Recommendations for Improving Identification and Quantification in Non-Targeted, GC-MS-Based Metabolomic Profiling of Human Plasma
Abstract
:1. Introduction
2. Results
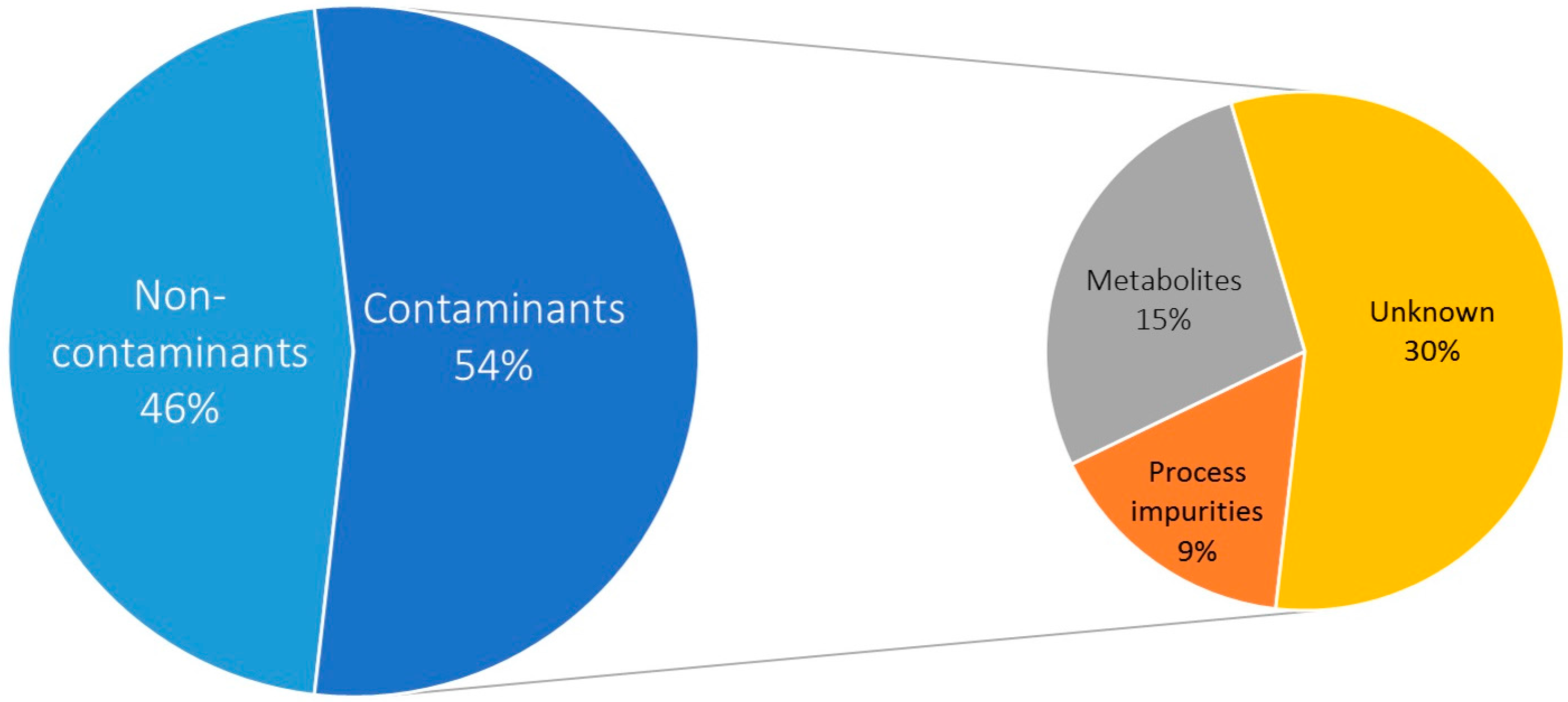
2.1. Selectivity: Contaminant Profile
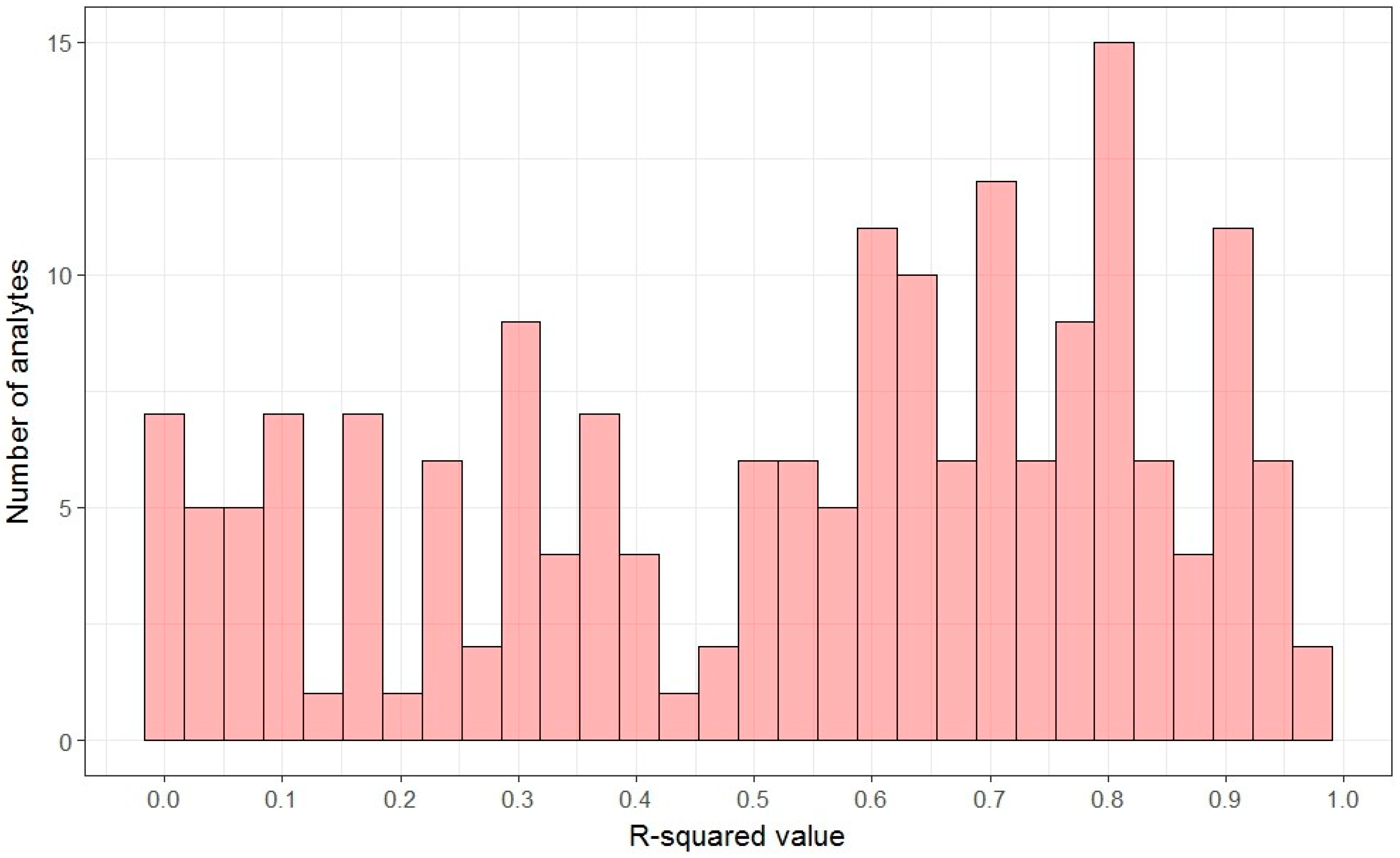
2.2. Linearity: Signal-Concentration Relationship
2.3. Linear Dynamic Range
2.4. Repeatability and Intermediate Precision
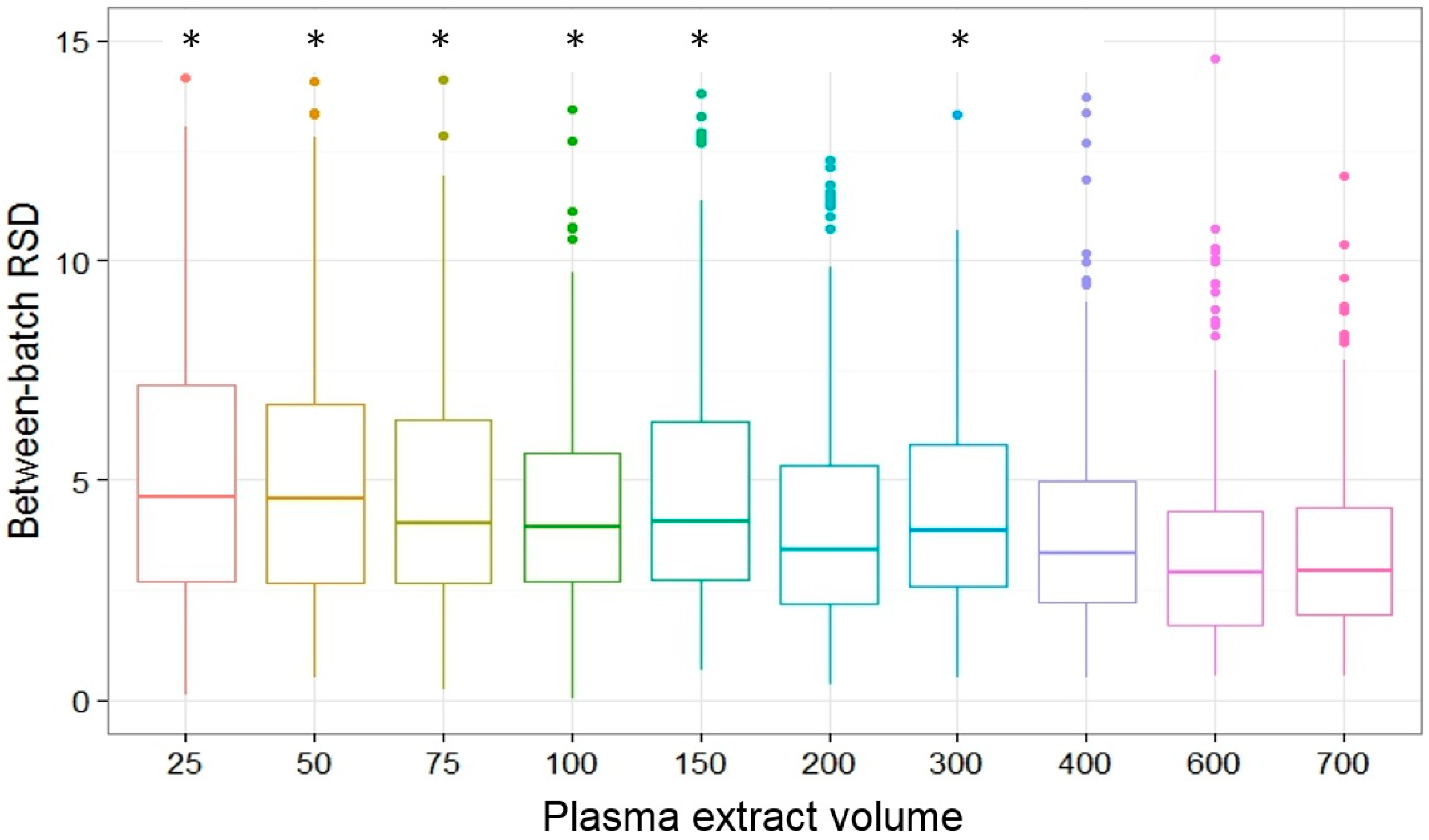
2.5. Concentration-Specific Study
3. Discussion
4. Materials and Methods
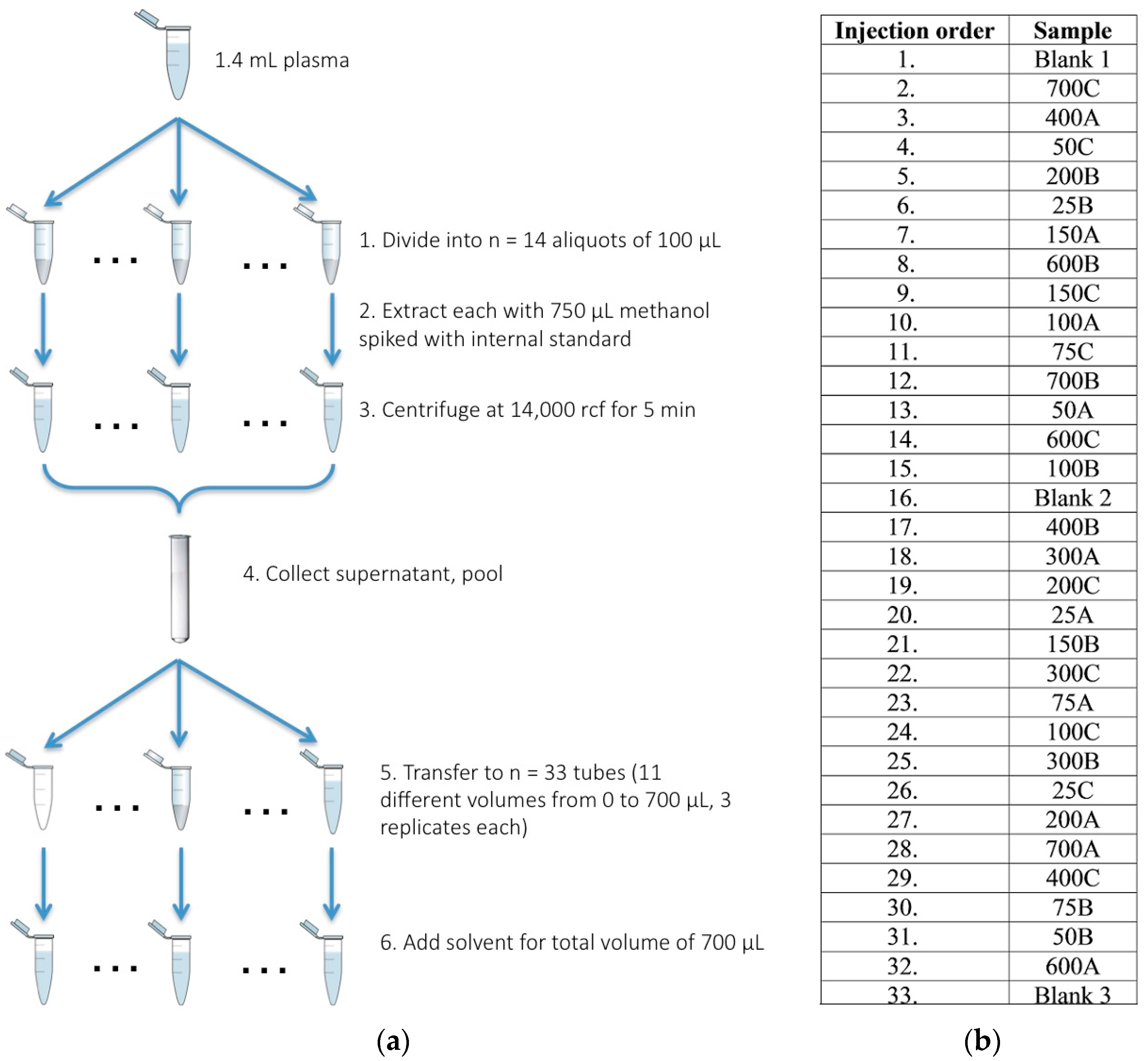
4.1. Sample Acquisition, Preparation, and Derivatization
4.2. GC-MS Analysis
4.3. Metabolite Identification and Quantification
4.4. Parameters Assessed for Method Development
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Beecher, C.W.W. The human metabolome. In Metabolic Profiling: Its Role in Biomarker Discovery and Gene Function Analysis; Springer: Berlin/Heidelberg, Germany, 2003; pp. 311–318. [Google Scholar]
- Wishart, D.S. Computational approaches to metabolomics. In Methods in Molecular Biology; Springer: Berlin/Heidelberg, Germany, 2010; pp. 263–282. [Google Scholar]
- Fiehn, O. Metabolomics—The link between genotypes and phenotypes. Plant Mol. Biol. 2002, 48, 155–171. [Google Scholar] [CrossRef] [PubMed]
- Barba, I.; Garcia-dorado, D. Metabolomics in cardiovascular disease : Towards clinical application. Coron. Artery Dis. 2012. Available online: https://cdn.intechopen.com/pdfs-wm/32774.pdf (accessed on 23 August 2017).
- Worley, B.; Powers, R. Multivariate analysis in metabolomics. Curr. Metabolom. 2013, 1, 92–107. [Google Scholar]
- Dunn, W.B.; Bailey, N.J.; Johnson, H.E. Measuring the metabolome: Current analytical technologies. Analyst 2005, 130, 606–625. [Google Scholar] [CrossRef] [PubMed]
- Naz, S.; Vallejo, M.; Garcia, A.; Barbas, C. Method validation strategies involved in non-targeted metabolomics. J. Chromatogr. A 2014, 1353, 99–105. [Google Scholar] [CrossRef] [PubMed]
- Koek, M.M.; Muilwijk, B.; Van Der Werf, M.J.; Hankemeier, T. Microbial metabolomics with gas chromatography/mass spectrometry. Anal. Chem. 2006, 78, 1272–1281. [Google Scholar] [CrossRef] [PubMed]
- Dunn, W.B.; Erban, A.; Weber, R.J.M.; Creek, D.J.; Brown, M.; Breitling, R.; Hankemeier, T.; Goodacre, R.; Neumann, S.; Kopka, J.; et al. Mass appeal: Metabolite identification in mass spectrometry-focused untargeted metabolomics. Metabolomics 2013, 9, 44–66. [Google Scholar] [CrossRef]
- Du, X.; Zeisel, S.H. Spectral deconvolution for gas chromatography mass spectrometry-based metabolomics: Current status and future perspectives. Comput. Struct. Biotechnol. J. 2013, 4, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Kopka, J. Current challenges and developments in GC-MS based metabolite profiling technology. J. Biotechnol. 2006, 124, 312–322. [Google Scholar] [CrossRef] [PubMed]
- Psychogios, N.; Hau, D.D.; Peng, J.; Guo, A.C.; Mandal, R.; Bouatra, S.; Sinelnikov, I.; Krishnamurthy, R.; Eisner, R.; Gautam, B.; et al. The human serum metabolome. PLoS ONE 2011, 6, e16957. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Jewison, T.; Guo, A.C.; Wilson, M.; Knox, C.; Liu, Y.; Djoumbou, Y.; Mandal, R.; Aziat, F.; Dong, E.; et al. HMDB 3.0-the human metabolome database in 2013. Nucleic Acids Res. 2013, 41, 801–807. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Klipfell, E.; Bennett, B.J.; Koeth, R.; Levison, B.S.; Dugar, B.; Feldstein, A.E.; Britt, E.B.; Fu, X.; Chung, Y.-M.; et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 2011, 472, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Brindle, J.T.; Antti, H.; Holmes, E.; Tranter, G.; Nicholson, J.K.; Bethell, H.W.L.; Clarke, S.; Schofield, P.M.; McKilligin, E.; Mosedale, D.E.; et al. Rapid and noninvasive diagnosis of the presence and severity of coronary heart disease using 1H-NMR-based metabonomics. Nat. Med. 2002, 8, 1439–1444. [Google Scholar] [CrossRef] [PubMed]
- Riedl, J.; Esslinger, S.; Fauhl-Hassek, C. Review of validation and reporting of non-targeted fingerprinting approaches for food authentication. Anal. Chim. Acta 2015, 885, 17–32. [Google Scholar] [CrossRef] [PubMed]
- Reza, P.R.; Masanori, M.S.; Correa, E.; Dayalan, S.; Tim, A.G. Data standards can boost metabolomics research, and if there is a will, there is a way. Metabolomics 2016, 12, 1–13. [Google Scholar]
- Lind, M.V.; Savolainen, O.I.; Ross, A.B. The use of mass spectrometry for analysing metabolite biomarkers in epidemiology: Methodological and statistical considerations for application to large numbers of biological samples. Eur. J. Epidemiol. 2016, 31, 717–733. [Google Scholar] [CrossRef] [PubMed]
- Kanani, H.; Chrysanthopoulos, P.K.; Klapa, M.I. Standardizing GC-MS metabolomics. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2008, 871, 191–201. [Google Scholar] [CrossRef] [PubMed]
- Gika, H.G.; Wilson, I.D.; Theodoridis, G.A. The Role of Mass Spectrometry in Nontargeted Metabolomics, 1st ed.; Elsevier: Philadelphia, PA, USA, 2014; Volume 63, pp. 213–233. [Google Scholar]
- Fiehn, O. Extending the breadth of metabolite profiling by gas chromatography coupled to mass spectrometry. TrAC Trends Anal. Chem. 2008, 27, 261–269. [Google Scholar] [CrossRef] [PubMed]
- Sumner, L.W.; Amberg, A.; Barrett, D.; Beale, M.H.; Beger, R.; Daykin, C.; Fan, T.W.M.; Fiehn, O.; Goodacre, R.; Griffin, J.L.; et al. Proposed minimum reporting standards for chemical analysis: Chemical analysis working group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics 2007, 3, 211–221. [Google Scholar] [CrossRef] [PubMed]
- Lisec, J.; Schauer, N.; Kopka, J.; Willmitzer, L.; Fernie, A.R. Gas chromatography mass spectrometry-based metabolite profiling in plants. Nat. Protoc. 2006, 1, 387–396. [Google Scholar] [CrossRef] [PubMed]
- Dunn, W.B.; Broadhurst, D.; Begley, P.; Zelena, E.; Francis-McIntyre, S.; Anderson, N.; Brown, M.; Knowles, J.D.; Halsall, A.; Haselden, J.N.; et al. Procedures for large-scale metabolic profiling of serum and plasma using gas chromatography and liquid chromatography coupled to mass spectrometry. Nat. Protoc. 2011, 6, 1060–1083. [Google Scholar] [CrossRef] [PubMed]
- Ammerlaan, W.; Trezzi, J.-P.; Lescuyer, P.; Mathay, C.; Hiller, K.; Betsou, F. Method validation for preparing serum and plasma samples from human blood for downstream proteomic, metabolomic, and circulating nucleic acid-based applications. Biopreserv. Biobank. 2014, 12, 269–280. [Google Scholar] [CrossRef] [PubMed]
- Godzien, J.; Alonso-Herranz, V.; Barbas, C.; Armitage, E.G. Controlling the quality of metabolomics data: New strategies to get the best out of the QC sample. Metabolomics 2014, 11, 518–528. [Google Scholar] [CrossRef]
- Dunn, W.B.; Wilson, I.D.; Nicholls, A.W.; Broadhurst, D. The importance of experimental design and QC samples in large-scale and MS-driven untargeted metabolomic studies of humans. Bioanalysis 2012, 4, 2249–2264. [Google Scholar] [CrossRef] [PubMed]
- Simon-Manso, Y.; Lowenthal, M.S.; Kilpatrick, L.E.; Sampson, M.L.; Telu, K.H.; Rudnick, P.A.; Mallard, W.G.; Bearden, D.W.; Schock, T.B.; Tchekhovskoi, D.V.; et al. Metabolite profiling of a NIST standard reference material for human plasma (SRM 1950): GC-MS, LC-MS, NMR, and clinical laboratory analyses, libraries, and web-based resources. Anal. Chem. 2013, 85, 11725–11731. [Google Scholar] [CrossRef] [PubMed]
- Sangster, T.; Major, H.; Plumb, R.; Wilson, A.J.; Wilson, I.D. A pragmatic and readily implemented quality control strategy for HPLC-MS and GC-MS-based metabonomic analysis. Analyst 2006, 131, 1075–1078. [Google Scholar] [CrossRef] [PubMed]
- Trutschel, D.; Schmidt, S.; Grosse, I.; Neumann, S. Experiment design beyond gut feeling: Statistical tests and power to detect differential metabolites in mass spectrometry data. Metabolomics 2015, 11, 851–860. [Google Scholar] [CrossRef]
- Sud, M.; Fahy, E.; Cotter, D.; Azam, K.; Vadivelu, I.; Burant, C.; Edison, A.; Fiehn, O.; Higashi, R.; Nair, K.S.; et al. Metabolomics workbench: An international repository for metabolomics data and metadata, metabolite standards, protocols, tutorials and training, and analysis tools. Nucleic Acids Res. 2016, 44, D463–D470. [Google Scholar] [CrossRef] [PubMed]
- Salek, R.M.; Arita, M.; Dayalan, S.; Ebbels, T.; Jones, A.R.; Neumann, S.; Rocca-Serra, P.; Viant, M.R.; Vizcaíno, J.A. Embedding standards in metabolomics: The metabolomics society data standards task group. Metabolomics 2015, 11, 782–783. [Google Scholar] [CrossRef]
- Lu, H.; Liang, Y.; Dunn, W.B.; Shen, H.; Kell, D.B. Comparative evaluation of software for deconvolution of metabolomics data based on GC-TOF-MS. TrAC Trends Anal. Chem. 2008, 27, 215–227. [Google Scholar] [CrossRef]
- Aa, J.; Trygg, J.; Gullberg, J.; Johansson, A.I.; Jonsson, P.; Antti, H.; Marklund, S.L.; Moritz, T. Extraction and GC/MS analysis of the human blood plasma metabolome. Anal. Chem. 2005, 77, 8086–8094. [Google Scholar] [CrossRef] [PubMed]
- Pasikanti, K.K.; Ho, P.C.; Chan, E.C.Y. Gas chromatography/mass spectrometry in metabolic profiling of biological fluids. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2008, 871, 202–211. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S. Computational strategies for metabolite identification in metabolomics. Bioanalysis 2009, 1, 1579–1596. [Google Scholar] [CrossRef] [PubMed]
- McNulty, N.P.; Yatsunenko, T.; Hsiao, A.; Faith, J.J.; Muegge, B.D.; Goodman, L.; Henrissat, B.; Oozeer, R.; Cools-Portier, S.; Gobert, G.; et al. The impact of a consortium of fermented milk strains on the gut microbiome of gnotobiotic mice and monozygotic twins. Sci. Transl. Med. 2011, 3. [Google Scholar] [CrossRef] [PubMed]
- Gustavo González, A.; Ángeles Herrador, M. A practical guide to analytical method validation, including measurement uncertainty and accuracy profiles. TrAC Trends Anal. Chem. 2007, 26, 227–238. [Google Scholar] [CrossRef]
- Christenson, R.H.; Duh, S.H. Methodological and analytic considerations for blood biomarkers. Prog. Cardiovasc. Dis. 2012, 55, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Styczynski, M.P.; Moxley, J.F.; Tong, L.V.; Walther, J.L.; Jensen, K.L.; Stephanopoulos, G.N. Systematic identification of conserved metabolites in GC/MS data for metabolomics and biomarker discovery. Anal. Chem. 2007, 79, 966–973. [Google Scholar] [CrossRef] [PubMed]
- Kind, T.; Wohlgemuth, G.; Lee, D.Y.; Lu, Y.; Palazoglu, M.; Shahbaz, S.; Fiehn, O. FiehnLib: Mass spectral and retention index libraries for metabolomics based on quadrupole and time-of-flight gas chromatography/mass spectrometry. Anal. Chem. 2009, 81, 10038–10048. [Google Scholar] [CrossRef] [PubMed]
- Kopka, J.; Schauer, N.; Krueger, S.; Birkemeyer, C.; Usadel, B.; Bergmüller, E.; Dörmann, P.; Weckwerth, W.; Gibon, Y.; Stitt, M.; Willmitzer, L.; et al. [email protected]: The Golm metabolome database. Bioinformatics 2005, 21, 1635–1638. [Google Scholar] [CrossRef] [PubMed]

| Type | Class | Metabolite | No. of Blanks (%) |
|---|---|---|---|
| Definite | Amino acids | Glycine | 6 (20%) |
| - | Benzene derivatives | Benzoic acid | 22 (73.3%) |
| - | Carbohydrates | Glucose and other aldohexoses | 20 (66.7%) |
| - | - | Sucrose and similar disaccharides | 10 (33.3%) |
| - | Fatty acids | Heptadecanoic acid or Octadecanol | 23 (76.7%) |
| - | - | Myristic acid or Pentadecanol | 27 (90%) |
| - | - | Nonanoic acid | 12 (40%) |
| - | - | Oleic acid | 12 (40%) |
| - | - | Palmitic acid | 27 (90%) |
| - | - | Pentadecanoic acid or Hexadecanol | 14 (46.7%) |
| - | - | Stearic acid | 27 (90%) |
| - | Lipids | alpha-Monopalmitin | 27 (90%) |
| - | - | beta-Monopalmitin | 27 (90%) |
| - | - | beta-Monostearin | 27 (90%) |
| - | - | Glycerol | 26 (86.7%) |
| - | - | Thymol | 15 (50%) |
| - | Organic acids | Pyruvic acid | 20 (66.7%) |
| - | - | Succinic acid | 7 (23.3%) |
| - | Other | Phosphoric acid | 23 (76.7%) |
| - | - | Uridine | 27 (90%) |
| Potential | Amino acids | Aspartic acid | 3 (10%) |
| - | Benzene derivatives | Gentisic acid | 4 (13.3%) |
| - | Phenol | 2 (6.7%) | |
| - | Carbohydrates | Fructose or similar ketohexose | 1 (3.3%) |
| - | Fatty acids | Arachidic acid or 1-Heneicosanol | 3 (10%) |
| - | - | Decanoic acid | 1 (3.3%) |
| - | - | Lauric acid | 4 (13.3%) |
| - | - | Methyl palmitate | 2 (6.7%) |
| - | - | Methyl stearate | 2 (6.7%) |
| - | Lipids | Gamma-Tocopherol | 2 (6.7%) |
| - | Organic acids | Acetoacetate or 2-Aminoisobutanoic acid | 3 (10%) |
| - | - | Glycolic acid | 2 (6.7%) |
| - | - | Lactic acid | 5 (16.7%) |
| - | - | Urea | 2 (6.7%) |
| - | Other | 1,2-Propanediol | 1 (3.3%) |
| - | - | 4-Hydroxypyridine or 3-Hydroxypyridine | 3 (10%) |
| - | - | Ethanolamine | 2 (6.7%) |
| - | - | O-Methylphosphate | 3 (10%) |
| - | - | Prunetin or similar isoflavone | 1 (3.3%) |
| Summary | No. (% ) | Known | Unknown |
|---|---|---|---|
| R2adj greater than 0.95 | 3 | 2 (1.6%) | 1 (1.8%) |
| R2adj (0.7, 0.95) | 64 | 52 (41.3%) | 12 (21.1%) |
| R2adj (0.5, 0.7) | 30 | 22 (29.7%) | 8 (21.1%) |
| R2adj less than 0.5 | 50 | 24 (32.5%) | 24 (63.2%) |
| Plasma Extract Volume (µL) | No. Analytes (%) |
|---|---|
| 75–100 | 8 (4.5%) |
| 100–150 | 100 (55.9%) |
| 150–200 | 62 (34.6%) |
| 200–300 | 6 (3.4%) |
| 300+ | 1 (0.6%) |
| Methanolic Plasma Extract Volume (µL) | Methanol/H2O Volume 1 (µL) | Equivalent Plasma Volume Injected 2 (nL) | Equivalent Plasma Concentration 3 (v/v) |
|---|---|---|---|
| 0 | 700 | 0 | 0 |
| 25 | 675 | 5.7 | 1.25 × 10−9 |
| 50 | 650 | 11.3 | 2.49 × 10−9 |
| 75 | 625 | 17.0 | 3.74 × 10−9 |
| 100 | 600 | 22.6 | 4.98 × 10−9 |
| 150 | 550 | 33.9 | 7.48 × 10−9 |
| 200 | 500 | 45.2 | 9.97 × 10−9 |
| 300 | 400 | 67.9 | 1.50 × 10−8 |
| 400 | 300 | 90.5 | 1.99 × 10−8 |
| 600 | 100 | 135.7 | 2.99 × 10−8 |
| 700 | 0 | 158.4 | 3.49 × 10−8 |
| Experimental Design | Recommendations |
|---|---|
| Establish method blanks | Include 3 blank samples in the beginning, middle and end of every sequence run |
| Use both blanks and manual curation for contaminant profiling | |
| Establish a list of highly reproducible and potential contaminants | |
| Linearity | Incorporate dilution into QC samples |
| Metabolites showing linearity can be used as targets to validate the methodology and monitor changes | |
| Lack of linearity may indicate contaminant effect or saturation effect | |
| Repeatability and intermediate precision | Batch should be included in reporting and analysis of non-targeted GC-MS profiling |
| Range | Linear dynamic range should be established through dilution studies |
| Optimal concentration established through dilution studies should be used for metabolic profiling | |
| Unknowns | Unknowns presenting as contaminants can be excluded from further analysis |
| Highly-linear unknowns may be biologically important metabolites | |
| Reproducible, highly linear and non-contaminant unknowns should be added to the library or databases for future references |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, H.; Muehlbauer, M.J.; O’Neal, S.K.; Newgard, C.B.; Hauser, E.R.; Bain, J.R.; Shah, S.H. Recommendations for Improving Identification and Quantification in Non-Targeted, GC-MS-Based Metabolomic Profiling of Human Plasma. Metabolites 2017, 7, 45. https://doi.org/10.3390/metabo7030045
Wang H, Muehlbauer MJ, O’Neal SK, Newgard CB, Hauser ER, Bain JR, Shah SH. Recommendations for Improving Identification and Quantification in Non-Targeted, GC-MS-Based Metabolomic Profiling of Human Plasma. Metabolites. 2017; 7(3):45. https://doi.org/10.3390/metabo7030045
Chicago/Turabian StyleWang, Hanghang, Michael J. Muehlbauer, Sara K. O’Neal, Christopher B. Newgard, Elizabeth R. Hauser, James R. Bain, and Svati H. Shah. 2017. "Recommendations for Improving Identification and Quantification in Non-Targeted, GC-MS-Based Metabolomic Profiling of Human Plasma" Metabolites 7, no. 3: 45. https://doi.org/10.3390/metabo7030045





