Influence of the RelA Activity on E. coli Metabolism by Metabolite Profiling of Glucose-Limited Chemostat Cultures
Abstract
:1. Introduction
2. Experimental Section
2.1. Bacterial Strains and Growth Conditions
2.2. Analytical Techniques
2.2.1. Quenching and Metabolite Extraction
2.2.2. Derivatization and GC-MS Analysis
2.3. Data Analysis
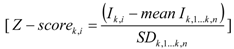
3. Results
3.1. Growth Parameters of E. coli Chemostat Cultures
| W3110 | ΔrelA mutant | |||||
|---|---|---|---|---|---|---|
| Dilution rate (h−1) | 0.05 | 0.10 | 0.20 | 0.05 | 0.10 | 0.20 |
| Biomass yield (gBiomass.gGlucose−1) | 0.36±0.056 | 0.44±0.15 | 0.55±0.10 | 0.46±0.063 | 0.46±0.064 | 0.67±0.3 |
| Biomass (gBiomass.L−1) | 1.8±0.28 | 2.2±0.34 | 2.7±0.43 | 2.3±0.31 | 2.3±0.32 | 3.3±0.45 |
| Glucose (gGlucose.L−1) | (1) | 0.029±0.0086 | 0.040±0.0033 | (1) | (1) | 0.023±0.010 |
| qGlucose (gGlucose.gBiomass−1.h−1) | 0.14±0.021 | 0.23±0.076 | 0.36±0.063 | 0.11±0.015 | 0.22±0.030 | 0.30±0.13 |
| Acetate (gAcetate.L−1) | (1) | (1) | 0.34 | (1) | (1) | 0.02 |
| qAcetate (×103)(gAcetate.gBiomass−1.h−1) | - | - | 25±3.8 | - | - | 1.1±0.15 |
3.2. Metabolite Profiling
| TCA intermediaries | Fatty acids | Amino acids | Others |
|---|---|---|---|
| alpha-ketoglutarate (akg) | Hexanoate (hxa, n-C6:0) | Aspartate (asp) | Benzoate* (bnz) |
| cis-Aconitate (acon-C) | Octanoate (octa, n-C8:0) | Isoleucine (ile) | NADP(H) |
| Citrate (cit) | Decanoate (dca, n-C10:0) | Lysine (lys) | Nicotinate (nac) |
| Fumarate (fum) | Tetradecanoate (ttdca, n-C14:0) | Threonine (thr) | Phosphoenolpyruvate (pep) |
| Malate (mal) | 10,13-Dimethyltetradecanoate (1013mlt) | Alanine (ala) | 5-oxo-D-proline*(pyrglu) |
| Succinate (succ) | Pentadecanoate (pdca, n-C15:0) | Leucine (leu) | Malonate* (ma) |
| 14-Methylpentadecanoate (14mpdca) | Valine (val) | Itaconate* (itcon) | |
| Octadecanoate (ocdca, n-C18:0) | Glycine (gly) | Lactate (lac) | |
| Octadecenoate (ocdcea, n-C18:1) | Serine (ser) | ||
| 9-cis,12-cis-Octadecadienoate (ocdcin, n-C18:2) | Glutamate (glu) | ||
| Proline (pro) | |||
| Phenylalanine (phe) | |||
| (2S)-2-isopropylmalate (3c3hmp) | |||
| N-Acetyl-L-glutamate (acglu) |

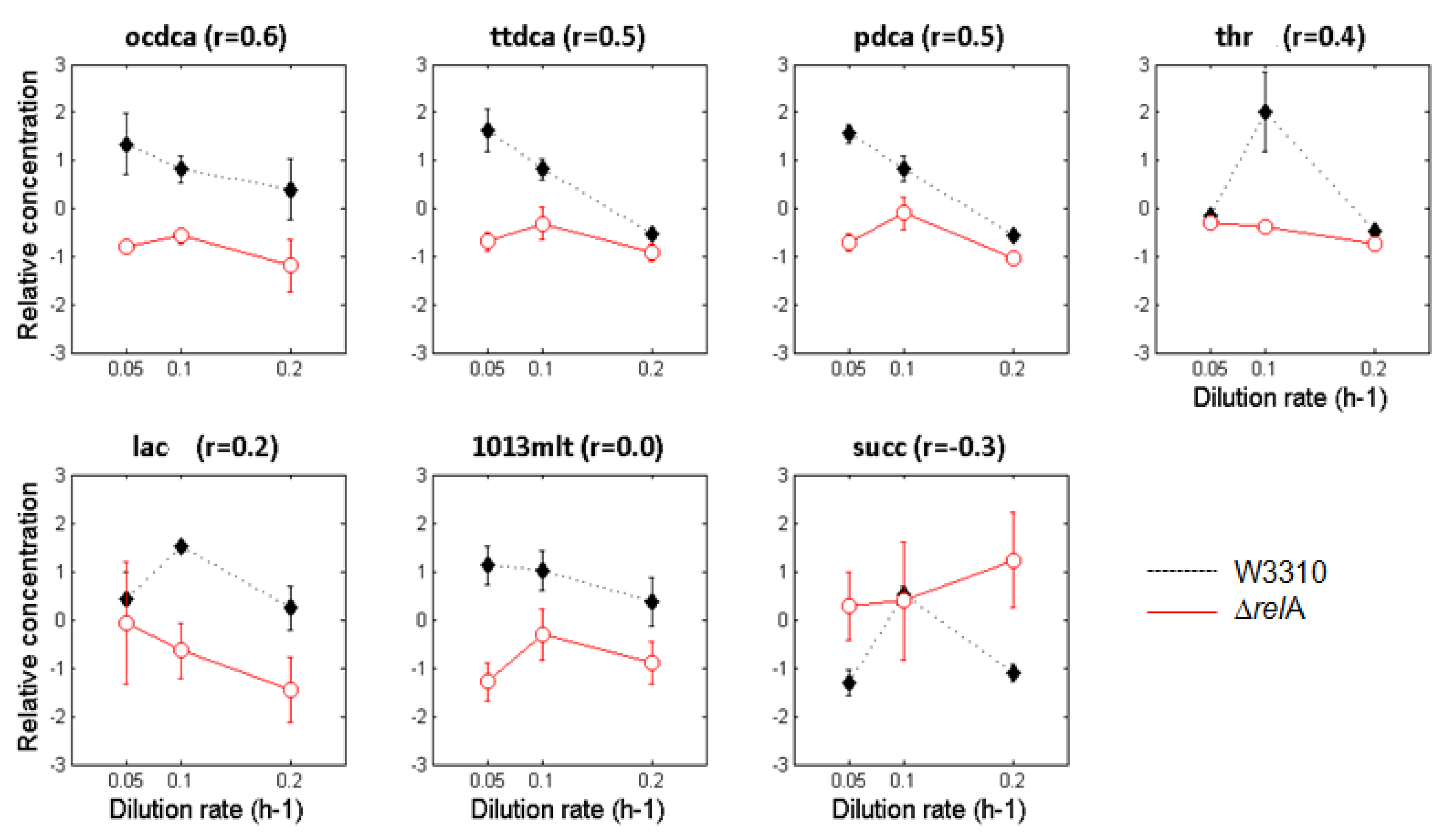
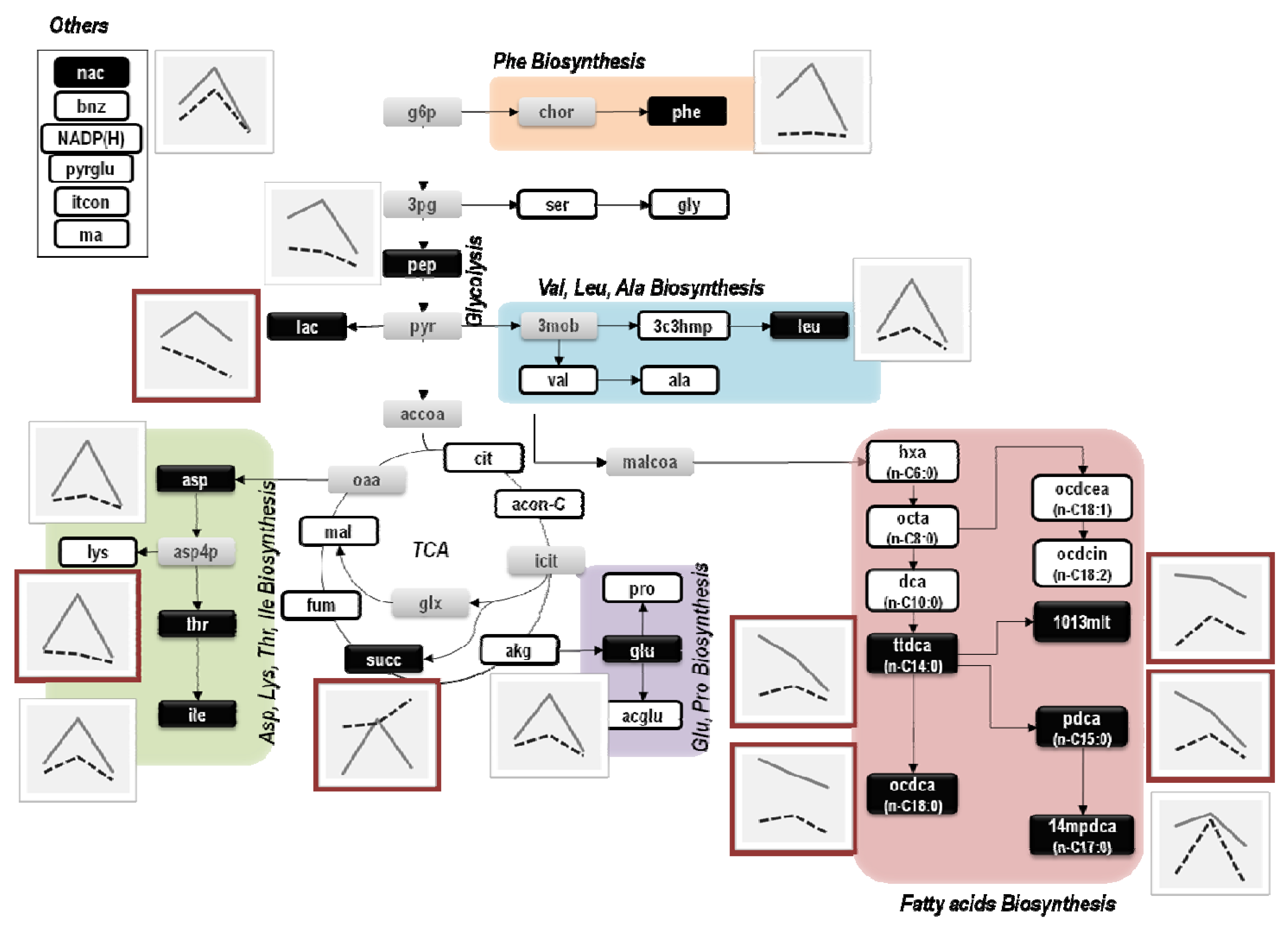
4. Discussion
5. Conclusions
Acknowledgments
Conflict of Interest
References
- Nanchen, A.; Schicker, A.; Sauer, U. Nonlinear dependency of intracellular fluxes on growth rate in miniaturized continuous cultures of Escherichia coli. Appl. Environ. Microbiol. 2006, 72, 1164–1172. [Google Scholar] [CrossRef]
- Ihssen, J.; Egli, T. Specific growth rate and not cell density controls the general stress response in Escherichia coli. Microbiology 2004, 150, 1637–1648. [Google Scholar] [CrossRef]
- Magnusson, L.U.; Farewell, A.; Nystrom, T. ppGpp: a global regulator in Escherichia coli. Trends Microbiol. 2005, 13, 236–242. [Google Scholar] [CrossRef]
- Jain, V.; Kumar, M.; Chatterji, D. ppGpp: Stringent response and survival. J. Microbiol. 2006, 44, 1–10. [Google Scholar]
- Chatterji, D.; Ojha, A.K. Revisiting the stringent response, ppGpp and starvation signaling. Curr. Opin. Microbiol. 2001, 4, 160–165. [Google Scholar] [CrossRef]
- Haseltin, W.A.; Block, R. Synthesis of guanosinetetraphosphate and pentaphosphate requires presence of a codon-specific, uncharged transfer ribonucleic acid in acceptor site of ribosomes - (Stringent control ppGpp (Msi) and pppGpp (Msii) protein synthesis Escherichia coli). Proc. Natl. Acad. Sci. USA 1973, 70, 1564–1568. [Google Scholar] [CrossRef]
- Torok, I.; Kari, C. Accumulation of ppGpp in a relA mutant of Escherichia coli during amino acid starvation. J. Biol. Chem. 1980, 255, 3838–3840. [Google Scholar]
- Traxler, M.F.; Chang, D.E.; Conway, T. Guanosine 3',5'-bispyrophosphate coordinates global gene expression during glucose-lactose diauxie in Escherichia coli. Proc. Natl. Acad. Sci. USA 2006, 103, 2374–2379. [Google Scholar] [CrossRef]
- Ostling, J.; Holmquist, L.; Kjelleberg, S. Global analysis of the carbon starvation response of a marine Vibrio species with disruptions in genes homologous to relA and spoT. J. Bacteriol. 1996, 178, 4901–4908. [Google Scholar]
- Ropers, D.; de Jong, H.; Page, M.; Schneider, D.; Geiselmann, J. Qualitative simulation of the carbon starvation response in Escherichia coli. Biosystems 2006, 84, 124–152. [Google Scholar] [CrossRef]
- Johnson, G.S.; Adler, C.R.; Collins, J.J.; Court, D. Role of the spoT gene product and manganese ion in the metabolism of guanosine 5'-diphosphate 3'-diphosphate in Escherichia coli. J. Biol. Chem. 1979, 254, 5483–5487. [Google Scholar]
- Murray, K.D.; Bremer, H. Control of spoT-dependent ppGpp synthesis and degradation in Escherichia coli. J. Mol. Biol. 1996, 259, 41–57. [Google Scholar] [CrossRef]
- Xiao, H.; Kalman, M.; Ikehara, K.; Zemel, S.; Glaser, G.; Cashel, M. Residual guanosine 3',5'-bispyrophosphate synthetic activity of relA null mutants can be eliminated by spoT null mutations. J. Biol. Chem. 1991, 266, 5980–5990. [Google Scholar]
- Dedhia, N.; Richins, R.; Mesina, A.; Chen, W. Improvement in recombinant protein production in ppGpp-deficient Escherichia coli. Biotechnol. Bioeng. 1997, 53, 380–386. [Google Scholar]
- Carneiro, S.; Villas-Boas, S.G.; Ferreira, E.C.; Rocha, I. Metabolic footprint analysis of recombinant Escherichia coli strains during fed-batch fermentations. Mol. Biosyst. 2011, 7, 899–910. [Google Scholar]
- Rocha, I.; Ferreira, E.C. On-line simultaneous monitoring of glucose and acetate with FIA during high cell density fermentation of recombinant E. coli. Anal. Chim. Acta 2002, 462, 293–304. [Google Scholar] [CrossRef]
- Smart, K.F.; Aggio, R.B.M.; van Houtte, J.R.; Villas-Boas, S.G. Analytical platform for metabolome analysis of microbial cells using methyl chloroformatederivatization followed by gas chromatography-mass spectrometry. Nat. Protocols 2010, 5, 1709–1729. [Google Scholar] [CrossRef]
- Miller, G.L. Use of dinitrosalicylic acid reagent for the determination of reducing sugar. Anal. Chem. 1959, 31, 426–428. [Google Scholar] [CrossRef]
- Villas-Boas, S.G.; Delicado, D.G.; Akesson, M.; Nielsen, J. Simultaneous analysis of amino and nonamino organic acids as methyl chloroformate derivatives using gas chromatography-mass spectrometry. Anal. Biochem. 2003, 322, 134–138. [Google Scholar] [CrossRef]
- Stein, S.E. An integrated method for spectrum extraction and compound identification from gas chromatography/mass spectrometry data. J. Am. Soc. Mass Spectrom. 1999, 10, 770–781. [Google Scholar] [CrossRef]
- Chagoyen, M.; Pazos, F. MBRole: enrichment analysis of metabolomic data. Bioinformatics 2011, 27, 730–731. [Google Scholar] [CrossRef]
- Aggio, R.B.; Ruggiero, K.; Villas-Boas, S.G. Pathway Activity Profiling (PAPi): from the metabolite profile to the metabolic pathway activity. Bioinformatics 2010, 26, 2969–2976. [Google Scholar] [CrossRef]
- Hua, Q.; Yang, C.; Oshima, T.; Mori, H.; Shimizu, K. Analysis of gene expression in Escherichia coli in response to changes of growth-limiting nutrient in chemostat cultures. Appl. Environ. Microbiol. 2004, 70, 2354–2366. [Google Scholar] [CrossRef]
- Nanchen, A.; Schicker, A.; Revelles, O.; Sauer, U. Cyclic AMP-dependent catabolite repression is the dominant control mechanism of metabolic fluxes under glucose limitation in Escherichia coli. J. Bacteriol. 2008, 190, 2323–2330. [Google Scholar] [CrossRef]
- Prasad, M.R.; Yu, P.L.; Seeto, S.; Ferenci, T. The role of isocitratelyase and the glyoxylate cycle in Escherichia coli growing under glucose limitation. Res. Microbiol. 2005, 156, 178–183. [Google Scholar] [CrossRef]
- Fischer, E.; Sauer, U. A novel metabolic cycle catalyzes glucose oxidation and anaplerosis in hungry Escherichia coli. J. Biol. Chem. 2003, 278, 46446–46451. [Google Scholar] [CrossRef]
- Sauer, U.; Eikmanns, B.J. The PEP-pyruvate-oxaloacetate node as the switch point for carbon flux distribution in bacteria. FEMS Microbiol. Rev. 2005, 29, 765–794. [Google Scholar] [CrossRef]
- Ferenci, T. Regulation by nutrient limitation. Curr. Opin. Microbiol. 1999, 2, 208–213. [Google Scholar] [CrossRef]
- Weber, J.; Kayser, A.; Rinas, U. Metabolic flux analysis of Escherichia coli in glucose-limited continuous culture. II. Dynamic response to famine and feast, activation of the methylglyoxal pathway and oscillatory behaviour. Microbiology 2005, 151, 707–716. [Google Scholar] [CrossRef]
- Kayser, A.; Weber, J.; Hecht, V.; Rinas, U. Metabolic flux analysis of Escherichia coli in glucose-limited continuous culture. I. Growth-rate-dependent metabolic efficiency at steady state. Microbiology 2005, 151, 693–706. [Google Scholar] [CrossRef]
- Wu, J.; Xie, J. Magic spot: (p) ppGpp. J. Cell Physiol. 2009, 220, 297–302. [Google Scholar] [CrossRef]
- Stephens, J.C.; Artz, S.W.; Ames, B.N. Guanosine 5'-diphosphate 3'-diphosphate (ppGpp) - Positive effector for histidineoperon transcription and general signal for amino acid deficiency. Proc. Natl. Acad. Sci. USA 1975, 72, 4389–4393. [Google Scholar] [CrossRef]
- Neubauer, P.; Ahman, M.; Tornkvist, M.; Larsson, G.; Enfors, S.O. Response of guanosinetetraphosphate to glucose fluctuations in fed-batch cultivations of Escherichia coli. J. Biotechnol. 1995, 43, 195–204. [Google Scholar]
- Mukherjee, T.K.; Raghavan, A.; Chatterji, D. Shortage of nutrients in bacteria: The stringent response. Curr. Sci. 1998, 75, 684–689. [Google Scholar]
- Traxler, M.F.; Summers, S.M.; Nguyen, H.T.; Zacharia, V.M.; Hightower, G.A.; Smith, J.T.; Conway, T. The global, ppGpp-mediated stringent response to amino acid starvation in Escherichia coli. Mol. Microbiol. 2008, 68, 1128–1148. [Google Scholar] [CrossRef]
- Renilla, S.; Bernal, V.; Fuhrer, T.; Castano-Cerezo, S.; Pastor, J.M.; Iborra, J.L.; Sauer, U.; Canovas, M. Acetate scavenging activity in Escherichia coli: interplay of acetyl-CoAsynthetase and the PEP-glyoxylate cycle in chemostat cultures. Appl. Microbiol. Biotechnol. 2012, 93, 2109–2124. [Google Scholar] [CrossRef]
- Valgepea, K.; Adamberg, K.; Nahku, R.; Lahtvee, P.J.; Arike, L.; Vilu, R. Systems biology approach reveals that overflow metabolism of acetate in Escherichia coli is triggered by carbon catabolite repression of acetyl-CoA synthetase. BMC Syst. Biol. 2010. [Google Scholar]
- Dong, T.; Schellhorn, H.E. Control of RpoS in global gene expression of Escherichia coli in minimal media. Mol. Genet. Genomics 2009, 281, 19–33. [Google Scholar] [CrossRef]
- Battesti, A.; Bouveret, E. Acyl carrier protein/SpoT interaction, the switch linking SpoT-dependent stress response to fatty acid metabolism. Mol. Microbiol. 2006, 62, 1048–1063. [Google Scholar] [CrossRef]
- Tuomanen, E.; Markiewicz, Z.; Tomasz, A. Autolysis-resistant peptidoglycan of anomalous composition in amino-acid-starved Escherichia coli. J. Bacteriol. 1988, 170, 1373–1376. [Google Scholar]
- Tuomanen, E.; Cozens, R. Changes in peptidoglycan composition and penicillin-binding proteins in slowly growing Escherichia coli. J. Bacteriol. 1987, 169, 5308–5310. [Google Scholar]
- Jones, P.G.; Cashel, M.; Glaser, G.; Neidhardt, F.C. Function of a relaxed-like state following temperature downshifts in Escherichia coli. J. Bacteriol. 1992, 174, 3903–3914. [Google Scholar]
- Schweder, T.; Hofmann, K.; Hecker, M. Escherichia coli K12 relA strains as safe hosts for expression of recombinant DNA. Appl. Environ. Microbiol. 1995, 42, 718–723. [Google Scholar]
- Lazzarin, R.A.; Cashel, M.; Gallant, J. On the regulation of guanosinetetraphosphate levels in stringent and relaxed strains of Escherichia coli. J. Biol. Chem. 1971, 246, 4381–4385. [Google Scholar]
- Traxler, M.F.; Zacharia, V.M.; Marquardt, S.; Summers, S.M.; Nguyen, H.T.; Stark, S.E.; Conway, T. Discretely calibrated regulatory loops controlled by ppGpp partition gene induction across the 'feast to famine' gradient in Escherichia coli. Mol. Microbiol. 2011, 79, 830–845. [Google Scholar] [CrossRef]
- Pao, C.C.; Dyess, B.T. Effect of unusual guanosine nucleotides on the activities of some Escherichia coli cellular enzymes. Biochim. Biophys. Acta 1981, 677, 358–362. [Google Scholar] [CrossRef]
- Bino, R.J.; Hall, R.D.; Fiehn, O.; Kopka, J.; Saito, K.; Draper, J.; Nikolau, B.J.; Mendes, P.; Roessner-Tunali, U.; Beale, M.H.; et al. Potential of metabolomics as a functional genomics tool. Trends Plant Sci. 2004, 9, 418–425. [Google Scholar] [CrossRef]
- Buchholz, A.; Hurlebaus, J.; Wandrey, C.; Takors, R. Metabolomics: quantification of intracellular metabolite dynamics. Biomol. Eng. 2002, 19, 5–15. [Google Scholar] [CrossRef]
- Coucheney, E.; Daniell, T.J.; Chenu, C.; Nunan, N. Gas chromatographic metabolic profiling: A sensitive tool for functional microbial ecology. J. Microbiol. Methods 2008, 75, 491–500. [Google Scholar] [CrossRef]
- Villas-Boas, S.G.; Moxley, J.F.; Akesson, M.; Stephanopoulos, G.; Nielsen, J. High-throughput metabolic state analysis: the missing link in integrated functional genomics of yeasts. Biochem. J. 2005, 388, 669–677. [Google Scholar] [CrossRef]
- Ferullo, D.J.; Lovett, S.T. The stringent response and cell cycle arrest in Escherichia coli. PLoS Genet. 2008, 4, e1000300. [Google Scholar] [CrossRef]
Supplementary Files
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Carneiro, S.; Villas-Bôas, S.G.; Ferreira, E.C.; Rocha, I. Influence of the RelA Activity on E. coli Metabolism by Metabolite Profiling of Glucose-Limited Chemostat Cultures. Metabolites 2012, 2, 717-732. https://doi.org/10.3390/metabo2040717
Carneiro S, Villas-Bôas SG, Ferreira EC, Rocha I. Influence of the RelA Activity on E. coli Metabolism by Metabolite Profiling of Glucose-Limited Chemostat Cultures. Metabolites. 2012; 2(4):717-732. https://doi.org/10.3390/metabo2040717
Chicago/Turabian StyleCarneiro, Sónia, Silas G. Villas-Bôas, Eugénio C. Ferreira, and Isabel Rocha. 2012. "Influence of the RelA Activity on E. coli Metabolism by Metabolite Profiling of Glucose-Limited Chemostat Cultures" Metabolites 2, no. 4: 717-732. https://doi.org/10.3390/metabo2040717
APA StyleCarneiro, S., Villas-Bôas, S. G., Ferreira, E. C., & Rocha, I. (2012). Influence of the RelA Activity on E. coli Metabolism by Metabolite Profiling of Glucose-Limited Chemostat Cultures. Metabolites, 2(4), 717-732. https://doi.org/10.3390/metabo2040717






