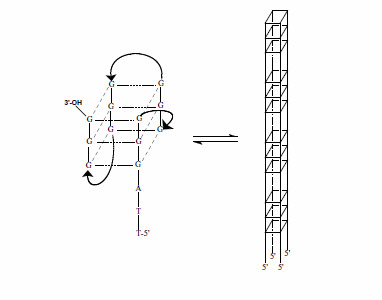
The Human Telomere Sequence, (TTAGGG)4, in the Absence and Presence of Cosolutes: A Spectroscopic Investigation
Abstract
:1. Introduction
2. Results and Discussion
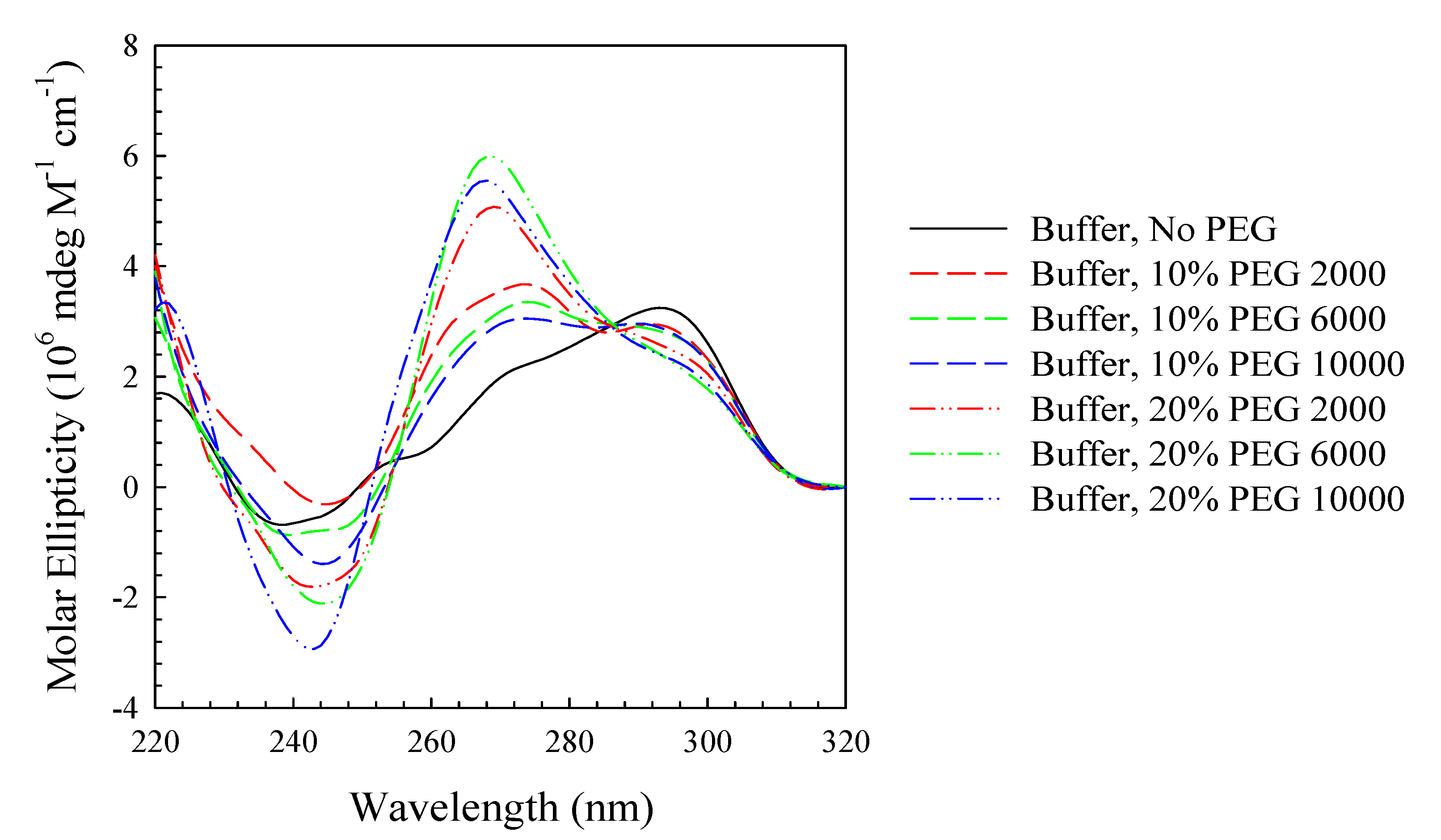
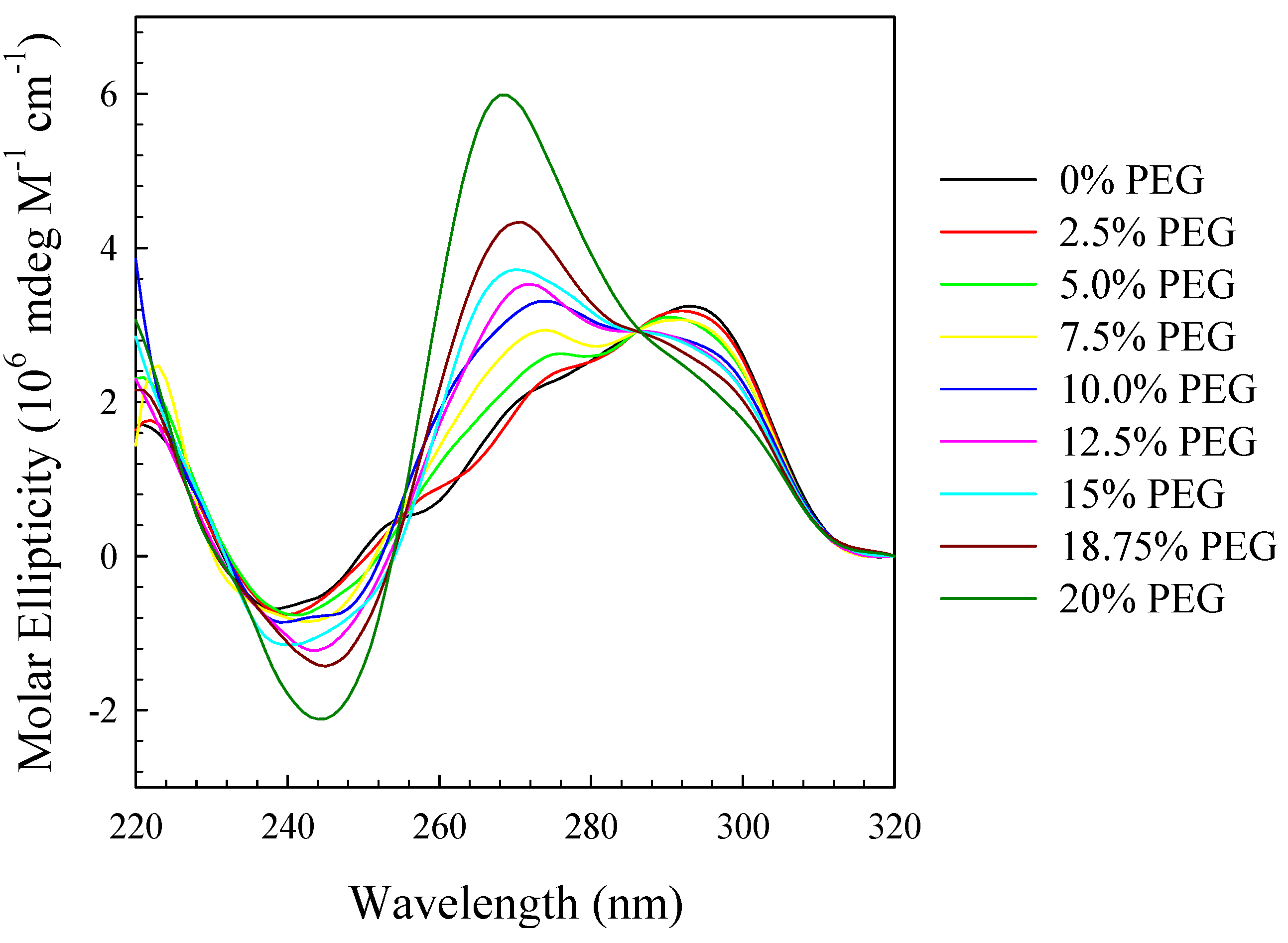


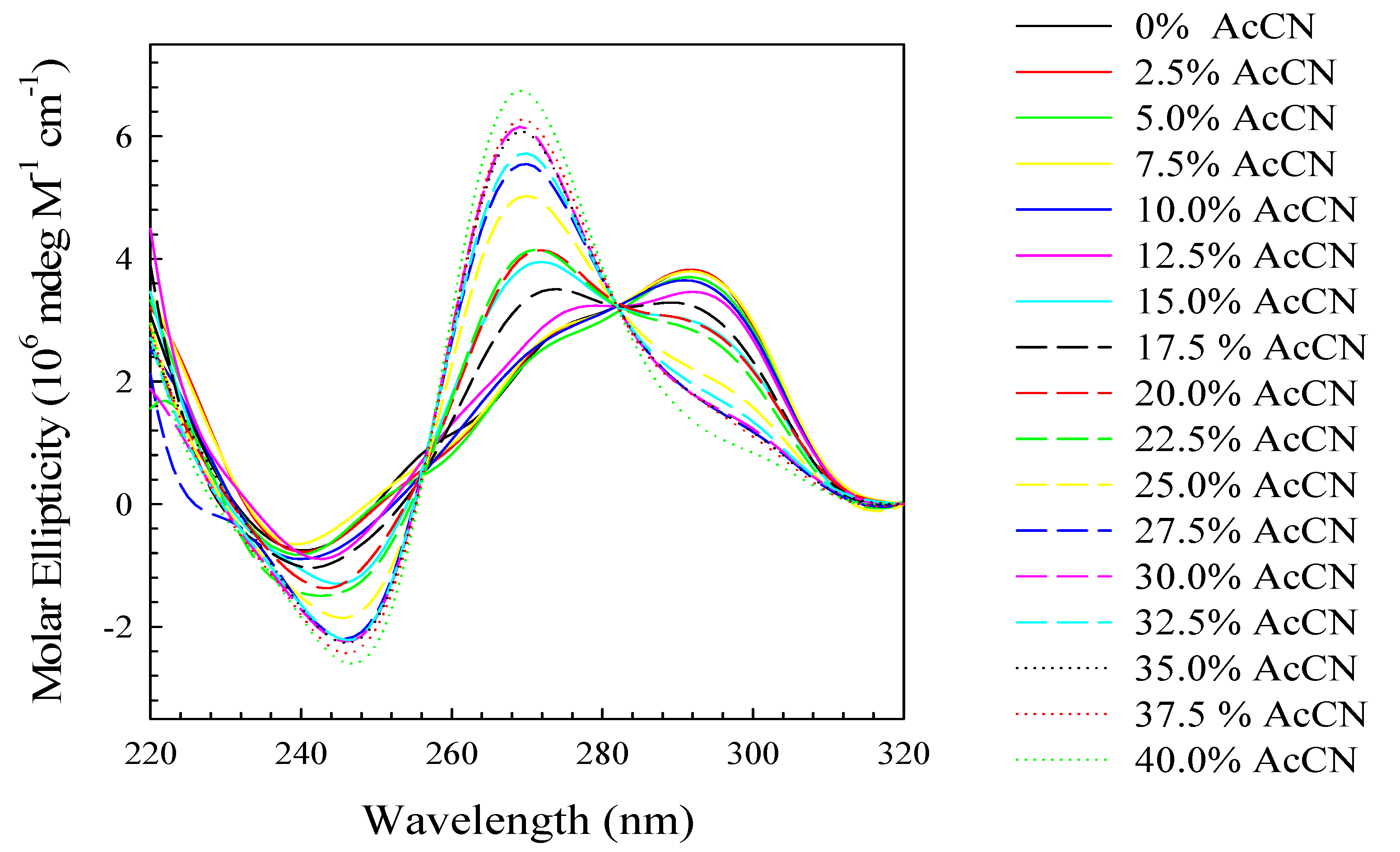

3. Experimental Section
3.1. Preparation of Buffer
3.2. DNA Oligomers
3.3. UV-Vis Spectroscopy
3.4. Circular Dichroism
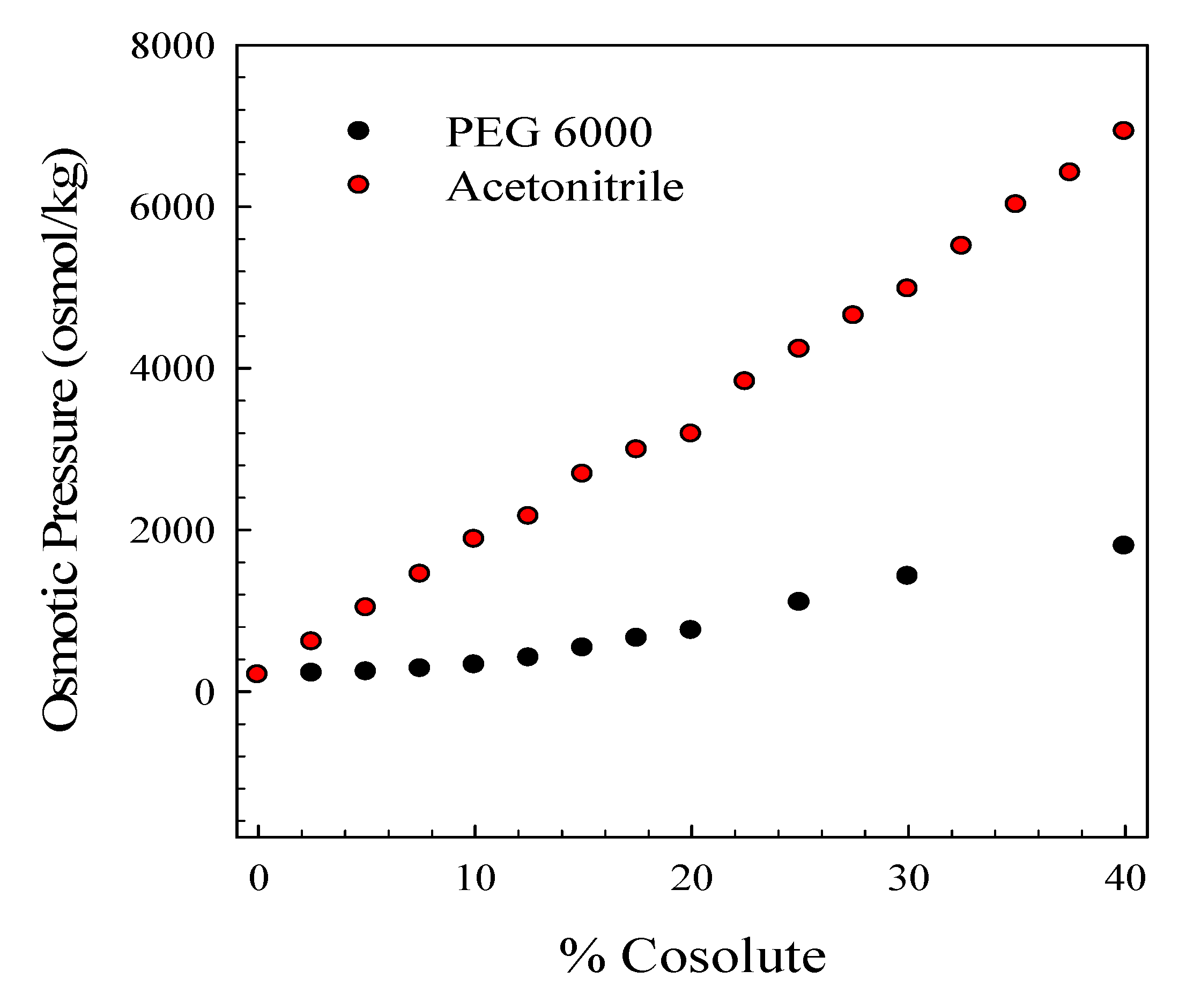
3.5. Osmotic Pressure Determinations
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Watson, J.D.; Crick, F.H.C. Molecular structure of nucleic acids—A structure for deoxyribose nucleic acid. Nature 1953, 171, 737–738. [Google Scholar] [CrossRef]
- Pohl, F.M.; Jovin, T.M. Salt-induced co-operative conformational change of synthetic DNA: Equilibrium and kinetic studies with poly(dG-dC). J. Mol. Biol. 1972, 67, 375–396. [Google Scholar] [CrossRef]
- Williamson, J.R. G-Tetrad structures in telomeric DNA. Ann. Rev. Biophys. Biomol. Struct. 1994, 23, 703–730. [Google Scholar] [CrossRef]
- Venczel, E.A.; Sen, D. Parallel and antiparallel G-DNA structures from a complex telomeric sequence. Biochemistry 1993, 32, 6220–6228. [Google Scholar] [CrossRef]
- Burge, S.; Parkinson, G.N.; Hazel, P.; Todd, A.K.; Neidle, S. Quadruplex DNA: Sequence, topology and structure. Nucleic Acid. Res. 2006, 34, 5402–5415. [Google Scholar] [CrossRef]
- Wang, Y.; Patel, D.J. Solution structure of the human telomeric repeat d[AG3(T2AG3)3] G-tetraplex. Structure 1993, 1, 263–282. [Google Scholar] [CrossRef]
- Ambrus, A.; Chen, D.; Dai, J.; Bialis, T.; Jones, R.; Yang, D. Human telomeric sequence forms a hybrid-type intramolecular G-quadruplex with mixed parallel/antiparallel strands in potassium solution. Nucleic Acid. Res. 2006, 34, 2723–2735. [Google Scholar] [CrossRef]
- Luu, K.N.; Phan, A.T.; Kuryavyi, V.; Lacroix, L.; Patel, D. Structure of the human telomere in K+solution: An intramolecular (3 + 1) G-quadruplex scaffold. J. Am. Chem. Soc. 2006, 128, 9963–9970. [Google Scholar]
- Xu, Y.; Noguchi, Y.; Sugiyama, H. The new models of the human telomere d [AGGG(TTAGGG)3] in K+ solution. Bio-Org. Med. Chem. 2006, 14, 5584–5591. [Google Scholar] [CrossRef]
- Phan, A.T.; Kuryavyi, V.; Luu, K.N.; Patel, D.J. Structure of two intramolecular G-quadruplexes formed by the natural human telomere sequence in K+ solution. Nucleic Acid. Res. 2007, 35, 6517–6525. [Google Scholar] [CrossRef]
- Lim, K.W.; Amrane, S.; Bouaziz, S.; Xu, W.; Mu, Y.; Patel, D.J.; Luu, K.N.; Kim, N.; Phan, A.T. Structure of the human telomere in K+ solution: A stable basket-type G-quadruplex with only two G-tetrad layers. J. Am. Chem. Soc. 2009, 131, 4301–4309. [Google Scholar] [CrossRef]
- Kuryavyi, V.; Patel, D. Solution structure of a unique G-quadruplex scaffold adopted by a guanosine-rich human intronic sequence. Structure 2010, 18, 73–82. [Google Scholar] [CrossRef]
- Zhang, Z.; Dai, J.; Veliath, E.; Jones, R.A.; Yang, D. Structure of a two-G-tetrad intramolecular G-quadruplex formed by a variant human telomeric sequence in K+ solution. Nucleic Acid. Res. 2010, 38, 1009–1021. [Google Scholar] [CrossRef]
- Viglasky, V.; Bauer, L.; Tluckova, K. Structural features of intra- and intermolecular G-quadruplexes derived from the human telomere. Biochemistry 2010, 49, 2110–2120. [Google Scholar] [CrossRef]
- Rachwal, P.A.; Fox, K.R. Quadruplex melting. Methods 2007, 43, 291–301. [Google Scholar] [CrossRef]
- Lane, A.N.; Chaires, J.B.; Gray, R.D.; Trent, J.O. Stability and kinetics of G-quadruplex structures. Nucleic Acid. Res. 2008, 36, 5482–5515. [Google Scholar] [CrossRef]
- Balkwill, G.D.; Garner, T.P.; Searle, M.S. Folding of single-stranded DNA quadruplexes containing an autonomously stable mini-hairpin loop. Mol. BioSyst. 2009, 5, 542–547. [Google Scholar] [CrossRef]
- Olsen, C.M.; Marky, L. Monitoring the temperature unfolding of G-quadruplexes by UV and circular dichroism spectroscopies and calorimetry techniques. Methods Mol. Biol. 2010, 608, 147–158. [Google Scholar] [CrossRef]
- Antonacci, C.; Chaires, J.B.; Sheardy, R.D. Biophysical characterization of the human telomeric repeat (TTAGGG)4 in potassium solution. Biochemistry 2007, 46, 4654–4660. [Google Scholar] [CrossRef]
- Yadav, D.; Sheardy, R.D. A single base permutation in any loop of a folded intramolecular quadruplex influences its structure and stability. J. Biophys. Chem. 2012, 3, 341–347. [Google Scholar] [CrossRef]
- Tucker, B.A.; Gabriel, S.; Sheardy, R.D. A CD Spectroscopic Investigation of Inter- and Intramolecular DNA Quadruplexes. In Frontiers in Nucleic Acids; Sheardy, R.D., Winkle, S.A., Eds.; ACS Symposium Books: Washington, DC, USA, 2011. [Google Scholar]
- Miyoshi, D.; Nakao, A.; Sugimoto, N. Molecular crowding regulates the structural switch of the DNA G-quadruplex. Biochemistry 2002, 41, 15017–15024. [Google Scholar] [CrossRef]
- Xue, Y.; Kan, Z.-Y.; Wang, Q.; Yao, Y.; Liu, J.; Hao, Y.-H.; Tan, Z. Human telomeric DNA forms parallel-stranded intramolecular G-quadruplex in K+ solution under molecular crowding condition. J. Am. Chem. Soc. 2007, 129, 11185–11191. [Google Scholar]
- Petraccone, L.; Malafronte, A.; Amato, J.; Giancola, C. G-Quadruplexes from human telomeric DNA: How many conformations in PEG containing solutions? J. Phys. Chem. 2012, 116, 2294–2305. [Google Scholar]
- Zheng, K.-W.; Chen, Z.; Hao, Y.-H.; Tan, Z. Molecular crowding creates an essential environment for the formation of stable G-quadruplexes in long double-stranded DNA. Nucleic Acid. Res. 2009, 38, 327–338. [Google Scholar] [CrossRef]
- Zhang, D.-H.; Fujimoto, T.; Saxena, S.; Yu, H.-Q.; Miyoshi, D.; Sugimoto, N. Monomorphic RNA G-quadruplex and polymorphic DNA G-quadruplex structures responding to cellular environmental factors. Biochemistry 2010, 49, 4554–4563. [Google Scholar] [CrossRef]
- Heddi, B.; Phan, A.T. Structure of human telomeric DNA in crowded solution. J. Am. Chem. Soc. 2011, 133, 9824–9833. [Google Scholar] [CrossRef]
- Fujimoto, T.; Nakano, S.-I.; Sugimoto, N.; Miyoshi, D. Thermodynamics-hydration relationships within loops that affect G-quadruplexes under molecular crowding conditions. J. Phys. Chem. 2012, 117, 963–972. [Google Scholar]
- Yu, H.; Gu, X.; Nakano, S.-I.; Miyoshi, D.; Sugimoto, N. Beads-on-a-string structure of long telomeric DNAs under molecular crowding condition. J. Am. Chem. Soc. 2012, 134, 20060–20069. [Google Scholar] [CrossRef]
- Knowles, D.B.; LaCroix, A.S.; Deines, N.F.; Shkel, I.; Record, M.T., Jr. Separation of preferential interaction and excluded volume effects on DNA duplex and hairpin stability. Proc. Natl. Acad. Sci. USA 2011, 108, 12699–12704. [Google Scholar] [CrossRef]
- Buscaglia, R.; Miller, M.C.; Dean, W.L.; Gray, R.D.; Lane, A.N.; Trent, J.O.; Chaires, J.B. Polyethylene glycol binding alters human telomere G-quadruplex structure by conformational selection. Nucleic Acids Res. 2013, 41, 7934–7946. [Google Scholar] [CrossRef]
- Zhou, H.-X.; Rivas, G.; Minton, A.P. Macromolecular crowding and confinement: Biochemical, biophysical, and potential physiological consequences. Ann. Rev. Biophys. 2008, 37, 375–397. [Google Scholar] [CrossRef]
- Elcock, A.H. Models of macromolecular crowding effects and the need for quantitative comparisons with experiments. Curr. Opin. Struct. Biol. 2010, 20, 196–206. [Google Scholar] [CrossRef]
- Hirano, A.; Shiraki, K.; Arakawa, T. Polyethylene glycol behaves like weak organic solvent. Biopolymers 2012, 97, 117–122. [Google Scholar] [CrossRef]
- Petraconne, L.; Pagano, B.; Giancola, C. Studying the effect of crowding and dehydration on DNA G-quadruplexes. Methods 2012, 57, 76–83. [Google Scholar] [CrossRef]
- Nakano, S.-I.; Yamaguchi, D.; Tateishi-Karimata, H.; Miyoshi, D.; Sugimoto, N. Hydration changes upon DNA folding studied by osmotic stress experiments. Biophys. J. 2012, 102, 2808–2817. [Google Scholar] [CrossRef]
- Ruggiero, N.J.; Pereira De, S.F.; Colombo, M.F. Hydration effects on DNA double helix stability modulates ligand binding to natural DNA in response to changes in water activity. Cell Mol. Biol. 2001, 47, 801–814. [Google Scholar]
- Priesler, R.S.; Chen, H.H.; Colombo, M.F.; Choe, Y.; Short, B.J., Jr.; Rau, D.C. The B form to Z form transition of poly(dG-m5dC) is sensitive to neutral solutes through osmotic stress. Biochemistry 1995, 34, 14400–14407. [Google Scholar] [CrossRef]
- Miller, M.C.; Buscaglia, R.; Chaires, J.B.; Lane, A.N.; Trent, J.O. Hydration is a major determinant of the G-quadruplex stability and conformation of the human telomere 3’sequence of d(AG3(TTAG3)3). J. Am. Chem. Soc. 2010, 132, 17105–17107. [Google Scholar] [CrossRef]
- Balasubramanian, S.; Hurley, L.H.; Neidle, S. Targeting G-quadruplexes in gene promoters: A novel anticancer strategy? Nat. Rev. Drug Dis. 2011, 10, 261–275. [Google Scholar] [CrossRef]
- Duchler, M. G-quadruplexes: Targets and tools in anticancer drug design. J. Drug Target. 2012, 20, 389–400. [Google Scholar] [CrossRef]
- Bidzinska, J.; Cimino-Reale, G.; Zaffaroni, N.; Folini, M. G-quadruplex structures in the human genome as novel therapeutic targets. Molecules 2012, 18, 12368–12395. [Google Scholar]
- Miller, K.M.; Rodriguez, R. G-quadruplexes: Selective DNA targeting for cancer therapeutics? Expert Rev. Clin. Pharmacol. 2011, 4, 139–142. [Google Scholar] [CrossRef]
- Wong, H.M.; Payet, L.; Huppert, J.L. Function and targeting of G-quadruplexes. Curr. Opin. Mol. Ther. 2009, 11, 146–155. [Google Scholar]
- Balagurumoorthy, P.; Brahmachari, S.K.; Mohanty, D.; Bansal, M.; Sassisekharan, V. Hairpin and parallel quartet structures for telomeric sequences. Nucleic Acids Res. 1992, 20, 4061–4067. [Google Scholar] [CrossRef]
- Marotta, S.P.; Tamburri, P.A.; Sheardy, R.D. Sequence and environmental effects on the self-assembly of DNA oligomers possessing GxT2Gy segments. Biochemistry 1996, 35, 10484–10492. [Google Scholar] [CrossRef]
- Dai, T.-Y.; Marotta, S.P.; Sheardy, R.D. Self-assembly of DNA oligomers into high molecular weight species. Biochemistry 1995, 34, 3655–3662. [Google Scholar] [CrossRef]
- Sen, D.; Gilbert, W. Superstructures formed by telomere like oligomers. Biochemistry 1992, 31, 65–70. [Google Scholar] [CrossRef]
- Marsh, T.C.; Henderson, E. G-Wires: Self-assembly of a telomeric oligonucleotide, d(GGGTTGGG), into large superstructures. Biochemistry 1994, 33, 10718–10724. [Google Scholar] [CrossRef]
- Shi, Y.; Luo, Q.; Li, N.B. A highly sensitive resonance Rayleigh scattering method to discriminate a parallel-stranded G-quadruplex from DNA with other topologies and structures. Chem. Commun. 2013, 49, 6209–6211. [Google Scholar] [CrossRef]
- Sample Availability: Samples of the DNA oligomers are available from the authors.
© 2014 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Sharma, V.R.; Sheardy, R.D. The Human Telomere Sequence, (TTAGGG)4, in the Absence and Presence of Cosolutes: A Spectroscopic Investigation. Molecules 2014, 19, 595-608. https://doi.org/10.3390/molecules19010595
Sharma VR, Sheardy RD. The Human Telomere Sequence, (TTAGGG)4, in the Absence and Presence of Cosolutes: A Spectroscopic Investigation. Molecules. 2014; 19(1):595-608. https://doi.org/10.3390/molecules19010595
Chicago/Turabian StyleSharma, Vishal R., and Richard D. Sheardy. 2014. "The Human Telomere Sequence, (TTAGGG)4, in the Absence and Presence of Cosolutes: A Spectroscopic Investigation" Molecules 19, no. 1: 595-608. https://doi.org/10.3390/molecules19010595