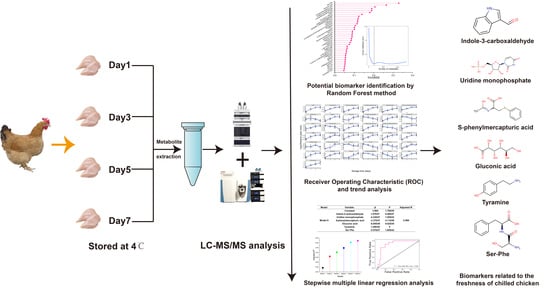
UHPLC–MS/MS-Based Nontargeted Metabolomics Analysis Reveals Biomarkers Related to the Freshness of Chilled Chicken
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Sample Collection and Storage
2.3. Metabolite Extraction
2.4. Liquid Chromatography–Mass Spectrometry Analysis
2.5. Quality Control
2.6. Data Preprocessing and Annotation
2.7. Differential Metabolites Analysis
2.8. Random Forest Regression Analysis
2.9. Stepwise Multivariate Linear Regression Analysis
2.10. Pathway Enrichment of Potential Metabolic Biomarkers
3. Results and Discussion
3.1. Quality Control
3.2. Metabolic Profiles Analysis of Chilled Chicken over Time
3.3. Identification of Differential Metabolites
3.4. Screening of Potential Biomarker Related to the Freshness of Chilled Chicken
3.5. Identification of Key Biomarkers Related to the Freshness of Chilled Chicken
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Patsias, A.; Badeka, A.V.; Savvaidis, I.N.; Kontominas, M.G. Combined effect of freeze chilling and MAP on quality parameters of raw chicken fillets. Food Microbiol. 2008, 25, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wang, H.; Li, N.; Ming, L.; Xu, X. High CO2-modified atmosphere packaging for extension of shelf-life of chilled yellow-feather broiler meat: A special breed in Asia. LWT Food Sci. Technol. 2015, 64, 1123–1129. [Google Scholar] [CrossRef]
- Wang, G.Y.; Wang, H.H.; Han, Y.W.; Xing, T.; Ye, K.P.; Xu, X.L.; Zhou, G.H. Evaluation of the spoilage potential of bacteria isolated from chilled chicken in vitro and in situ. Food Microbiol. 2017, 63, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Nowak, A.; Rygala, A.; Oltuszak-Walczak, E.; Walczak, P. The prevalence and some metabolic traits of Brochothrix thermosphacta in meat and meat products packaged in different ways. J. Sci. Food Agric. 2012, 92, 1304–1310. [Google Scholar] [CrossRef] [PubMed]
- Casaburi, A.; Piombino, P.; Nychas, G.-J.; Villani, F.; Ercolini, D. Bacterial populations and the volatilome associated to meat spoilage. Food Microbiol. 2015, 45, 83–102. [Google Scholar] [CrossRef]
- Cevallos-Cevallos, J.M.; Reyes-De-Corcuera, J.I.; Etxeberria, E.; Danyluk, M.D.; Rodrick, G.E. Metabolomic analysis in food science: A review. Trends Food Sci. Technol. 2009, 20, 557–566. [Google Scholar] [CrossRef]
- Cubero-Leon, E.; Peñalver, R.; Maquet, A. Review on metabolomics for food authentication. Food Res. Int. 2014, 60, 95–107. [Google Scholar] [CrossRef]
- Dunn, W.B.; Ellis, D.I. Metabolomics: Current analytical platforms and methodologies. Trac Trends Anal. Chem. 2005, 24, 285–294. [Google Scholar] [CrossRef]
- Johnson, A.E.; Sidwick, K.L.; Pirgozliev, V.R.; Edge, A.; Thompson, D.F. Metabonomic Profiling of Chicken Eggs during Storage Using High-Performance Liquid Chromatography-Quadrupole Time-of-Flight Mass Spectrometry. Anal. Chem. 2018, 90, 7489–7494. [Google Scholar] [CrossRef] [Green Version]
- Jääskeläinen, E.; Jakobsen, L.M.A.; Hultman, J.; Eggers, N.; Bertram, H.C.; Björkroth, J. Metabolomics and bacterial diversity of packaged yellowfin tuna (Thunnus albacares) and salmon (Salmo salar) show fish species-specific spoilage development during chilled storage. Int J. Food Microbiol. 2019, 293, 44–52. [Google Scholar] [CrossRef]
- Mabuchi, R.; Adachi, M.; Ishimaru, A.; Zhao, H.; Tanimoto, S. Changes in Metabolic Profiles of Yellowtail (Seriola Quinqueradiata) Muscle during Cold Storage as a Freshness Evaluation Tool Based on GC-MS Metabolomics. Foods 2019, 8, 511. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aru, V.; Pisano, M.B.; Savorani, F.; Engelsen, S.B.; Cosentino, S.; Marincola, F.C. Metabolomics analysis of shucked mussels’ freshness. Food Chem. 2016, 205, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Syukri, D.; Thammawong, M.; Naznin, H.A.; Kuroki, S.; Tsuta, M.; Yoshida, M.; Nakano, K. Identification of a freshness marker metabolite in stored soybean sprouts by comprehensive mass-spectrometric analysis of carbonyl compounds. Food Chem. 2018, 269, 588–594. [Google Scholar] [CrossRef] [PubMed]
- Luan, H.; Liu, L.F.; Tang, Z.; Zhang, M.; Chua, K.K.; Song, J.X.; Mok, V.C.; Li, M.; Cai, Z. Comprehensive urinary metabolomic profiling and identification of potential noninvasive marker for idiopathic Parkinsonh’s disease. Sci. Rep. 2015, 5, 13888. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Zhang, X.; Wu, D.; Huang, Z.; Hou, T.; Jian, C.; Yu, P.; Lu, F.; Zhang, R.; Sun, T.; et al. Mitochondrial flashes regulate ATP homeostasis in the heart. eLife 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Phonchai, A.; Wilairat, P.; Chantiwas, R. Development of a solid-phase extraction method with simple MEKC-UV analysis for simultaneous detection of indole metabolites in human urine after administration of indole dietary supplement. Talanta 2017, 174, 314–319. [Google Scholar] [CrossRef] [PubMed]
- Roager, H.M.; Licht, T.R. Microbial tryptophan catabolites in health and disease. Nat. Commun. 2018, 9, 3294. [Google Scholar] [CrossRef] [Green Version]
- Cevallos-Cevallos, J.M.; Reyes-De-Corcuera, J.I. Metabolomics in food science. Adv. Food Nutr. Res. 2012, 67, 1–24. [Google Scholar] [CrossRef]
- Johanningsmeier, S.D.; Harris, G.K.; Klevorn, C.M. Metabolomic Technologies for Improving the Quality of Food: Practice and Promise. Annu. Rev. Food Sci. Technol. 2016, 7, 413–438. [Google Scholar] [CrossRef] [Green Version]
- Lin, Y.; Li, W.; Sun, L.; Lin, Z.; Jiang, Y.; Ling, Y.; Lin, X. Comparative metabolomics shows the metabolic profiles fluctuate in multi-drug resistant Escherichia coli strains. J. Proteom. 2019, 207, 103468. [Google Scholar] [CrossRef]
- Chong, J.; Wishart, D.S.; Xia, J. Using MetaboAnalyst 4.0 for Comprehensive and Integrative Metabolomics Data Analysis. Curr. Protoc. Bioinform. 2019, 68, e86. [Google Scholar] [CrossRef] [PubMed]
- Kamarudin, A.N.; Cox, T.; Kolamunnage-Dona, R. Time-dependent ROC curve analysis in medical research: Current methods and applications. BMC Med. Res. Methodol. 2017, 17, 53. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, X.; Liu, L.; Lan, X.; Cohen, D.; Zhang, Y.; Ravindran, A.V.; Yuan, S.; Zheng, P.; Coghill, D.; Yang, L.; et al. Polyunsaturated fatty acids metabolism, purine metabolism and inosine as potential independent diagnostic biomarkers for major depressive disorder in children and adolescents. Mol. Psychiatry 2019, 24, 1478–1488. [Google Scholar] [CrossRef] [PubMed]
- Wen, D.; Liu, Y.; Yu, Q. Metabolomic approach to measuring quality of chilled chicken meat during storage. Poult. Sci. 2020, 99, 2543–2554. [Google Scholar] [CrossRef] [PubMed]
- Gram, L.; Ravn, L.; Rasch, M.; Bruhn, J.B.; Christensen, A.B.; Givskov, M. Food spoilage—interactions between food spoilage bacteria. Int. J. Food Microbiol. 2002, 78, 79–97. [Google Scholar] [CrossRef]
- Wojnowski, W.; Majchrzak, T.; Dymerski, T.; Gebicki, J.; Namiesnik, J. Poultry meat freshness evaluation using electronic nose technology and ultra-fast gas chromatography. Mon. Chem. 2017, 148, 1631–1637. [Google Scholar] [CrossRef] [Green Version]
- Rigatti, S.J. Random Forest. J. Insur. Med. 2017, 47, 31–39. [Google Scholar] [CrossRef] [Green Version]
- Szabo de Edelenyi, F.; Goumidi, L.; Bertrais, S.; Phillips, C.; Macmanus, R.; Roche, H.; Planells, R.; Lairon, D. Prediction of the metabolic syndrome status based on dietary and genetic parameters, using Random Forest. Genes Nutr. 2008, 3, 173–176. [Google Scholar] [CrossRef] [Green Version]
- Fathi, F.; Majari-Kasmaee, L.; Mani-Varnosfaderani, A.; Kyani, A.; Rostami-Nejad, M.; Sohrabzadeh, K.; Naderi, N.; Zali, M.R.; Rezaei-Tavirani, M.; Tafazzoli, M.; et al. 1H NMR based metabolic profiling in Crohn’s disease by random forest methodology. Magn. Reson. Chem. 2014, 52, 370–376. [Google Scholar] [CrossRef] [Green Version]
- Worachartcheewan, A.; Shoombuatong, W.; Pidetcha, P.; Nopnithipat, W.; Prachayasittikul, V.; Nantasenamat, C. Predicting Metabolic Syndrome Using the Random Forest Method. Sci. World J. 2015, 2015, 581501. [Google Scholar] [CrossRef] [Green Version]
- Oza, V.H.; Aicher, J.K.; Reed, L.K. Random Forest Analysis of Untargeted Metabolomics Data Suggests Increased Use of Omega Fatty Acid Oxidation Pathway in Drosophila Melanogaster Larvae Fed a Medium Chain Fatty Acid Rich High-Fat Diet. Metabolites 2018, 9, 5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, T.; Cao, Y.; Zhang, Y.; Liu, J.; Bao, Y.; Wang, C.; Jia, W.; Zhao, A. Random forest in clinical metabolomics for phenotypic discrimination and biomarker selection. Evid. Based Complement. Altern. Med. Ecam 2013, 2013, 298183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Broughton-Neiswanger, L.E.; Rivera-Velez, S.M.; Suarez, M.A.; Slovak, J.E.; Hwang, J.K.; Villarino, N.F. Pharmacometabolomics with a combination of PLS-DA and random forest algorithm analyses reveal meloxicam alters feline plasma metabolite profiles. J. Vet. Pharmacol. Ther. 2020. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Vicente Gonçalves, C.M.; Dai, L.; Lu, H.M.; Huang, J.H.; Ji, H.; Wang, D.S.; Yi, L.Z.; Liang, Y.Z. Exploring metabolic syndrome serum profiling based on gas chromatography mass spectrometry and random forest models. Anal. Chim. Acta 2014, 827, 22–27. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Peng, Q. A deep ensemble model to predict miRNA-disease association. Sci. Rep. 2017, 7, 14482. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, Q.J.; Zhao, J.R.; Hao, J.; Li, B.; Huo, Y.; Han, Y.L.; Wan, L.L.; Li, J.; Huang, J.; Lu, J.; et al. Serum and urine metabolomics study reveals a distinct diagnostic model for cancer cachexia. J. Cachexia Sarcopenia Muscle 2018, 9, 71–85. [Google Scholar] [CrossRef]
- Acharjee, A.; Kloosterman, B.; de Vos, R.C.; Werij, J.S.; Bachem, C.W.; Visser, R.G.; Maliepaard, C. Data integration and network reconstruction with ~omics data using Random Forest regression in potato. Anal. Chim. Acta 2011, 705, 56–63. [Google Scholar] [CrossRef]
- Staats, R.; Barros, I.; Fernandes, D.; Grencho, D.; Reis, C.; Matos, F.; Valença, J.; Marôco, J.; de Almeida, A.B.; Bárbara, C. The Importance of Sleep Fragmentation on the Hemodynamic Dipping in Obstructive Sleep Apnea Patients. Front. Physiol. 2020, 11, 104. [Google Scholar] [CrossRef] [Green Version]
- Lázaro, C.A.; Junior, C.A.C.; Canto, A.C.; Monteiro, M.L.G.; Franco, R.M. Biogenic amines as bacterial quality indicators in different poultry meat species. LWT Food Sci. Technol. 2015, 60, 15–21. [Google Scholar] [CrossRef]
- Pierini, G.D.; Robledo, S.N.; Zon, M.A.; Di Nezio, M.S.; Granero, A.M.; Fernández, H. Development of an electroanalytical method to control quality in fish samples based on an edge plane pyrolytic graphite electrode. Simultaneous determination of hypoxanthine, xanthine and uric acid. Microchem. J. 2018, 138, 58–64. [Google Scholar] [CrossRef]
- Zhang, L.S.; Davies, S.S. Microbial metabolism of dietary components to bioactive metabolites: Opportunities for new therapeutic interventions. Genome Med. 2016, 8, 46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wishart, D.S.; Feunang, Y.D.; Marcu, A.; Guo, A.C.; Liang, K.; Vázquez-Fresno, R.; Sajed, T.; Johnson, D.; Li, C.; Karu, N.; et al. HMDB 4.0: The human metabolome database for 2018. Nucleic Acids Res. 2018, 46, D608–D617. [Google Scholar] [CrossRef] [PubMed]
- Nychas, G.J.E.; Arkoudelos, J.S. Microbiological and physicochemical changes in minced meats under carbon dioxide, nitrogen or air at 3°C. Int. J. Food Ence Technol. 2010, 25, 389–398. [Google Scholar] [CrossRef]






| Model | N a | MSE d | VE (%) e |
|---|---|---|---|
| Random forest model b | 500 | 0.86 | 81.98 |
| Parsimonious model c | 500 | 0.49 | 89.86 |
| Model | Variable | Β b | P c | Adjusted R2 | Durbin–Watson |
|---|---|---|---|---|---|
| Model 1 | Constant a | −1.324 | 4.00 × 10−3 | 0.857 | 2.067 |
| Indole-3-carboxaldehyde | 3.87 × 10−7 | 1.99 × 10−14 | |||
| Model 2 | Constant a | 1.767 | 4.70 × 10−2 | 0.904 | |
| Indole-3-carboxaldehyde | 2.56 × 10−7 | 7.06 × 1−7 | |||
| Uridine monophosphate | −5.96 × 10−7 | 0 | |||
| Model 3 | Constant a | 2.014 | 1.50 × 10−2 | 0.921 | |
| Indole-3-carboxaldehyde | 2.87 × 10−7 | 4.39 × 10−8 | |||
| Uridine monophosphate | −7.47 × 10−7 | 2.41 × 10−5 | |||
| S-phenylmercapturic acid | −2.30 × 10−7 | 1.20 × 10−2 | |||
| Model 4 | Constant a | 3.356 | 0 | 0.939 | |
| Indole-3-carboxaldehyde | 2.46 × 10−7 | 3.07 × 10−7 | |||
| Uridine monophosphate | −7.56 × 10−7 | 3.22 × 10−6 | |||
| S-phenylmercapturic acid | −2.41 × 10−7 | 3.00 × 10−3 | |||
| Gluconic acid | −7.87 × 10−8 | 5.00 × 10−3 | |||
| Model 5 | Constant a | 2.796 | 0 | 0.962 | |
| Indole-3-carboxaldehyde | 2.37 × 10−7 | 1.20 × 10−8 | |||
| Uridine monophosphate | −5.16 × 10−7 | 0 | |||
| S-phenylmercapturic acid | −3.23 × 10−7 | 2.47 × 10−5 | |||
| Gluconic acid | −9.42 × 10−8 | 0 | |||
| Tyramine | 1.27 × 10−8 | 0 | |||
| Model 6 | Constant a | 3.964 | 1.76 × 10−5 | 0.969 | |
| Indole-3-carboxaldehyde | 1.97 × 10−7 | 9.48 × 10−7 | |||
| Uridine monophosphate | −4.22 × 10−7 | 1.00 × 10−3 | |||
| S-phenylmercapturic acid | −3.37 × 10−7 | 4.11 × 10−6 | |||
| Gluconic acid | −8.80 × 10−8 | 8.62 × 10−5 | |||
| Tyramine | 1.26 × 10−8 | 0 | |||
| Serylphenylalanine | −5.57 × 10−7 | 1.60 × 10−2 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, T.; Zhang, S.; Chen, L.; Ding, H.; Wu, P.; Zhang, G.; Xie, K.; Dai, G.; Wang, J. UHPLC–MS/MS-Based Nontargeted Metabolomics Analysis Reveals Biomarkers Related to the Freshness of Chilled Chicken. Foods 2020, 9, 1326. https://doi.org/10.3390/foods9091326
Zhang T, Zhang S, Chen L, Ding H, Wu P, Zhang G, Xie K, Dai G, Wang J. UHPLC–MS/MS-Based Nontargeted Metabolomics Analysis Reveals Biomarkers Related to the Freshness of Chilled Chicken. Foods. 2020; 9(9):1326. https://doi.org/10.3390/foods9091326
Chicago/Turabian StyleZhang, Tao, Shanshan Zhang, Lan Chen, Hao Ding, Pengfei Wu, Genxi Zhang, Kaizhou Xie, Guojun Dai, and Jinyu Wang. 2020. "UHPLC–MS/MS-Based Nontargeted Metabolomics Analysis Reveals Biomarkers Related to the Freshness of Chilled Chicken" Foods 9, no. 9: 1326. https://doi.org/10.3390/foods9091326
APA StyleZhang, T., Zhang, S., Chen, L., Ding, H., Wu, P., Zhang, G., Xie, K., Dai, G., & Wang, J. (2020). UHPLC–MS/MS-Based Nontargeted Metabolomics Analysis Reveals Biomarkers Related to the Freshness of Chilled Chicken. Foods, 9(9), 1326. https://doi.org/10.3390/foods9091326