Fluorescent Probes for Nucleic Acid Visualization in Fixed and Live Cells
Abstract
:1. Introduction
- high affinity and specificity for the target nucleic acid sequence,
- good cellular penetration and ability to reach its nucleic acid target in cells,
- stability of probes under physiological conditions,
- minimal cytotoxicity,
- minimal distortion of cellular functions,
- simple detection in living cells after excitation with non-harmful visible light,
- modulation of fluorescence spectra upon interaction with a target,
- high signal to background ratio.
2. Probes for Double-Stranded DNA Imaging by Fluorescence
2.1. DNA Imaging in Fixed Cells and Method FISH
2.2. Non-Specific DNA Detection and Staining
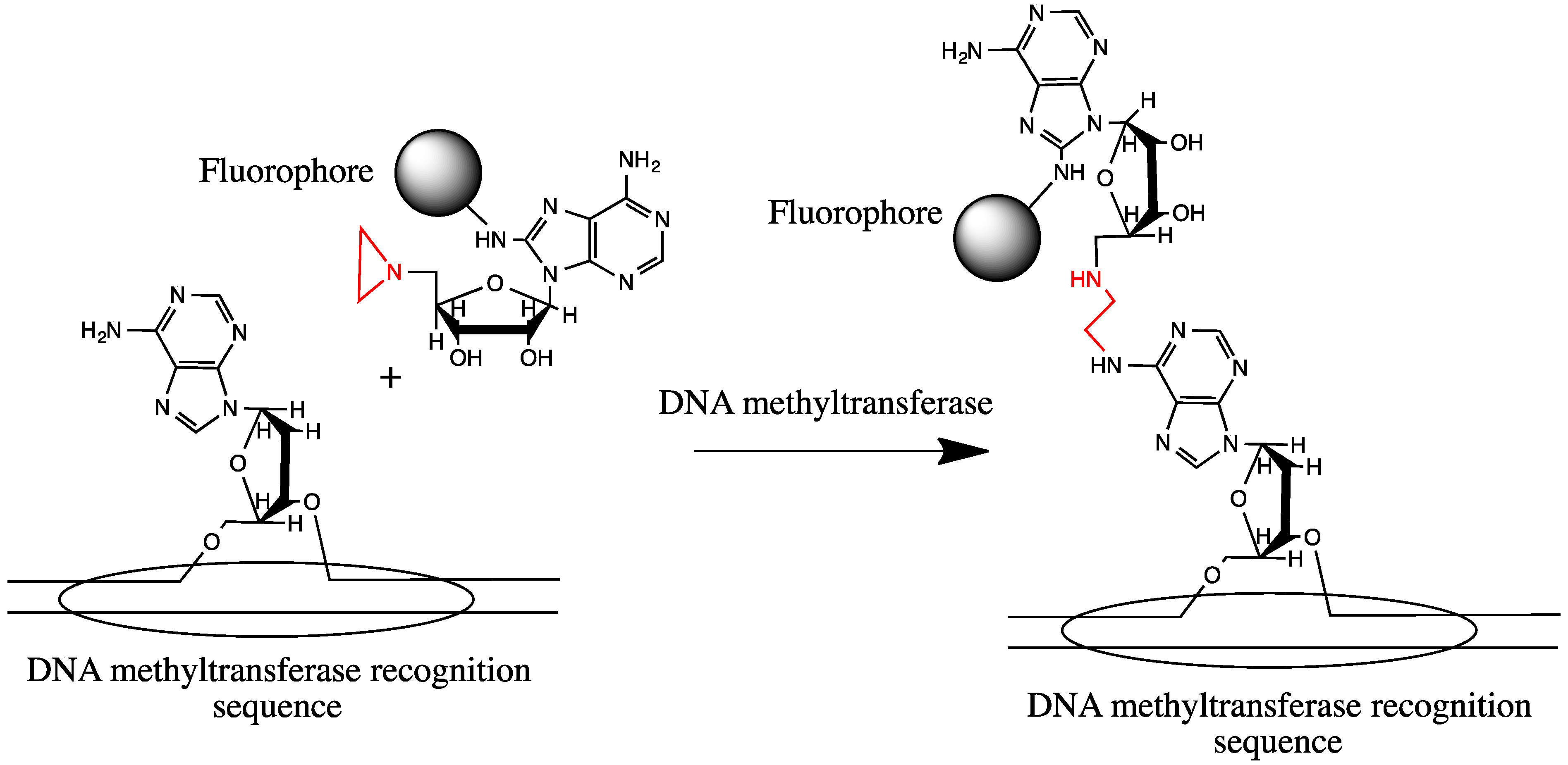
2.3. Sequence-Specific DNA Labeling
2.4. Fluorescent Antibodies for DNA Imaging
2.5. Fused GFP and Other Color Proteins as DNA-Specific Probes
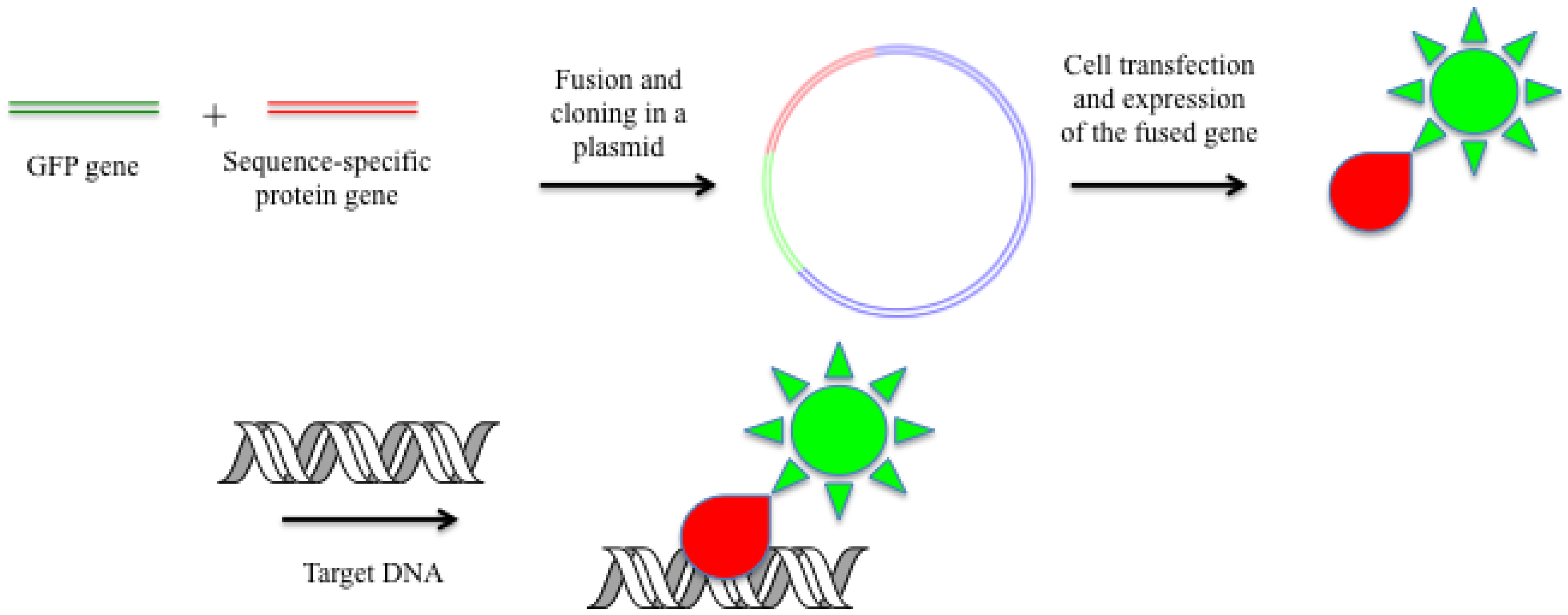
2.6. Hybridization with Oligonucleotide Analogs in Living Cells
2.7. Sequence-Specific Proteins Recognizing Double-Stranded DNA (Zinc Fingers, TALE)
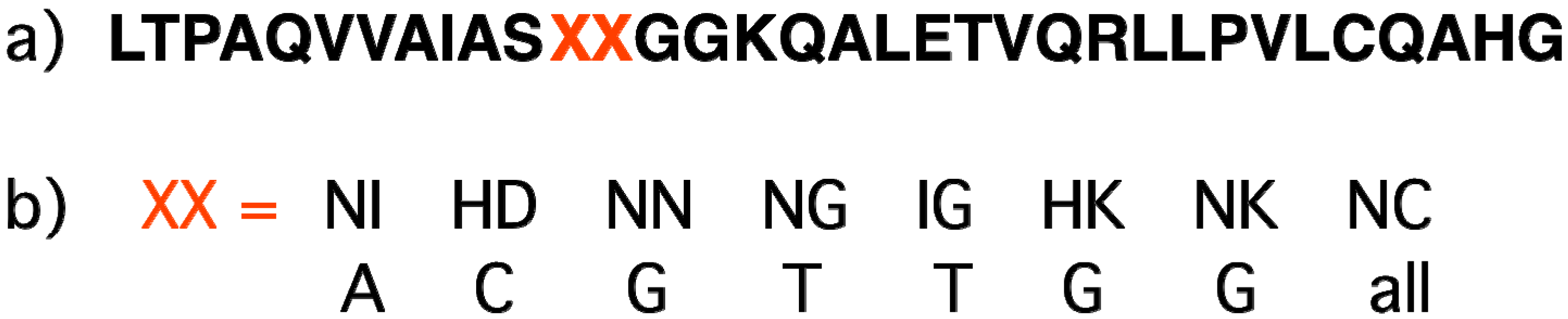
2.8. Triplex-Forming Oligonucleotides

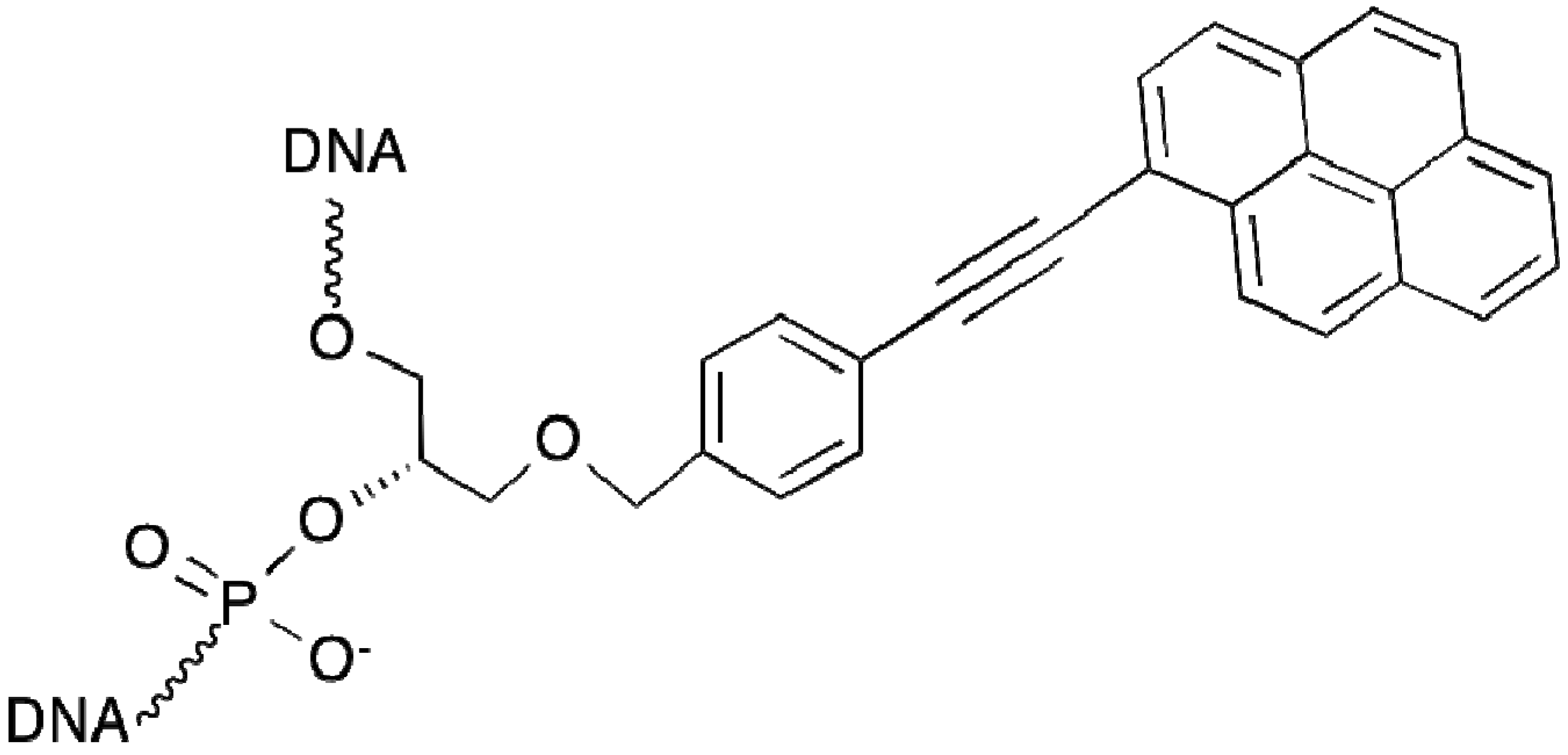
2.9. Polyamide N-methylpyrrole—N-methylimidazole Minor Groove Binders
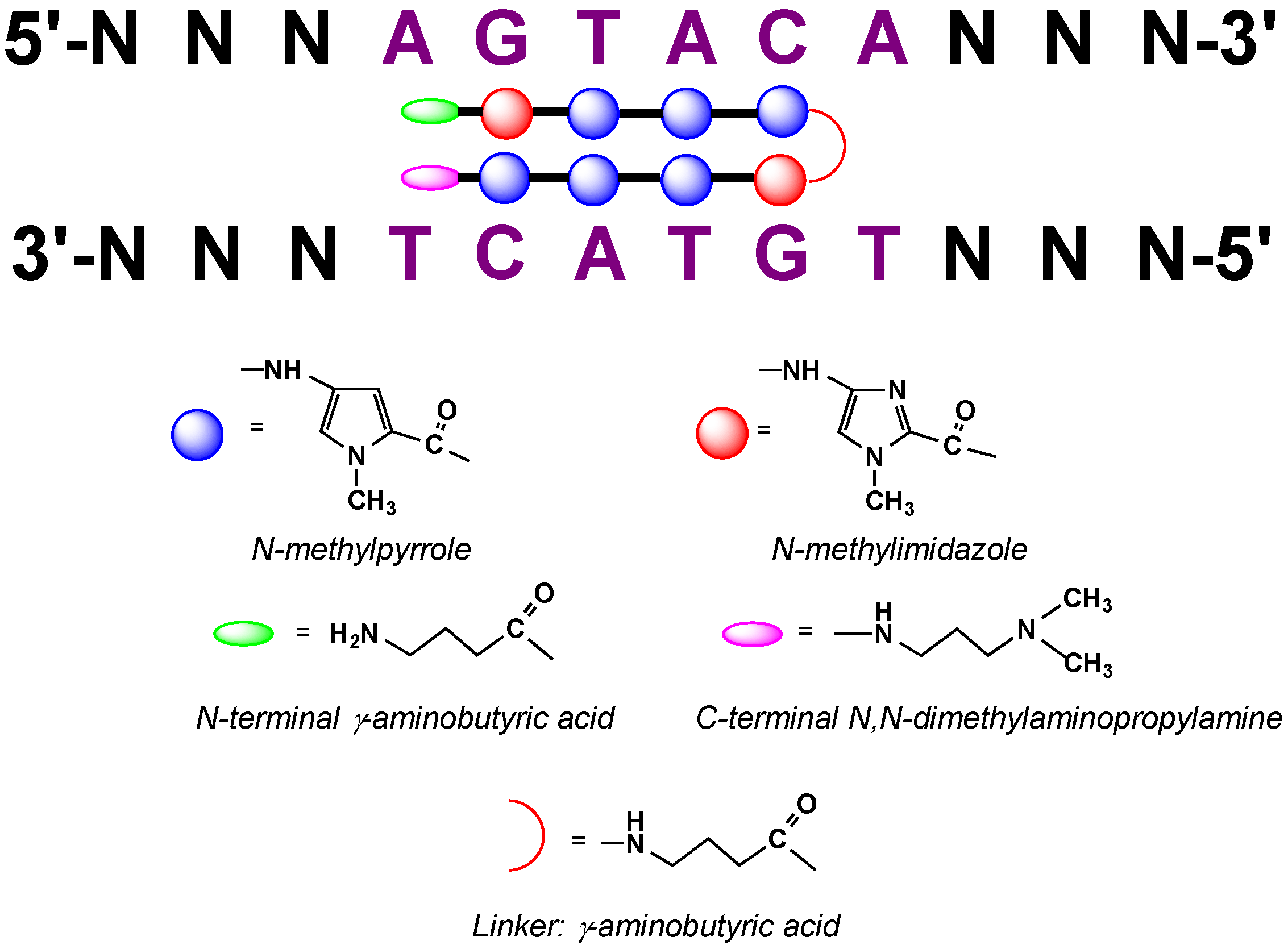
3. Choice of Approaches and Fluorophores for Live Cell Applications


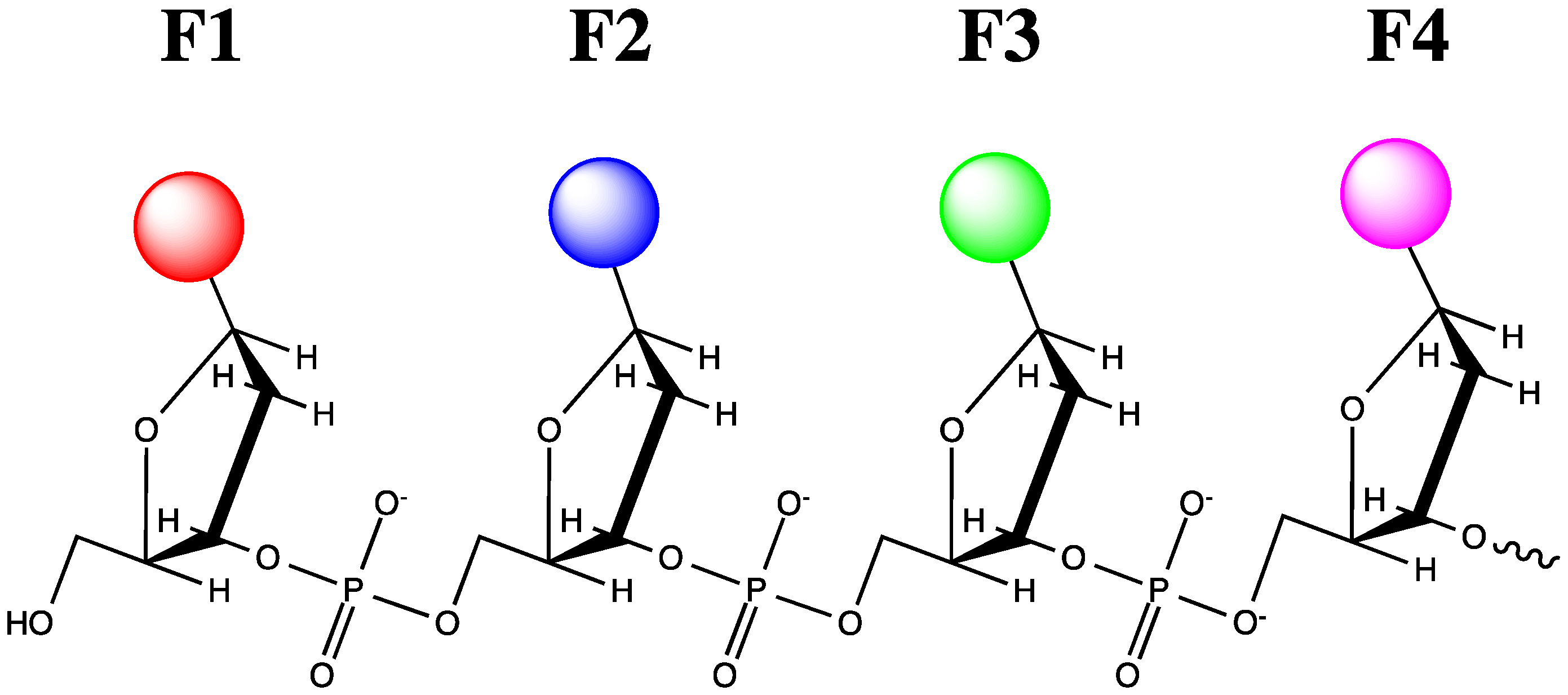
4. Probes for RNA and Single-Stranded DNA Imaging by Fluorescence
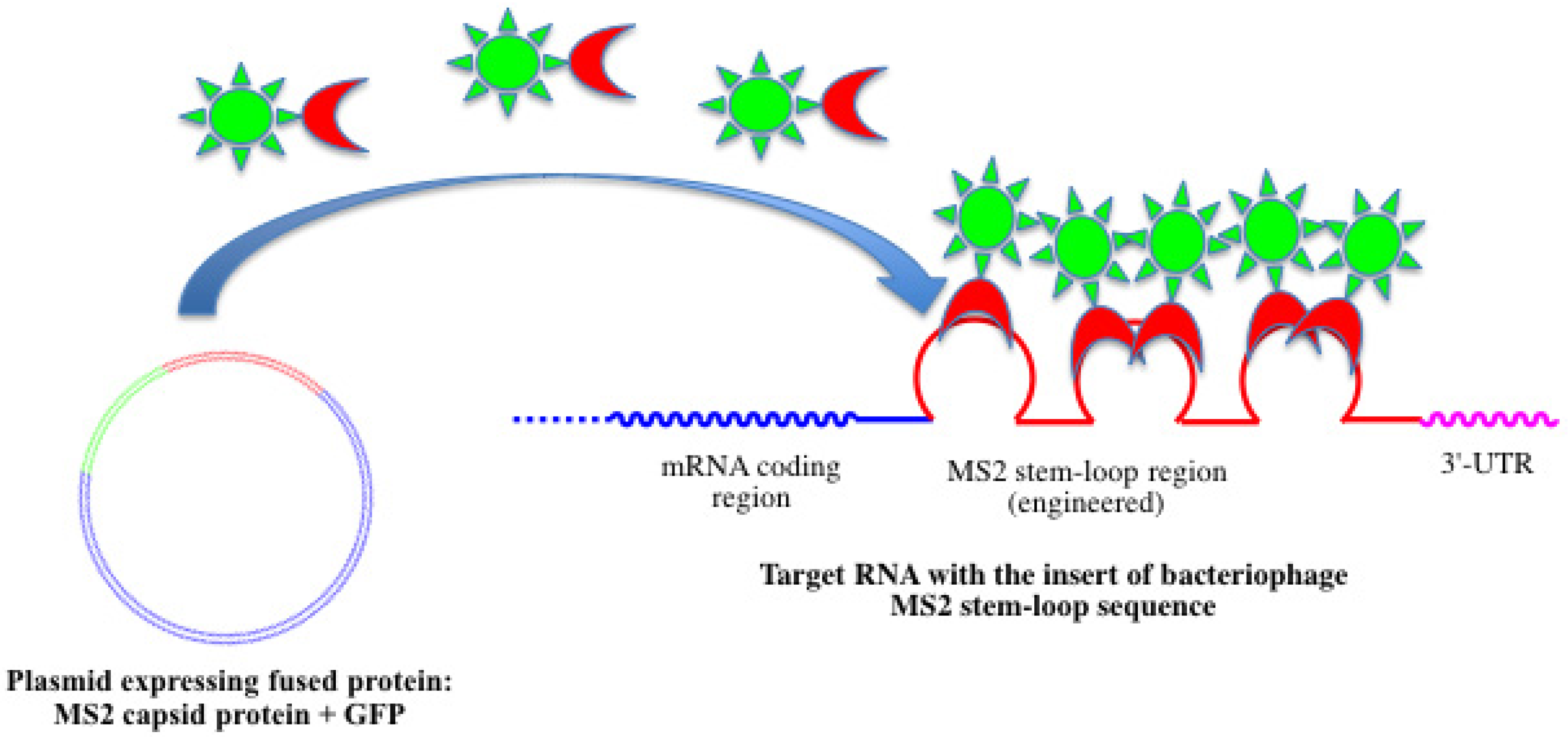
4.1. Fused Fluorescent Proteins
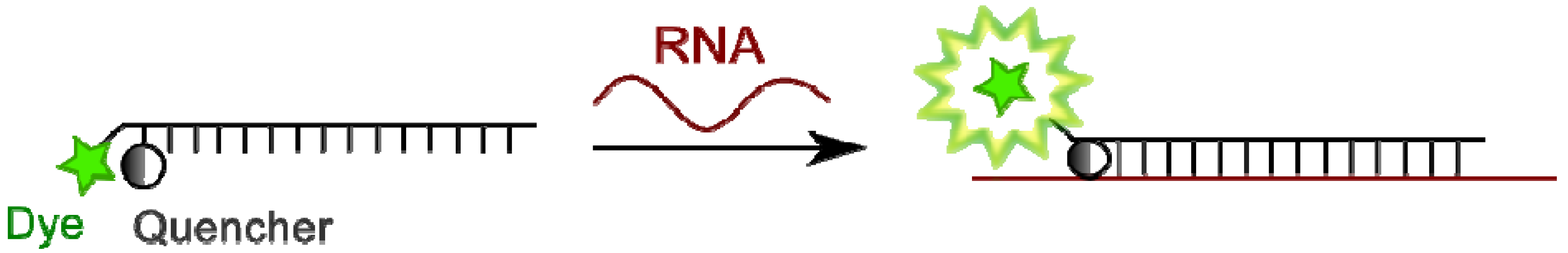
4.2. Linear Fluorescent Oligonucleotide Probes

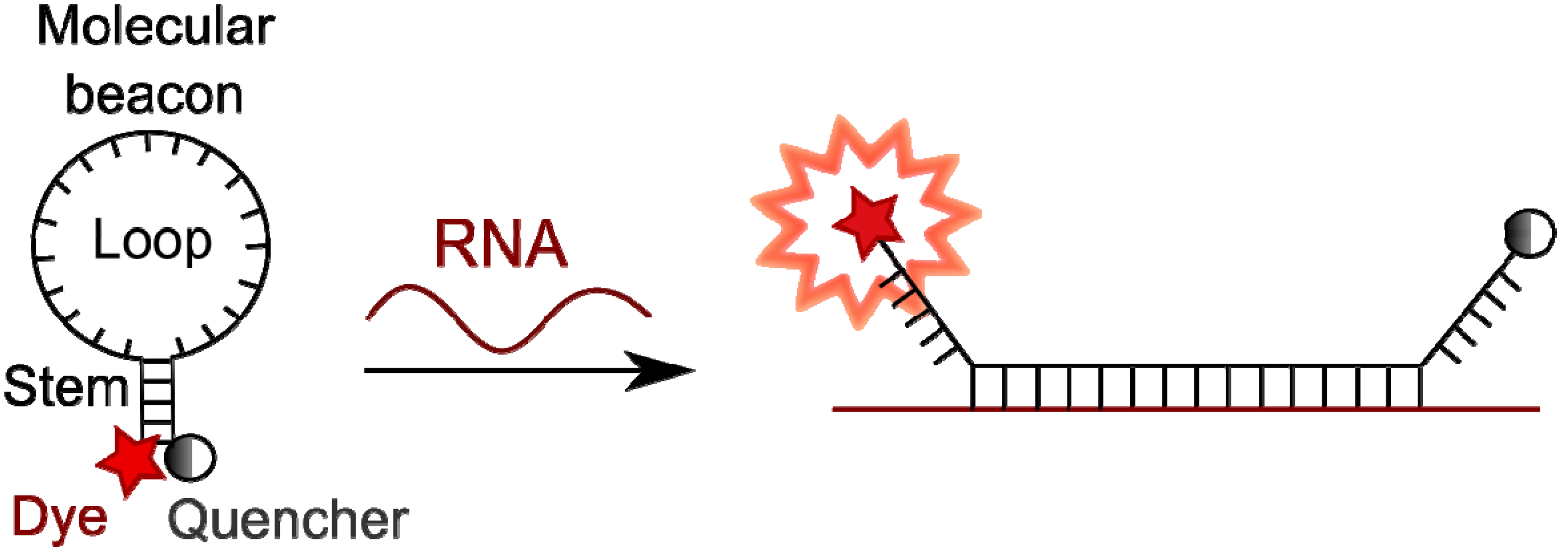
4.3. Molecular Beacons

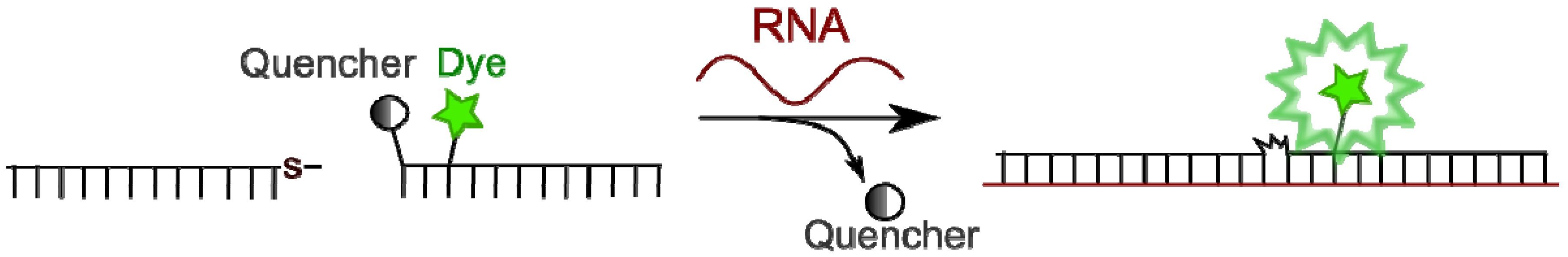
4.4. Binary Probes
4.4.1. Fluorescence Resonance Energy Transfer (FRET) and Excimer Formation
4.4.2. Template-Directed Chemical Reactions with Activation or Formation of a Fluorophore

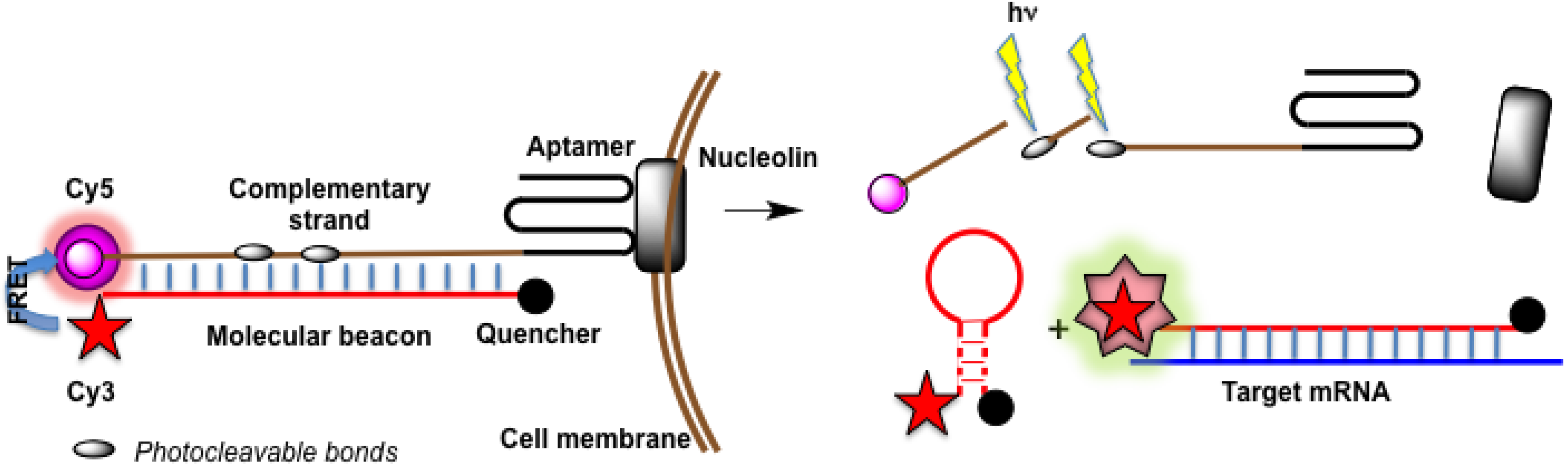
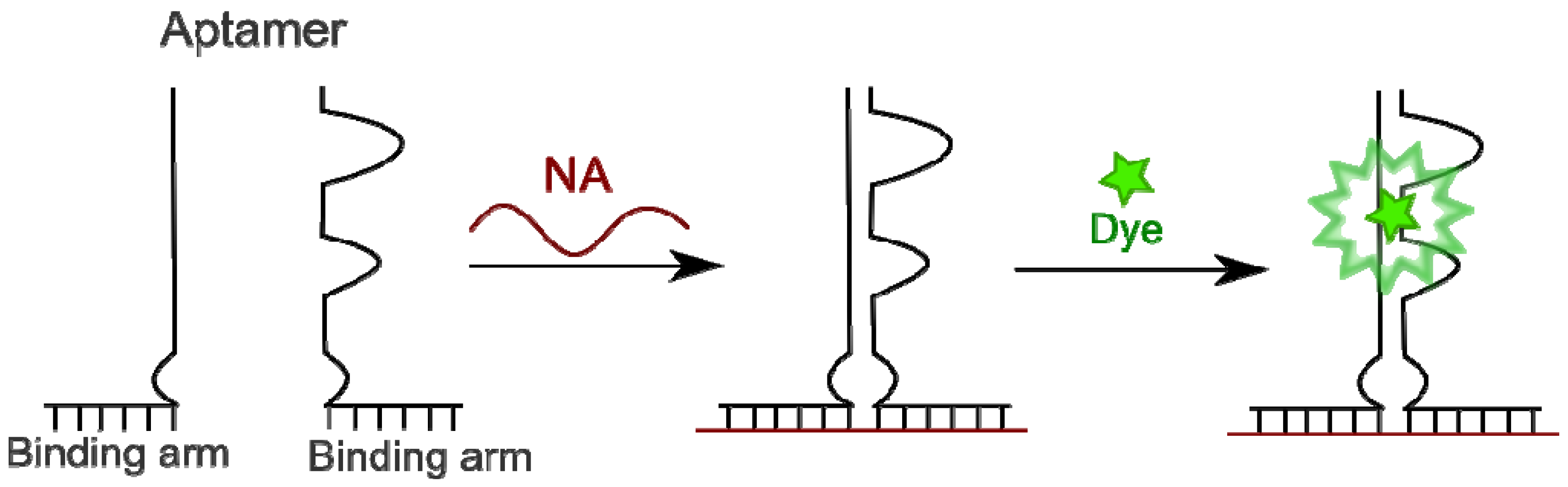
4.4.3. Aptamers as Binary Probes
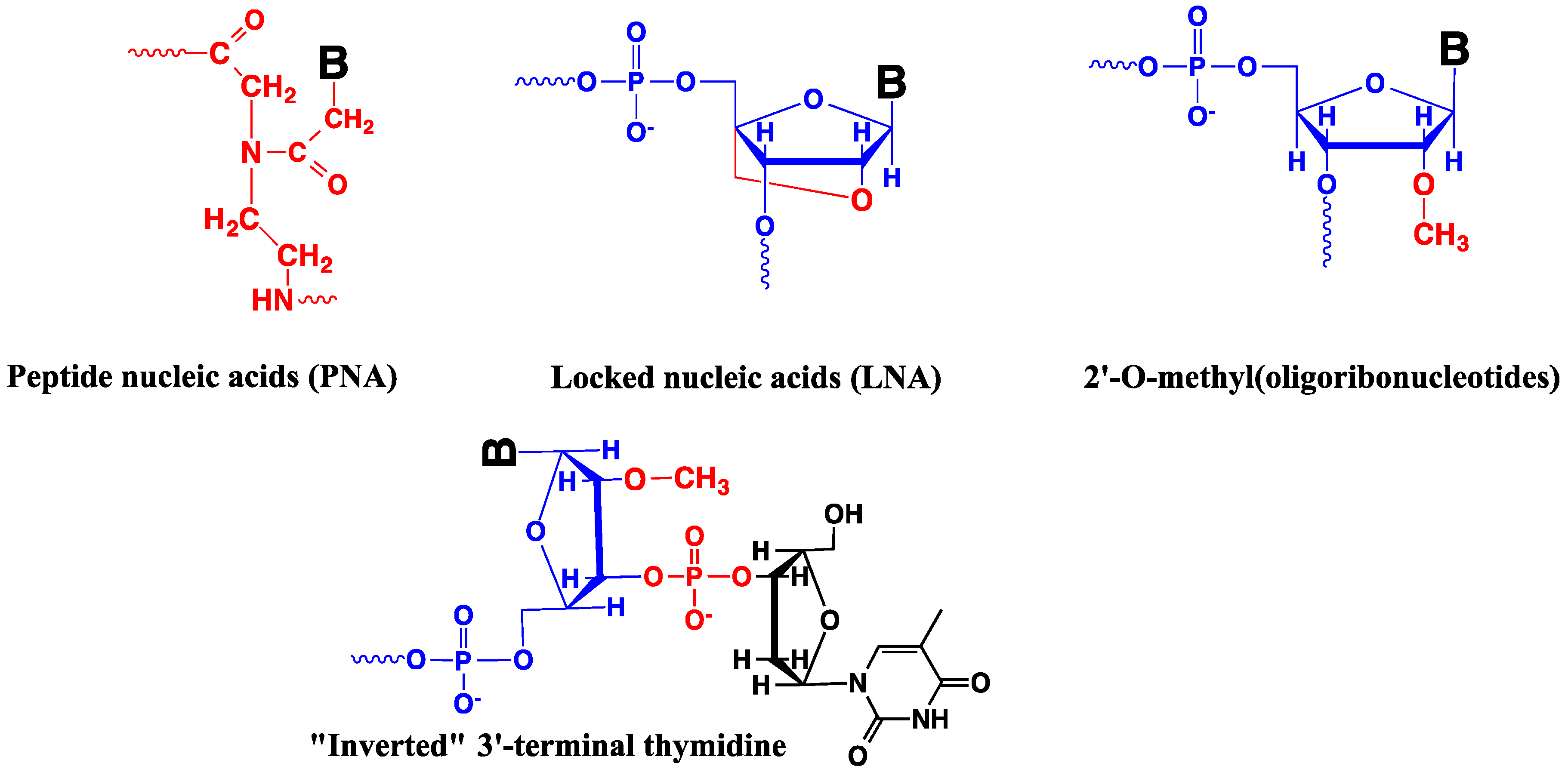
4.5. Modified Oligonucleotides in Design of Nucleic Acid Probes
5. Intracellular Delivery of Oligonucleotides and Their Analogs
6. Conclusions
Acknowledgments
Conflicts of Interest
Abbreviations
| BHQ | black hole quencher |
| BO, Cy3, Cy5, DRAQ5, TAMRA, TOTO | commercial names of fluorophores |
| DAPI | 4',6-diamidino-2-phenylindole |
| ELISA | enzyme-linked immunosorbent assay |
| FRET | fluorescence resonance energy transfer |
| GFP | green fluorescent protein |
| FISH | fluorescence in situ hybridization |
| LNA | locked nucleic acid |
| MGB | minor groove binder |
| PCR | polymerase chain reaction |
| PET | positron emission tomography |
| PNA | peptide nucleic acid |
| QUAL | quenched autoligation probe |
| SSB | single strand DNA-binding protein |
| TALE | transcription activator-like effector |
| TALEN | artificial nuclease based on TALE |
| TFO | triplex-forming oligonucleotide |
| TINA | twisted intercalating nucleic acid |
| TISH | triplex in situ hybridization |
| TO | thiazole orange |
| TR | thiazole red |
References
- Dirks, R.W.; Tanke, H.J. Advances in fluorescent tracking of nucleic acids in living cells. BioTechniques 2006, 40, 489–496. [Google Scholar] [CrossRef]
- Silverman, A.P.; Kool, E.T. Quenched probes for highly specific detection of cellular RNAs. Trends Biotechnol. 2005, 23, 225–230. [Google Scholar] [CrossRef]
- Weigert, R.; Porat-Shliom, N.; Amornphimoltham, P. Imaging cell biology in live animals: Ready for prime time. J. Cell Biol. 2013, 201, 969–979. [Google Scholar] [CrossRef]
- Soon, W.W.; Hariharan, M.; Snyder, M.P. High-throughput sequencing for biology and medicine. Mol. Syst. Biol. 2013, 9, 640. [Google Scholar]
- Bernstein, B.E.; Birney, E.; Dunham, I.; Green, E.D.; Gunter, C.; Snyder, M. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar]
- Klein, K.; Gigler, A.M.; Aschenbrenne, T.; Monetti, R.; Bunk, W.; Jamitzky, F.; Morfill, G.; Stark, R.W.; Schlegel, J. Label-free live-cell imaging with confocal Raman microscopy. Biophys. J. 2012, 102, 360–368. [Google Scholar]
- Palonpon, A.F.; Ando, J.; Yamakoshi, H.; Dodo, K.; Sodeoka, M.; Kawata, S.; Fujita, K. Raman and SERS microscopy for molecular imaging of live cells. Nat. Protoc. 2013, 8, 677–692. [Google Scholar] [CrossRef]
- Lendvai, G.; Estrada, S.; Bergstrom, M. Radiolabelled oligonucleotides for imaging of gene expression with PET. Curr. Med. Chem. 2009, 16, 4445–4461. [Google Scholar] [CrossRef]
- Harki, D.A.; Satyamurthy, N.; Stout, D.B.; Phelps, M.E.; Dervan, P.B. In vivo imaging of pyrrole-imidazole polyamides with positron emission tomography. Proc. Nat. Acad. Sci. USA 2008, 105, 13039–13044. [Google Scholar]
- Fernández-Suárez, M.; Ting, A.Y. Fluorescent probes for super-resolution imaging in living cells. Nat. Rev. Mol. Cell. Biol. 2008, 9, 929–943. [Google Scholar] [CrossRef]
- Tsien, R.Y. The geen fluorescent protein. Annu. Rev. Biochem. 1998, 67, 509–544. [Google Scholar] [CrossRef]
- Heim, R.; Tsien, R.Y. Engineering green fluorescent protein for improved brightness, longer wavelengths and fluorescence resonance energy transfer. Curr. Biol. 1996, 6, 178–182. [Google Scholar] [CrossRef]
- Zimmer, M. Green fluorescent protein (GFP): Applications, structure, and related photophysical behavior. Chem. Rev. 2002, 102, 759–782. [Google Scholar] [CrossRef]
- Lippincott-Schwartz, J.; Patterson, G.H. Development and use of fluorescent protein markers in living cells. Science 2003, 300, 87–91. [Google Scholar]
- Choo, K.H.A. The Centromere; Oxford University Press: Oxford, UK, New York, NY, USA, Tokyo, Japan, 1997; p. 318. [Google Scholar]
- Oulton, R.; Harrington, L. Telomeres, telomerase, and cancer: Life on the edge of genomic stability. Curr. Opin. Oncol. 2000, 12, 74–81. [Google Scholar] [CrossRef]
- Rezler, E.M.; Bearss, D.J.; Hurley, L.H. Telomere inhibition and telomere disruption as processes for drug targeting. Annu. Rev. Pharmacol. Toxicol. 2003, 43, 359–379. [Google Scholar] [CrossRef]
- O'Sullivan, R.J.; Karlseder, J. Telomeres: Protecting chromosomes against genome instability. Nat. Rev. Mol. Cell. Biol. 2010, 11, 171–181. [Google Scholar]
- Nath, J.; Johnson, K.L. A review of Fluorescence in Situ Hybridization (FISH): Current status and future prospects. Biotech. Histochem. 2000, 75, 54–78. [Google Scholar] [CrossRef]
- Levsky, J.M.; Singer, R.H. Fluorescence in Situ hybridization: Past, present and future. J. Cell Sci. 2003, 116, 2833–2838. [Google Scholar] [CrossRef]
- Smolina, I.; Lee, C.; Frank-Kamenetskii, M. Detection of low-copy-number genomic DNA sequences in individual bacterial cells by using peptide nucleic acid-assisted rolling-circle amplification and fluorescence in situ hybridization. Appl. Environ. Microbiol. 2007, 73, 2324–2328. [Google Scholar] [CrossRef]
- Jaco, I.; Canela, A.; Vera, E.; Blasco, M.A. Centromere mitotic recombination in mammalian cells. J. Cell Biol. 2008, 181, 885–892. [Google Scholar] [CrossRef]
- Paulasova, P.; Pellestor, F. The peptide nucleic acids (PNAs): A new generation of probes for genetic and cytogenetic analyses. Ann. Genet. Paris 2004, 47, 349–358. [Google Scholar] [CrossRef]
- Pellestor, F.; Paulasova, P. The peptide nucleic acids, efficient tools for molecular diagnosis (Review). Int. J. Mol. Med. 2004, 13, 521–525. [Google Scholar]
- Pellestor, F.; Paulasova, P. The peptide nucleic acids (PNAs), powerful tools for molecular genetics and cytogenetics. Eur. J. Hum. Genet. 2004, 12, 694–700. [Google Scholar] [CrossRef]
- Astakhova, I.V.; Ustinov, A.V.; Korshun, V.A.; Wengel, J. LNA for optimization of fluorescent oligonucleotide probes: Improved spectral properties and target binding. Bioconjug. Chem. 2011, 22, 533–539. [Google Scholar] [CrossRef]
- Silahtaroglu, A.; Pfundheller, H.; Koshkin, A.; Tommerup, N.; Kauppinen, S. LNA-modified oligonucleotides are highly efficient as FISH probes. Cytogenet. Genome Res. 2004, 107, 32–37. [Google Scholar] [CrossRef]
- Silahtaroglu, A.N.; Tommerup, N.; Vissing, H. FISHing with locked nucleic acids (LNA): Evaluation of different LNA/DNA mixmers. Mol. Cell Probes 2003, 17, 165–169. [Google Scholar] [CrossRef]
- Thomsen, R.; Nielsen, P.S.; Jensen, T.H. Dramatically improved RNA in situ hybridization signals using LNA-modified probes. RNA 2005, 11, 1745–1748. [Google Scholar] [CrossRef]
- Bridger, J.M.; Volpi, E.V.; Schmitt, E.; Schwarz-Finsterle, J.; Stein, S.; Boxler, C.; Müller, P.; Mokhir, A.; Krämer, R.; Cremer, C.; et al. Combinatorial Oligo FISH: Directed Labeling of Specific Genome Domains in Differentially Fixed Cell Material and Live Cells. In Methods in Molecular Biology: Fluorescence in situ Hybridization (FISH); Humana Press: New York, NY, USA, 2010; Volume 659, pp. 185–202. [Google Scholar]
- Hausmann, M.; Winkler, R.; Hildenbrand, G.; Finsterle, J.; Weisel, A.; Rapp, A.; Schmitt, E.; Janz, S.; Cremer, C. COMBO-FISH: Specific labeling of nondenatured chromatin targets by computer-selected DNA oligonucleotide probe combinations. BioTechniques 2003, 35, 564–577. [Google Scholar]
- Schwarz-Finsterle, J.; Stein, S.; Grossmann, C.; Schmitt, E.; Trakhtenbrot, L.; Rechavi, G.; Amariglio, N.; Cremer, C.; Hausmann, M. Comparison of triple helical COMBO-FISH and standard FISH by means of quantitative microscopic image analysis of ABL/BCR positions in cell nuclei. J. Biochem. Biophys. Methods 2007, 70, 397–406. [Google Scholar] [CrossRef]
- Stevens, N.; O’Connor, N.; Vishwasrao, H.; Samaroo, D.; Kandel, E.R.; Akins, D.L.; Drain, C.M.; Turro, N.J. Two Color RNA intercalating probe for cell imaging applications. J. Am. Chem. Soc. 2008, 130, 7182–7183. [Google Scholar]
- Martin, R.M.; Leonhardt, H.; Cardoso, M.C. DNA labeling in living cells. Cytometry Part A 2005, 67A, 45–52. [Google Scholar] [CrossRef]
- Kapuscinski, J. DAPI: A DNA-specific fluorescent probe. Biotech. Histochem. 1995, 70, 220–233. [Google Scholar] [CrossRef]
- Pljevaljčić, G.; Pignot, M.; Weinhold, E. Design of a new fluorescent cofactor for DNA methyltransferases and sequence-specific labeling of DNA. J. Am. Chem. Soc. 2003, 125, 3486–3492. [Google Scholar] [CrossRef]
- Pljevaljcic, G.; Schmidt, F.; Peschlow, A.; Weinhold, E. Sequence-specific DNA labeling using methyltransferases. Methods Mol. Biol. 2004, 283, 145–161. [Google Scholar]
- Pljevaljčić, G.; Schmidt, F.; Scheidig, A.J.; Lurz, R.; Weinhold, E. Quantitative labeling of long plasmid DNA with nanometer precision. ChemBioChem 2007, 8, 1516–1519. [Google Scholar] [CrossRef]
- Maya-Mendoza, A.; Olivares-Chauvet, P.; Kohlmeier, F.; Jackson, D.A. Visualising chromosomal replication sites and replicons in mammalian cells. Methods 2012, 57, 140–148. [Google Scholar] [CrossRef]
- Brickner, D.G.; Light, W.; Brickner, J.H. Chapter 22—Quantitative Localization of Chromosomal Loci by Immunofluorescence. In Methods in Enzymology; Jonathan, W., Christine, G., Gerald, R.F., Eds.; Academic Press: New York, NY, USA, 2010; Volume 470, pp. 569–580. [Google Scholar]
- Biffi, G.; Tannahill, D.; McCafferty, J.; Balasubramanian, S. Quantitative visualization of DNA G-quadruplex structures in human cells. Nat. Chem. 2013, 5, 182–186. [Google Scholar] [CrossRef]
- Sugimoto, K.; Senda-Murata, K.; Oka, S. Construction of three quadruple-fluorescent MDA435 cell lines that enable monitoring of the whole chromosome segregation process in the living state. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2008, 657, 56–62. [Google Scholar] [CrossRef]
- Wang, X.; Reyes-Lamothe, R.; Sherratt, D.J. Visualizing genetic loci and molecular machines in living bacteria (Biochemical Society Linked Focused Meetings). Biochem. Soc. Trans. 2008, 36, 749–753. [Google Scholar] [CrossRef]
- De Vos, W.H.; Hoebe, R.A.; Joss, G.H.; Haffmans, W.; Baatout, S.; van Oostveldt, P.; Manders, E.M.M. Controlled light exposure microscopy reveals dynamic telomere microterritories throughout the cell cycle. Cytom. Part A 2009, 75A, 428–439. [Google Scholar] [CrossRef]
- De Vos, W.H.; Joss, G.H.; Haffmans, W.; Hoebe, R.A.; Manders, E.M.M.; van Oostveldt, P. Four-dimensional telomere analysis in recordings of living human cells acquired with Controlled Light Exposure Microscopy. J. Microsc. 2010, 238, 254–264. [Google Scholar]
- Molenaar, C.; Wiesmeijer, K.; Verwoerd, N.P.; Khazen, S.; Eils, R.; Tanke, H.J.; Dirks, R.W. Visualizing telomere dynamics in living mammalian cells using PNA probes. EMBO J. 2003, 22, 6631–6641. [Google Scholar] [CrossRef]
- Flierl, A.; Jackson, C.; Cottrell, B.; Murdock, D.; Seibel, P.; Wallace, D.C. Targeted delivery of DNA to the mitochondrial compartment via import sequence-conjugated peptide nucleic acid. Mol. Ther. 2003, 7, 550–557. [Google Scholar] [CrossRef]
- Garvie, C.W.; Wolberger, C. Recognition of specific DNA sequences. Mol. Cell 2001, 8, 937–946. [Google Scholar] [CrossRef]
- Tachikawa, K.; Briggs, S.P. Targeting the human genome. Curr. Opin. Biotechnol. 2006, 17, 659–665. [Google Scholar] [CrossRef]
- Ghosh, I.; Stains, C.I.; Ooi, A.T.; Segal, D.J. Direct detection of double-stranded DNA: Molecular methods and applications for DNA diagnostics. Mol. Biosyst. 2006, 2, 551–560. [Google Scholar] [CrossRef]
- Rodriguez-Martinez, J.A.; Peterson-Kaufman, K.J.; Ansari, A.Z. Small-molecule regulators that mimic transcription factors. Biochim. Biophys. Acta-Gene Regul. Mech. 2010, 1799, 768–774. [Google Scholar] [CrossRef]
- Klug, A. The discovery of zinc fingers and their applications in gene regulation and genome manipulation. Annu. Rev. Biochem. 2010, 79, 213–231. [Google Scholar] [CrossRef]
- Urnov, F.D.; Rebar, E.J.; Holmes, M.C.; Zhang, H.S.; Gregory, P.D. Genome editing with engineered zinc finger nucleases. Nat. Rev. Genet. 2010, 11, 636–646. [Google Scholar] [CrossRef]
- Lindhout, B.I.; Fransz, P.; Tessadori, F.; Meckel, T.; Hooykaas, P.J.J.; van der Zaal, B.J. Live cells imaging of repetitive DNA sequences via GFP-tagged polydactyl zinc finger proteins. Nucleic Acids Res. 2007, 35, e107. [Google Scholar] [CrossRef]
- Boch, J.; Scholze, H.; Schornack, S.; Landgraf, A.; Hahn, S.; Kay, S.; Lahaye, T.; Nickstadt, A.; Bonas, U. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 2009, 326, 1509–1512. [Google Scholar] [CrossRef]
- Mak, A.N.-S.; Bradley, P.; Cernadas, R.A.; Bogdanove, A.J.; Stoddard, B.L. The crystal structure of TAL effector PthXo1 bound to its DNA target. Science 2012, 335, 716–719. [Google Scholar] [CrossRef]
- Christian, M.; Cermak, T.; Doyle, E.L.; Schmidt, C.; Zhang, F.; Hummel, A.; Bogdanove, A.J.; Voytas, D.F. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 2010, 186, 757–761. [Google Scholar] [CrossRef]
- Miller, J.C.; Tan, S.; Qiao, G.; Barlow, K.A.; Wang, J.; Xia, D.F.; Meng, X.; Paschon, D.E.; Leung, E.; Hinkley, S.J.; et al. A TALE nuclease architecture for efficient genome editing. Nat. Biotech. 2011, 29, 143–148. [Google Scholar] [CrossRef]
- Bedell, V.M.; Wang, Y.; Campbell, J.M.; Poshusta, T.L.; Starker, C.G.; Krug Ii, R.G.; Tan, W.; Penheiter, S.G.; Ma, A.C.; Leung, A.Y.H.; et al. In vivo genome editing using a high-efficiency TALEN system. Nature 2012, 491, 114–118. [Google Scholar] [CrossRef]
- Praseuth, D.; Guieysse, A.L.; Hélène, C. Triple helix formation and the antigene strategy for sequence-specific control of gene expression. Biochim. Biophys. Acta Gene Struct. Express. 1999, 1489, 181–206. [Google Scholar]
- Faria, M.; Giovannangeli, C. Triplex-forming molecules: From concepts to applications. J. Gene Med. 2001, 3, 299–310. [Google Scholar] [CrossRef]
- Dervan, P.B.; Bürli, R.W. Sequence-specific DNA recognition by polyamides. Curr. Opin. Chem. Biol. 1999, 3, 688–693. [Google Scholar] [CrossRef]
- Dervan, P.B. Molecular recognition of DNA by small molecules. Bioorgan. Med. Chem. 2001, 9, 2215–2235. [Google Scholar] [CrossRef]
- Sun, J.S.; Hélène, C. Oligonucleotide-directed triple-helix formation. Curr. Opin. Struct. Biol. 1993, 3, 345–356. [Google Scholar] [CrossRef]
- Fox, K.R. Targeting DNA with triplexes. Curr. Med. Chem. 2000, 7, 17–37. [Google Scholar] [CrossRef]
- Casey, B.P.; Glazer, P.M. Gene Targeting via Triple-Helix Formation. In Progress in Nucleic Acid Research and Molecular Biology; Moldave, K., Ed.; Academic Press Inc: San Diego, CA, USA, 2001; Volume 67, pp. 163–192. [Google Scholar]
- Knauert, M.P.; Glazer, P.M. Triplex forming oligonucleotides: Sequence-specific tools for gene targeting. Hum. Mol. Genet. 2001, 10, 2243–2251. [Google Scholar] [CrossRef]
- Vasquez, K.M.; Glazer, P.M. Triplex-forming oligonucleotides: Principles and applications. Quart. Rev. Biophys. 2002, 35, 89–107. [Google Scholar]
- Duca, M.; Vekhoff, P.; Oussedik, K.; Halby, L.; Arimondo, P.B. The triple helix: 50 years later, the outcome. Nucleic Acids Res. 2008, 36, 5123–5138. [Google Scholar]
- Mergny, J.L.; Boutorine, A.S.; Garestier, T.; Belloc, F.; Rougée, M.; Bulychev, N.V.; Koshkin, A.A.; Bourson, J.; Lebedev, A.V.; Valeur, B.; et al. Fliorescence energy transfer as a probe for nucleic acid structure and sequence. Nucleic Acids Res. 1994, 22, 920–928. [Google Scholar] [CrossRef]
- Grimm, G.N.; Boutorine, A.S.; Lincoln, P.; Nordén, B.; Hélène, C. Formation of DNA triple helices by an oligonucleotide conjugated to a fluorescent ruthenium complex. ChemBioChem 2002, 3, 324–331. [Google Scholar] [CrossRef]
- Filichev, V.V.; Pedersen, E.B. Stable and selective formation of Hoogsteen-type triplexes and duplexes using twisted intercalating nucleic acids (TINA) prepared via postsynthetic sonogashira solid-phase coupling reactions. J. Am. Chem. Soc. 2005, 127, 14849–14858. [Google Scholar] [CrossRef]
- Filichev, V.V.; Nielsen, M.C.; Bomholt, N.; Jessen, C.H.; Pedersen, E.B. High thermal stability of 5'-5'-linked alternate Hoogsteen triplexes at physiological pH. Angew. Chem. Int. Ed. 2006, 45, 5311–5315. [Google Scholar] [CrossRef]
- Géci, I.; Filichev, V.; Pedersen, E. Stabilization of parallel triplexes by Twisted Intercalating Nucleic Acids (TINAs) Incorporating 1,2,3-triazole units and prepared by microwave-accelerated click chemistry. Chem. Eur. J. 2007, 13, 6379–6386. [Google Scholar] [CrossRef]
- Doluca, O.; Boutorine, A.S.; Filichev, V.V. Triplex-forming Twisted Intercalating Nucleic Acids (TINAs): Design rules, stabilization of antiparallel DNA triplexes and inhibition of G-quartet-dependent self-association. ChemBioChem 2011, 12, 2365–2374. [Google Scholar] [CrossRef]
- Van Daele, I.; Bomholt, N.; Filichev, V.V.; van Calenbergh, S.; Pedersen, E.B. Triplex formation by pyrene-labelled probes for nucleic acid detection in fluorescence assays. ChemBioChem 2008, 9, 791–801. [Google Scholar] [CrossRef]
- Renard, B.L.; Lartia, R.; Asseline, U. Targeting DNA with “light-up” pyrimidine triple-helical forming oligonucleotides conjugated to stabilizing fluorophores (LU-TFOs). Org. Biomol. Chem. 2008, 6, 4413–4425. [Google Scholar] [CrossRef]
- Johnson, M.D., III; Fresco, J.R. Third-strand in situ hybridization (TISH) to non-denatured metaphase spreads and interphase nuclei. Chromosoma 1999, 108, 181–189. [Google Scholar] [CrossRef]
- Dervan, P.B.; Edelson, B.S. Recognition of the DNA minor groove by pyrrole-imidazole polyamides. Curr. Opin. Struct. Biol. 2003, 13, 284–299. [Google Scholar] [CrossRef]
- Heckel, A.; Dervan, P.B. U-Pin polyamide motif for recognition of the DNA minor groove. Chem. Eur. J. 2003, 9, 3353–3366. [Google Scholar] [CrossRef]
- Farkas, M.E.; Tsai, S.M.; Dervan, P.B. α-Diaminobutyric acid-linked hairpin polyamides. Bioorg. Med. Chem. 2007, 15, 6927–6936. [Google Scholar]
- Meier, J.L.; Montgomery, D.C.; Dervan, P.B. Enhancing the cellular uptake of Py-Im polyamides through next-generation aryl turns. Nucleic Acids Res. 2012, 40, 2345–2356. [Google Scholar] [CrossRef]
- Marques, M.A.; Doss, R.M.; Foister, S.; Dervan, P.B. Expanding the repertoire of heterocycle ring pairs for programmable minor groove DNA recognition. J. Am. Chem. Soc. 2004, 126, 10339–10349. [Google Scholar] [CrossRef]
- Krutzik, P.O.; Chamberlin, A.R. Rapid solid-phase synthesis of DNA-binding pyrrole-imidazole polyamides. Bioorg. Med. Chem. Lett. 2002, 12, 2129–2132. [Google Scholar] [CrossRef]
- Krutzik, P.O.; Chamberlin, A.R. Synthesis of DNA-binding polyamides. Robust solid-phase methods for coupling heterocyclic aromatic amino acids. Methods Mol. Biol. 2002, 201, 77–92. [Google Scholar]
- Chenoweth, D.M.; Harki, D.A.; Dervan, P.B. Solution-phase synthesis of pyrrole-imidazole polyamides. J. Am. Chem. Soc. 2009, 131, 7175–7181. [Google Scholar] [CrossRef]
- Burnett, R.; Melander, C.; Puckett, J.W.; Son, L.S.; Wells, R.B.; Dervan, P.B.; Gottesfeld, J.M. DNA sequence-specific polyamides alleviate transcription inhibition associated with long GAA center dot TTC repeats in Friedreich’s ataxia. Proc. Nat. Acad. Sci. USA 2006, 103, 11497–11502. [Google Scholar]
- Dudouet, B.; Burnett, R.; Dickinson, L.A.; Wood, M.R.; Melander, C.; Belitsky, J.M.; Edelson, B.; Wurtz, N.; Briehn, C.; Dervan, P.B.; et al. Accessibility of nuclear chromatin by DNA binding polyamides. Chem. Biol. 2003, 10, 859–867. [Google Scholar] [CrossRef]
- Nickols, N.G.; Dervan, P.B. Suppression of androgen receptor-mediated gene expression by a sequence-specific DNA-binding polyamide. Proc. Nat. Acad. Sci. USA 2007, 104, 10418–10423. [Google Scholar] [CrossRef]
- Nickols, N.G.; Jacobs, C.S.; Farkas, M.E.; Dervan, P.B. Improved nuclear localization of DNA-binding polyamides. Nucleic Acids Res. 2007, 35, 363–370. [Google Scholar]
- Dervan, P.B.; Doss, R.M.; Marques, M.A. Programmable DNA binding oligomers for control of transcription. Curr. Med. Chem. Anti-Cancer Agents 2005, 5, 373–387. [Google Scholar] [CrossRef]
- Szewczyk, J.W.; Baird, E.E.; Dervan, P.B. Sequence-specific recognition of DNA by a major and minor groove binding ligands. Angew. Chem. Int. Ed. 1996, 35, 1487–1489. [Google Scholar] [CrossRef]
- Szewczyk, J.W.; Baird, E.E.; Dervan, P.B. Cooperative triple-helix formation via a minor groove dimerization domain. J. Am. Chem. Soc. 1996, 118, 6778–6779. [Google Scholar] [CrossRef]
- Trauger, J.W.; Baird, E.E.; Dervan, P.B. Recognition of 16 base pairs in the minor groove of DNA by a pyrrole-imidazole polyamide dimer. J. Am. Chem. Soc. 1998, 120, 3534–3535. [Google Scholar] [CrossRef]
- Herman, D.M.; Baird, E.E.; Dervan, P.B. Tandem hairpin motif for recognition in the minor groove of DNA by pyrrole - imidazole polyamides. Chem. Eur. J. 1999, 5, 975–983. [Google Scholar] [CrossRef]
- Kers, I.; Dervan, P.B. Search for the optimal linker in tandem hairpin polyamides. Bioorg. Med. Chem. 2002, 10, 3339–3349. [Google Scholar] [CrossRef]
- Halby, L.; Ryabinin, V.A.; Sinyakov, A.N.; Boutorine, A.S. Functionalized head-to-head hairpin polyamides: Synthesis, double-stranded DNA-binding activity and affinity. Bioorg. Med. Chem. Lett. 2005, 15, 3720–3724. [Google Scholar] [CrossRef]
- Bhattacharya, S.; Thomas, M. DNA binding properties of novel dansylated distamycin analogues in which the fluorophore is directly conjugated to the N-methylpyrrole carboxamide backbone. J. Biomol. Struct. Dyn. 2002, 19, 935–945. [Google Scholar] [CrossRef]
- Rucker, V.C.; Foister, S.; Melander, C.; Dervan, P.B. Sequence-specific fluorescence detection of double strand DNA. J. Am. Chem. Soc. 2003, 125, 1195–1202. [Google Scholar]
- Fechter, E.J.; Olenyuk, B.; Dervan, P.B. Sequence-specific fluorescence detection of DNA by polyamide-thiazole orange conjugates. J. Am. Chem. Soc. 2005, 127, 16685–16691. [Google Scholar] [CrossRef]
- Bando, T.; Fujimoto, J.; Minoshima, M.; Shinohara, K.I.; Sasaki, S.; Kashiwazaki, G.; Mizumura, M.; Sugiyama, H. Detection of CAG repeat DNA sequences by pyrene-functionalized pyrrole-imidazole polyamides. Bioorg. Med. Chem. 2007, 15, 6937–6942. [Google Scholar] [CrossRef]
- Chenoweth, D.M.; Viger, A.; Dervan, P.B. Fluorescent sequence-specific dsDNA binding oligomers. J. Am. Chem. Soc. 2007, 129, 2216–2217. [Google Scholar] [CrossRef]
- Fujimoto, J.; Bando, T.; Minoshima, M.; Kashiwazaki, G.; Nishijima, S.; Shinohara, K.; Sugiyama, H. Perylene-conjugated pyrrole polyamide as a sequence-specific fluorescent probe. Bioorg. Med. Chem. 2008, 16, 9741–9744. [Google Scholar] [CrossRef]
- Hsu, C.F.; Dervan, P.B. Quantitating the concentration of Py-Im polyamide-fluorescein conjugates in live cells. Bioorg. Med. Chem. Lett. 2008, 18, 5851–5855. [Google Scholar] [CrossRef]
- Nishijima, S.; Shinohara, K.; Bando, T.; Minoshima, M.; Kashiwazaki, G.; Sugiyama, H. Cell permeability of Py-Im-polyamide-fluorescein conjugates: Influence of molecular size and Py/Im content. Bioorg. Med. Chem. 2010, 18, 978–983. [Google Scholar] [CrossRef]
- Dupureur, C.M.; Bashkin, J.K.; Aston, K.; Koeller, K.J.; Gaston, K.R.; He, G. Fluorescence assay of polyamide-DNA interactions. Anal. Biochem. 2012, 423, 178–183. [Google Scholar] [CrossRef]
- Maeshima, K.; Janssen, S.; Laemmli, U.K. Specific targeting of insect and vertebrate telomeres with pyrrole and imidazole polyamides. EMBO J. 2001, 20, 3218–3228. [Google Scholar] [CrossRef]
- Vaijayanthi, T.; Bando, T.; Pandian, G.N.; Sugiyama, H. Progress and prospects of pyrrole-imidazole polyamide–fluorophore conjugates as sequence-selective DNA probes. ChemBioChem 2012, 13, 2170–2185. [Google Scholar] [CrossRef]
- Dickinson, L.A.; Burnett, R.; Melander, C.; Edelson, B.S.; Arora, P.S.; Dervan, P.B.; Gottesfeld, J.M. Arresting cancer proliferation by small-molecule gene regulation. Chem. Biol. 2004, 11, 1583–1594. [Google Scholar] [CrossRef]
- Peng, X.; Wu, T.; Fan, J.; Wang, J.; Zhang, S.; Song, F.; Sun, S. An effective minor groove binder as a red fluorescent marker for live-cell DNA imaging and quantification. Angew. Chem. Int. Ed. 2011, 50, 4180–4183. [Google Scholar] [CrossRef]
- Melander, C.; Burnett, R.; Gottesfeld, J.M. Regulation of gene expression with pyrrole-imidazole polyamides. J. Biotechnol. 2004, 112, 195–220. [Google Scholar] [CrossRef]
- Minoshima, M.; Chou, J.C.; Lefebvre, S.; Bando, T.; Shinohara, K.; Gottesfeld, J.M.; Sugiyama, H. Potent activity against K562 cells by polyamide-seco-CBI conjugates targeting histone H4 genes. Bioorg. Med. Chem. 2010, 18, 168–174. [Google Scholar] [CrossRef]
- Jacobs, C.S.; Dervan, P.B. Modifications at the C-terminus to improve pyrrole-imidazole polyamide activity in cell culture. J. Med. Chem. 2009, 52, 7380–7388. [Google Scholar] [CrossRef]
- Synold, T.W.; Xi, B.X.; Wu, J.; Yen, Y.; Li, B.C.; Yang, F.; Phillips, J.W.; Nickols, N.G.; Dervan, P.B. Single-dose pharmacokinetic and toxicity analysis of pyrrole-imidazole polyamides in mice. Cancer Chemother. Pharmacol. 2012, 70, 617–625. [Google Scholar] [CrossRef]
- Raskatov, J.A.; Hargrove, A.E.; So, A.Y.; Dervan, P.B. Pharmacokinetics of Py-Im polyamides depend on architecture: Cyclic versus linear. J. Am. Chem. Soc. 2012, 134, 7995–7999. [Google Scholar] [CrossRef]
- Kawamoto, Y.; Bando, T.; Kamada, F.; Li, Y.; Hashiya, K.; Maeshima, K.; Sugiyama, H. Development of a new method for synthesis of tandem hairpin pyrrole–imidazole polyamide probes targeting human telomeres. J. Am. Chem. Soc. 2013, 135, 16468–16477. [Google Scholar] [CrossRef]
- Svanvik, N.; Nygren, J.; Westman, G.; Kubista, M. Free-probe fluorescence of light-up probes. J. Am. Chem. Soc. 2001, 123, 803–809. [Google Scholar] [CrossRef]
- Svanvik, N.; Westman, G.; Wang, D.; Kubista, M. Light-Up Probes: Thiazole orange-conjugated peptide nucleic acid for detection of target nucleic acid in homogeneous solution. Anal. Biochem. 2000, 281, 26–35. [Google Scholar] [CrossRef]
- Kricka, L.J. Stains, labels and detection strategies for nucleic acids assays. Ann. Clin. Biochem. 2002, 39, 114–129. [Google Scholar] [CrossRef]
- Hilal, H.; Taylor, J.A. Cyanine dyes for the detection of double stranded DNA. J. Biochem. Biophys. Meth. 2008, 70, 1104–1108. [Google Scholar] [CrossRef]
- Cullander, C. Imaging in the far-red with electronic light microscopy: Requirements and limitations. J. Microsc. 1994, 176, 281–286. [Google Scholar] [CrossRef]
- Ohulchanskyy, T.Y.; Pudavar, H.E.; Yarmoluk, S.M.; Yashchuk, V.M.; Bergey, E.J.; Prasad, P.N. A monomethine cyanine dye Cyan 40 for two-photon-excited fluorescence detection of nucleic acids and their visualization in live cells. Photochem. Photobiol. 2003, 77, 138–145. [Google Scholar] [CrossRef]
- Akbay, N.; Losytskyy, M.; Kovalska, V.; Balanda, A.; Yarmoluk, S. The mechanism of benzothiazole styrylcyanine dyes binding with dsDNA: Studies by spectral-luminescent methods. J. Fluoresc. 2008, 18, 139–147. [Google Scholar] [CrossRef]
- Losytskyy, M.Y.; Volkova, K.D.; Kovalska, V.B.; Makovenko, I.E.; Slominskii, Y.L.; Tolmachev, O.I.; Yarmoluk, S.M. Fluorescent properties of pentamethine cyanine dyes with cyclopentene and cyclohexene group in presence of biological molecules. J. Fluoresc. 2005, 15, 849–857. [Google Scholar] [CrossRef]
- Kovalska, V.B.; Kryvorotenko, D.V.; Balanda, A.O.; Losytskyy, M.Y.; Tokar, V.P.; Yarmoluk, S.M. Fluorescent homodimer styrylcyanines: Synthesis and spectral-luminescent studies in nucleic acids and protein complexes. Dyes Pigments 2005, 67, 47–54. [Google Scholar] [CrossRef]
- Yarmoluk, S.; Kovalska, V.; Losytskyy, M. Symmetric cyanine dyes for detecting nucleic acids. Biotech. Histochem. 2008, 83, 131–145. [Google Scholar] [CrossRef]
- Didenko, V.V. DNA probes using fluorescence resonance energy transfer (FRET): Designs and applications. BioTechniques 2001, 31, 1106–1121. [Google Scholar]
- Foister, S. Shape Selective Recognition of the DNA Minor Groove by Hairpin Polyamides. In Thesis for the Degree of Doctor of Philosophy; California Institute of Technology: Pasadena, CA, USA, 2003; pp. 129–127. [Google Scholar]
- Stadler, A.L.; Delos Santos, J.O.; Stensrud, E.S.; Dembska, A.; Silva, G.L.; Liu, S.P.; Shank, N.I.; Kunttas-Tatli, E.; Sobers, C.J.; Gramlich, P.M.E.; et al. Fluorescent DNA nanotags featuring covalently attached intercalating dyes: Synthesis, antibody conjugation, and intracellular imaging. Bioconjug. Chem. 2011, 22, 1491–1502. [Google Scholar] [CrossRef]
- Kim, M.Y.; Kim, J.; Hah, S.S. Poly(A)-targeting molecular beacons: Fluorescence resonance energy transfer-based in vitro quantitation and time-dependent imaging in live cells. Anal. Biochem. 2012, 429, 92–98. [Google Scholar] [CrossRef]
- Duhamel, J. New insights in the study of pyrene excimer fluorescence to characterize macromolecules and their supramolecular assemblies in solution. Langmuir 2012, 28, 6527–6538. [Google Scholar] [CrossRef]
- Krasheninina, O.A.; Novopashina, D.S.; Venyaminova, A.G. Oligo(2'-O-methylribonucleotides) containing insertions of 2'-bispyrenylmethylphosphoro-diamidate nucleoside derivatives as prospective fluorescent probes for RNA detection. Russ. J. Bioorg. Chem. 2011, 37, 244–248. [Google Scholar] [CrossRef]
- Novopashina, D.S.; Meschaninova, M.I.; Kholodar, S.A.; Lomzov, A.A.; Venyaminova, A.G. New eximer-based tandem systems for SNP detection. Nucleic Acids Symp. Ser. 2008, 52, 229–230. [Google Scholar] [CrossRef]
- Kholodar, S.A.; Novopashina, D.S.; Meschaninova, M.I.; Lomzov, A.A.; Venyaminova, A.G. Multipyrene tandem probes for detection of C677T polymorphism in MTHFR gene. Nucleic Acids Symp. Ser. 2009, 53, 143–144. [Google Scholar]
- Kolpashchikov, D.M. Binary probes for nucleic acid analysis. Chem. Rev. 2010, 110, 4709–4723. [Google Scholar] [CrossRef]
- Novikova, I.V.; Afonin, K.A.; Leontis, N.B. New Ideas for in Vivo Detection of RNA. In Biosensors; Serra, P.A., Ed.; InTech: Rijeka, Croatia, 2010; pp. 127–150. [Google Scholar]
- Guo, J.; Ju, J.; Turro, N.J. Fluorescent hybridization probes for nucleic acid detection. Anal. Bioanal. Chem. 2012, 402, 3115–3125. [Google Scholar] [CrossRef]
- Teo, Y.N.; Wilson, J.N.; Kool, E.T. Polyfluorophores on a DNA backbone: A multicolor set of labels excited at one wavelength. J. Am. Chem. Soc. 2009, 131, 3923–3933. [Google Scholar] [CrossRef]
- Wang, S.; Guo, J.; Ono, T.; Kool, E.T. DNA polyfluorophores for real-time multicolor tracking of dynamic biological systems. Angew. Chem. Int. Ed. 2012, 51, 7176–7180. [Google Scholar] [CrossRef]
- Guo, J.; Wang, S.; Dai, N.; Teo, Y.N.; Kool, E.T. Multispectral labeling of antibodies with polyfluorophores on a DNA backbone and application in cellular imaging. Proc. Nat. Acad. Sci. USA 2011, 108, 3493–3498. [Google Scholar] [CrossRef]
- Teo, Y.N.; Kool, E.T. DNA-multichromophore systems. Chem. Rev. 2012, 112, 4221–4245. [Google Scholar] [CrossRef]
- Ponting, C.P.; Oliver, P.L.; Reik, W. Evolution and functions of long noncoding RNAs. Cell 2009, 136, 629–641. [Google Scholar] [CrossRef]
- Kruger, K.; Grabowski, P.J.; Zaug, A.J.; Sands, J.; Gottschling, D.E.; Cech, T.R. Self-splicing RNA: Autoexcision and autocyclization of the ribosomal RNA intervening sequence of tetrahymena. Cell 1982, 31, 147–157. [Google Scholar] [CrossRef]
- Guerrier-Takada, C.; Gardiner, K.; Marsh, T.; Pace, N.; Altman, S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 1983, 35, 849–857. [Google Scholar] [CrossRef]
- Green, P.J.; Pines, O.; Inouye, M. The role of antisense RNA in gene regulation. Annu. Rev. Biochem. 1986, 55, 569–597. [Google Scholar] [CrossRef]
- The FANTOM Consortium. The transcriptional landscape of the mammalian genome. Science 2005, 309, 1559–1563. [Google Scholar] [CrossRef]
- Bushati, N.; Cohen, S.M. MicroRNA functions. Ann. Rev. Cell Dev. Biol. 2007, 23, 175–205. [Google Scholar] [CrossRef]
- Wang, Y.; Stricker, H.M.; Gou, D.M.; Liu, L. MicroRNA: Past and present. Front. Biosci. 2007, 12, 2316–2329. [Google Scholar] [CrossRef]
- Bouzinba-Segard, H.; Guais, A.; Francastel, C. Accumulation of small murine minor satellite transcripts leads to impaired centromeric architecture and function. Proc. Nat. Acad. Sci. USA 2006, 103, 8709–8714. [Google Scholar] [CrossRef]
- Thorsen, M.; Hansen, H.; Venturi, M.; Holmberg, S.; Thon, G. Mediator regulates non-coding RNA transcription at fission yeast centromeres. Epigenetics Chromatin 2012, 5, 19. [Google Scholar] [CrossRef]
- Carlsten, J.O.; Szilagyi, Z.; Liu, B.; Lopez, M.D.; Szászi, E.; Djupedal, I.; Nyström, T.; Ekwall, K.; Gustafsson, C.M.; Zhu, X. Mediator promotes CENP-a incorporation at fission yeast centromeres. Mol. Cell. Biol. 2012, 32, 4035–4043. [Google Scholar] [CrossRef]
- Enukashvily, N.I.; Malashicheva, A.B.; Waisertreiger, I.S.R. Satellite DNA spatial localization and transcriptional activity in mouse embryonic E-14 and IOUD2 stem cells. Cytogenet. Genome Res. 2009, 124, 277–287. [Google Scholar] [CrossRef]
- Chan, F.L.; Wong, L.H. Transcription in the maintenance of centromere chromatin identity. Nucleic Acids Res. 2012, 40, 11178–11188. [Google Scholar] [CrossRef]
- Chan, F.L.; Marshall, O.J.; Saffery, R.; Won Kim, B.; Earle, E.; Choo, K.H.A.; Wong, L.H. Active transcription and essential role of RNA polymerase II at the centromere during mitosis. Proc. Nat. Acad. Sci. USA 2012, 109, 1979–1984. [Google Scholar]
- Gent, J.I.; Dawe, R.K. RNA as a structural and regulatory component of the centromere. Annu. Rev. Genet. 2012, 46, 443–453. [Google Scholar] [CrossRef]
- Hall, L.E.; Mitchell, S.E.; O’Neill, R.J. Pericentric and centromeric transcription: A perfect balance required. Chromosome Res. 2012, 20, 535–546. [Google Scholar] [CrossRef]
- Dictenberg, J. Genetic encoding of fluorescent RNA ensures a bright future for visualizing nucleic acid dynamics. Trends Biotechnol. 2012, 30, 621–626. [Google Scholar] [CrossRef]
- Tyagi, S. Imaging intracellular RNA distribution and dynamics in living cells. Nat. Methods 2009, 6, 331–338. [Google Scholar] [CrossRef]
- Bertrand, E.; Chartrand, P.; Schaefer, M.; Shenoy, S.M.; Singer, R.H.; Long, R.M. Localization of ASH1 mRNA particles in living yeast. Mol. Cell 1998, 2, 437–445. [Google Scholar] [CrossRef]
- Beach, D.L.; Salmon, E.D.; Bloom, K. Localization and anchoring of mRNA in budding yeast. Curr. Biol. 1999, 9, S569–S561. [Google Scholar] [CrossRef]
- Haim, L.; Zipor, G.; Aronov, S.; Gerst, J.E. A genomic integration method to visualize localization of endogenous mRNAs in living yeast. Nat. Methods 2007, 4, 409–412. [Google Scholar]
- Bao, G.; Rhee, W.J.; Tsourkas, A. Fluorescent probes for live-cell RNA detection. Ann. Rev. Biomed. Eng. 2009, 11, 25–47. [Google Scholar] [CrossRef]
- Dirks, R.W.; Molenaar, C.; Tanke, H.J. Visualizing RNA molecules inside the nucleus of living cells. Methods 2003, 29, 51–57. [Google Scholar] [CrossRef]
- Bao, G.; Santangelo, P.; Nitin, N.; Rhee, W.J. Nanostructured Probes for in Vivo Gene Detection. In Nanotechnology; Fuchs, H., Grätzel, M., Krug, H., Schmid, G., Vogel, V., Waser, R., Eds.; Wiley-VCH Verlag GmbH & Co. KGaA: Chichester, UK, 2010; Volume 5, (Nanomedicine); pp. 143–165. [Google Scholar]
- Ranasinghe, R.T.; Brown, L.J.; Brown, T. Linear fluorescent oligonucleotide probes with an acridine quencher generate a signal upon hybridisation. Chem. Commun. 2001, 1480–1481. [Google Scholar] [CrossRef]
- Waki, R.; Yamayoshi, A.; Kobori, A.; Murakami, A. Development of a system to sensitively and specifically visualize c-fos mRNA in living cells using bispyrene-modified RNA probes. Chem. Commun. 2011, 47, 4204–4206. [Google Scholar]
- Menacher, F.; Rubner, M.; Berndl, S.; Wagenknecht, H.-A. Thiazole orange and Cy3: Improvement of fluorescent DNA probes with use of short range electron transfer. J. Org. Chem. 2008, 73, 4263–4266. [Google Scholar] [CrossRef]
- Bethge, L.; Singh, I.; Seitz, O. Designed thiazole orange nucleotides for the synthesis of single labelled oligonucleotides that fluoresce upon matched hybridization. Org. Biomol. Chem. 2010, 8, 2439–2448. [Google Scholar] [CrossRef]
- Holzhauser, C.; Wagenknecht, H.A. “DNA traffic lights”: Concept of wavelength-shifting DNA probes and application in an aptasensor. ChemBioChem 2012, 13, 1136–1138. [Google Scholar] [CrossRef]
- Holzhauser, C.; Berndl, S.; Menacher, F.; Breunig, M.; Göpferich, A.; Wagenknecht, H.-A. Synthesis and optical properties of cyanine dyes as fluorescent DNA base substitutions for live cell imaging. Eur. J. Org. Chem. 2010, 2010, 1239–1248. [Google Scholar]
- Holzhauser, C.; Liebl, R.; Goepferich, A.; Wagenknecht, H.-A.; Breunig, M. RNA “Traffic lights”: An analytical tool to monitor siRNA integrity. ACS Chem. Biol. 2013, 8, 890–894. [Google Scholar] [CrossRef]
- Kashida, H.; Takatsu, T.; Sekiguchi, K.; Asanuma, H. An efficient fluorescence resonance energy transfer (FRET) between pyrene and perylene assembled in a DNA duplex and its potential for discriminating single-base changes. Chem. Eur. J. 2010, 16, 2479–2486. [Google Scholar] [CrossRef]
- Kato, T.; Kashida, H.; Kishida, H.; Yada, H.; Okamoto, H.; Asanuma, H. Development of a robust model system of FRET using base surrogates tethering fluorophores for strict control of their position and orientation within DNA duplex. J. Am. Chem. Soc. 2012, 135, 741–750. [Google Scholar]
- Ikeda, S.; Okamoto, A. Hybridization-sensitive on–off DNA probe: Application of the exciton coupling effect to effective fluorescence quenching. Chem. Asian J. 2008, 3, 958–968. [Google Scholar] [CrossRef]
- Kubota, T.; Ikeda, S.; Yanagisawa, H.; Yuki, M.; Okamoto, A. Hybridization-sensitive fluorescent probe for long-term monitoring of intracellular RNA. Bioconjug. Chem. 2009, 20, 1256–1261. [Google Scholar] [CrossRef]
- Köhler, O.; Seitz, O. Thiazole orange as fluorescent universal base in peptide nucleic acids. Chem. Commun. 2003, 2938–2939. [Google Scholar]
- Köhler, O.; Jarikote, D.V.; Seitz, O. Forced intercalation probes (FIT probes): Thiazole orange as a fluorescent base in peptide nucleic acids for homogeneous single-nucleotide-polymorphism detection. ChemBioChem 2005, 6, 69–77. [Google Scholar] [CrossRef]
- Kummer, S.; Knoll, A.; Socher, E.; Bethge, L.; Herrmann, A.; Seitz, O. Fluorescence imaging of influenza H1N1 mRNA in living infected cells using single-chromophore FIT-PNA. Angew. Chem. Int. Ed. 2011, 50, 1931–1934. [Google Scholar]
- Kummer, S.; Knoll, A.; Socher, E.; Bethge, L.; Herrmann, A.; Seitz, O. PNA FIT-probes for the dual color imaging of two viral mRNA targets in influenza H1N1 infected live cells. Bioconjug. Chem. 2012, 23, 2051–2060. [Google Scholar]
- Tyagi, S.; Kramer, F.R. Molecular beacons: Probes that fluoresce upon hybridization. Nat. Biotechnol. 1996, 14, 303–308. [Google Scholar] [CrossRef]
- Tan, W.; Wang, K.; Drake, T.J. Molecular beacons. Curr. Opin. Chem. Biol. 2004, 8, 547–553. [Google Scholar] [CrossRef]
- Huang, K.; Martí, A. Recent trends in molecular beacon design and applications. Anal. Bioanal. Chem. 2012, 402, 3091–3102. [Google Scholar] [CrossRef]
- Tsourkas, A.; Behlke, M.A.; Bao, G. Hybridization of 2'-O-methyl and 2'-deoxy molecular beacons to RNA and DNA targets. Nucleic Acids Res. 2002, 30, 5168–5174. [Google Scholar] [CrossRef]
- Tsourkas, A.; Behlke, M.A.; Bao, G. Structure–function relationships of shared-stem and conventional molecular beacons. Nucleic Acids Res. 2002, 30, 4208–4215. [Google Scholar] [CrossRef]
- Tsourkas, A.; Behlke, M.A.; Rose, S.D.; Bao, G. Hybridization kinetics and thermodynamics of molecular beacons. Nucleic Acids Res. 2003, 31, 1319–1330. [Google Scholar] [CrossRef]
- Rhee, W.J.; Bao, G. Slow non-specific accumulation of 2'-deoxy and 2'-O-methyl oligonucleotide probes at mitochondria in live cells. Nucleic Acids Res. 2010, 38, e109. [Google Scholar] [CrossRef]
- Rhee, W.J.; Bao, G. Simultaneous detection of mRNA and protein stem cell markers in live cells. BMC Biotechnol. 2009, 9, 30. [Google Scholar] [CrossRef]
- Chen, A.K.; Davydenko, O.; Behlke, M.A.; Tsourkas, A. Ratiometric bimolecular beacons for the sensitive detection of RNA in single living cells. Nucleic Acids Res. 2010, 38, e148. [Google Scholar] [CrossRef]
- Molenaar, C.; Marras, S.A.; Slats, J.C.M.; Truffert, J.-C.; Lemaïtre, M.; Raap, A.K.; Dirks, R.W.; Tanke, H.J. Linear 2'-O-Methyl RNA probes for the visualization of RNA in living cells. Nucleic Acids Res. 2001, 29, e89. [Google Scholar] [CrossRef]
- Liu, L.; Tang, Z.; Wang, K.; Tan, W.; Li, J.; Guo, Q.; Meng, X.; Ma, C. Using molecular beacon to monitor activity of E. coli DNA ligase. Analyst 2005, 130, 350–357. [Google Scholar] [CrossRef]
- Tang, Z.; Wang, K.; Tan, W.; Li, J.; Liu, L.; Guo, Q.; Meng, X.; Ma, C.; Huang, S. Real-time monitoring of nucleic acid ligation in homogenous solutions using molecular beacons. Nucleic Acids Res. 2003, 31, e148. [Google Scholar]
- Ma, C.; Tang, Z.; Wang, K.; Tan, W.; Yang, X.; Li, W.; Li, Z.; Lv, X. Real-time monitoring of restriction endonuclease activity using molecular beacon. Anal. Biochem. 2007, 363, 294–296. [Google Scholar] [CrossRef]
- Ma, C.; Tang, Z.; Wang, K.; Tan, W.; Yang, X.; Li, W.; Li, Z.; Li, H.; Lv, X. Real-time monitoring of nucleic acid dephosphorylation by using molecular beacons. ChemBioChem 2007, 8, 1487–1490. [Google Scholar] [CrossRef]
- Zhang, C.; Su, X.; Liang, Y.; Zhu, X.; Song, C.; Zhao, M. A transformer of molecular beacon for sensitive and real-time detection of phosphatases with effective inhibition of the false positive signals. Biosens. Bioelectron. 2011, 28, 13–16. [Google Scholar] [CrossRef]
- Ma, C.; Tang, Z.; Wang, K.; Tan, W.; Li, J.; Li, W.; Li, Z.; Yang, X.; Li, H.; Liu, L. Real-time monitoring of DNA polymerase activity using molecular beacon. Anal. Biochem. 2006, 353, 141–143. [Google Scholar] [CrossRef]
- Li, J.; Yan, H.; Wang, K.; Tan, W.; Zhou, X. Hairpin fluorescence DNA probe for real-time monitoring of DNA methylation. Anal. Chem. 2007, 79, 1050–1056. [Google Scholar] [CrossRef]
- Tang, Z.; Wang, K.; Tan, W.; Ma, C.; Li, J.; Liu, L.; Guo, Q.; Meng, X. Real-time investigation of nucleic acids phosphorylation process using molecular beacons. Nucleic Acids Res. 2005, 33, e97. [Google Scholar] [CrossRef]
- Ma, C.; Yang, X.; Wang, K.; Tang, Z.; Li, W.; Tan, W.; Lv, X. A novel kinase-based ATP assay using molecular beacon. Anal. Biochem. 2008, 372, 131–133. [Google Scholar] [CrossRef]
- Fujimoto, K.; Shimizu, H.; Inouye, M. Unambiguous detection of target DNAs by excimer-monomer switching molecular beacons. J. Org. Chem. 2004, 69, 3271–3275. [Google Scholar]
- Huang, J.; Wu, Y.; Chen, Y.; Zhu, Z.; Yang, X.; Yang, C.J.; Wang, K.; Tan, W. Pyrene-excimer probes based on the hybridization chain reaction for the detection of nucleic acids in complex biological fluids. Angew. Chem. Int. Ed. 2011, 50, 401–404. [Google Scholar] [CrossRef]
- Yamana, K.; Ohshita, Y.; Fukunaga, Y.; Nakamura, M.; Maruyama, A. Bis-pyrene-labeled molecular beacon: A monomer–excimer switching probe for the detection of DNA base alteration. Bioorg. Med. Chem. 2008, 16, 78–83. [Google Scholar]
- Biner, S.M.; Häner, R. A two-color, self-controlled molecular beacon. ChemBioChem 2011, 12, 2733–2736. [Google Scholar] [CrossRef]
- Xiang, D.; Zhang, C.; Chen, L.; Ji, X.; He, Z. Tricolour fluorescence detection of sequence-specific DNA with a new molecular beacon and a nucleic acid dye TOTO-3. Analyst 2012, 137, 5898–5905. [Google Scholar] [CrossRef]
- Yang, C.J.; Martinez, K.; Lin, H.; Tan, W. Hybrid molecular probe for nucleic acid analysis in biological samples. J. Am. Chem. Soc. 2006, 128, 9986–9987. [Google Scholar] [CrossRef]
- Zhu, G.; Zhang, S.; Song, E.; Zheng, J.; Hu, R.; Fang, X.; Tan, W. Building fluorescent DNA nanodevices on target living cell surfaces. Angew. Chem. Int. Ed. 2013, 52, 5490–5496. [Google Scholar] [CrossRef]
- Varghese, R.; Wagenknecht, H.-A. Red-white-blue emission switching molecular beacons: Ratiometric multicolour DNA hybridization probes. Org. Biomol. Chem. 2010, 8, 526–528. [Google Scholar] [CrossRef]
- Urano, Y.; Asanuma, D.; Hama, Y.; Koyama, Y.; Barrett, T.; Kamiya, M.; Nagano, T.; Watanabe, T.; Hasegawa, A.; Choyke, P.L.; et al. Selective molecular imaging of viable cancer cells with pH-activatable fluorescence probes. Nat. Med. 2009, 15, 104–109. [Google Scholar] [CrossRef]
- Kashida, H.; Yamaguchi, K.; Hara, Y.; Asanuma, H. Quencher-free molecular beacon tethering 7-hydroxycoumarin detects targets through protonation/deprotonation. Bioorg. Med. Chem. 2012, 20, 4310–4315. [Google Scholar] [CrossRef]
- Su, X.; Zhang, C.; Zhao, M. Discrimination of the false-positive signals of molecular beacons by combination of heat inactivation and using single walled carbon nanotubes. Biosens. Bioelectron. 2011, 26, 3596–3601. [Google Scholar] [CrossRef]
- Petersen, K.; Vogel, U.; Rockenbauer, E.; Vang Nielsen, K.; Kølvraa, S.; Bolund, L.; Nexø, B. Short PNA molecular beacons for real-time PCR allelic discrimination of single nucleotide polymorphisms. Mol. Cell. Probes 2004, 18, 117–122. [Google Scholar] [CrossRef]
- Kim, Y.; Yang, C.J.; Tan, W. Superior structure stability and selectivity of hairpin nucleic acid probes with an L-DNA stem. Nucleic Acids Res. 2007, 35, 7279–7287. [Google Scholar] [CrossRef]
- Yang, C.J.; Wang, L.; Wu, Y.; Kim, Y.; Medley, C.D.; Lin, H.; Tan, W. Synthesis and investigation of deoxyribonucleic acid/locked nucleic acid chimeric molecular beacons. Nucleic Acids Res. 2007, 35, 4030–4041. [Google Scholar] [CrossRef]
- Wang, L.; Yang, C.J.; Medley, C.D.; Benner, S.A.; Tan, W. Locked nucleic acid molecular beacons. J. Am. Chem. Soc. 2005, 127, 15664–15665. [Google Scholar]
- Martinez, K.; Estevez, M.C.; Wu, Y.; Phillips, J.A.; Medley, C.D.; Tan, W. Locked nucleic acid based beacons for surface interaction studies and biosensor development. Anal. Chem. 2009, 81, 3448–3454. [Google Scholar]
- Chen, A.K.; Behlke, M.A.; Tsourkas, A. Sub-cellular trafficking and functionality of 2'-O-methyl and 2'-O-methyl-phosphorothioate molecular beacons. Nucleic Acids Res. 2009, 37, e149. [Google Scholar] [CrossRef]
- Qiu, L.; Wu, C.; You, M.; Han, D.; Chen, T.; Zhu, G.; Jiang, J.; Yu, R.; Tan, W. A targeted, self-delivered, and photocontrolled molecular meacon for mRNA detection in living cells. J. Am. Chem. Soc. 2013, 135, 12952–12955. [Google Scholar] [CrossRef]
- Okabe, K.; Harada, Y.; Zhang, J.; Tadakuma, H.; Tani, T.; Funatsu, T. Real time monitoring of endogenous cytoplasmic mRNA using linear antisense 2'-O-methyl RNA probes in living cells. Nucleic Acids Res. 2011, 39, e20. [Google Scholar] [CrossRef]
- Martí, A.A.; Li, X.; Jockusch, S.; Li, Z.; Raveendra, B.; Kalachikov, S.; Russo, J.J.; Morozova, I.; Puthanveettil, S.V.; Ju, J.; et al. Pyrene binary probes for unambiguous detection of mRNA using time-resolved fluorescence spectroscopy. Nucleic Acids Res. 2006, 34, 3161–3168. [Google Scholar] [CrossRef]
- Silverman, A.P.; Abe, H.; Kool, E.T. Quenched Autoligation (QUAL) Probes. In Molecular Beacons: Signalling Nucleic Acid Probes, Methods, and Protocols; Marx, A., Seitz, O., Eds.; Humana Press: Totowa, NJ, USA, 2008; Volume 249, pp. 161–170. [Google Scholar]
- Sando, S.; Kool, E.T. Imaging of RNA in bacteria with self-ligating quenched probes. J. Am. Chem. Soc. 2002, 124, 9686–9687. [Google Scholar] [CrossRef]
- Abe, H.; Kool, E.T. Destabilizing universal linkers for signal amplification in self-ligating probes for RNA. J. Am. Chem. Soc. 2004, 126, 13980–13986. [Google Scholar] [CrossRef]
- Abe, H.; Kool, E.T. Flow cytometric detection of specific RNAs in native human cells with quenched autoligating FRET probes. Proc. Nat. Acad. Sci. USA 2006, 103, 263–268. [Google Scholar] [CrossRef]
- Silverman, A.P.; Baron, E.J.; Kool, E.T. RNA-templated chemistry in cells: Discrimination of Escherichia, Shigella and Salmonella bacterial strains with a new two-color FRET strategy. ChemBioChem 2006, 7, 1890–1894. [Google Scholar] [CrossRef]
- Silverman, A.P.; Kool, E.T. Quenched autoligation probes allow discrimination of live bacterial species by single nucleotide differences in rRNA. Nucleic Acids Res. 2005, 33, 4978–4986. [Google Scholar] [CrossRef]
- Miller, G.P.; Silverman, A.P.; Kool, E.T. New, stronger nucleophiles for nucleic acid-templated chemistry: Synthesis and application in fluorescence detection of cellular RNA. Bioorg. Med. Chem. 2008, 16, 56–64. [Google Scholar] [CrossRef]
- Franzini, R.M.; Kool, E.T. Efficient nucleic acid detection by templated reductive quencher release. J. Am. Chem. Soc. 2009, 131, 16021–16023. [Google Scholar]
- Kleinbaum, D.J.; Kool, E.T. Sandwich probes: Two simultaneous reactions for templated nucleic acid detection. Chem. Commun. 2010, 46, 8154–8156. [Google Scholar] [CrossRef]
- Franzini, R.M.; Kool, E.T. Two successive reactions on a DNA template: A strategy for improving background fluorescence and specificity in nucleic acid detection. Chem. Eur. J. 2011, 17, 2168–2175. [Google Scholar] [CrossRef]
- Franzini, R.M.; Kool, E.T. Improved templated fluorogenic probes enhance the analysis of closely related pathogenic bacteria by microscopy and flow cytometry. Bioconjug. Chem. 2011, 22, 1869–1877. [Google Scholar] [CrossRef]
- Röthlingshöfer, M.; Gorska, K.; Winssinger, N. Nucleic acid-templated energy transfer leading to a photorelease reaction and its application to a system displaying a nonlinear response. J. Am. Chem. Soc. 2011, 133, 18110–18113. [Google Scholar] [CrossRef]
- Röthlingshöfer, M.; Gorska, K.; Winssinger, N. Nucleic acid templated uncaging of fluorophores using Ru-catalyzed photoreduction with visible light. Org. Lett. 2011, 14, 482–485. [Google Scholar]
- Sadhu, K.K.; Winssinger, N. Detection of miRNA in live cells by using templated Ru(II)-catalyzed unmasking of a fluorophore. Chem. Eur. J. 2013, 19, 8182–8189. [Google Scholar]
- Grossmann, T.N.; Seitz, O. DNA-catalyzed transfer of a reporter group. J. Am. Chem. Soc. 2006, 128, 15596–15597. [Google Scholar] [CrossRef]
- Grossmann, T.N.; Roglin, L.; Seitz, O. Target-catalyzed transfer reactions for the amplified detection of RNA. Angew. Chem. Int. Ed. 2008, 47, 7119–7122. [Google Scholar] [CrossRef]
- Chen, X.H.; Roloff, A.; Seitz, O. Consecutive signal amplification for DNA detection based on de novo fluorophore synthesis and host-guest chemistry. Angew. Chem. Int. Ed. 2012, 51, 4479–4483. [Google Scholar] [CrossRef]
- Syed, M.A.; Pervaiz, S. Advances in aptamers. Oligonucleotides 2010, 20, 215–224. [Google Scholar] [CrossRef]
- Ni, X.; Castanares, M.; Mukherjee, A.; Lupold, S.E. Nucleic acid aptamers: Clinical applications and promising new horizons. Curr. Med. Chem. 2011, 18, 4206–4214. [Google Scholar] [CrossRef]
- Mascini, M.; Palchetti, I.; Tombelli, S. Nucleic acid and peptide aptamers: Fundamentals and bioanalytical aspects. Angew. Chem. Int. Ed. 2012, 51, 1316–1332. [Google Scholar] [CrossRef]
- Vendrell, M.; Zhai, D.; Er, J.C.; Chang, Y.-T. Combinatorial strategies in fluorescent probe development. Chem. Rev. 2012, 112, 4391–4420. [Google Scholar] [CrossRef]
- Kolpashchikov, D.M. Binary malachite green aptamer for fluorescent detection of nucleic acids. J. Am. Chem. Soc. 2005, 127, 12442–12443. [Google Scholar] [CrossRef]
- Sando, S.; Narita, A.; Aoyama, Y. Light-up hoechst–DNA aptamer pair: Generation of an aptamer-selective fluorophore from a conventional DNA-staining dye. ChemBioChem 2007, 8, 1795–1803. [Google Scholar] [CrossRef]
- Eydeler, K.; Magbanua, E.; Werner, A.; Ziegelmüller, P.; Hahn, U. Fluorophore binding aptamers as a tool for RNA visualization. Biophys. J. 2009, 96, 3703–3707. [Google Scholar]
- Paige, J.S.; Wu, K.Y.; Jaffrey, S.R. RNA Mimics of green fluorescent protein. Science 2011, 333, 642–646. [Google Scholar] [CrossRef]
- Sassolas, A.; Blum, L.J.; Leca-Bouvier, B.D. Homogeneous assays using aptamers. Analyst 2011, 136, 257–274. [Google Scholar] [CrossRef]
- Astakhova, I.V.; Korshun, V.A.; Jahn, K.; Kjems, J.; Wengel, J. Perylene attached to 2'-amino-LNA: Synthesis, incorporation into oligonucleotides, and remarkable fluorescence properties in vitro and in cell culture. Bioconjug. Chem. 2008, 19, 1995–2007. [Google Scholar] [CrossRef]
- Cummins, L.L.; Owens, S.R.; Risen, L.M.; Lesnik, E.A.; Freier, S.M.; McGee, D.; Guinosso, C.J.; Cook, P.D. Characterization of fully 2'-modified oligoribonucleotide hetero-and homoduplex hybridization and nuclease sensitivity. Nucleic Acids Res. 1995, 23, 2019–2024. [Google Scholar] [CrossRef]
- Novopashina, D.; Sinyakov, A.; Ryabinin, V.; Venyaminova, A.; Boutorine, A. Conjugates of oligo(2'-O-methylribonucleotides) with minor groove binders as new sequence-specific agents recognizing both grooves of double-stranded DNA. Nucleosides Nucleotides Nucleic Acids 2003, 22, 1179–1182. [Google Scholar] [CrossRef]
- Novopashina, D.S.; Sinyakov, A.N.; Ryabinin, V.A.; Venyaminova, A.G.; Halby, L.; Sun, J.S.; Boutorine, A.S. Sequence-specific conjugates of oligo(2'-O-methylribonucleotides) and hairpin oligocarboxamide minor-groove binders: Design, synthesis, and binding studies with double-stranded DNA. Chem. Biodivers. 2005, 2, 936–952. [Google Scholar] [CrossRef]
- Boutorine, A.S.; Venyaminova, A.G.; Repkova, M.N.; Sergueyeva, Z.A.; Pyshnyi, D.V. Effect of derivatization of ribophosphate backbone and terminal ribophosphate groups in oligoribonucleotides an their stability and interaction with eukaryotic cells. Biochimie 1994, 76, 23–32. [Google Scholar] [CrossRef]
- Novopashina, D.; Kuznetsova, M.; Venyaminova, A. 2'-O-modified oligoribonucleotides with terminal 3'-3'-internucleotide linkage and their derivatives. Nucleosides Nucleotides Nucleic Acids 2001, 20, 903–907. [Google Scholar] [CrossRef]
- Novopashina, D.S.; Totskaya, O.S.; Lomzov, A.A.; Venyaminova, A.G. 3'-modified oligo(2'-O-methylribonucleotides) as improved probes for hybridization with RNA. Nucleosides Nucleotides Nucleic Acids 2005, 24, 527–531. [Google Scholar] [CrossRef]
- Majlessi, M.; Nelson, N.C.; Becker, M.M. Advantages of 2'-O-methyl oligoribonucleotide probes for detecting RNA targets. Nucleic Acids Res. 1998, 26, 2224–2229. [Google Scholar] [CrossRef]
- Bratu, D.P.; Cha, B.J.; Mhlanga, M.M.; Kramer, F.R.; Tyagi, S. Visualizing the distribution and transport of mRNAs in living cells. Proc. Nat. Acad. Sci. USA 2003, 100, 13308–13313. [Google Scholar] [CrossRef]
- Malik, R.; Roy, I. Making sense of therapeutics using antisense technology. Expert Opin. Drug Discov. 2011, 6, 507–526. [Google Scholar] [CrossRef]
- Juliano, R.L.; Ming, X.; Nakagawa, O. Cellular uptake and intracellular trafficking of antisense and siRNA oligonucleotides. Bioconjug. Chem. 2011, 23, 147–157. [Google Scholar] [CrossRef]
- Lochmann, D.; Jauk, E.; Zimmer, A. Drug delivery of oligonucleotides by peptides. Eur. J. Pharm. Biopharm. 2004, 58, 237–251. [Google Scholar] [CrossRef]
- Elsabahy, M.; Nazarali, A.; Foldvari, M. Non-viral nucleic acid delivery: Key challenges and future directions. Curr. Drug Deliv. 2011, 8, 235–244. [Google Scholar] [CrossRef]
- Lacerda, L.; Bianco, A.; Prato, M.; Kostarelos, K. Carbon nanotube cell translocation and delivery of nucleic acids in vitro and in vivo. J. Mater. Chem. 2008, 18, 17–22. [Google Scholar] [CrossRef]
- Campidelli, S.; Ballesteros, B.; Filoramo, A.; Díaz, D.D.; de la Torre, G.; Torres, T.; Rahman, G.M.A.; Ehli, C.; Kiessling, D.; Werner, F.; et al. Facile decoration of functionalized single-wall carbon nanotubes with phthalocyanines via “click chemistry”. J. Am. Chem. Soc. 2008, 130, 11503–11509. [Google Scholar] [CrossRef]
- Brunetti, F.G.; Herrero, M.A.; Munoz, J.D.; Diaz-Ortiz, A.; Alfonsi, J.; Meneghetti, M.; Prato, M.; Vazquez, E. Microwave-induced multiple functionalization of carbon nanotubes. J. Am. Chem. Soc. 2008, 130, 8094–8100. [Google Scholar] [CrossRef]
- Prato, M.; Kostarelos, K.; Bianco, A. Functionalized carbon nanotubes in drug design and discovery. Acc. Chem. Res. 2008, 41, 60–68. [Google Scholar] [CrossRef]
- Apartsin, E.K.; Novopashina, D.S.; Nastaushev, Y.V.; Venyaminova, A.G. Fluorescently labeled single-walled carbon nanotubes and their hybrids with oligonucleotides. Nanotechnol. Russ. 2012, 7, 99–109. [Google Scholar] [CrossRef]
- Apartsin, E.K.; Buyanova, M.Y.; Novopashina, D.S.; Ryabchikova, E.I.; Venyaminova, A.G. Non-Covalent Immobilization of Oligonucleotides on Single-Walled Carbon Nanotubes. In Nanomaterials Imaging Techniques, Surface Studies, and Applications; Fesenko, O., Yatsenko, L., Brodin, M., Eds.; Springer: New York, NY, USA, 2013; Volume 146, pp. 291–307. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Boutorine, A.S.; Novopashina, D.S.; Krasheninina, O.A.; Nozeret, K.; Venyaminova, A.G. Fluorescent Probes for Nucleic Acid Visualization in Fixed and Live Cells. Molecules 2013, 18, 15357-15397. https://doi.org/10.3390/molecules181215357
Boutorine AS, Novopashina DS, Krasheninina OA, Nozeret K, Venyaminova AG. Fluorescent Probes for Nucleic Acid Visualization in Fixed and Live Cells. Molecules. 2013; 18(12):15357-15397. https://doi.org/10.3390/molecules181215357
Chicago/Turabian StyleBoutorine, Alexandre S., Darya S. Novopashina, Olga A. Krasheninina, Karine Nozeret, and Alya G. Venyaminova. 2013. "Fluorescent Probes for Nucleic Acid Visualization in Fixed and Live Cells" Molecules 18, no. 12: 15357-15397. https://doi.org/10.3390/molecules181215357