, s, OMe-6, -6'), 3.91 (6H, s, OMe-2, -2'), 5.45 (2H, brs, OH-1, -1'), 6.38 (2H, s, H-5, -5'). 13C-NMR (CDCl3, 100 MHz): δC 31.4 (C-7, -7'), 56.4 (OMe-3, -3'), 60.7 (OMe-6, -6'), 61.0 (OMe-2, -2'), 107.2 (C-5, -5'), 125.2 (C-4, -4'), 137.3 (C-1, -1'), 140.4 (C-2, -2'), 143.2 (C-6, -6'), 145.2 (C-3, -3'). ESIMS: m/z = 395 [M+H]+. HRESIMS: m/z = 395.1710 [M+H]+ (calcd. for C20H27O8, 395.1706).
3.4. Cells and Viability Assay
Murine colorectal cancer CT26 cells were cultured in RPMI1640 medium (Gibco, Grand Island, NY, USA) supplemented with 10% heat-inactivated fetal bovine serum (FBS; Hyclone, Logan, UT) and 2 mM l-glutamine at 37 °C in a humidified 5% CO2 incubator. The viability of CT26 cells was determined by using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT, Sigma) colorimetric assay. Human chronic myeloid leukemia K562 cells were cultured in RPMI 1640 medium, 10% FBS and 2 mM l-glutamine at 37 °C in an incubator. The viability of K562 cells with compound treatments for one day were measured using the Trypan blue dye exclusion test. The IC50 values of compounds in CT26 and K562 cells were determined by using SigmaStat software (Jandel Scientific, San Rafael, CA, USA).