Yeasts from Different Habitats and Their Potential as Biocontrol Agents
Abstract
:1. Introduction
2. Materials and Methods
2.1. Yeasts, Fungi, and Culture Conditions
2.2. Preparation of Fungal Spore Suspensions
2.3. Enzymatic Activites
2.3.1. β-Glucosidase Activity
2.3.2. Cellulase Activity
2.3.3. Amylase Activity
2.3.4. Protease Activity
2.3.5. Chitinase Activity
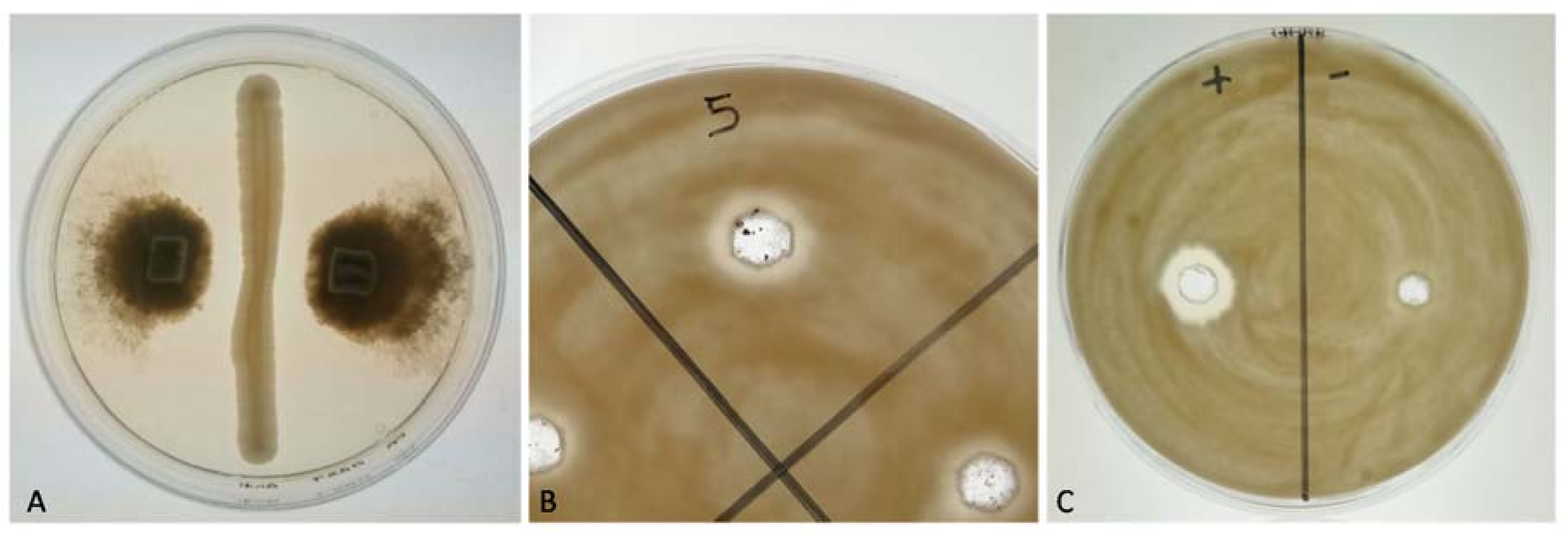
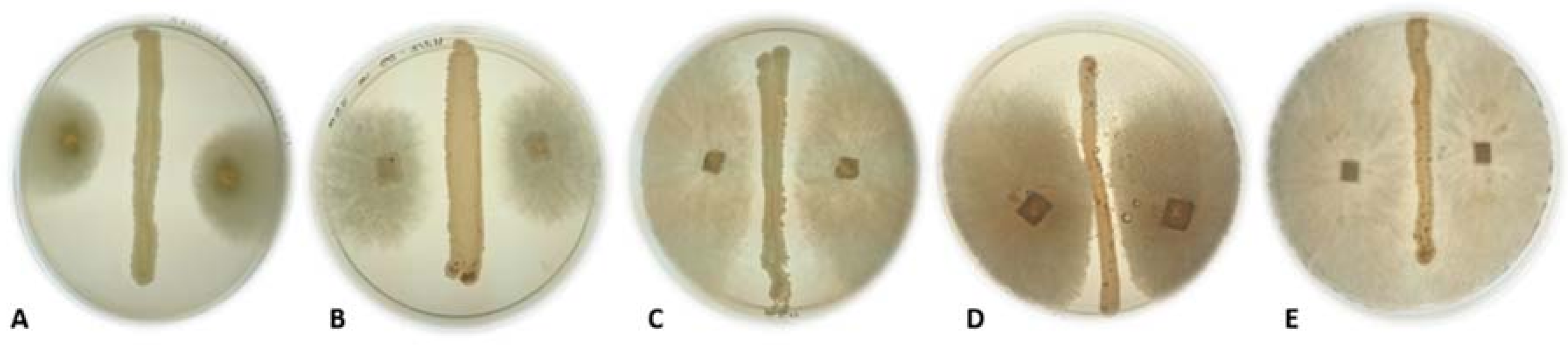
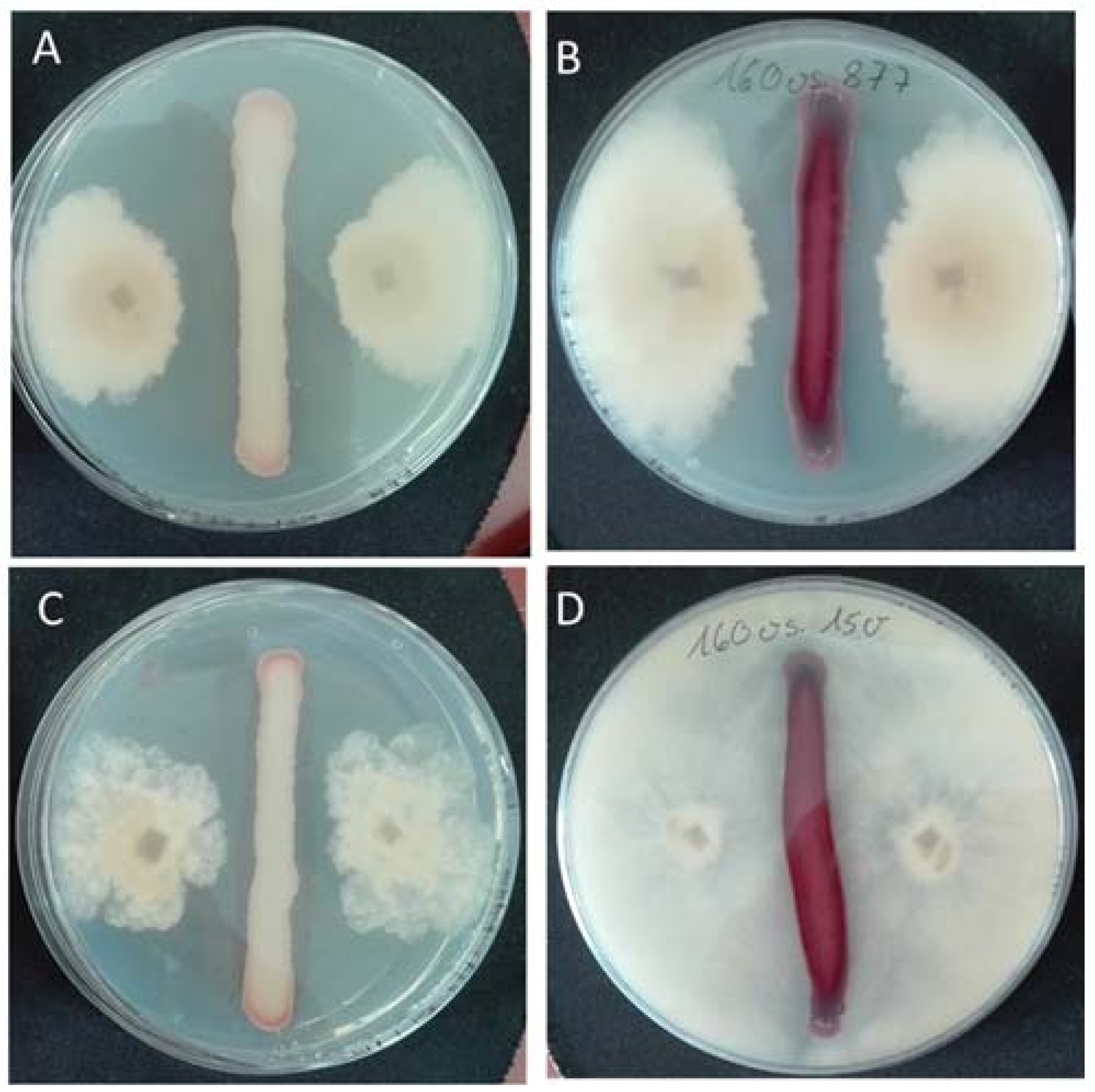
2.4. Antagonistic Activity of Yeasts against Phytopathogenic Fungi on Agar Plates
2.5. Preparation of Fungal Cell Wall Components
2.6. Preparation of Cell-Free Culture Filtrates
2.6.1. Lyophilization
2.6.2. Ultrafiltration
2.6.3. Extraction with Ethyl Acetate
2.7. Agar Diffusion Assay
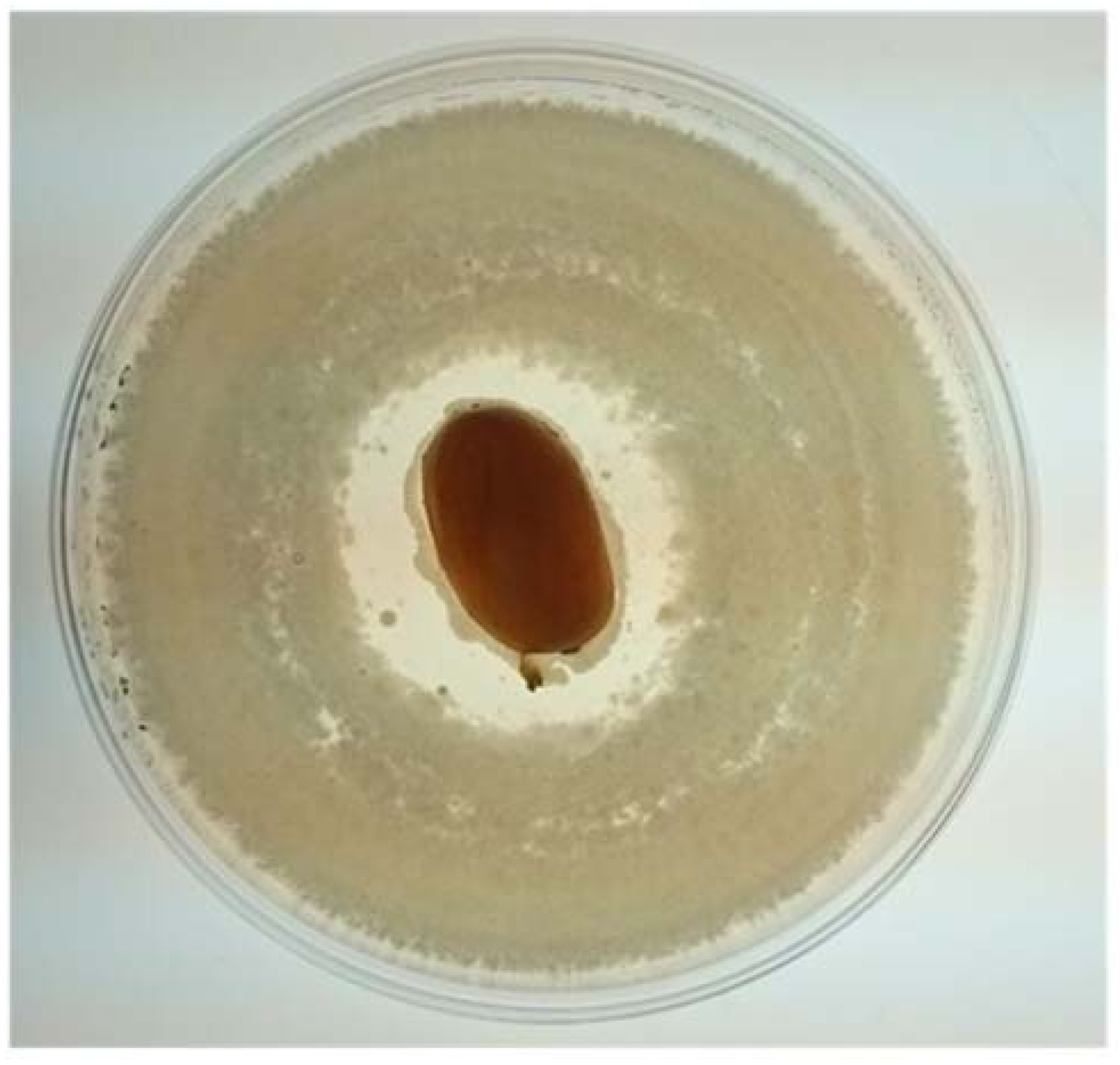
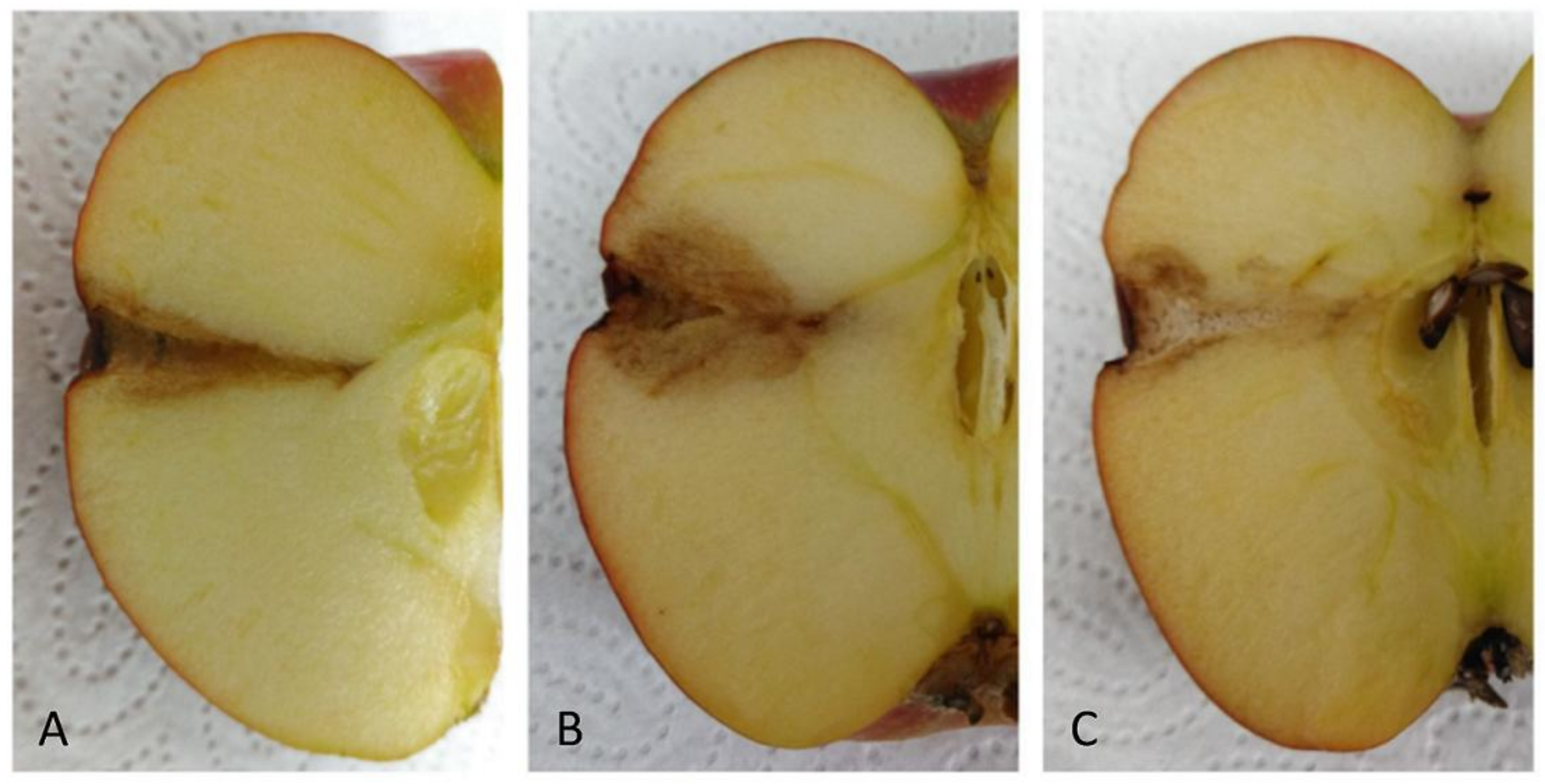
2.8. Antagonistic Activities on Fruits
3. Results and Discussion
4. Conclusions
Author Contributions
Conflicts of Interest
References
- Pozo-Bayón, M.Á.; Monagas, M.; Bartolomé, B.; Morena-Arribas, M.V. Wine features related to safety and consumer health: An integrated perspective. Crit. Rev. Food Sci. Nutr. 2012, 52, 31–54. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fillinger, S.; Elad, Y. Botrytis—The Fungus, the Pathogen and Its Management in Agricultural Systems; Springer International Publishing AG: Basel, Switzerland, 2016. [Google Scholar]
- Kassemeyer, H.H. Fungi of grapes. In Biology of Microorganisms on Grapes, in Must and in Wine; König, H., Unden, F., Fröhlich, J., Eds.; Springer International Publishing AG: Basel, Switzerland, 2017; pp. 103–132. [Google Scholar]
- Kretschmer, M.; Hahn, M. Fungicide resistance and genetic diversity of Botrytis cinerea isolates from a vineyard in Germany. J. Plant Dis. Prot. 2008, 115, 214–219. [Google Scholar] [CrossRef]
- Rupp, S.; Weber, R.W.S.; Rieger, D.; Detzel, P.; Hahn, M. Spread of Botrytis cinerea strains with multiple fungicide resistance in German horticulture. Front. Microbiol. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Neuhauser, S.; Huber, L.; Kirchmair, M. Is Roesleria subterranea a primary pathogen or a minor parasite of grapevines? Risk assessment and a diagnostic decision scheme. Eur. J. Plant Pathol. 2011, 130, 503–510. [Google Scholar] [CrossRef] [PubMed]
- Fischer, J.; Thines, E. Secondary metabolites of fungal vine pathogens. In Biology of Microorganisms on Grapes, in Must and in Wine; König, H., Unden, F., Fröhlich, J., Eds.; Springer International Publishing AG: Basel, Switzerland, 2017; pp. 165–185. [Google Scholar]
- Moisy, C.; Berger, G.; Flutre, T.; Cunff, L.L.; Péros, J. Quantitative assessment of grapevine wood colonization by the dieback fungus Eutypa lata. J. Fungi 2017, 3, 21. [Google Scholar] [CrossRef] [PubMed]
- Dean, R.; van Kan, J.A.L.; Pretorius, Z.A.; Hammond-Kosack, K.E.; di Pietro, A.; Spanu, P.D.; Rudd, J.J.; Dickman, M.; Kahmann, R.; Ellis, J.; et al. The top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 2012, 13, 414–430. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Sui, Y.; Wisniewski, M.; Droby, S.; Liu, Y. Review: Utilization of antagonistic yeasts to manage postharvest diseases in fruit. Int. J. Food Microbiol. 2013, 167, 153–160. [Google Scholar] [CrossRef] [PubMed]
- Muccilli, S.; Restuccia, C. Bioprotective role of yeasts. Microorganisms 2015, 3, 588–611. [Google Scholar] [CrossRef] [PubMed]
- Nicot, P.C.; Stewart, A.; Bardin, M.; Elad, Y. Biological control and biopesticide suppression of Botrytis-incited diseases. In Botrytis—The Fungus, the Pathogen and Its Management in Agricultural Systems; Fillinger, S., Elad, Y., Eds.; Springer International Publishing: Basel, Switzerlad, 2016; pp. 165–188. [Google Scholar]
- Rotolo, C.; De Miccolis Angelini, R.M.; Dongiovanni, C.; Pollastro, S.; Fumarola, G.; Di Carolo, M.; Perrelli, D.; Natal, P.; Faretra, F. Use of biocontrol agents and botanicals in integrated management of Botrytis cinerea in table grape vineyards. Pest Manag. Sci. 2018, 74, 715–725. [Google Scholar] [CrossRef] [PubMed]
- Walker, G.M.; McLeod, A.H.; Hodgson, V.J. Interactions between killer yeasts and pathogenic fungi. FEMS Microbiol. Lett. 1995, 127, 213–222. [Google Scholar] [CrossRef] [PubMed]
- Druvefors, Ä.U.; Schnürer, J. Mold-inhibitory activity of different yeast species during airtight storage of wheat grain. FEMS Yeast Res. 2005, 5, 373–378. [Google Scholar] [CrossRef] [PubMed]
- Rosa-Magri, M.M.; Tauk-Tornisielo, S.M.; Ceccato-Antonini, S.R. Bioprospection of yeasts as biocontrol agents phytopathogenic molds. Braz. Arch. Biol. Biotechnol. 2011, 54, 1–5. [Google Scholar] [CrossRef]
- Sundh, I.; Melin, P. Safety and regulation of yeasts used for biocontrol or biopreservation in the food and feed chain. Antonie Van Leeuwenhoek 2011, 99, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Oro, L.; Feliziani, E.; Ciani, M. Biocontrol of postharvest brown rot of sweet cherries and population dynamic of Saccharomyces cerevisiae Disva599, Metschnikowia pulcherrima Disva267 and Wickerhamomyces anomalus Disva2 strains. Postharvest Biol. Technol. 2014, 96, 64–68. [Google Scholar] [CrossRef]
- Parafati, L.; Vitale, A.; Restuccia, C.; Circilleri, G. Biocontrol ability and action mechanism of food-isolated yeast strains against Botrytis cinerea causing post-harvest bunch rot of table grape. Food Microbiol. 2015, 47, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Cordero-Bueso, G.; Mangieri, N.; Maghradze, D.; Foschino, R.; Valdetara, F.; Cantoral, J.M.; Vigentini, I. Wild grape-associated yeasts as promising agents against Vitis vinifera fungal pathogens. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Grzegorczyk, M.; Żarowska, B.; Restuccia, C.; Cirvilleri, G. Postharvest biocontrol ability of killer yeasts against Monilinia fructigena and Monilinia fructicola on stone fruit. Food Microbiol. 2017, 61, 93–101. [Google Scholar] [CrossRef] [PubMed]
- Bisson, L.F.; Joseph, C.M.L.; Domizio, P. Yeasts. In Biology of Microorganisms on Grapes, in Must and in Wine; König, H., Unden, F., Fröhlich, J., Eds.; Springer International Publishing AG: Basel, Switzerland, 2017; pp. 65–101. [Google Scholar]
- São-José, C.; Santos, M.A.; Schmitt, M.J. Viruses of wine-associated yeasts and bacteria. In Biology of Microorganisms on Grapes, in Must and in Wine; König, H., Unden, G., Fröhlich, J., Eds.; Springer International Publishing AG: Basel, Switzerland, 2017; pp. 133–153. [Google Scholar]
- Hua, S.S.; Beck, J.J.; Sarreal, S.B.; Gee, W. The major volatile compound 2-phenylethanol from the biocontrol yeast, Pichia anomala, inhibits growth and expression of aflatoxin biosynthetic genes of Aspergillus flavus. Mycotoxin Res. 2014, 30, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Jijakli, M.H. Pichia anomala in biocontrol for apples: 20 years of fundamental research and practical applications. Antonie Van Leeuwenhoek 2011, 99, 93–105. [Google Scholar]
- Prillinger, H.; Messner, R.; König, H.; Bauer, R.; Lopandic, K.; Molnar, O.; Dangel, P.; Weigang, F.; Kiristis, T.; Nakase, T.; et al. Yeasts associated with termites: A phenotypic and genotypic characterization and use of coevolution for dating evolutionary radiations in Asco- and Basidiomycetes. Syst. Appl. Microbiol. 1996, 19, 265–283. [Google Scholar] [CrossRef]
- Bäumlisberger, M.; Moellecken, U.; König, H.; Claus, H. The potential of the yeast Debaryomyces hansenii H525 to degrade biogenic amines in food. Microorganisms 2015, 3, 839–850. [Google Scholar] [CrossRef] [PubMed]
- Radler, F.; Pfeiffer, P.; Dennert, M. Killer toxins in new isolates of the yeasts Hanseniaspora uvarum and Pichia kluyveri. FEMS Microbiol. Lett. 1985, 29, 269–272. [Google Scholar]
- Zorg, J.; Kilian, S.; Radler, F. Killer toxin producing strains of the yeasts Hanseniaspora uvarum and Pichia kluyveri. Arch. Microbiol. 1988, 149, 261–267. [Google Scholar] [CrossRef]
- Schlander, M.; Distler, U.; Tenzer, S.; Thines, E.; Claus, H. Purification and properties of yeast proteases secreted by Wickerhamomyces anomalus 227 and Metschnikowia pulcherrima 446 during growth in a white grape juice. Fermentation 2017, 3, 2. [Google Scholar] [CrossRef]
- Sabel, A.; Martens, S.; Petri, A.; König, H.; Claus, H. Wickerhamomyces anomalus AS1: A new strain with potential to improve wine aroma. Ann. Microbiol. 2014, 64, 483–491. [Google Scholar] [CrossRef]
- Schwentke, J.; Sabel, A.; Petri, A.; König, H.; Claus, H. The wine yeast Wickerhamomyces anomalus AS1 secretes a multifunctional exo-ß-1,3 glucanse with implications for winemaking. Yeast 2014, 31, 349–359. [Google Scholar] [CrossRef] [PubMed]
- Pfannebecker, J.; Schiffer-Hetz, C.; Fröhlich, J.; Becker, B. Culture medium optimization for osmotolerant yeasts by use of a papallel fermenter system and rapid microbiological testing. J. Microbiol. Methods 2016, 130, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Molnar, O.; Schatzmayr, G.; Fuchs, E.; Prillinger, H. Trichosporon mycotoxinivorans sp. nov., a new yeast species useful in biological detoxification of various mycotoxins. Syst. Appl. Microbiol. 2004, 27, 661–671. [Google Scholar] [CrossRef] [PubMed]
- Handel, S.; Wang, T.; Yurkov, A.M.; König, H. Sugiyamaella mastotermitis sp. nov. and Papiliotrema odontotermitis f.a., sp. nov. from the gut of the temites Mastotermes darwiniensis and Odontotermes obesus. Int. J. Syst. Evol. Microbiol. 2016, 66, 4600–4608. [Google Scholar] [PubMed]
- Charoenchai, C.; Fleet, F.H.; Henschke, P.A.; Todd, B.E.N. Screening of non-Saccharomyces wine yeasts for the presence of extracellular hydrolytic enzymes. Aust. J. Grape Wine Res. 1997, 3, 2–8. [Google Scholar] [CrossRef]
- Fernández, M.; Úbeda, J.F.; Briones, A.I. Typing of non-Saccharomyces yeasts with enzymatic activities of interest in wine-making. Int. J. Food Microbiol. 2000, 59, 29–36. [Google Scholar] [CrossRef]
- Strauss, M.L.A.; Jolly, N.P.; Lambrechts, M.G.; van Rensburg, P. Screening for the production of extracellular hydrolytic enzymes by non-Saccharomyces wine yeasts. J. Appl. Microbiol. 2001, 91, 182–190. [Google Scholar] [CrossRef] [PubMed]
- Raspor, P.; Miklič-Milek, D.; Avbely, M.; Čadež, N. Biocontrol of grey mould disease on grape caused by Botrytis cinerea with autochthonous wine yeasts. Food Technol. Biotechnol. 2010, 48, 336–343. [Google Scholar]
- Sipiczki, M. Metschnikowia strains isolated from botrytized grapes antagonize fungal and bacterial growth by iron depletion. Appl. Environ. Microbiol. 2006, 72, 6716–6724. [Google Scholar] [CrossRef] [PubMed]
- Saravanakumar, D.; Ciavorella, A.; Spadaro, D.; Garibaldi, A.; Gullino, M.L. Metschnikowia pulcherrima strain MACH1 outcompetes Botrytis cinerea, Alternaria alternata and Penicillium expansum in apples through iron depletion. Postharvest Biol. Technol. 2008, 49, 121–128. [Google Scholar] [CrossRef]
- Chan, Z.; Tian, S. Interaction of antagonistic yeasts against postharvest pathogens of apple fruit and possible mode of action. Postharvest Biol. Technol. 2005, 36, 215–223. [Google Scholar] [CrossRef]
- Zhang, D.; Spadaro, D.; Garibaldi, A.; Gullino, M.L. Efficacy of the antagonist Aureobasidium pullulans PL5 against postharvest pathogens of peach, apple and plum and its modes of action. Biol. Control 2010, 54, 172–180. [Google Scholar] [CrossRef]
- Walker, A.S. Diversity within and between species of Botrytis. In Botrytis—The Fungus, the Pathogen and Its Management in Agricultural Systems; Fillinger, S., Elad, Y., Eds.; Springer International Publishing: Basel, Switzerland, 2016; pp. 91–125. [Google Scholar]
- Sláviková, E.; Vadkertiková, R.; Vránová, D. Yeasts colonizing the leaf surfaces. J. Basic Microbiol. 2007, 47, 344–350. [Google Scholar] [CrossRef] [PubMed]
- Janisiewicz, W.J.; Jurick, W.M.; Peter, K.A.; Kurtzman, C.; Buyer, J. Yeast associated with plum and their potential for controlling brown rot after harvest. Yeast 2014, 31, 207–218. [Google Scholar] [CrossRef] [PubMed]
- Molnárova, J.; Vadkertiová, R.; Stratilová, E. Extracellular enzymatic activities and physiological profiles of yeasts colonizing fruit trees. J. Basic Microbiol. 2014, 54, S74–S84. [Google Scholar] [CrossRef] [PubMed]
- Varela, C.; Borneman, A.R. Yeasts found in vineyards and wineries. Yeast 2017, 34, 111–128. [Google Scholar] [CrossRef] [PubMed]
- Spadaro, D.; Vola, R.; Piano, S.; Gullino, M.L. Mechanisms of action and efficacy of four isolates of the yeast Metschnikowia pulcherrima active against postharvest pathogens on apples. Postharvest Biol. Technol. 2002, 24, 123–134. [Google Scholar] [CrossRef]
- Oro, L.; Feliziani, E.; Ciani, M.; Romanazzi, G.; Comitini, F. Volatile organic compounds from Wickerhamomyces anomalus, Metschnikowia pulcherrima and Saccharomyces cerevisiae inhibit growth of decay causing fungi and control postharvest diseases of strawberries. Int. J. Food Microbiol. 2018, 265, 18–22. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.Q.; Li, W.; Linag, Z.T.; Jing, M.M.; Shao, Y.Z. The preservation effect of Metschnikowia pulcherrima yeast on anthracnose of postharvest mango fruits and the possible mechanism. Food Sci. Biotechnol. 2018, 27, 95–105. [Google Scholar] [CrossRef]
- Sisti, M.; Savani, V. Antifungal properties of the human Metschnikowia strain IHEM 25107. Folia Microbiol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Hershkovitz, V.; Sela, N.; Taha-Salaime, L.; Liu, J.; Rafael, G.; Kessler, C.; Aly, R.; Levy, M.; Wisniewski, M.; Droby, S. De-novo assembly and characterization of the transcriptome of Metschnikowia fructicola reveals differences in gene expression following interaction with Penicillium digitatum and grapefruit peel. MMC Genom. 2013, 14, 168. [Google Scholar] [CrossRef] [PubMed]
- Spadaro, D.; Lorè, A.; Garibaldi, A.; Gullino, M.L. A new strain of Metschnikowia fructicola for postharvest control of Penicillium expansum and patulin accumulation on four cultivars of apple. Postharvest Biol. Technol. 2013, 75, 1–8. [Google Scholar] [CrossRef]
- Kurtzman, C.P.; Droby, S. Metschnikowia fructicola, a new ascosporic yeast with potential for biocontrol of postharvest fruit rots. Syst. Appl. Microbiol. 2001, 24, 395–399. [Google Scholar] [CrossRef] [PubMed]
- Lachance, M.A. Metschnikowia: Half tetrads, a regicide and the fountain of youth. Yeast 2016, 33, 563–574. [Google Scholar] [CrossRef] [PubMed]
- Piombo, E.; Sela, N.; Wisniewski, M.; Hoffmann, M.; Gullino, M.L.; Allard, M.W.; Levin, E.; Spadaro, D.; Droby, S. Genome sequence, assembly and characterization of two Metschnikowia fructicola strains used as biocontrol agents of postharvest diseases. Front. Microbiol. 2018, 9, 593. [Google Scholar] [CrossRef] [PubMed]
- Türkel, S.; Ener, B. Isolation and characterization of new Metschnikowia pulcherrima strains as producers of the antimicrobial pigment pulcherrimin. Z. Naturforsch. 2009, 64, 405–410. [Google Scholar] [CrossRef]
- Banani, H.; Spadaro, D.; Zhang, D.; Matic, S.; Garibaldi, A.; Gullino, M.L. Postharvest application of a novel chitinase from Metschnikowia fructicola and overexpressed in Pichia pastoris to control brown rot of peaches. Int. J. Food Microbiol. 2015, 199, 54–61. [Google Scholar] [CrossRef] [PubMed]
- Saravanakumar, D.; Spadaro, D.; Garibaldi, A.; Gullino, M.L. Detection of enzymatic activity and partial sequence of a chitinase in Metschnikowia pulcherrima strain MACH1 used as postharvest biocontrol agent. Eur. J. Plant Pathol. 2009, 123, 183–193. [Google Scholar] [CrossRef]
- Grevesse, C.; Lepoivre, P.; Jijakli, M.H. Characterization of the exoglucanase-encoding gene PaEXG2 and study of its role in the biocontrol activity of Pichia anomala strain K. Phytopathology 2003, 93, 1145–1152. [Google Scholar] [CrossRef] [PubMed]
- Friel, D.; Gomez Pessoa, N.M.; Vandenbol, M.; Jijakli, M.H. Separate and combined disruptions of two exo-ß-1,3-glucanase genes decrease the efficiency of Pichia anomala (Strain K) biocontrol against Botrytis cinerea on apple. Mol. Plant Microbe Interact. 2007, 20, 371–379. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Spadaro, D.; Valente, S.; Garibaldi, A.; Gullino, M.L. Cloning, characterization and expression of an exo-1,3-ß-glucanase gene from the antagonistic yeast, Pichia guilliermondii strain M8 against grey mold on apples. Biol. Control 2011, 59, 284–293. [Google Scholar] [CrossRef]
- Zhang, D.; Spadaro, D.; Valente, S.; Garibaldi, A.; Gullino, M.L. Cloning, characterization, expression and antifungal activity of an alkaline serine protease of Aureobasidium pullulans PL5 involved in the biological control of pathogens. J. Food Microbiol. 2012, 153, 453–464. [Google Scholar] [CrossRef] [PubMed]
- Salazar, O.; Asenjo, J.A. Enzymatic lysis of microbial cells. Biotechnol. Lett. 2007, 29, 985–994. [Google Scholar] [CrossRef] [PubMed]
- König, H.; Li, L.; Fröhlich, J. The cellulolytic system of the termite gut. Appl. Microbiol. Biotechnol. 2013, 97, 7943–7962. [Google Scholar] [CrossRef] [PubMed]
- Khalel, A.S.; Khaled, J.M.; Kandeal, S.A. Enzymatic activity and some molecular properties of Trichosporon mycotoxinovorans yeast and their effect on liver function in mice. Afr. J. Microbiol. Res. 2012, 6, 2567–2573. [Google Scholar]
- Bourdichon, F.; Casarogola, S.; Farrokh, C.; Frisvad, J.C.; Gerds, M.L.; Hammes, W.P.; Harnett, J.; Huys, G.; Laulund, S.; Ouwehand, A.; et al. Food fermentations: Microorganisms with technological beneficial use. Int. J. Food Microbiol. 2012, 154, 87–97. [Google Scholar] [CrossRef] [PubMed]


| Species/Strain | Habitat | References |
|---|---|---|
| Candida sake 2/42 | roach | [26] |
| Debaryomyces hansenii var. fabryii 525 | cherry | [27] |
| Hanseniapora uvarum 469 | pear | [28] |
| Hanseniapora uvarum 470 (ATCC 64295) | raspberry | [28] |
| Hanseniapora uvarum 471 | peach | [28] |
| Hanseniapora uvarum 473 | apple | |
| Hanseniapora uvarum 486 | grape | |
| Hanseniapora uvarum 527 | grape | [29] |
| Kluyveromyces marxianus 118 | grape | |
| Kluyveromyces thermotolerans 76 | grape (Crete) | |
| Metschnikowia pulcherrima 152 | apple | |
| Metschnikowia pulcherrima 160 | grape | |
| Metschnikowia pulcherrima 192 | unknown | |
| Metschnikowia pulcherrima 2305 | unknown | |
| Metschnikowia pulcherrima 446 | grape | [30] |
| Metschnikowia pulcherrima 523 | grape | |
| Metschnikowia pulcherrima 648 | grape | |
| Pichia anomala 457 (NCYC 435) | unknown | [15] |
| Pichia kluyveri 2143 | apple | |
| Pichia kluyveri var. kluyveri 395 (CBS 7145, ATCC 66811) | grape | [29] |
| Pichia methanolica H1/3-1 | cherry | |
| Torulaspora delbrueckii 3/40 | cherry | |
| Wickerhamomyces anomalus 15 | apple | |
| Wickerhamomyces anomalus 227 | grape | [30] |
| Wickerhamomyces anomalus AS1 (DSM 28943) | grape | [31,32] |
| Wickerhamomyces anomalus H.3.2 | wheat flour (Spain) | |
| Wickerhamomyces anomalus WH 1021 | praline | [33] |
| Williopsis californica 3/62 | cherry | |
| Williopsis saturnus var. makrii 458 (NCYC 500) | soil (Papua New Guinea) | [15] |
| Zygosaccharomyces bailii 412 (CBS 6708) | orange (Brasil) | |
| Zygosaccharomyces bailii 550 | grape | |
| Apiotrichum mycotoxinovorans MYG | termite | [34,35] |
| Apiotrichum mycotoxinovorans MD123D | termite | [34,35] |
| Naganishia albida OO1 | termite | [35] |
| Papiliotrema odontotermitis OO5 (DSM 100791T, CBS 14181T) | termite | [35] |
| Saitozyma flava OO2 | termite | [35] |
| Sugiyamaella mastotermitis MD39V (DSM 100793T, CBS 14182T) | termite | [35] |
| Sugiyamaella smithiae NM1 | termite | [35] |
| Fungal Strain | Habitat | Source |
|---|---|---|
| Botrytis cinerea V15 | Vitis vinifera | ISA |
| Botrytis cinerea V19 | Vitis vinifera | ISA |
| Botrytis cinerea V27 | Vitis vinifera | ISA |
| Botrytis cinerea B16-14 | Vitis vinifera | ISA |
| Botrytis cinerea DSM 877 | unknown | DSMZ |
| Monilinia fructigena DSM 2677 | Malus | DSMZ |
| Magnaporthe oryzae 7015 | Oryza sativa | IBWF |
| Eutypa lata 1190 | Vitis vinifera | IMWF |
| Eutypa lata 16012 | Vitis vinfera | IBWF |
| Roesleria subterranea CBS 201.25 | Vitis vinifera | CBS |
| Roesleria subterranea CBS 271.82 | Populus | CBS |
| Roesleria subterranea CBS 320.33 | Malus | CBS |
| Roesleria subterranea CBS 339.96 | shrub | CBS |
| Fungal Strain | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Yeast Strain | B. cinerea V15 | B. cinerea V19 | B. cinerea V27 | B. cinerea 16-14 | B. cinerea DSM 877 | M. fructigena DSM 2677 | M. oryzae 7015 | E. lata 1190 | E. lata 16012 | R. subterranea CBS 271.82 | R. subterranea CBS 320.33 | R. subterranea CBS 339.96 |
| M. pulcherrima 160 | + | − | − | ++ | ++ | ++ | +++ | + | nd | ++ | ++ | ++ |
| M. pulcherrima 446 | ++ | − | − | ++ | +++ | ++ | +++ | + | nd | +++ | ++ | +++ |
| M. pulcherrima 648 | +++ | − | + | ++ | +++ | ++ | +++ | − | nd | + | + | + |
| M. pulcherrima 152 | +++ | − | + | +++ | +++ | ++ | ++ | − | nd | +++ | +++ | +++ |
| M. pulcherrima 192 | +++ | − | + | ++ | +++ | ++ | ++ | − | nd | ++ | − | − |
| M. pulcherrima 523 | + | − | − | + | ++ | + | ++ | − | nd | + | − | − |
| M. pulcherrima 2305 | + | − | − | ++ | ++ | + | +++ | + | nd | +++ | ++ | + |
| C. sake 2/42 | - | − | + | + | +++ | + | − | − | − | ++ | + | − |
| P. anomala 457 | +++ | − | + | ++ | − | +++ | ++ | ++ | +++ | nd | nd | − |
| P. kluyveri 2143 | ++ | − | + | ++ | +++ | − | +++ | − | − | − | − | − |
| P. kluyveri 395 | ++ | − | − | ++ | − | − | +++ | − | − | + | − | + |
| P. methanolica H1/3-1 | + | − | +++ | + | ++ | − | +++ | − | − | + | nd | − |
| Z. bailii 412 | + | − | − | + | +++ | + | +++ | − | − | + | − | + |
| Z. bailii 550 | − | − | − | − | + | − | +++ | + | − | + | − | − |
| W. anomalus AS1 | − | − | +++ | ++ | +++ | ++ | +++ | − | − | ++ | +++ | +++ |
| W. anomalus H.3.2 | − | − | +++ | +++ | +++ | + | +++ | − | − | + | ++ | + |
| W. anomalus WH 1021 | − | − | +++ | ++ | +++ | + | +++ | − | − | + | + | +++ |
| W. anomalus 15 | + | − | +++ | ++ | +++ | + | +++ | − | − | ++ | ++ | +++ |
| W. anomalus 227 | + | + | − | ++ | +++ | − | +++ | − | − | ++ | +++ | +++ |
| W. californica 3/62 | − | − | ++ | + | + | ++ | − | − | − | − | ++ | ++ |
| W. saturnus 458 | − | − | − | + | ++ | ++ | − | − | − | + | + | − |
| T. delbrueckii 3/40 | − | − | +++ | + | ++ | − | +++ | − | − | − | − | − |
| K. thermotolerans 76 | − | − | − | − | − | − | − | − | − | − | − | − |
| K. marxianus 118 | − | − | + | − | + | + | − | − | − | − | + | − |
| H. uvarum 469 | − | − | ++ | + | + | + | ++ | ++ | + | ++ | − | − |
| H. uvarum 470 | − | − | + | − | + | ++ | − | − | − | − | − | − |
| H. uvarum 471 | − | − | ++ | − | ++ | ++ | ++ | + | ++ | − | − | − |
| H. uvarum 473 | − | − | ++ | + | + | + | ++ | + | ++ | + | − | − |
| H. uvarum 486 | − | − | − | − | ++ | + | − | + | + | − | − | − |
| H. uvarum 527 | − | − | + | − | ++ | + | − | + | + | − | − | − |
| D. hansenii 525 | − | − | + | − | + | − | − | − | − | − | − | − |
| A. mycotoxinovorans MYG | − | + | + | − | + | − | − | − | − | − | − | − |
| A. mycotoxinovorans MD123D | − | − | + | − | + | − | − | − | − | − | − | − |
| N. albida OO1 | − | − | + | + | + | + | − | − | − | − | − | + |
| P. odontotermitis OO5 | + | − | + | ++ | + | + | − | − | − | − | − | − |
| S. flava OO2 | + | − | + | + | + | − | − | − | − | − | − | − |
| S. mastotemitis MD39V | − | − | + | + | + | − | − | − | − | − | − | + |
| S. smithiae NM1 | − | − | + | − | + | − | + | − | − | − | − | − |
| Secreted Enzymatic Activity | |||||
|---|---|---|---|---|---|
| Yeast | Amylase | Cellulase | β-Glucosidase | Chitinase | Protease |
| M. pulcherrima 160 | ++ | − | +++ | + | ++ |
| M. pulcherrima 446 | + | − | +++ | + | ++ |
| M. pulcherrima 648 | +++ | − | +++ | + | +++ |
| M. pulcherrima 152 | + | − | +++ | + | + |
| M. pulcherrima 192 | + | − | +++ | + | ++ |
| M. pulcherrima 523 | + | − | +++ | + | +++ |
| M. pulcherrima 2305 | ++ | − | +++ | + | ++ |
| C. sake 2/42 | +++ | − | ++ | − | + |
| P. anomala 457 | +++ | − | ++ | + | − |
| P. kluyveri 2143 | ++ | − | ++ | − | + |
| P. kluyveri 395 | ++ | + | + | − | ++ |
| P. methanolica H1/3-1 | +++ | − | +++ | +++ | +++ |
| Z. bailii 412 | ++ | + | ++ | − | ++ |
| Z. bailii 550 | + | − | + | − | − |
| W. anomalus AS1 | + | + | + | + | +++ |
| W. anomalus H.3.2 | + | + | + | + | ++ |
| W. anomalus WH 1021 | + | + | + | − | ++ |
| W. anomalus 15 | + | + | + | +++ | ++ |
| W. anomalus 227 | + | + | + | ++ | +++ |
| W. californica 3/62 | + | + | + | − | + |
| W. saturnus 458 | + | + | + | ++ | − |
| T. delbrueckii 3/40 | + | + | + | +++ | + |
| K. thermotolerans 76 | + | + | + | + | + |
| K. marxianus 118 | + | − | + | + | − |
| H. uvarum 469 | − | + | + | + | + |
| H. uvarum 470 | − | + | + | − | + |
| H. uvarum 471 | − | + | + | + | − |
| H. uvarum 473 | − | + | + | + | + |
| H. uvarum 486 | − | − | + | − | + |
| H. uvarum 527 | − | + | + | + | + |
| D. hansenii 525 | ++ | + | + | − | + |
| A. mycotoxinovorans MYG | +++ | − | ++ | − | ++ |
| A. mycotoxinovorans MD123D | +++ | − | +++ | − | + |
| N. albida OO1 | +++ | + | ++ | − | + |
| P. odontotermitis OO5 | +++ | + | ++ | + | ++ |
| S. flava OO2 | − | − | + | + | − |
| S. mastotermitis MD39V | +++ | + | ++ | − | + |
| S. smithiae NM1 | + | + | + | + | + |
| Fungus | |||||
| B. cinerea V15 | + | − | + | + | − |
| B. cinerea V19 | + | − | + | + | − |
| B. cinerea V27 | + | − | − | + | + |
| B. cinerea 16-14 | + | − | + | + | + |
| B. cinerea 877 | + | − | + | + | + |
| M. fructigena DSM 2677 | + | − | + | + | + |
| M. oryzae 7015 | + | − | + | + | + |
| E. lata 1190 | + | − | + | + | + |
| E. lata 16012 | + | − | + | + | + |
| R. subterranea CBS 271.82 | + | − | + | + | + |
| R. subterranea CBS 320.33 | + | − | + | + | − |
| R. subterranea CBS 339.96 | + | − | + | + | − |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pretscher, J.; Fischkal, T.; Branscheidt, S.; Jäger, L.; Kahl, S.; Schlander, M.; Thines, E.; Claus, H. Yeasts from Different Habitats and Their Potential as Biocontrol Agents. Fermentation 2018, 4, 31. https://doi.org/10.3390/fermentation4020031
Pretscher J, Fischkal T, Branscheidt S, Jäger L, Kahl S, Schlander M, Thines E, Claus H. Yeasts from Different Habitats and Their Potential as Biocontrol Agents. Fermentation. 2018; 4(2):31. https://doi.org/10.3390/fermentation4020031
Chicago/Turabian StylePretscher, Julia, Tilman Fischkal, Sina Branscheidt, Lucas Jäger, Susann Kahl, Martina Schlander, Eckhard Thines, and Harald Claus. 2018. "Yeasts from Different Habitats and Their Potential as Biocontrol Agents" Fermentation 4, no. 2: 31. https://doi.org/10.3390/fermentation4020031





