The Use of Hoechst Dyes for DNA Staining and Beyond
Abstract
:1. Introduction
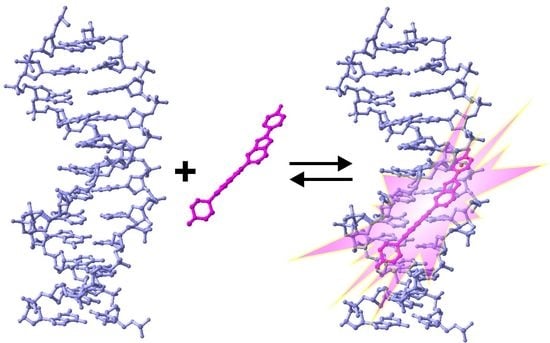
2. Properties of Hoechst Dyes
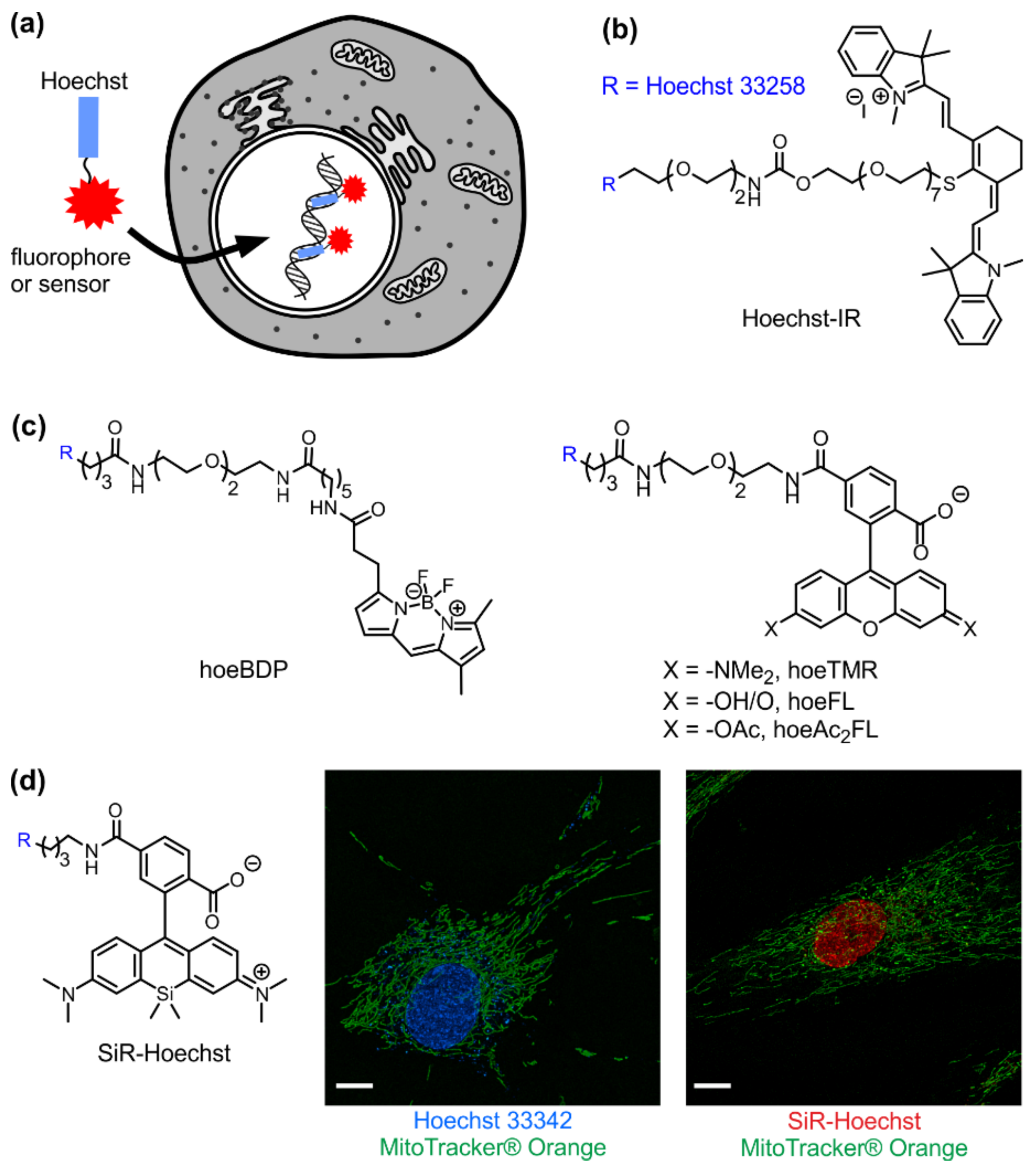
3. Visible/Infrared DNA Probes
4. Sensors of Nucleus Microenvironment
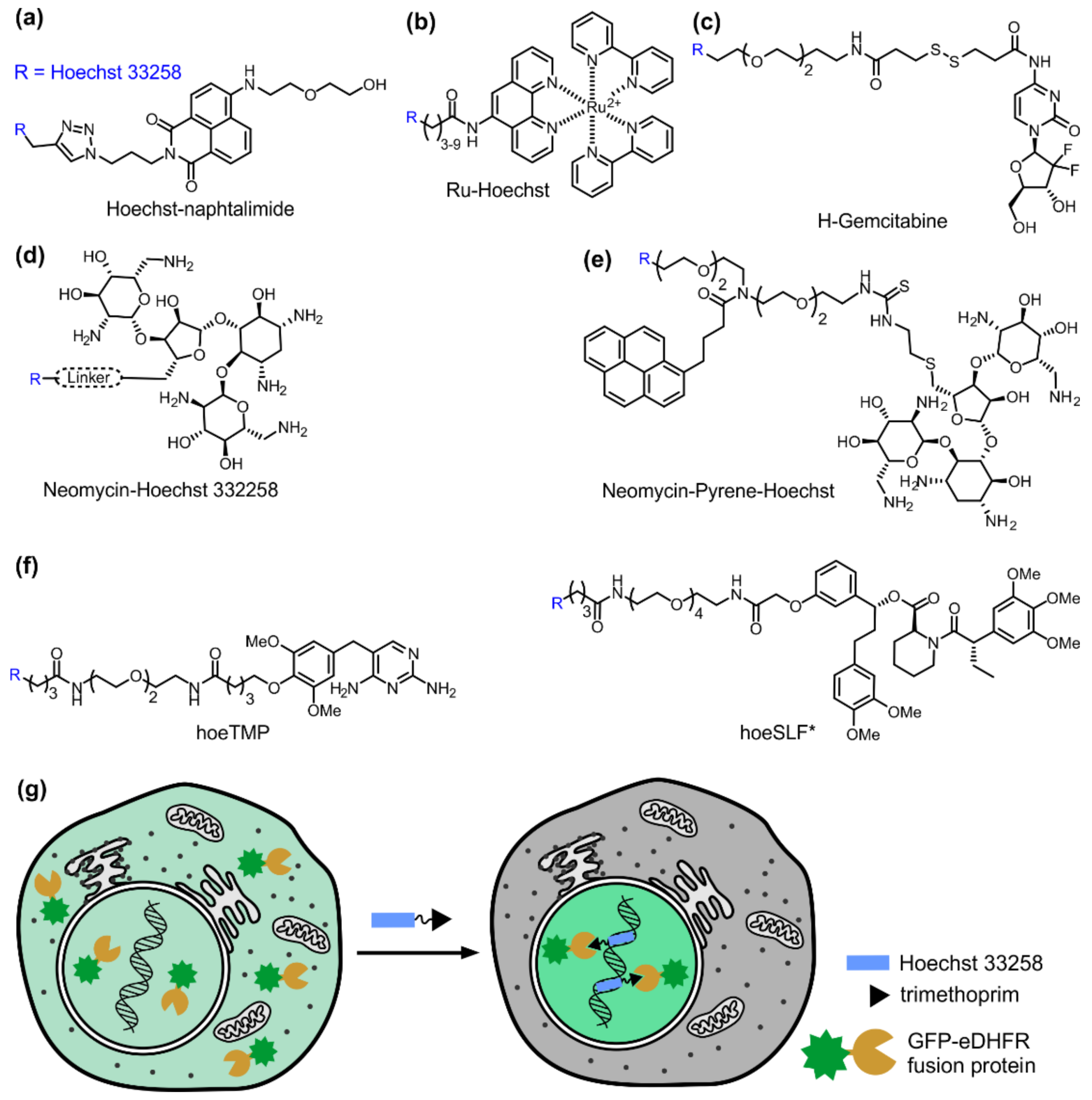
5. Hoechst as a Targeting Module for Drug Delivery
6. Targeting Proteins to the Nucleus
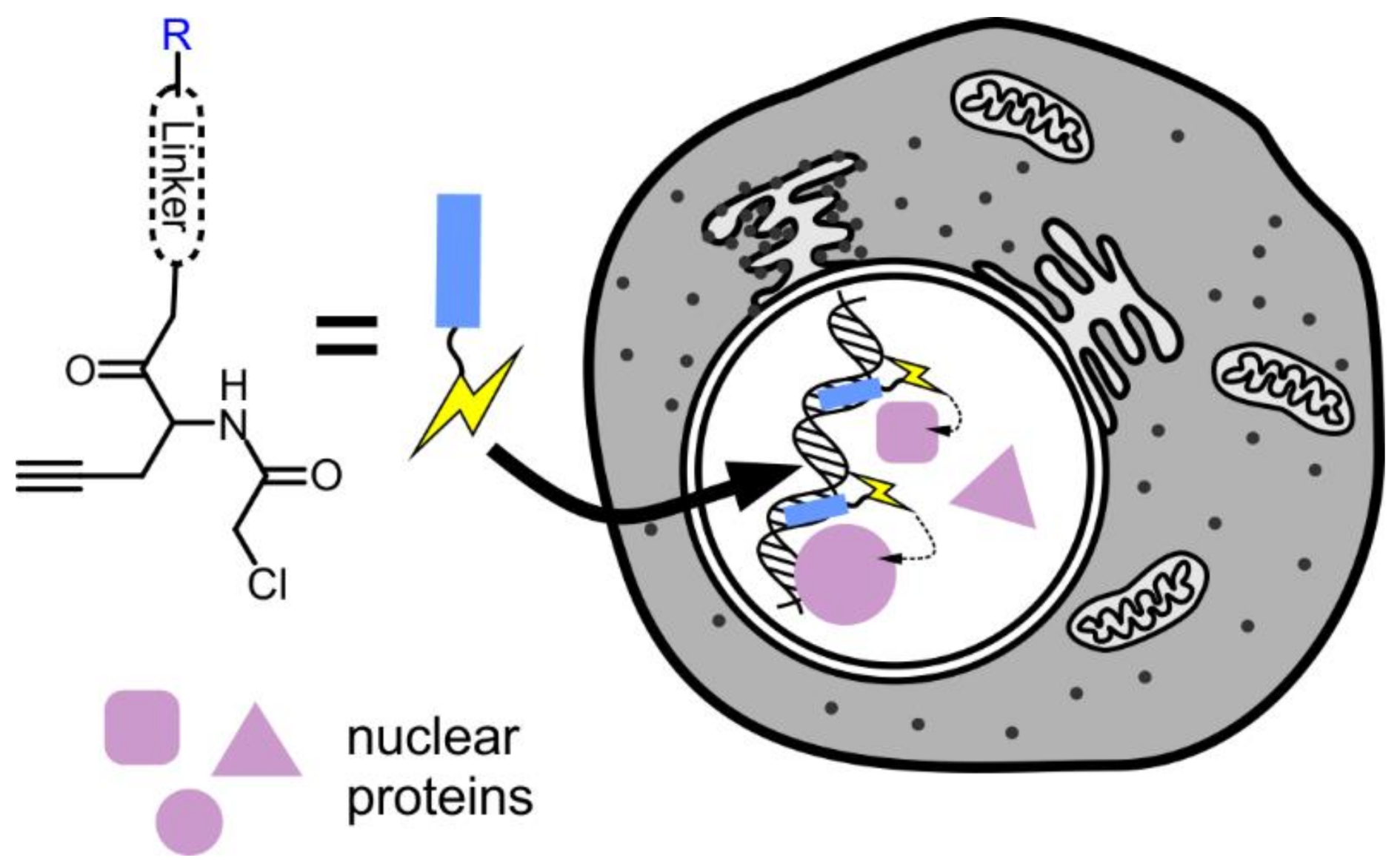
7. Proteomics of the Nucleus
8. Hoechst–DNA Interaction as a Read-Out in Aptamer-Based Sensors
9. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Mazzini, G.; Danova, M. Fluorochromes for DNA staining and quantitation. Methods Mol. Biol. 2017, 1560, 239–259. [Google Scholar] [PubMed]
- Gottfried, A.; Weinhold, E. Sequence-specific covalent labelling of DNA. Biochem. Soc. Trans. 2011, 39, 623–628. [Google Scholar] [CrossRef] [PubMed]
- Bontemps, J.; Houssier, C.; Fredericq, E. Physico-chemical study of the complexes of “33258 hoechst” with DNA and nucleohistone. Nucleic Acids Res. 1975, 2, 971–984. [Google Scholar] [CrossRef] [PubMed]
- Weisblum, B.; Haenssler, E. Fluorometric properties of the bibenzimidazole derivative hoechst 33258, a fluorescent probe specific for at concentration in chromosomal DNA. Chromosoma 1974, 46, 255–260. [Google Scholar] [CrossRef] [PubMed]
- Han, F.; Taulier, N.; Chalikian, T.V. Association of the minor groove binding drug hoechst 33258 with d(cgcgaattcgcg)2: Volumetric, calorimetric, and spectroscopic characterizations. Biochemistry 2005, 44, 9785–9794. [Google Scholar] [CrossRef] [PubMed]
- Pal, S.K.; Zhao, L.; Zewail, A.H. Water at DNA surfaces: Ultrafast dynamics in minor groove recognition. Proc. Natl. Acad. Sci. USA 2003, 100, 8113–8118. [Google Scholar] [CrossRef] [PubMed]
- Adhikary, A.; Buschmann, V.; Muller, C.; Sauer, M. Ensemble and single-molecule fluorescence spectroscopic study of the binding modes of the bis-benzimidazole derivative hoechst 33258 with DNA. Nucleic Acids Res. 2003, 31, 2178–2186. [Google Scholar] [CrossRef] [PubMed]
- Watkins, A.M.; Chan, P.J.; Kalugdan, T.H.; Patton, W.C.; Jacobson, J.D.; King, A. Analysis of the flow cytometer stain hoechst 33342 on human spermatozoa. Mol. Hum. Reprod. 1996, 2, 709–712. [Google Scholar] [CrossRef] [PubMed]
- Loontiens, F.G.; Regenfuss, P.; Zechel, A.; Dumortier, L.; Clegg, R.M. Binding characteristics of hoechst 33258 with calf thymus DNA, poly[d(a-t)], and d(ccggaattccgg): Multiple stoichiometries and determination of tight binding with a wide spectrum of site affinities. Biochemistry 1990, 29, 9029–9039. [Google Scholar] [CrossRef] [PubMed]
- Sandhu, L.C.; Warters, R.L.; Dethlefsen, L.A. Fluorescence studies of hoechst 33342 with supercoiled and relaxed plasmid pbr322 DNA. Cytometry 1985, 6, 191–194. [Google Scholar] [CrossRef] [PubMed]
- Dasari, M.; Lee, S.; Sy, J.; Kim, D.; Lee, S.; Brown, M.; Davis, M.; Murthy, N. Hoechst-ir: An imaging agent that detects necrotic tissue in vivo by binding extracellular DNA. Org. Lett. 2010, 12, 3300–3303. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, A.; Bianco, A.C.; Zhang, C.Y.; Lowell, B.B.; Frangioni, J.V. Quantitation of brown adipose tissue perfusion in transgenic mice using near-infrared fluorescence imaging. Mol. Imaging 2003, 2, 37–49. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, A.; Takigawa, K.; Kurishita, Y.; Kuwata, K.; Ishida, M.; Shimoda, Y.; Hamachi, I.; Tsukiji, S. Hoechst tagging: A modular strategy to design synthetic fluorescent probes for live-cell nucleus imaging. Chem. Commun. 2014, 50, 6149–6152. [Google Scholar] [CrossRef] [PubMed]
- Lukinavičius, G.; Blaukopf, C.; Pershagen, E.; Schena, A.; Reymond, L.; Derivery, E.; Gonzalez-Gaitan, M.; D’Este, E.; Hell, S.W.; Gerlich, D.W.; et al. Sir-hoechst is a far-red DNA stain for live-cell nanoscopy. Nat. Commun. 2015, 6, 8497. [Google Scholar] [CrossRef] [PubMed]
- Bentivoglio, M.; Kuypers, H.G.; Catsman-Berrevoets, C.E.; Loewe, H.; Dann, O. Two new fluorescent retrograde neuronal tracers which are transported over long distances. Neurosci. Lett. 1980, 18, 25–30. [Google Scholar] [CrossRef]
- Lee, H.S.; Mihailoff, G.A. Fluorescent double-label study of lateral reticular nucleus projections to the spinal cord and periaqueductal gray in the rat. Anat. Rec. 1999, 256, 91–98. [Google Scholar] [CrossRef]
- Katoh, Y.Y.; Arai, R.; Benedek, G. Bifurcating projections from the cerebellar fastigial neurons to the thalamic suprageniculate nucleus and to the superior colliculus. Brain Res. 2000, 864, 308–311. [Google Scholar] [CrossRef]
- Nagatomo, F.; Ishihara, A.; Ohira, Y. Effects of hindlimb unloading at early postnatal growth on cell body size in spinal motoneurons innervating soleus muscle of rats. Int. J. Dev. Neurosci. 2009, 27, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Karg, T.J.; Golic, K.G. Photoconversion of dapi and hoechst dyes to green and red-emitting forms after exposure to uv excitation. Chromosoma 2017. [Google Scholar] [CrossRef] [PubMed]
- Zurek-Biesiada, D.; Kedracka-Krok, S.; Dobrucki, J.W. Uv-activated conversion of hoechst 33258, dapi, and vybrant dyecycle fluorescent dyes into blue-excited, green-emitting protonated forms. Cytometry A 2013, 83, 441–451. [Google Scholar] [CrossRef] [PubMed]
- Szczurek, A.T.; Prakash, K.; Lee, H.K.; Zurek-Biesiada, D.J.; Best, G.; Hagmann, M.; Dobrucki, J.W.; Cremer, C.; Birk, U. Single molecule localization microscopy of the distribution of chromatin using hoechst and dapi fluorescent probes. Nucleus 2014, 5, 331–340. [Google Scholar] [CrossRef] [PubMed]
- Guan, Y.; Shi, R.; Li, X.; Zhao, M.; Li, Y. Multiple binding modes for dicationic hoechst 33258 to DNA. J. Phys. Chem. B 2007, 111, 7336–7344. [Google Scholar] [CrossRef] [PubMed]
- Pjura, P.E.; Grzeskowiak, K.; Dickerson, R.E. Binding of hoechst 33258 to the minor groove of B-DNA. J. Mol. Biol. 1987, 197, 257–271. [Google Scholar] [CrossRef]
- Aymami, J.; Nunn, C.M.; Neidle, S. DNA minor groove recognition of a non-self-complementary at-rich sequence by a tris-benzimidazole ligand. Nucleic Acids Res. 1999, 27, 2691–2698. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Li, T.K.; Kim, J.S.; LaVoie, E.J.; Breslauer, K.J.; Liu, L.F.; Pilch, D.S. DNA minor groove binding-directed poisoning of human DNA topoisomerase i by terbenzimidazoles. Biochemistry 1998, 37, 3558–3566. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Carrasco, C.; Kumar, A.; Stephens, C.E.; Bailly, C.; Boykin, D.W.; Wilson, W.D. Evaluation of the influence of compound structure on stacked-dimer formation in the DNA minor groove. Biochemistry 2001, 40, 2511–2521. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Kumar, A.; Boykin, D.W.; Bailly, C.; Wilson, W.D. Comparative thermodynamics for monomer and dimer sequence-dependent binding of a heterocyclic dication in the DNA minor groove. J. Mol. Biol. 2002, 317, 361–374. [Google Scholar] [CrossRef] [PubMed]
- Amirbekyan, K.; Duchemin, N.; Benedetti, E.; Joseph, R.; Colon, A.; Markarian, S.A.; Bethge, L.; Vonhoff, S.; Klussmann, S.; Cossy, J.; et al. Design, synthesis, and binding affinity evaluation of hoechst 33258 derivatives for the development of sequence-specific DNA-based asymmetric catalysts. ACS Catal. 2016, 6, 3096–3105. [Google Scholar] [CrossRef]
- Libbus, B.L.; Perreault, S.D.; Johnson, L.A.; Pinkel, D. Incidence of chromosome aberrations in mammalian sperm stained with hoechst 33342 and uv-laser irradiated during flow sorting. Mutat. Res. 1987, 182, 265–274. [Google Scholar] [CrossRef]
- Wagner, M.; Weber, P.; Bruns, T.; Strauss, W.S.; Wittig, R.; Schneckenburger, H. Light dose is a limiting factor to maintain cell viability in fluorescence microscopy and single molecule detection. Int. J. Mol. Sci. 2010, 11, 956–966. [Google Scholar] [CrossRef] [PubMed]
- Waldchen, S.; Lehmann, J.; Klein, T.; van de Linde, S.; Sauer, M. Light-induced cell damage in live-cell super-resolution microscopy. Sci. Rep. 2015, 5, 15348. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Wang, C.; Wang, L.; Ye, Z.-W.; Song, X.-B.; Xiao, Y. Hoechst-naphthalimide dyad with dual emissions as specific and ratiometric sensor for nucleus DNA damage. Chin. Chem. Lett. 2017, 28, 2019–2022. [Google Scholar] [CrossRef]
- Schoor, S.; Lung, S.-C.; Sigurdson, D.; Chuong, S.D.X. Fluorescent staining of living plant cells. In Plant Microtechniques and Protocols; Yeung, E.C.T., Stasolla, C., Sumner, M.J., Huang, B.Q., Eds.; Springer International Publishing: Cham, Switzerland, 2015; pp. 153–165. [Google Scholar]
- Nakamura, A.; Tsukiji, S. Ratiometric fluorescence imaging of nuclear ph in living cells using hoechst-tagged fluorescein. Bioorg. Med. Chem. Lett. 2017, 27, 3127–3130. [Google Scholar] [CrossRef] [PubMed]
- Tanabe, K.; Hara, D.; Umehara, Y.; Son, A.; Asahi, W.; Misu, S.; Kurihara, R.; Kondo, T. Tracking the oxygen status in the cell nucleus by using a hoechst-tagged phosphorescent ruthenium complex. Chembiochem 2018. [Google Scholar] [CrossRef]
- Dasari, M.; Acharya, A.P.; Kim, D.; Lee, S.; Lee, S.; Rhea, J.; Molinaro, R.; Murthy, N. H-gemcitabine: A new gemcitabine prodrug for treating cancer. Bioconj. Chem. 2013, 24, 4–8. [Google Scholar] [CrossRef] [PubMed]
- Kaul, M.; Pilch, D.S. Thermodynamics of aminoglycoside-rrna recognition: The binding of neomycin-class aminoglycosides to the a site of 16s rrna. Biochemistry 2002, 41, 7695–7706. [Google Scholar] [CrossRef] [PubMed]
- Arya, D.P.; Willis, B. Reaching into the major groove of b-DNA: Synthesis and nucleic acid binding of a neomycin-hoechst 33258 conjugate. J. Am. Chem. Soc. 2003, 125, 12398–12399. [Google Scholar] [CrossRef] [PubMed]
- Willis, B.; Arya, D.P. Recognition of b-DNA by neomycin—Hoechst 33258 conjugates. Biochemistry 2006, 45, 10217–10232. [Google Scholar] [CrossRef] [PubMed]
- Thamban Chandrika, N.; Shrestha, S.K.; Ranjan, N.; Sharma, A.; Arya, D.P.; Garneau-Tsodikova, S. New application of neomycin b-bisbenzimidazole hybrids as antifungal agents. ACS Infect. Dis. 2018, 4, 196–207. [Google Scholar] [CrossRef] [PubMed]
- Willis, B.; Arya, D.P. Triple recognition of B-DNA by a neomycin-hoechst 33258-pyrene conjugate. Biochemistry 2010, 49, 452–469. [Google Scholar] [CrossRef] [PubMed]
- Willis, B.; Arya, D.P. Recognition of rna duplex by a neomycin-hoechst 33258 conjugate. Bioorg. Med. Chem. 2014, 22, 2327–2332. [Google Scholar] [CrossRef] [PubMed]
- Ranjan, N.; Arya, D.P. Linker dependent intercalation of bisbenzimidazole-aminosugars in an rna duplex; selectivity in RNA vs. DNA binding. Bioorg. Med. Chem. Lett. 2016, 26, 5989–5994. [Google Scholar] [CrossRef] [PubMed]
- Ishida, M.; Watanabe, H.; Takigawa, K.; Kurishita, Y.; Oki, C.; Nakamura, A.; Hamachi, I.; Tsukiji, S. Synthetic self-localizing ligands that control the spatial location of proteins in living cells. J. Am. Chem. Soc. 2013, 135, 12684–12689. [Google Scholar] [CrossRef] [PubMed]
- Yasueda, Y.; Tamura, T.; Hamachi, I. Nucleus-selective chemical proteomics using hoechst-tagged reactive molecules. Chem. Lett. 2016, 45, 265–267. [Google Scholar] [CrossRef]
- Ranjan, N.; Kellish, P.; King, A.; Arya, D.P. Impact of linker length and composition on fragment binding and cell permeation: Story of a bisbenzimidazole dye fragment. Biochemistry 2017, 56, 6434–6447. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Yang, C.; Zhou, X.; Qin, J. Label-free aptamer-based sensors for l-argininamide by using nucleic acid minor groove binding dyes. Chem. Commun. 2011, 47, 3192–3194. [Google Scholar] [CrossRef] [PubMed]
- Ozaki, H.; Nishihira, A.; Wakabayashi, M.; Kuwahara, M.; Sawai, H. Biomolecular sensor based on fluorescence-labeled aptamer. Bioorg. Med. Chem. Lett. 2006, 16, 4381–4384. [Google Scholar] [CrossRef] [PubMed]
- Sarpong, K.; Datta, B. Nucleic-acid-binding chromophores as efficient indicators of aptamer-target interactions. J. Nucleic Acids 2012, 2012, 247280. [Google Scholar] [CrossRef] [PubMed]
- Le, H.-N.; Jiang, X.-Q.; Zhang, M.; Ye, B.-C. Label-free fluorescent assay of atp based on an aptamer-assisted light-up of hoechst dyes. Anal. Methods 2014, 6, 2028–2030. [Google Scholar] [CrossRef]
- Zhang, M.; Le, H.N.; Jiang, X.Q.; Ye, B.C. “Molecular beacon”-directed fluorescence of hoechst dyes for visual detection of hg(II) and biothiols and its application for a logic gate. Chem. Commun. 2013, 49, 2133–2135. [Google Scholar] [CrossRef] [PubMed]
- Paramanik, B.; Bain, D.; Patra, A. Making and breaking of DNA-metal base pairs: Hg2+ and au nanocluster based off/on probe. J. Phys. Chem. C 2016, 120, 17127–17135. [Google Scholar] [CrossRef]


| Probe | λabs, nm | λem, nm | QY 2, % | Kd, nM | F/F0 | References |
|---|---|---|---|---|---|---|
| Hoechst 33342 | 358 | 460 | 38 | 1–10 | 30–40 | [9,10] |
| Hoechst-IR 1 | 768 | 803 | 3.3 | 0.21 | – | [11,12] |
| hoeFL | 500 | 520 | 44 | 2500 | 94 | [13] |
| hoeBDP | 503 | 514 | 23 | 28 | 4.9 | [13] |
| hoeTMR | 550 | 580 | 8.9 | 1800 | 27 | [13] |
| SiR-Hoechst | 652 | 674 | 17 | 8400 | 50 | [14] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bucevičius, J.; Lukinavičius, G.; Gerasimaitė, R. The Use of Hoechst Dyes for DNA Staining and Beyond. Chemosensors 2018, 6, 18. https://doi.org/10.3390/chemosensors6020018
Bucevičius J, Lukinavičius G, Gerasimaitė R. The Use of Hoechst Dyes for DNA Staining and Beyond. Chemosensors. 2018; 6(2):18. https://doi.org/10.3390/chemosensors6020018
Chicago/Turabian StyleBucevičius, Jonas, Gražvydas Lukinavičius, and Rūta Gerasimaitė. 2018. "The Use of Hoechst Dyes for DNA Staining and Beyond" Chemosensors 6, no. 2: 18. https://doi.org/10.3390/chemosensors6020018