Comparative Characterization of Pseudoroegneria libanotica and Pseudoroegneria tauri Based on Their Repeatome Peculiarities
Abstract
:1. Introduction
2. Results
2.1. Repeatome Characterization
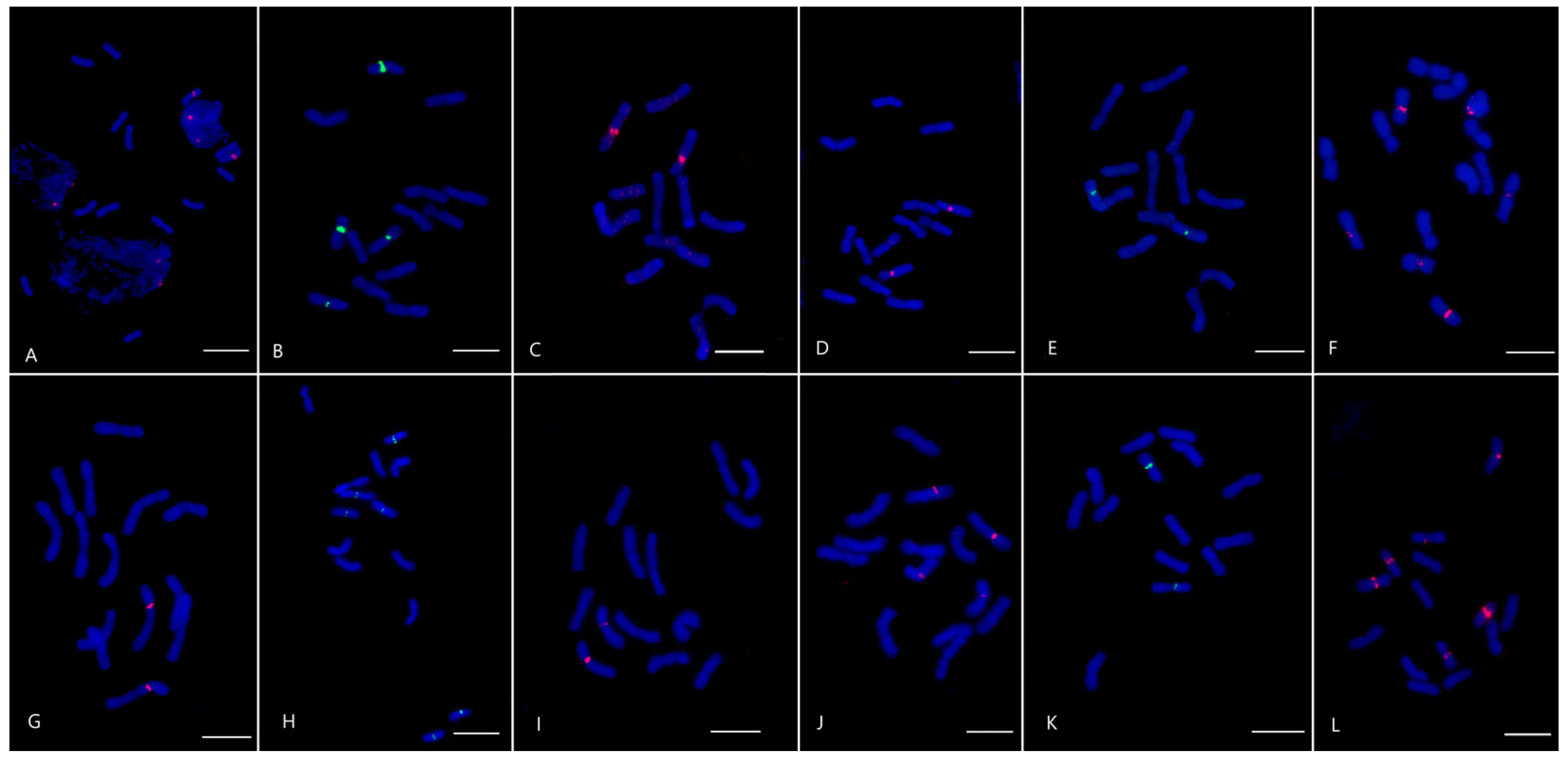
2.2. Satellite Repeats Characterization and Their Chromosomal Localization in Ps. libanotica and Ps. tauri
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Sequencing and Bioinformatics Analysis
4.3. Fluorescence In Situ Hybridization (FISH)
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Dewey, D.R. The Genomic System of Classification as a Guide to Intergeneric Hybridization with the Perennial Triticeae. In Gene Manipulation in Plant Improvement; Springer: Boston, MA, USA, 1984; pp. 209–279. [Google Scholar]
- Prive, K.; Orr, M.R.; Kilkenny, F.F.; Reuter, R.J.; Prendeville, H.R. Phenological Variation in Bluebunch Wheatgrass (Pseudoroegneria spicata): Implications for Seed Sourcing, Harvest, and Restoration. Land 2021, 10, 1064. [Google Scholar] [CrossRef]
- Kolb, P.F.; Robberecht, R. Pinus ponderosa Seedling Establishment and the Influence of Competition with the Bunchgrass Agropyron Spicatum. Int. J. Plant Sci. 1996, 157, 509–515. [Google Scholar] [CrossRef]
- Fraser, L.H.; Greenall, A.; Carlyle, C.; Turkington, R.; Friedman, C.R. Adaptive Phenotypic Plasticity of Pseudoroegneria spicata: Response of Stomatal Density, Leaf Area and Biomass to Changes in Water Supply and Increased Temperature. Ann. Bot. 2009, 103, 769–775. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Zhai, X.; Chen, C.; Yang, X.; Cheng, S.; Sha, L.; Cheng, Y.; Fan, X.; Kang, H.; Wang, Y.; et al. A Chromosome Level Genome Assembly of Pseudoroegneria libanotica Reveals a Key Kcs Gene Involves in the Cuticular Wax Elongation for Drought Resistance. Authorea 2023, 1–26. [Google Scholar] [CrossRef]
- Zhang, L.; Zhu, X.; Zhao, Y.; Guo, J.; Zhang, T.; Huang, W.; Huang, J.; Hu, Y.; Huang, C.-H.; Ma, H. Phylotranscriptomics Resolves the Phylogeny of Pooideae and Uncovers Factors for Their Adaptive Evolution. Mol. Biol. Evol. 2022, 39, msac026. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Fan, X.; Zhang, C.; Ding, C.; Wang, X.; Zhou, Y. Phylogenetic Relationships of Species in Pseudoroegneria (Poaceae: Triticeae) and Related Genera Inferred from Nuclear RDNA ITS (Internal Transcribed Spacer) Sequences. Biologia 2008, 63, 498–505. [Google Scholar] [CrossRef]
- Chen, N.; Chen, W.-J.; Yan, H.; Wang, Y.; Kang, H.-Y.; Zhang, H.-Q.; Zhou, Y.-H.; Sun, G.-L.; Sha, L.-N.; Fan, X. Evolutionary Patterns of Plastome Uncover Diploid-Polyploid Maternal Relationships in Triticeae. Mol. Phylogenet. Evol. 2020, 149, 106838. [Google Scholar] [CrossRef] [PubMed]
- Mahelka, V.; Kopecký, D.; Paštová, L. On the Genome Constitution and Evolution of Intermediate Wheatgrass (Thinopyrum intermedium: Poaceae, Triticeae). BMC Evol. Biol. 2011, 11, 127. [Google Scholar] [CrossRef]
- Fan, X.; Sha, L.-N.; Dong, Z.-Z.; Zhang, H.-Q.; Kang, H.-Y.; Wang, Y.; Wang, X.-L.; Zhang, L.; Ding, C.-B.; Yang, R.-W.; et al. Phylogenetic Relationships and Y Genome Origin in Elymus L. Sensu Lato (Triticeae; Poaceae) Based on Single-Copy Nuclear Acc1 and Pgk1 Gene Sequences. Mol. Phylogenet. Evol. 2013, 69, 919–928. [Google Scholar] [CrossRef]
- Wang, L.; Jiang, Y.; Shi, Q.; Wang, Y.; Sha, L.; Fan, X.; Kang, H.; Zhang, H.; Sun, G.; Zhang, L.; et al. Genome Constitution and Evolution of Elytrigia lolioides Inferred from Acc1, EF-G, ITS, TrnL-F Sequences and GISH. BMC Plant Biol. 2019, 19, 158. [Google Scholar] [CrossRef]
- Liu, Q.; Liu, L.; Ge, S.; Fu, L.; Bai, S.; Lv, X.; Wang, Q.; Chen, W.; Wang, F.; Wang, L.; et al. Endo-allopolyploidy of Autopolyploids and Recurrent Hybridization—A Possible Mechanism to Explain the Unresolved Y-genome Donor in Polyploid Elymus Species (Triticeae: Poaceae). J. Syst. Evol. 2022, 60, 344–360. [Google Scholar] [CrossRef]
- Agafonov, A.V.; Shabanova, E.V.; Emtseva, M.V.; Asbaganov, S.V.; Dorogina, O.V. Phylogenetic Relationships among Different Morphotypes of StY-Genomic Species Elymus Ciliaris and E. Amurensis (Poaceae) as a Unified Macroevolutional Complex. Bot. Pacifica 2021, 10, 19–28. [Google Scholar] [CrossRef]
- Mahelka, V.; Kopecký, D.; Baum, B.R. Contrasting Patterns of Evolution of 45S and 5S RDNA Families Uncover New Aspects in the Genome Constitution of the Agronomically Important Grass Thinopyrum intermedium (Triticeae). Mol. Biol. Evol. 2013, 30, 2065–2086. [Google Scholar] [CrossRef] [PubMed]
- Bieniek, W.; Mizianty, M.; Szklarczyk, M. Sequence Variation at the Three Chloroplast Loci (MatK, RbcL, TrnH-PsbA) in the Triticeae tribe (Poaceae): Comments on the Relationships and Utility in DNA Barcoding of Selected Species. Plant Syst. Evol. 2015, 301, 1275–1286. [Google Scholar] [CrossRef]
- Gao, G.; Tang, Z.; Wang, Q.; Gou, X.; Ding, C.; Zhang, L.; Zhou, Y.; Yang, R. Phylogeny and Maternal Donor of Kengyilia (Triticeae: Poaceae) Based on Chloroplast TrnT–TrnL Sequences. Biochem. Syst. Ecol. 2014, 57, 102–107. [Google Scholar] [CrossRef]
- Gao, G.; Gou, X.; Wang, Q.; Zhang, Y.; Deng, J.; Ding, C.; Zhang, L.; Zhou, Y.; Yang, R. Phylogenetic Relationships and Y Genome Origin in Chinese Elymus (Triticeae: Poaceae) Based on Single Copy Gene DMC1. Biochem. Syst. Ecol. 2014, 57, 420–426. [Google Scholar] [CrossRef]
- Wang, R.R.-C.; Larson, S.R.; Jensen, K.B.; Bushman, B.S.; DeHaan, L.R.; Wang, S.; Yan, X. Genome Evolution of Intermediate Wheatgrass as Revealed by EST-SSR Markers Developed from Its Three Progenitor Diploid Species. Genome 2015, 58, 63–70. [Google Scholar] [CrossRef]
- Gao, G.; Deng, J.; Gou, X.; Wang, Q.; Ding, C.; Zhang, L.; Zhou, Y.; Yang, R. Phylogenetic Relationships among Elymus and Related Diploid Genera (Triticeae: Poaceae) Based on Nuclear RDNA ITS Sequences. Biologia 2015, 70, 183–189. [Google Scholar] [CrossRef]
- Yang, Y.; Fan, X.; Wang, L.; Zhang, H.-Q.; Sha, L.-N.; Wang, Y.; Kang, H.-Y.; Zeng, J.; Yu, X.-F.; Zhou, Y.-H. Phylogeny and Maternal Donors of Elytrigia Desv. Sensu Lato (Triticeae; Poaceae) Inferred from Nuclear Internal-Transcribed Spacer and TrnL-F Sequences. BMC Plant Biol. 2017, 17, 207. [Google Scholar] [CrossRef]
- Lucía, V.; Martínez-Ortega, M.M.; Rico, E.; Anamthawat-Jónsson, K. Discovery of the Genus Pseudoroegneria (Triticeae, Poaceae) in the Western Mediterranean on Exploring the Generic Boundaries of Elymus. J. Syst. Evol. 2019, 57, 23–41. [Google Scholar] [CrossRef]
- Song, H.; Nan, Z.B.; Tian, P. Phylogenetic Analysis of Elymus (Poaceae) in Western China. Genet. Mol. Res. 2015, 14, 12228–12239. [Google Scholar] [CrossRef] [PubMed]
- Gamache, J.; Sun, G. Phylogenetic Analysis of the Genus Pseudoroegneria and the Triticeae tribe Using the RbcL Gene. Biochem. Syst. Ecol. 2015, 62, 73–81. [Google Scholar] [CrossRef]
- Gao, G.; Deng, J.; Zhang, Y.; Li, Y.; Li, W.; Zhou, Y.; Yang, R. Phylogeny and Maternal Donor of Chinese Elymus (Triticeae: Poaceae) Inferred from Chloroplast TrnH-PsbA Sequences. Biochem. Syst. Ecol. 2016, 68, 128–134. [Google Scholar] [CrossRef]
- Lei, Y.-X.; Liu, J.; Fan, X.; Sha, L.-N.; Wang, Y.; Kang, H.-Y.; Zhou, Y.-H.; Zhang, H.-Q. Phylogeny and Maternal Donor of Roegneria and Its Affinitive Genera (Poaceae: Triticeae) Based on Sequence Data for Two Chloroplast DNA Regions (NdhF and TrnH-PsbA). J. Syst. Evol. 2018, 56, 105–119. [Google Scholar] [CrossRef]
- Tang, C.; Qi, J.; Chen, N.; Sha, L.-N.; Wang, Y.; Zeng, J.; Kang, H.-Y.; Zhang, H.-Q.; Zhou, Y.-H.; Fan, X. Genome Origin and Phylogenetic Relationships of Elymus Villosus (Triticeae: Poaceae) Based on Single-Copy Nuclear Acc1, Pgk1, DMC1 and Chloroplast TrnL-F Sequences. Biochem. Syst. Ecol. 2017, 70, 168–176. [Google Scholar] [CrossRef]
- Lei, Y.X.; Fan, X.; Sha, L.N.; Wang, Y.; Kang, H.Y.; Zhou, Y.H.; Zhang, H.Q. Phylogenetic Relationships and the Maternal Donor of Roegneria (Triticeae: Poaceae) Based on Three Nuclear DNA Sequences (ITS, Acc1, and Pgk1) and One Chloroplast Region (TrnL-F). J. Syst. Evol. 2022, 60, 305–318. [Google Scholar] [CrossRef]
- Zeng, J.; Fan, X.; Zhang, L.; Wang, X.; Zhang, H.; Kang, H.; Zhou, Y. Molecular Phylogeny and Maternal Progenitor Implication in the Genus Kengyilia (Triticeae: Poaceae): Evidence from COXII Intron Sequences. Biochem. Syst. Ecol. 2010, 38, 202–209. [Google Scholar] [CrossRef]
- Dong, Z.-Z.; Fan, X.; Sha, L.-N.; Zeng, J.; Wang, Y.; Chen, Q.; Kang, H.-Y.; Zhang, H.-Q.; Zhou, Y.-H. Phylogeny and Molecular Evolution of the RbcL Gene of St Genome in Elymus Sensu Lato (Poaceae: Triticeae). Biochem. Syst. Ecol. 2013, 50, 322–330. [Google Scholar] [CrossRef]
- Hu, Q.; Yan, C.; Sun, G. Phylogenetic Analysis Revealed Reticulate Evolution of Allotetraploid Elymus Ciliaris. Mol. Phylogenet. Evol. 2013, 69, 805–813. [Google Scholar] [CrossRef]
- Liao, J.-Q.; Ross, L.; Fan, X.; Sha, L.-N.; Kang, H.-Y.; Zhang, H.-Q.; Wang, Y.; Liu, J.; Wang, X.-L.; Yu, X.-F.; et al. Phylogeny and Maternal Donors of the Tetraploid Species with St Genome (Poaceae: Triticeae) Inferred from CoxII and ITS Sequences. Biochem. Syst. Ecol. 2013, 50, 277–285. [Google Scholar] [CrossRef]
- Yan, C.; Hu, Q.; Sun, G. Nuclear and Chloroplast DNA Phylogeny Reveals Complex Evolutionary History of Elymus pendulinus. Genome 2014, 57, 97–109. [Google Scholar] [CrossRef]
- Dong, Z.-Z.; Fan, X.; Sha, L.-N.; Wang, Y.; Zeng, J.; Kang, H.-Y.; Zhang, H.-Q.; Wang, X.-L.; Zhang, L.; Ding, C.-B.; et al. Phylogeny and Differentiation of the St Genome in Elymus L. Sensu Lato (Triticeae; Poaceae) Based on One Nuclear DNA and Two Chloroplast Genes. BMC Plant Biol. 2015, 15, 179. [Google Scholar] [CrossRef] [PubMed]
- Hu, Q.; Sun, D.; Sun, G. Molecular Phylogeny Revealed Distinct Origin of the Y and St Genome in Elymus longearistatus (Triticeae: Poaceae). Mol. Phylogenet. Evol. 2015, 85, 141–149. [Google Scholar] [CrossRef]
- Mason-Gamer, R.J.; Burns, M.M.; Naum, M. Reticulate Evolutionary History of a Complex Group of Grasses: Phylogeny of Elymus StStHH Allotetraploids Based on Three Nuclear Genes. PLoS ONE 2010, 5, e10989. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.R.-C.; Li, X.; Robbins, M.D.; Larson, S.R.; Bushman, S.B.; Jones, T.A.; Thomas, A. DNA Sequence-Based Mapping and Comparative Genomics of the St Genome of Pseudoroegneria spicata (Pursh) Á. Löve versus Wheat (Triticum aestivum L.) and Barley (Hordeum vulgare L.). Genome 2020, 63, 445–457. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Xia, M.; Liu, D.; Jiang, L.; Shen, J.; Chen, W. Analysis of the Maternal Genome of Elymus nutans from the Qinghai-Tibet Plateau Based on Chloroplast Genomes. Grassl. Sci. 2022, 68, 114–123. [Google Scholar] [CrossRef]
- Qi, F.; Liang, S.; Xing, P.; Bao, Y.; Wang, R.R.-C.; Li, X. Genome Analysis of Thinopyrum intermedium and Its Potential Progenitor Species Using Oligo-FISH. Plants 2023, 12, 3705. [Google Scholar] [CrossRef]
- Harun, A.; Liu, H.; Song, S.; Asghar, S.; Wen, X.; Fang, Z.; Chen, C. Oligonucleotide Fluorescence In Situ Hybridization: An Efficient Chromosome Painting Method in Plants. Plants 2023, 12, 2816. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; Liu, H.; Miao, L.; He, L.; Xie, W.; Lan, H.; Yu, C.; Yan, W.; Wu, Y.; Wen, X.; et al. Molecular Cytogenetic Map Visualizes the Heterozygotic Genome and Identifies Translocation Chromosomes in Citrus sinensis. J. Genet. Genom. 2023, 50, 410–421. [Google Scholar] [CrossRef] [PubMed]
- Cai, B.; Wang, T.; Yue, F.; Harun, A.; Zhu, B.; Qian, W.; Ge, X.; Li, Z. Production and Cytology of Brassica Autoallohexaploids with Two and Four Copies of Two Subgenomes. Theor. Appl. Genet. 2022, 135, 2641–2653. [Google Scholar] [CrossRef]
- Yin, L.; Zhu, Z.; Luo, X.; Huang, L.; Li, Y.; Mason, A.S.; Yang, J.; Ge, X.; Long, Y.; Wang, J.; et al. Genome-Wide Duplication of Allotetraploid Brassica Napus Produces Novel Characteristics and Extensive Ploidy Variation in Self-Pollinated Progeny. G3 2020, 10, 3687–3699. [Google Scholar] [CrossRef] [PubMed]
- Divashuk, M.G.; Karlov, G.I.; Kroupin, P.Y. Copy Number Variation of Transposable Elements in Thinopyrum intermedium and Its Diploid Relative Species. Plants 2019, 9, 15. [Google Scholar] [CrossRef] [PubMed]
- Kroupin, P.Y.; Ulyanov, D.S.; Karlov, G.I.; Divashuk, M.G. The Launch of Satellite: DNA Repeats as a Cytogenetic Tool in Discovering the Chromosomal Universe of Wild Triticeae. Chromosoma 2023, 132, 65–88. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Shi, Q.; Su, H.; Wang, Y.; Sha, L.; Fan, X.; Kang, H.; Zhang, H.; Zhou, Y. St2-80: A New FISH Marker for St Genome and Genome Analysis in Triticeae. Genome 2017, 60, 553–563. [Google Scholar] [CrossRef]
- Linc, G.; Gaal, E.; Molnar, I.; Icsoa, D.; Badaeva, E.; Molnaar-Lang, M. Molecular Cytogenetic (FISH) and Genome Analysis of Diploid Wheatgrasses and Their Phylogenetic Relationship. PLoS ONE 2017, 12, e0173623. [Google Scholar] [CrossRef]
- Liu, R.; Yu, F.; Wei, L.; Liu, B.; Liu, D.; Dou, Q. Development and Application of Transposable Element-Based Chromosomal Markers for the St Genome in Triticeae. Cytogenet. Genome Res. 2020, 159, 215–224. [Google Scholar] [CrossRef]
- Divashuk, M.G.; Khuat, T.M.L.; Kroupin, P.Y.; Kirov, I.V.; Romanov, D.V.; Kiseleva, A.V.; Khrustaleva, L.I.; Alexeev, D.G.; Zelenin, A.S.; Klimushina, M.V.; et al. Variation in Copy Number of Ty3/Gypsy Centromeric Retrotransposons in the Genomes of Thinopyrum intermedium and Its Diploid Progenitors. PLoS ONE 2016, 11, e0154241. [Google Scholar] [CrossRef]
- Zhao, J.; Hao, W.; Tang, C.; Yao, H.; Li, B.; Zheng, Q.; Li, Z.; Zhang, X. Plasticity in Triticeae Centromere DNA Sequences: A Wheat × Tall Wheatgrass (Decaploid) Model. Plant J. 2019, 100, 314–327. [Google Scholar] [CrossRef]
- Wu, D.; Yang, N.; Xiang, Q.; Zhu, M.; Fang, Z.; Zheng, W.; Lu, J.; Sha, L.; Fan, X.; Cheng, Y.; et al. Pseudorogneria Libanotica Intraspecific Genetic Polymorphism Revealed by Fluorescence In Situ Hybridization with Newly Identified Tandem Repeats and Wheat Single-Copy Gene Probes. Int. J. Mol. Sci. 2022, 23, 14818. [Google Scholar] [CrossRef]
- Kroupin, P.Y.; Badaeva, E.D.; Sokolova, V.M.; Chikida, N.N.; Belousova, M.K.; Surzhikov, S.A.; Nikitina, E.A.; Kocheshkova, A.A.; Ulyanov, D.S.; Ermolaev, A.S.; et al. Aegilops Crassa Boiss. Repeatome Characterized Using Low-Coverage NGS as a Source of New FISH Markers: Application in Phylogenetic Studies of the Triticeae. Front. Plant Sci. 2022, 13, 980764. [Google Scholar] [CrossRef]
- Yen, C.; Yang, J.; Baum, B.R. Systematics of Triticeae; Agriculture Press: Bejing, China, 2007; Volume 4. [Google Scholar]
- Wang, R.R.; Dewey, D.R.; Hsiao, C. Genome Analysis of the Tetraploid Pseudoroegneri Tauri. Crop Sci. 1986, 26, 723–727. [Google Scholar] [CrossRef]
- Yang, Y.; Zhang, H.; Fan, X.; Sha, L.; Kang, H.; Wang, Y.; Zhou, Y. Polymorphism of Gliadin and Glutelin and Systematics Studies in Elytrigia. Chin. Bull. Bot. 2017, 52, 579–589. [Google Scholar] [CrossRef]
- Yan, C.; Sun, G. Nucleotide Divergence and Genetic Relationships of Pseudoroegneria Species. Biochem. Syst. Ecol. 2011, 39, 309–319. [Google Scholar] [CrossRef]
- Mason-Gamer, R.J. Phylogeny of a Genomically Diverse Group of Elymus (Poaceae) Allopolyploids Reveals Multiple Levels of Reticulation. PLoS ONE 2013, 8, e78449. [Google Scholar] [CrossRef] [PubMed]
- Yan, C.; Sun, G.; Sun, D. Distinct Origin of the Y and St Genome in Elymus Species: Evidence from the Analysis of a Large Sample of St Genome Species Using Two Nuclear Genes. PLoS ONE 2011, 6, e26853. [Google Scholar] [CrossRef] [PubMed]
- Markova, D.N.; Mason-Gamer, R.J. Diversity, Abundance, and Evolutionary Dynamics of Pong-like Transposable Elements in Triticeae. Mol. Phylogenet. Evol. 2015, 93, 318–330. [Google Scholar] [CrossRef] [PubMed]
- Tsujimoto, H.; Gill, B.S. Repetitive DNA Sequences from Polyploid Elymus trachycaulus and the Diploid Progenitor Species: Detection and Genomic Affinity of Elymus Chromatin Added to Wheat. Genome 1991, 34, 782–789. [Google Scholar] [CrossRef]
- Lei, J.; Zhou, J.; Sun, H.; Wan, W.; Xiao, J.; Yuan, C.; Karafiátová, M.; Doležel, J.; Wang, H.; Wang, X. Development of Oligonucleotide Probes for FISH Karyotyping in Haynaldia villosa, a Wild Relative of Common Wheat. Crop J. 2020, 8, 676–681. [Google Scholar] [CrossRef]
- Yu, Z.; Wang, H.; Jiang, W.; Jiang, C.; Yuan, W.; Li, G.; Yang, Z. Karyotyping Dasypyrum breviaristatum Chromosomes with Multiple Oligonucleotide Probes Reveals the Genomic Divergence in Dasypyrum. Genome 2021, 64, 789–800. [Google Scholar] [CrossRef]
- Jiang, C.; Liu, X.; Yang, Z.; Li, G. Chromosome Rearrangement in Elymus dahuricus Revealed by ND-FISH and Oligo-FISH Painting. Plants 2023, 12, 3268. [Google Scholar] [CrossRef]
- Nikitina, E.; Kuznetsova, V.; Kroupin, P.; Karlov, G.I.; Divashuk, M.G. Development of Specific Thinopyrum Cytogenetic Markers for Wheat-Wheatgrass Hybrids Using Sequencing and QPCR Data. Int. J. Mol. Sci. 2020, 21, 4495. [Google Scholar] [CrossRef] [PubMed]
- Tang, S.; Tang, Z.; Qiu, L.; Yang, Z.; Li, G.; Lang, T.; Zhu, W.; Zhang, J.; Fu, S. Developing New Oligo Probes to Distinguish Specific Chromosomal Segments and the A, B, D Genomes of Wheat (Triticum aestivum L.) Using ND-FISH. Front. Plant Sci. 2018, 9, 1104. [Google Scholar] [CrossRef] [PubMed]
- Komuro, S.; Endo, R.; Shikata, K.; Kato, A. Genomic and Chromosomal Distribution Patterns of Various Repeated DNA Sequences in Wheat Revealed by a Fluorescence in Situ Hybridization Procedure. Genome 2013, 56, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Tang, Y.; Yao, L.; Wu, H.; Tu, X.; Zhuang, L.; Qi, Z. Cytological and Molecular Characterization of Thinopyrum bessarabicum Chromosomes and Structural Rearrangements Introgressed in Wheat. Mol. Breed. 2019, 39, 146. [Google Scholar] [CrossRef]
- Wu, D.; Ruban, A.; Fuchs, J.; Macas, J.; Novák, P.; Vaio, M.; Zhou, Y.; Houben, A. Nondisjunction and Unequal Spindle Organization Accompany the Drive of Aegilops Speltoides B Chromosomes. New Phytol. 2019, 223, 1340–1352. [Google Scholar] [CrossRef] [PubMed]
- Klemme, S.; Banaei-Moghaddam, A.M.; Macas, J.; Wicker, T.; Novák, P.; Houben, A. High-copy Sequences Reveal Distinct Evolution of the Rye B Chromosome. New Phytol. 2013, 199, 550–558. [Google Scholar] [CrossRef] [PubMed]
- Han, H.; Liu, W.; Lu, Y.; Zhang, J.; Yang, X.; Li, X.; Hu, Z.; Li, L. Isolation and Application of P Genome-Specific DNA Sequences of Agropyron Gaertn. In Triticeae. Planta 2017, 245, 425–437. [Google Scholar] [CrossRef]
- Salina, E.A.; Adonina, I.G.; Vatolina, T.Y.; Kurata, N. A Comparative Analysis of the Composition and Organization of Two Subtelomeric Repeat Families in Aegilops speltoides Tausch. and Related Species. Genetica 2004, 122, 227–237. [Google Scholar] [CrossRef]
- Evtushenko, E.V.; Vershinin, A.V. Heterogeneous Organization of a Tandem Repeat Family in Subtelomeric Heterochromatin of Rye. Russ. J. Genet. 2010, 46, 1074–1076. [Google Scholar] [CrossRef]
- Appels, R.; Dennis, E.S.; Smyth, D.R.; Peacock, W.J. Two Repeated DNA Sequences from the Heterochromatic Regions of Rye (Secale cereale) Chromosomes. Chromosoma 1981, 84, 265–277. [Google Scholar] [CrossRef]
- Du, P.; Zhuang, L.; Wang, Y.; Yuan, L.; Wang, Q.; Wang, D.; Dawadondup; Tan, L.; Shen, J.; Xu, H.; et al. Development of Oligonucleotides and Multiplex Probes for Quick and Accurate Identification of Wheat and Thinopyrum bessarabicum Chromosomes. Genome 2017, 60, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Badaeva, E.D.; Zoshchuk, S.A.; Paux, E.; Gay, G.; Zoshchuk, N.V.; Roger, D.; Zelenin, A.V.; Bernard, M.; Feuillet, C. Fat Element—A New Marker for Chromosome and Genome Analysis in the Triticeae. Chromosome Res. 2010, 18, 697–709. [Google Scholar] [CrossRef] [PubMed]
- Said, M.; Hřibová, E.; Danilova, T.V.; Karafiátová, M.; Čížková, J.; Friebe, B.; Doležel, J.; Gill, B.S.; Vrána, J. The Agropyron Cristatum Karyotype, Chromosome Structure and Cross-Genome Homoeology as Revealed by Fluorescence In Situ Hybridization with Tandem Repeats and Wheat Single-Gene Probes. Theor. Appl. Genet. 2018, 131, 2213–2227. [Google Scholar] [CrossRef] [PubMed]
- Khuat, T.M.L. Analysis of the Organization of Repeated DNA Sequences in the Genomes of Wild Wheat Relatives. Ph.D. Thesis, Russian State Agrarian University—Moscow Timiryazev Agricultural Academy, Moscow, Russia, 2015. (In Russian). [Google Scholar]
- Kroupin, P.; Kuznetsova, V.; Romanov, D.; Kocheshkova, A.; Karlov, G.; Dang, T.X.; Khuat, T.M.L.; Kirov, I.; Alexandrov, O.; Polkhovskiy, A.; et al. Pipeline for the Rapid Development of Cytogenetic Markers Using Genomic Data of Related Species. Genes 2019, 10, 113. [Google Scholar] [CrossRef]
- Tang, Z.; Yang, Z.; Fu, S. Oligonucleotides Replacing the Roles of Repetitive Sequences PAs1, PSc119. 2, PTa-535, PTa71, CCS1, and PAWRC. 1 for FISH Analysis. J. Appl. Genet. 2014, 55, 313–318. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Song, X.; Xiao, J.; Sun, H.; Dai, K.; Lan, C.; Singh, P.; Yuan, C.; Zhang, S.; Singh, R.; et al. Development and Characterization of a Complete Set of Triticum aestivum—Roegneria ciliaris Disomic Addition Lines. Theor. Appl. Genet. 2018, 131, 1793–1806. [Google Scholar] [CrossRef]
- Grewal, S.; Yang, C.; Edwards, S.H.; Scholefield, D.; Ashling, S.; Burridge, A.J.; King, I.P.; King, J. Characterisation of Thinopyrum bessarabicum Chromosomes through Genome-Wide Introgressions into Wheat. Theor. Appl. Genet. 2018, 131, 389–406. [Google Scholar] [CrossRef]
- Kroupin, P.Y.; Kuznetsova, V.M.; Nikitina, E.A.; Martirosyan, Y.T.; Karlov, G.I.; Divashuk, M.G. Development of New Cytogenetic Markers for Thinopyrum ponticum (Podp.) Z.-W. Liu & R.-C. Wang. Comp. Cytogenet. 2019, 13, 231–243. [Google Scholar] [CrossRef]
- Villasante, A.; Abad, J.P.; Méndez-Lago, M. Centromeres Were Derived from Telomeres during the Evolution of the Eukaryotic Chromosome. Proc. Natl. Acad. Sci. USA 2007, 104, 10542–10547. [Google Scholar] [CrossRef]
- Ruiz-Herrera, A.; Nergadze, S.G.; Santagostino, M.; Giulotto, E. Telomeric Repeats Far from the Ends: Mechanisms of Origin and Role in Evolution. Cytogenet. Genome Res. 2008, 122, 219–228. [Google Scholar] [CrossRef]
- Li, G.; Zhang, T.; Yu, Z.; Wang, H.; Yang, E.; Yang, Z. An Efficient Oligo-FISH Painting System for Revealing Chromosome Rearrangements and Polyploidization in Triticeae. Plant J. 2021, 105, 978–993. [Google Scholar] [CrossRef] [PubMed]
- Badaeva, E.D.; Ruban, A.S.; Zoshchuk, S.A.; Surzhikov, S.A.; Knüpffer, H.; Kilian, B. Molecular cytogenetic characterization of Triticum timopheevii chromosomes provides new insight on genome evolution of T. zhukovskyi. Plant Syst. Evol. 2016, 302, 943–956. [Google Scholar] [CrossRef]
- Ruban, A.S.; Badaeva, E.D. Evolution of the S-genomes in Triticum-Aegilops alliance: Evidences from chromosome analysis. Front. Plant Sci. 2018, 9, 1756. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Friebe, B.; Gill, B.S. Fate of Aegilops speltoides-derived, repetitive DNA sequences in diploid Aegilops species, wheat-Aegilops amphiploids and derived chromosome addition lines. Cytogenet. Genome Res. 2010, 129, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Dai, Y.; Huang, S.; Sun, G.; Li, H.; Chen, S.; Gao, Y.; Chen, J. Origins and chromosome differentiation of Thinopyrum elongatum revealed by PepC and Pgk1 genes and ND-FISH. Genome 2021, 64, 901–913. [Google Scholar] [CrossRef]
- Li, J.; Bao, Y.; Han, R.; Wang, X.; Xu, W.; Li, G.; Yang, Z.; Zhang, X.; Li, X.; Liu, A.; et al. Molecular and cytogenetic identification of stem rust resistant wheat–Thinopyrum intermedium introgression lines. Plant Dis. 2021, 106, 2447–2454. [Google Scholar] [CrossRef]
- Li, G.; Gao, D.; Zhang, H.; Li, J.; Wang, H.; La, S.; Ma, J.; Yang, Z. Molecular cytogenetic characterization of Dasypyrum breviaristatum chromosomes in wheat background revealing the genomic divergence between Dasypyrum species. Mol. Cytogenet. 2016, 9, 1–9. [Google Scholar] [CrossRef]
- Chen, C.; Han, Y.; Xiao, H.; Zou, B.; Wu, D.; Sha, L.; Yang, C.; Liu, S.; Cheng, Y.; Wang, Y.; et al. Chromosome-specific painting in Thinopyrum species using bulked oligonucleotides. Theor. Appl. Genet. 2023, 136, 177. [Google Scholar] [CrossRef]

| Type of DNA Repeat | Ps. libanotica | Ps. tauri | Ps. spicata | |||
|---|---|---|---|---|---|---|
| Reads | Summarized% | Reads | Summarized% | Reads | Summarized% | |
| Unclassified_repeat (conflicting evidences) | 0 | 43.38 | 0 | 39.82 | 966 | 48.76 |
| |--rDNA | 0 | 0.41 | 0 | 0.22 | 0 | 0.25 |
| | |--45S_rDNA | 0 | 0.37 | 0 | 0.18 | 2385 | 0.18 |
| | | |--18S_rDNA | 3710 | 0.13 | 2684 | 0.11 | 0 | 0 |
| | | |--25S_rDNA | 7109 | 0.24 | 1532 | 0.07 | 1538 | 0.07 |
| | ′--5S_rDNA | 1267 | 0.04 | 1062 | 0.04 | 1551 | 0.07 |
| |--satellite | 157,704 | 5.35 | 55,132 | 2.36 | 122,341 | 5.42 |
| ′--mobile element | 0 | 37.62 | 0 | 37.24 | 0 | 43.05 |
| |--Class_I | 0 | 35.31 | 0 | 35.01 | 0 | 40.03 |
| | |--LTR | 187,564 | 35.25 | 108,460 | 34.96 | 137,616 | 39.98 |
| | | |--Ty1/Copia | 110 | 7.92 | 0 | 8.34 | 0 | 9.30 |
| | | | |--Ale | 493 | 0.02 | 0 | 0 | 0 | 0 |
| | | | |--Angela | 143,057 | 4.85 | 124,821 | 5.33 | 104,690 | 4.64 |
| | | | |--Bianca | 166 | 0.01 | 0 | 0 | 0 | 0 |
| | | | |--Ikeros | 1825 | 0.06 | 794 | 0.03 | 1172 | 0.05 |
| | | | |--SIRE | 80,047 | 2.71 | 64,755 | 2.77 | 99,682 | 4.41 |
| | | | |--TAR | 7413 | 0.25 | 4935 | 0.21 | 4685 | 0.2 |
| | | | |--Tork | 617 | 0.02 | 0 | 0 | 0 | 0 |
| | | ′--Ty3/Gypsy | 0 | 20.98 | 0 | 21.99 | 0 | 24.58 |
| | | |--non-chromovirus | 0 | 14.06 | 0 | 15.89 | 0 | 17.39 |
| | | | |--Athila | 305,327 | 10.35 | 288,182 | 12.31 | 263,871 | 11.69 |
| | | | |--Ogre | 3420 | 0.12 | 2465 | 0.1 | 23,767 | 1.05 |
| | | | ′--Retand | 105,876 | 3.59 | 81,421 | 3.48 | 104,927 | 4.65 |
| | | ′--chromovirus | 0 | 6.92 | 0 | 6.1 | 0 | 7.19 |
| | | |--CRM | 30,327 | 1.03 | 17,216 | 0.73 | 17,452 | 0.77 |
| | | |--Tekay | 173,721 | 5.89 | 125,519 | 5.37 | 144,859 | 6.42 |
| | ′--LINE | 1825 | 0.06 | 1276 | 0.05 | 1267 | 0.05 |
| ′--Class_II | 0 | 2.31 | 0 | 2.23 | 0 | 3.02 |
| | |--EnSpm/CACTA | 59,033 | 2 | 45,535 | 1.95 | 62,034 | 2.75 |
| | |--MuDR/Mutator | 5277 | 0.18 | 1668 | 0.07 | 5504 | 0.25 |
| | |--PIF/Harbinger | 3464 | 0.12 | 4840 | 0.21 | 306 | 0.02 |
| ′--Helitron | 237 | 0.01 | 0 | 0 | 0 | 0 |
| |--plastid | 38,982 | - | 40,246 | - | 16,336 | - |
| ′--mitochondria | 6774 | - | 0 | - | 4362 | - |
| Unclassified repeat (No evidence) | 335,904 | - | 251,051 | - | 206,323 | - |
| Repeat | Species of Origin | NCBI Accession | Identity to New Satellites, % | ||
|---|---|---|---|---|---|
| CL69 | CL101 | CL119 | |||
| Sc26c38_V112 | S. cereale | KC243240.1 | xxx ** | xxx | 74.2 |
| AesTR-183 | Ae. speltoides | MK283667.1 | xxx | xxx | 75.4 |
| pTa-465 | T. aestivum | KC290905.1 | xxx | xxx | 77.8 |
| CL131 | Ae. crassa | ON872663.1 | xxx | xxx | 79.0 |
| pAcPR5 | A. cristatum | KX390696.1 | xxx | xxx | 82.5 |
| BSCL156-3 | Th. bessarabicum | n/a * | xxx | xxx | 84.8 |
| CL149 | Th. bessarabicum | ON872689.1 | xxx | xxx | 85.0 |
| BSCL156-1 | Th. bessarabicum | n/a | xxx | xxx | 86.5 |
| 18-158 | Th. ponticum | n/a | xxx | xxx | 86.9 |
| BSCL156-2 | Th. bessarabicum | n/a | xxx | xxx | 89.7 |
| Oligo-1AL | T. aestivum | n/a | xxx | xxx | 90.0 |
| CL232 | Ae. crassa | ON872668.1 | xxx | xxx | 94.7 |
| CL211 | Th. bessarabicum | ON872686.1 | 71.9 | xxx | xxx |
| CL239 | Ae. crassa | ON872677.1 | 82.4 | 71.4 | xxx |
| oligo-6VS-57 | D. villosum | n/a | 92.5 | xxx | xxx |
| oligo-7E-744 | Th. elongatum | n/a | 98.2 | 79.5 | xxx |
| Spelt1 | Ae. speltoides | AY117402.1 | xxx | 68.3 | xxx |
| pSp1B16.1 | Ae. speltoides | FJ594248.1 | xxx | 69.7 | xxx |
| pSp1B16.3 | Ae. speltoides | FJ617549.1 | xxx | 76.3 | xxx |
| pSp1B16.4 | Ae. speltoides | FJ617550.1 | xxx | 69.0 | xxx |
| Tri-MS-6 | T. aestivum | EF469549.1 | xxx | 69.9 | xxx |
| Repeat | Species of Origin | NCBI Accession | Identity to New Satellites, % | |||||
|---|---|---|---|---|---|---|---|---|
| CL82 | CL89 | CL168 | CL185 | CL192 | CL207 | |||
| Oligo-1AS | A. speltoides | n/a * | xxx ** | 67.7 | xxx | xxx | xxx | 67.7 |
| StLIB98 | Ps. libanotica | OL685354.1 | 76.8 | xxx | xxx | xxx | xxx | xxx |
| oligo-7E-430 | Th. elongatum | n/a | 78.2 | xxx | xxx | xxx | xxx | xxx |
| oligo-5D151 | T. aestivum | n/a | 81.1 | xxx | xxx | xxx | xxx | xxx |
| S5 | Ps. stipifolia | n/a | 81.3 | xxx | xxx | xxx | 72.7 | xxx |
| pTa-451 | T. aestivum | KC290912.1 | 87.5 | xxx | xxx | xxx | xxx | xxx |
| CL149 | Th. bessarabicum | ON872689.1 | 94.4 | xxx | xxx | xxx | xxx | xxx |
| pAcPR3 | A. cristatum | KX390694.1 | xxx | xxx | xxx | xxx | 82.1 | xxx |
| FAT | T. aestivum | DX374230.1 | 81.2 | 74.6 | 91.7 | 82.9 | 73.0 | 90.5 |
| pAs1-4, oligo-pAs1-1, pAs1 | A. speltoides | n/a | xxx | 85,7 | xxx | xxx | 83.3 | xxx |
| RcAfa | Roegneria ciliaris | n/a | xxx | xxx | xxx | xxx | 82.9 | xxx |
| oligo-pTa535-1 | T. aestivum | n/a | xxx | xxx | xxx | xxx | 76.2 | xxx |
| CL3 | Ae. crassa | ON872662.1 | 84.4 | 85.7 | xxx | xxx | 74.4 | xxx |
| CL18 | Ae. crassa | n/a | 84.2 | 75.0 | 91.7 | 91.7 | xxx | 90.5 |
| ACRI_CL80 | A. cristatum | MG323513.1 | 94.4 | 70.4 | 78.0 | 74.1 | xxx | 67.1 |
| CL193 | Ae. crassa | ON872676.1 | 73.5 | 76.8 | 80.4 | 82.6 | xxx | xxx |
| CL148 | Th. bessarabicum | ON872688.1 | xxx | 72.1 | 76.4 | 78.5 | xxx | 68.75 |
| P631 | Ae. tauschii | MK256651.1 | 85.2 | 100,0 | 82.9 | 95.5 | xxx | xxx |
| P523 | Ae. tauschii | MK256655.1 | 77.2 | xxx | xxx | xxx | 100.0 | xxx |
| P720 | Ae. tasuchii | MK256649.1 | 94.4 | 80.6 | xxx | xxx | 72.9 | xxx |
| Repeat | Primers | Monomer Length, bp |
|---|---|---|
| CL69 | F: 5′-ACTACCTTTTCAAGCCACCGT-3′ R: 5′-GGAGGTCATATATGGAGACCTATTT-3′ | 178 |
| CL82 | F: 5′-TGACACCATGCCAAGTTTCAT-3′ R: 5′-GTGCATGTTTAGGTCCCATGC-3′ | 503 |
| CL89 | F: 5′-CACTGGGCACAACCAAAGTT-3′ R: 5′-ACAAAAGGGCTCCATGCACA-3′ | 658 |
| CL101 | F: 5′-TTAAGGATGGTTTGGGCAGC-3′ R: 5′-ACCACACGTCACTCTGAAACA-3′ | 177 |
| CL119 | F: 5′-CCTTTGACTTTCGCCGGAC-3′ R: 5′- CGACACGGAGGGAATCTTGC-3′ | 668 |
| CL168 | F: 5′-TTTTTGTGAAGCAAGTGCCAT-3′ R: 5′-TAGAGCACACTTGCAGTTCA-3′ | 476 |
| CL185 | F: 5′-CACATGGGATGCCAACTGC-3′ R: 5′-TGGTCGAAACTAGAGCACACT-3′ | 659 |
| CL192 | F: 5′-TATACGCCATTGGAAGCCCC-3′ R: 5′-ACTCGTTAGCACGCCCAAAT-3′ | 339 |
| CL207 | F: 5′-TTGGATGGCCACTGACCAAG-3′ R: 5′-TGGCAATTTTCAGGACCAAACT-3′ | 657 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kroupin, P.Y.; Yurkina, A.I.; Ulyanov, D.S.; Karlov, G.I.; Divashuk, M.G. Comparative Characterization of Pseudoroegneria libanotica and Pseudoroegneria tauri Based on Their Repeatome Peculiarities. Plants 2023, 12, 4169. https://doi.org/10.3390/plants12244169
Kroupin PY, Yurkina AI, Ulyanov DS, Karlov GI, Divashuk MG. Comparative Characterization of Pseudoroegneria libanotica and Pseudoroegneria tauri Based on Their Repeatome Peculiarities. Plants. 2023; 12(24):4169. https://doi.org/10.3390/plants12244169
Chicago/Turabian StyleKroupin, Pavel Yu., Anna I. Yurkina, Daniil S. Ulyanov, Gennady I. Karlov, and Mikhail G. Divashuk. 2023. "Comparative Characterization of Pseudoroegneria libanotica and Pseudoroegneria tauri Based on Their Repeatome Peculiarities" Plants 12, no. 24: 4169. https://doi.org/10.3390/plants12244169






