Long Non-Coding RNAs Dysregulation and Function in Glioblastoma Stem Cells
Abstract
:1. Introduction
2. GSCs-Definitions, Characteristics and Biomarkers
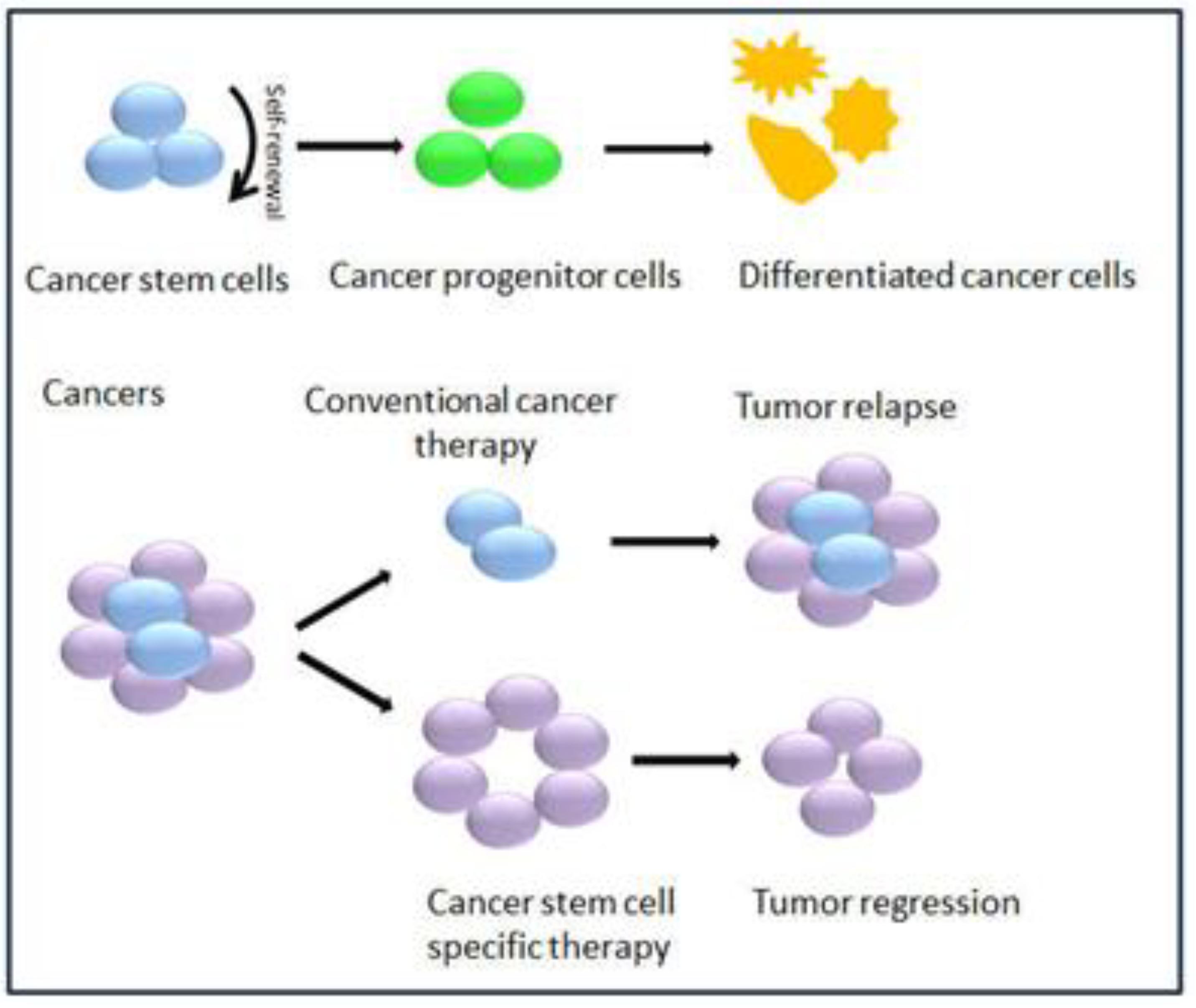
3. LncRNAs Dysregulated in GSCs
| Authors 1 | Year | Samples | No. of | No. of | Ref. |
|---|---|---|---|---|---|
| Dysregulated LncRNAs between GSCs and Differentiated GBM Cells Comparison | |||||
| Araki et al. | 2013 | GSC (sphere) vs. differentiated GBM cells | 6: LOC100127888, H19, RP11-112J3.16, et al. | 28: DLX6-AS, LOC643763, FLJ39609, et al. | [47] |
| Aldaz et al. | 2013 | GSC (sphere) vs. differentiated GBM cells | 28: H19, MIAT, LOC150622, LOC100127888, XIST, RP11-112J3.16, et al. | 11: RP11-346D6.6, C6orf155, HCG4, FLJ39609, et al. | [42] |
| Dysregulated LncRNAs between GSCs with Different Subtypes | |||||
| Beier et al. | 2007 | CD133 + GSCs vs. CD133 − GSCs | 38: XIST, H19, HOTAIR, LOC100192378, AC006213.1, MIAT, et al. | 34: CTC-231O11.1, RP11-745C15.2, LOC100130776, C14orf139, et al. | [40] |
| Gunther et al. | 2008 | CD133+ GSCs vs. CD133 − GSCs | 51: H19, RP11-331K15.1, RP11-547I7.2, LOC100192378, MIAT, HOTAIR, et al. | 10: C14orf139, DLX6-AS, MIR155HG, LOC100130776, et al. | [41] |
| Dysregulated LncRNAs between GSCs and NSCs Comparison | |||||
| Rheinbay, et al. | 2013 | GSCs vs. NSCs | 173: LOC399959, LOC645323, HOTAIRM1, H19, MALAT1, SOX2ot, et al. | 19: HYMAI, AL133167.1, FLJ31485, et al. | [58] |
3.1. Dysregulated LncRNAs during GSC Differentiation
3.2. Dysregulated LncRNAs between GSCs with Different Subtypes
3.3. Dysregulated LncRNAs between GSCs and Neural Stem Cells (NSCs)
4. Functional Roles and Molecular Mechanism of LncRNAs in GSCs
4.1. Interaction with Stem Cell Transcription Factors (TFs)

| LncRNAs 2 | Interactive TFs 3 (Number of Binding Sites) | Interactive miRNAs 3 | Interactive RBPs 3 |
|---|---|---|---|
| H19 | NFKB (39), E2F (30), c-Myc (45), CTCF (60) | miR-29a, miR-29b, miR-29c, miR-18a, miR-19a, miR-20a, miR-19b, et al. | NA |
| MIAT | NFKB (45), E2F (43), Nanog (27), SMAD (20), c-Myc (15), Oct-4 (5), CTCF (20) | miR-29a, miR-29b, miR-29c, and miR-150 | NA |
| XIST | NFKB (18), TAF1 (14), c-Myc (8), CTCF (8), Nanog (2), HNF4A(6) | miR-124, miR-34a, miR-137, miR-146a, miR-326, miR-7 and miR-425, miR-152, let-7, et al. | LIN28, IGF2BP |
| LOC100127888 | NA | NA | NA |
| RP11-112J3.16 | HNF4A (1) | NA | NA |
| LOC643763 | CTCF (3), c-Myc (1), NFKB (1), Nanog (1) | NA | NA |
| RP11-346D6.6 | CDX2(2), GATA6 (3), HNF4A (3), Nanog (1) | NA | NA |
| FLJ39609 | CDX2(2), c-Myc (1), USF-1 (4) | NA | NA |
| C6orf155 | CDX2(5), E2F (10), HNF4A (4), Nanog (5), SMAD (5) | NA | NA |
| HCG4 | NA | NA | NA |
| DLX6-AS | NA | NA | NA |
4.2. Interaction with MiRNAs
4.3. Interaction with NFKB Pathways
4.4. Interaction with Other Molecules or Pathways
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lobo, N.A.; Shimono, Y.; Qian, D.; Clarke, M.F. The biology of cancer stem cells. Ann. Rev. Cell Dev. Biol. 2007, 23, 675–699. [Google Scholar] [CrossRef] [PubMed]
- Clevers, H. The cancer stem cell: Premises, promises and challenges. Nat. Med. 2011, 17, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Frank, N.Y.; Schatton, T.; Frank, M.H. The therapeutic promise of the cancer stem cell concept. J. Clin. Investig. 2010, 120, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Huang, Y.H.; Chen, J.L. Understanding and targeting cancer stem cells: Therapeutic implications and challenges. Acta Pharmacol. Sin. 2013, 34, 732–740. [Google Scholar] [CrossRef] [PubMed]
- Hemmati, H.D.; Nakano, I.; Lazareff, J.A.; Masterman-Smith, M.; Geschwind, D.H.; Bronner-Fraser, M.; Kornblum, H.I. Cancerous stem cells can arise from pediatric brain tumors. PNAS 2003, 100, 15178–15183. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.K.; Hawkins, C.; Clarke, I.D.; Squire, J.A.; Bayani, J.; Hide, T.; Henkelman, R.M.; Cusimano, M.D.; Dirks, P.B. Identification of human brain tumour initiating cells. Nature 2004, 432, 396–401. [Google Scholar] [CrossRef] [PubMed]
- Galli, R.; Binda, E.; Orfanelli, U.; Cipelletti, B.; Gritti, A.; de Vitis, S.; Fiocco, R.; Foroni, C.; Dimeco, F.; Vescovi, A. Isolation and characterization of tumorigenic, stem-like neural precursors from human glioblastoma. Cancer Res. 2004, 64, 7011–7021. [Google Scholar] [CrossRef] [PubMed]
- Rich, J.N.; Eyler, C.E. Cancer stem cells in brain tumor biology. Cold Spring Harb. Symp. Quant. Biol. 2008, 73, 411–420. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, A.U.; Auffinger, B.; Lesniak, M.S. Understanding glioma stem cells: Rationale, clinical relevance and therapeutic strategies. Exp. Rev. Neurother. 2013, 13, 545–555. [Google Scholar] [CrossRef] [PubMed]
- Cho, D.Y.; Lin, S.Z.; Yang, W.K.; Lee, H.C.; Hsu, D.M.; Lin, H.L.; Chen, C.C.; Liu, C.L.; Lee, W.Y.; Ho, L.H. Targeting cancer stem cells for treatment of glioblastoma multiforme. Cell Transplant. 2013, 22, 731–739. [Google Scholar] [CrossRef] [PubMed]
- Frosina, G. Frontiers in targeting glioma stem cells. Eur. J. Cancer 2011, 47, 496–507. [Google Scholar] [CrossRef] [PubMed]
- Stupp, R.; Hegi, M.E. Targeting brain-tumor stem cells. Nat. Biotechnol. 2007, 25, 193–194. [Google Scholar] [CrossRef] [PubMed]
- Ciechomska, I.A.; Kocyk, M.; Kaminska, B. Glioblastoma stem-like cells-isolation, biology and mechanisms of chemotherapy resistance. Curr. Signal Transduct. Ther. 2013, 8, 256–267. [Google Scholar] [CrossRef]
- Yamada, K.; Tso, J.; Ye, F.; Choe, J.; Liu, Y.; Liau, L.M.; Tso, C.L. Essential gene pathways for glioblastoma stem cells: Clinical implications for prevention of tumor recurrence. Cancers 2011, 3, 1975–1995. [Google Scholar] [CrossRef] [PubMed]
- Gutschner, T.; Diederichs, S. The hallmarks of cancer: A long non-coding rna point of view. RNA Biol. 2012, 9, 703–719. [Google Scholar] [CrossRef] [PubMed]
- Gibb, E.A.; Brown, C.J.; Lam, W.L. The functional role of long non-coding RNA in human carcinomas. Mol. Cancer 2011, 10, 38. [Google Scholar] [CrossRef] [PubMed]
- Cao, J. The functional role of long non-coding RNAs and epigenetics. Biol. Proced. Online 2014, 16, 11. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.C.; Chang, H.Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Ren, Z.; Sun, P. Overexpression of the long non-coding RNA MEG3 impairs in vitro glioma cell proliferation. J. Cell. Biochem. 2012, 113, 1868–1874. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Sun, X.; Zhou, X.; Han, L.; Chen, L.; Shi, Z.; Zhang, A.; Ye, M.; Wang, Q.; Liu, C.; et al. Long non-coding RNA hotair promotes glioblastoma cell cycle progression in an EZH2 dependent manner. Oncotarget 2015, 6, 537–546. [Google Scholar] [PubMed]
- Shi, Y.; Wang, Y.; Luan, W.; Wang, P.; Tao, T.; Zhang, J.; Qian, J.; Liu, N.; You, Y. Long non-coding RNA H19 promotes glioma cell invasion by deriving MIR-675. PLoS ONE 2014, 9, e86295. [Google Scholar] [CrossRef] [PubMed]
- Ma, K.X.; Wang, H.J.; Li, X.R.; Li, T.; Su, G.; Yang, P.; Wu, J.W. Long noncoding RNA MALAT1 associates with the malignant status and poor prognosis in glioma. Tumour Biol. 2015, 36, 3355–3359. [Google Scholar] [CrossRef] [PubMed]
- Park, J.Y.; Lee, J.E.; Park, J.B.; Yoo, H.; Lee, S.H.; Kim, J.H. Roles of long non-coding RNAs on tumorigenesis and glioma development. Brain Tumor Res. Treat. 2014, 2, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Ma, J.; Xue, Y.; Wang, P.; Li, Z.; Liu, J.; Chen, L.; Xi, Z.; Teng, H.; Wang, Z.; et al. Knockdown of long non-coding RNA xist exerts tumor-suppressive functions in human glioblastoma stem cells by up-regulating MIR-152. Cancer Lett. 2015, 359, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Clarke, M.F.; Dick, J.E.; Dirks, P.B.; Eaves, C.J.; Jamieson, C.H.; Jones, D.L.; Visvader, J.; Weissman, I.L.; Wahl, G.M. Cancer stem cells-perspectives on current status and future directions: AACR workshop on cancer stem cells. Cancer Res. 2006, 66, 9339–9344. [Google Scholar] [CrossRef] [PubMed]
- Sampetrean, O.; Saya, H. Characteristics of glioma stem cells. Brain Tumor Pathol. 2013, 30, 209–214. [Google Scholar] [CrossRef] [PubMed]
- Gursel, D.B.; Shin, B.J.; Burkhardt, J.K.; Kesavabhotla, K.; Schlaff, C.D.; Boockvar, J.A. Glioblastoma stem-like cells-biology and therapeutic implications. Cancers 2011, 3, 2655–2666. [Google Scholar] [CrossRef] [PubMed]
- Eramo, A.; Ricci-Vitiani, L.; Zeuner, A.; Pallini, R.; Lotti, F.; Sette, G.; Pilozzi, E.; Larocca, L.M.; Peschle, C.; De Maria, R. Chemotherapy resistance of glioblastoma stem cells. Cell Death Differ. 2006, 13, 1238–1241. [Google Scholar] [CrossRef] [PubMed]
- Bao, S.; Wu, Q.; McLendon, R.E.; Hao, Y.; Shi, Q.; Hjelmeland, A.B.; Dewhirst, M.W.; Bigner, D.D.; Rich, J.N. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature 2006, 444, 756–760. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Kotliarova, S.; Kotliarov, Y.; Li, A.G.; Su, Q.; Donin, N.M.; Pastorino, S.; Purow, B.W.; Christopher, N.; Zhang, W.; et al. Tumor stem cells derived from glioblastomas cultured in BFGF and EGF more closely mirror the phenotype and genotype of primary tumors than do serum-cultured cell lines. Cancer Cell 2006, 9, 391–403. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Nguyen, D.H.; Dong, Q.; Shitaku, P.; Chung, K.; Liu, O.Y.; Tso, J.L.; Liu, J.Y.; Konkankit, V.; Cloughesy, T.F.; et al. Molecular properties of CD133 + glioblastoma stem cells derived from treatment-refractory recurrent brain tumors. J. Neuro-Oncol. 2009, 94, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Pointer, K.B.; Clark, P.A.; Zorniak, M.; Alrfaei, B.M.; Kuo, J.S. Glioblastoma cancer stem cells: Biomarker and therapeutic advances. Neurochem. Int. 2014, 71, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Stiles, C.D.; Rowitch, D.H. Glioma stem cells: A midterm exam. Neuron 2008, 58, 832–846. [Google Scholar] [CrossRef] [PubMed]
- Chu, P.M.; Ma, H.I.; Chen, L.H.; Chen, M.T.; Huang, P.I.; Lin, S.Z.; Chiou, S.H. Deregulated microRNAs identified in isolated glioblastoma stem cells: An overview. Cell Transplant. 2013, 22, 741–753. [Google Scholar] [CrossRef] [PubMed]
- Mizrak, D.; Brittan, M.; Alison, M. CD133: Molecule of the moment. J. Pathol. 2008, 214, 3–9. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.J.; Wu, P.Y. CD133 as a marker for cancer stem cells: Progresses and concerns. Stem. Cells Dev. 2009, 18, 1127–1134. [Google Scholar] [CrossRef] [PubMed]
- Yuan, X.; Curtin, J.; Xiong, Y.; Liu, G.; Waschsmann-Hogiu, S.; Farkas, D.L.; Black, K.L.; Yu, J.S. Isolation of cancer stem cells from adult glioblastoma multiforme. Oncogene 2004, 23, 9392–9400. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Yuan, X.; Zeng, Z.; Tunici, P.; Ng, H.; Abdulkadir, I.R.; Lu, L.; Irvin, D.; Black, K.L.; Yu, J.S. Analysis of gene expression and chemoresistance of CD133 + cancer stem cells in glioblastoma. Mol. Cancer 2006, 5, 67. [Google Scholar] [CrossRef] [PubMed]
- Beier, D.; Hau, P.; Proescholdt, M.; Lohmeier, A.; Wischhusen, J.; Oefner, P.J.; Aigner, L.; Brawanski, A.; Bogdahn, U.; Beier, C.P. CD133(+) and CD133(−) glioblastoma-derived cancer stem cells show differential growth characteristics and molecular profiles. Cancer Res. 2007, 67, 4010–4015. [Google Scholar] [CrossRef] [PubMed]
- Gunther, H.S.; Schmidt, N.O.; Phillips, H.S.; Kemming, D.; Kharbanda, S.; Soriano, R.; Modrusan, Z.; Meissner, H.; Westphal, M.; Lamszus, K. Glioblastoma-derived stem cell-enriched cultures form distinct subgroups according to molecular and phenotypic criteria. Oncogene 2008, 27, 2897–2909. [Google Scholar] [CrossRef] [PubMed]
- Aldaz, B.; Sagardoy, A.; Nogueira, L.; Guruceaga, E.; Grande, L.; Huse, J.T.; Aznar, M.A.; Diez-Valle, R.; Tejada-Solis, S.; Alonso, M.M.; et al. Involvement of miRNAs in the differentiation of human glioblastoma multiforme stem-like cells. PLoS ONE 2013, 8, e77098. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Sun, S.; Pu, J.K.; Tsang, A.C.; Lee, D.; Man, V.O.; Lui, W.M.; Wong, S.T.; Leung, G.K. Long non-coding RNA expression profiles predict clinical phenotypes in glioma. Neurobiol. Dis. 2012, 48, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.Q.; Sun, S.; Lam, K.F.; Kiang, K.M.; Pu, J.K.; Ho, A.S.; Lui, W.M.; Fung, C.F.; Wong, T.S.; Leung, G.K. A long non-coding RNA signature in glioblastoma multiforme predicts survival. Neurobiol. Dis. 2013, 58, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Michelhaugh, S.K.; Lipovich, L.; Blythe, J.; Jia, H.; Kapatos, G.; Bannon, M.J. Mining affymetrix microarray data for long non-coding RNAs: Altered expression in the nucleus accumbens of heroin abusers. J. Neurochem. 2011, 116, 459–466. [Google Scholar] [CrossRef] [PubMed]
- Johnson, R. Long non-coding RNAs in huntington’s disease neurodegeneration. Neurobiol. Dis. 2012, 46, 245–254. [Google Scholar] [CrossRef] [PubMed]
- Araki, N.; Niibori, A.; Midorikawa, U. Expression profiling of glioma initiating cells (GICs) in the sphere and differentiation conditions. Unpublished results. The raw microarray data is accessible at public NCBI GEO database. Accession No. GSE43762.
- Venkatraman, A.; He, X.C.; Thorvaldsen, J.L.; Sugimura, R.; Perry, J.M.; Tao, F.; Zhao, M.; Christenson, M.K.; Sanchez, R.; Yu, J.Y.; et al. Maternal imprinting at the H19-IGF2 locus maintains adult haematopoietic stem cell quiescence. Nature 2013, 500, 345–349. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.Q.; Leung, G.K. Functional roles of non-coding RNAs in glioma and their clinical perspectives. Neurochem. Int. 2014, 77, 78–85. [Google Scholar] [CrossRef] [PubMed]
- Luo, M.; Li, Z.; Wang, W.; Zeng, Y.; Liu, Z.; Qiu, J. Long non-coding RNA H19 increases bladder cancer metastasis by associating with EZH2 and inhibiting E-cadherin expression. Cancer Lett. 2013, 333, 213–221. [Google Scholar] [CrossRef] [PubMed]
- Matouk, I.J.; Raveh, E.; Abu-lail, R.; Mezan, S.; Gilon, M.; Gershtain, E.; Birman, T.; Gallula, J.; Schneider, T.; Barkali, M.; et al. Oncofetal H19 RNA promotes tumor metastasis. BBA 2014, 1843, 1414–1426. [Google Scholar] [PubMed]
- Gupta, R.A.; Shah, N.; Wang, K.C.; Kim, J.; Horlings, H.M.; Wong, D.J.; Tsai, M.C.; Hung, T.; Argani, P.; Rinn, J.L.; et al. Long non-coding RNA hotair reprograms chromatin state to promote cancer metastasis. Nature 2010, 464, U1071–U1148. [Google Scholar] [CrossRef] [PubMed]
- Morfouace, M.; Lalier, L.; Bahut, M.; Bonnamain, V.; Naveilhan, P.; Guette, C.; Oliver, L.; Gueguen, N.; Reynier, P.; Vallette, F.M. Comparison of spheroids formed by rat glioma stem cells and neural stem cells reveals differences in glucose metabolism and promising therapeutic applications. J. Biol. Chem. 2012, 287, 33664–33674. [Google Scholar] [CrossRef] [PubMed]
- Lang, M.F.; Yang, S.; Zhao, C.N.; Sun, G.Q.; Murai, K.; Wu, X.W.; Wang, J.H.; Gao, H.L.; Brown, C.E.; Liu, X.X.; et al. Genome-wide profiling identified a set of miRNAs that are differentially expressed in glioblastoma stem cells and normal neural stem cells. PLoS ONE 2012, 7, e36248. [Google Scholar] [CrossRef] [PubMed]
- Sanai, N.; Alvarez-Buylla, A.; Berger, M.S. Mechanisms of disease: Neural stem cells and the origin of gliomas. N. Engl. J. Med. 2005, 353, 811–822. [Google Scholar] [CrossRef] [PubMed]
- Engstrom, P.G.; Tommei, D.; Stricker, S.H.; Ender, C.; Pollard, S.M.; Bertone, P. Digital transcriptome profiling of normal and glioblastoma-derived neural stem cells identifies genes associated with patient survival. Genome Med. 2012, 4, 76. [Google Scholar] [CrossRef] [PubMed]
- Sandberg, C.J.; Altschuler, G.; Jeong, J.; Stromme, K.K.; Stangeland, B.; Murrell, W.; Grasmo-Wendler, U.H.; Myklebost, O.; Helseth, E.; Vik-Mo, E.O.; et al. Comparison of glioma stem cells to neural stem cells from the adult human brain identifies dysregulated Wnt-signaling and a fingerprint associated with clinical outcome. Exp. Cell Res. 2013, 319, 2230–2243. [Google Scholar] [CrossRef] [PubMed]
- Rheinbay, E.; Suva, M.L.; Gillespie, S.M.; Wakimoto, H.; Patel, A.P.; Shahid, M.; Oksuz, O.; Rabkin, S.D.; Martuza, R.L.; Rivera, M.N.; et al. An aberrant transcription factor network essential for Wnt signaling and stem cell maintenance in glioblastoma. Cell Rep. 2013, 3, 1567–1579. [Google Scholar] [CrossRef] [PubMed]
- Hou, Z.B.; Zhao, W.; Zhou, J.; Shen, L.; Zhan, P.; Xu, C.H.; Chang, C.J.; Bi, H.; Zou, J.; Yao, X.; et al. A long noncoding RNA Sox2ot regulates lung cancer cell proliferation and is a prognostic indicator of poor survival. Int. J. Biochem. Cell Biol. 2014, 53, 380–388. [Google Scholar] [CrossRef] [PubMed]
- Askarian-Amiri, M.E.; Seyfoddin, V.; Smart, C.E.; Wang, J.L.; Kim, J.E.; Hansji, H.; Baguley, B.C.; Finlay, G.J.; Leung, E.Y. Emerging role of long non-coding RNA Sox2ot in Sox2 regulation in breast cancer. PLoS ONE 2014, 9, e102140. [Google Scholar] [CrossRef] [PubMed]
- Gutschner, T.; Hammerle, M.; Eissmann, M.; Hsu, J.; Kim, Y.; Hung, G.; Revenko, A.; Arun, G.; Stentrup, M.; Gross, M.; et al. The noncoding RNA malat1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013, 73, 1180–1189. [Google Scholar] [CrossRef] [PubMed]
- Matouk, I.J.; Mezan, S.; Mizrahi, A.; Ohana, P.; Abu-Lail, R.; Fellig, Y.; Degroot, N.; Galun, E.; Hochberg, A. The oncofetal H19 RNA connection: Hypoxia, p53 and cancer. Biochim. Biophys. Acta 2010, 1803, 443–451. [Google Scholar] [CrossRef] [PubMed]
- Rinn, J.L.; Chang, H.Y. Genome regulation by long noncoding RNAs. Ann. Rev. Biochem. 2012, 81, 145–166. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Fu, H.; Wu, Y.; Zheng, X. Function of lncrnas and approaches to lncRNA-protein interactions. Sci. China Life Sci. 2013, 56, 876–885. [Google Scholar] [CrossRef] [PubMed]
- Li, J.H.; Liu, S.; Zhou, H.; Qu, L.H.; Yang, J.H. Starbase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014, 42, D92–D97. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.H.; Li, J.H.; Jiang, S.; Zhou, H.; Qu, L.H. Chipbase: A database for decoding the transcriptional regulation of long non-coding RNA and microrna genes from CHIP-Seq data. Nucleic Acids Res. 2013, 41, D177–D187. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.L.; Wang, H.; Li, Z.Z.; Wu, Q.L.; Lathia, J.D.; McLendon, R.E.; Hjelmeland, A.B.; Rich, J.N. c-Myc is required for maintenance of glioma cancer stem cells. PLoS ONE 2008, 3, e3769. [Google Scholar] [CrossRef] [PubMed]
- Chiou, S.H.; Wang, M.L.; Chou, Y.T.; Chen, C.J.; Hong, C.F.; Hsieh, W.J.; Chang, H.T.; Chen, Y.S.; Lin, T.W.; Hsu, H.S.; et al. Coexpression of Oct4 and nanog enhances malignancy in lung adenocarcinoma by inducing cancer stem cell-like properties and epithelial-mesenchymal transdifferentiation. Cancer Res. 2010, 70, 10433–10444. [Google Scholar] [CrossRef] [PubMed]
- Jeter, C.R.; Liu, B.; Liu, X.; Chen, X.; Liu, C.; Calhoun-Davis, T.; Repass, J.; Zaehres, H.; Shen, J.J.; Tang, D.G. Nanog promotes cancer stem cell characteristics and prostate cancer resistance to androgen deprivation. Oncogene 2011, 30, 3833–3845. [Google Scholar] [CrossRef] [PubMed]
- Moon, J.H.; Kwon, S.; Jun, E.K.; Kim, A.; Whang, K.Y.; Kim, H.; Oh, S.; Yoon, B.S.; You, S. Nanog-induced dedifferentiation of p53-deficient mouse astrocytes into brain cancer stem-like cells. Biochem. Biophys. Res. Commun. 2011, 412, 175–181. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.M.; Liu, S.; Lu, H.; Zhang, H.; Zhang, P.J.; Gimotty, P.A.; Guerra, M.; Guo, W.; Xu, X. Acquired cancer stem cell phenotypes through Oct4-mediated dedifferentiation. Oncogene 2012, 31, 4898–4911. [Google Scholar] [CrossRef] [PubMed]
- Samardzija, C.; Quinn, M.; Findlay, J.K.; Ahmed, N. Attributes of Oct4 in stem cell biology: Perspectives on cancer stem cells of the ovary. J. Ovarian Res. 2012, 5, 37. [Google Scholar] [CrossRef] [PubMed]
- Morfouace, M.; Lalier, L.; Oliver, L.; Cheray, M.; Pecqueur, C.; Cartron, P.F.; Vallette, F.M. Control of glioma cell death and differentiation by PKM2-Oct4 interaction. Cell Death Dis. 2014, 5, e1036. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, B.A.; Bar-Nur, O.; Silva, J.C.R.; Hochedlinger, K. Nanog is dispensable for the generation of induced pluripotent stem cells. Curr. Biol. 2014, 24, 347–350. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, J.S.; Gaughwin, P.M.; Lim, B.; Robson, P.; Lipovich, L. Conserved long noncoding RNAs transcriptionally regulated by Oct4 and nanog modulate pluripotency in mouse embryonic stem cells. RNA 2010, 16, 324–337. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Qureshi, I.A.; Gokhan, S.; Dinger, M.E.; Li, G.Y.; Mattick, J.S.; Mehler, M.F. Long noncoding RNAs in neuronal-glial fate specification and oligodendrocyte lineage maturation. BMC Neurosci. 2010, 11, 14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ratajczak, M.Z. IGF2-H19, an imprinted tandem gene, is an important regulator of embryonic development, a guardian of proliferation of adult pluripotent stem cells, a regulator of longevity, and a “passkey” to cancerogenesis. Folia Histochem. Cytobiol. 2012, 50, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Gabory, A.; Jammes, H.; Dandolo, L. The h19 locus: Role of an imprinted non-coding RNA in growth and development. Bioessays 2010, 32, 473–480. [Google Scholar] [CrossRef] [PubMed]
- Barsyte-Lovejoy, D.; Lau, S.K.; BoutroS, P.C.; Khosravi, F.; Jurisica, I.; Andrulis, I.L.; Tsao, M.S.; Penn, L.Z. The c-Myc oncogene directly induces the H19 noncoding RNA by allele-specific binding to potentiate tumorigenesis. Cancer Res. 2006, 66, 5330–5337. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Wang, H.; Liu, K.; Wang, F.; Ye, X.; Liu, M.; Xiang, R.; Liu, N.; Liu, L. Knockdown of H19 enhances differentiation capacity to epidermis of parthenogenetic embryonic stem cells. Curr. Mol. Med. 2014, 14, 737–748. [Google Scholar] [CrossRef] [PubMed]
- Holwerda, S.J.; de Laat, W. CTCF: The protein, the binding partners, the binding sites and their chromatin loops. Philo. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368, 20120369. [Google Scholar] [CrossRef] [PubMed]
- Herold, M.; Bartkuhn, M.; Renkawitz, R. CTCF: Insights into insulator function during development. Development 2012, 139, 1045–1057. [Google Scholar] [CrossRef] [PubMed]
- Plasschaert, R.N.; Vigneau, S.; Tempera, I.; Gupta, R.; Maksimoska, J.; Everett, L.; Davuluri, R.; Mamorstein, R.; Lieberman, P.M.; Schultz, D.; et al. CTCF binding site sequence differences are associated with unique regulatory and functional trends during embryonic stem cell differentiation. Nucleic Acids Res. 2014, 42, 774–789. [Google Scholar] [CrossRef] [PubMed]
- Sone, M.; Hayashi, T.; Tarui, H.; Agata, K.; Takeichi, M.; Nakagawa, S. The mRNA-like noncoding RNA gomafu constitutes a novel nuclear domain in a subset of neurons. J. Cell Sci. 2007, 120, 2498–2506. [Google Scholar] [CrossRef] [PubMed]
- Hirayama, T.; Tarusawa, E.; Yoshimura, Y.; Galjart, N.; Yagi, T. CTCF is required for neural development and stochastic expression of clustered Pcdh genes in neurons. Cell Rep. 2012, 2, 345–357. [Google Scholar] [CrossRef] [PubMed]
- Koesters, C.; Unger, B.; Bilic, I.; Schmidt, U.; Bluml, S.; Lichtenberger, B.; Schreiber, M.; Stockl, J.; Ellmeier, W. Regulation of dendritic cell differentiation and subset distribution by the zinc finger protein CTCF. Immunol. Lett. 2007, 109, 165–174. [Google Scholar] [CrossRef] [PubMed]
- Kurukuti, S.; Tiwari, V.K.; Tavoosidana, G.; Pugacheva, E.; Murrell, A.; Zhao, Z.H.; Lobanenkov, V.; Reik, W.; Ohlsson, R. CTCF binding at the H19 imprinting control region mediates maternally inherited higher-order chromatin conformation to restrict enhancer access to IGF2. PNAS 2006, 103, 10684–10689. [Google Scholar] [CrossRef] [PubMed]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A ceRNA hypothesis: The rosetta stone of a hidden RNA language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Kartha, R.V.; Subramanian, S. Competing endogenous RNAs (ceRNAs): New entrants to the intricacies of gene regulation. Front. Genet. 2014, 5, 8. [Google Scholar] [CrossRef] [PubMed]
- Chi, S.W.; Zang, J.B.; Mele, A.; Darnell, R.B. Argonaute hits-clip decodes microRNA-mRNA interaction maps. Nature 2009, 460, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Ebert, M.S.; Neilson, J.R.; Sharp, P.A. Microrna sponges: Competitive inhibitors of small RNAs in mammalian cells. Nat. Method. 2007, 4, 721–726. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.Y.; Liu, X.F.; Wu, H.C.; Ni, P.H.; Gu, Z.D.; Qiao, Y.X.; Chen, N.; Sun, F.Y.; Fan, Q.S. Creb up-regulates long non-coding RNA, hulc expression through interaction with microRNA-372 in liver cancer. Nucleic Acids Res. 2010, 38, 5366–5383. [Google Scholar] [CrossRef] [PubMed]
- Cai, X.Z.; Cullen, B.R. The imprinted H19 noncoding RNA is a primary microRNA precursor. RNA 2007, 13, 313–316. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Dutta, A.; Abounader, R. The role of microRNAs in glioma initiation and progression. Front. Biosci. 2012, 17, 700–712. [Google Scholar] [CrossRef]
- Yu, F.; Yao, H.; Zhu, P.; Zhang, X.; Pan, Q.; Gong, C.; Huang, Y.; Hu, X.; Su, F.; Lieberman, J.; et al. Let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 2007, 131, 1109–1123. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.C.; Mei, P.J.; Chen, C.; Miao, F.A.; Zhang, H.; Li, Z.L. Mir-29c inhibits glioma cell proliferation, migration, invasion and angiogenesis. J. Neuro-Oncol. 2013, 115, 179–188. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, Y.; Sun, J.; Wang, Q.; Sun, C.; Yan, Y.; Yu, L.; Cheng, D.; An, T.; Shi, C.; et al. Tumor-suppressive effects of miR-29c on gliomas. Neuroreport 2013, 24, 637–645. [Google Scholar] [CrossRef] [PubMed]
- Ernst, A.; Campos, B.; Meier, J.; Devens, F.; Liesenberg, F.; Wolter, M.; Reifenberger, G.; Herold-Mende, C.; Lichter, P.; Radlwimmer, B. De-repression of CTGF via the miR-17-92 cluster upon differentiation of human glioblastoma spheroid cultures. Oncogene 2010, 29, 3411–3422. [Google Scholar] [CrossRef] [PubMed]
- Atkinson, G.P.; Nozell, S.E.; Benveniste, E.T. NF-kappaB and STAT3 signaling in glioma: Targets for future therapies. Exp. Rev. Neurother. 2010, 10, 575–586. [Google Scholar] [CrossRef] [PubMed]
- Garner, J.M.; Fan, M.; Yang, C.H.; Du, Z.; Sims, M.; Davidoff, A.M.; Pfeffer, L.M. Constitutive activation of signal transducer and activator of transcription 3 (STAT3) and nuclear factor kappaB signaling in glioblastoma cancer stem cells regulates the notch pathway. J. Biol. Chem. 2013, 288, 26167–26176. [Google Scholar] [CrossRef] [PubMed]
- Hjelmeland, A.B.; Wu, Q.; Wickman, S.; Eyler, C.; Heddleston, J.; Shi, Q.; Lathia, J.D.; Macswords, J.; Lee, J.; McLendon, R.E.; et al. Targeting a20 decreases glioma stem cell survival and tumor growth. PLoS Biol. 2010, 8, e1000319. [Google Scholar] [CrossRef] [PubMed]
- Iliopoulos, D.; Hirsch, H.A.; Struhl, K. An epigenetic switch involving NF-kappaB, Lin28, Let-7 microRNA, and IL6 links inflammation to cell transformation. Cell 2009, 139, 693–706. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Vodyanik, M.A.; Smuga-Otto, K.; Antosiewicz-Bourget, J.; Frane, J.L.; Tian, S.; Nie, J.; Jonsdottir, G.A.; Ruotti, V.; Stewart, R.; et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 2007, 318, 1917–1920. [Google Scholar] [CrossRef] [PubMed]
- Moss, E.G.; Lee, R.C.; Ambros, V. The cold shock domain protein Lin28 controls developmental timing in c-Elegans and is regulated by the Lin-4 RNA. Cell 1997, 88, 637–646. [Google Scholar] [CrossRef]
- Shyh-Chang, N.; Zhu, H.; Yvanka de Soysa, T.; Shinoda, G.; Seligson, M.T.; Tsanov, K.M.; Nguyen, L.; Asara, J.M.; Cantley, L.C.; Daley, G.Q. Lin28 enhances tissue repair by reprogramming cellular metabolism. Cell 2013, 155, 778–792. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Ng, S.B.; Chng, W.J. Lin28/lin28b: An emerging oncogenic driver in cancer stem cells. Int. J. Biochem. Cell Biol. 2013, 45, 973–978. [Google Scholar] [CrossRef] [PubMed]
- Qin, R.; Zhou, J.X.; Chen, C.; Xu, T.; Yan, Y.; Ma, Y.S.; Zheng, Z.L.; Shen, Y.P.; Lu, Y.C.; Fu, D.; et al. Lin28 is involved in glioma carcinogenesis and predicts outcomes of glioblastoma multiforme patients. PLoS ONE 2014, 9, e86446. [Google Scholar] [CrossRef] [PubMed]
- Viswanathan, S.R.; Daley, G.Q.; Gregory, R.I. Selective blockade of microrna processing by Lin28. Science 2008, 320, 97–100. [Google Scholar] [CrossRef] [PubMed]
- Ali, P.S.S.; Ghoshdastider, U.; Hoffmann, J.; Brutschy, B.; Filipek, S. Recognition of the Let-7g miRNA precursor by human Lin28b. FEBS Lett. 2012, 586, 3986–3990. [Google Scholar] [PubMed]
- Guo, L.; Chen, C.; Shi, M.; Wang, F.; Chen, X.; Diao, D.; Hu, M.; Yu, M.; Qian, L.; Guo, N. STAT3-coordinated Lin-28-Let-7-Hmga2 and Mir-200-Zeb1 circuits initiate and maintain oncostatin m-Driven epithelial-mesenchymal transition. Oncogene 2013, 32, 5272–5282. [Google Scholar] [CrossRef] [PubMed]
- Derynck, R.; Zhang, Y.E. Smad-dependent and smad-independent pathways in TGF-beta family signalling. Nature 2003, 425, 577–584. [Google Scholar] [CrossRef] [PubMed]
- Miyazono, K.; ten Dijke, P.; Heldin, C.H. TGF-β signaling by smad proteins. Adv. Immunol. 2000, 75, 115–157. [Google Scholar] [PubMed]
- Moustakas, A.; Souchelnytskyi, S.; Heldin, C.H. Smad regulation in TGF-β signal transduction. J. Cell Sci. 2001, 114, 4359–4369. [Google Scholar] [PubMed]
- Mendjan, S.; Mascetti, V.L.; Ortmann, D.; Ortiz, M.; Karjosukarso, D.W.; Ng, Y.; Moreau, T.; Pedersen, R.A. Nanog and CDX2 pattern distinct subtypes of human mesoderm during exit from pluripotency. Cell Stem Cell 2014, 15, 310–325. [Google Scholar] [CrossRef] [PubMed]
- Muller, H.; Helin, K. The E2f transcription factors: Key regulators of cell proliferation. Biochim. Biophys. Acta 2000, 1470, M1–M12. [Google Scholar] [CrossRef]
- Ren, B.; Cam, H.; Takahashi, Y.; Volkert, T.; Terragni, J.; Young, R.A.; Dynlacht, B.D. E2f integrates cell cycle progression with DNA repair, replication, and g(2)/m checkpoints. Genes Dev. 2002, 16, 245–256. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Timmers, C.; Maiti, B.; Saavedra, H.I.; Sang, L.; Chong, G.T.; Nuckolls, F.; Giangrande, P.; Wright, F.A.; Field, S.J.; et al. The E2f1-3 transcription factors are essential for cellular proliferation. Nature 2001, 414, 457–462. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, X.; Kiang, K.M.; Zhang, G.P.; Leung, G.K. Long Non-Coding RNAs Dysregulation and Function in Glioblastoma Stem Cells. Non-Coding RNA 2015, 1, 69-86. https://doi.org/10.3390/ncrna1010069
Zhang X, Kiang KM, Zhang GP, Leung GK. Long Non-Coding RNAs Dysregulation and Function in Glioblastoma Stem Cells. Non-Coding RNA. 2015; 1(1):69-86. https://doi.org/10.3390/ncrna1010069
Chicago/Turabian StyleZhang, Xiaoqin, Karrie Meiyee Kiang, Grace Pingde Zhang, and Gilberto Kakit Leung. 2015. "Long Non-Coding RNAs Dysregulation and Function in Glioblastoma Stem Cells" Non-Coding RNA 1, no. 1: 69-86. https://doi.org/10.3390/ncrna1010069






