Ordered and Ushered; the Assembly and Translocation of the Adhesive Type I and P Pili
Abstract
:1. Introduction
2. The Pilus and Its Subunits
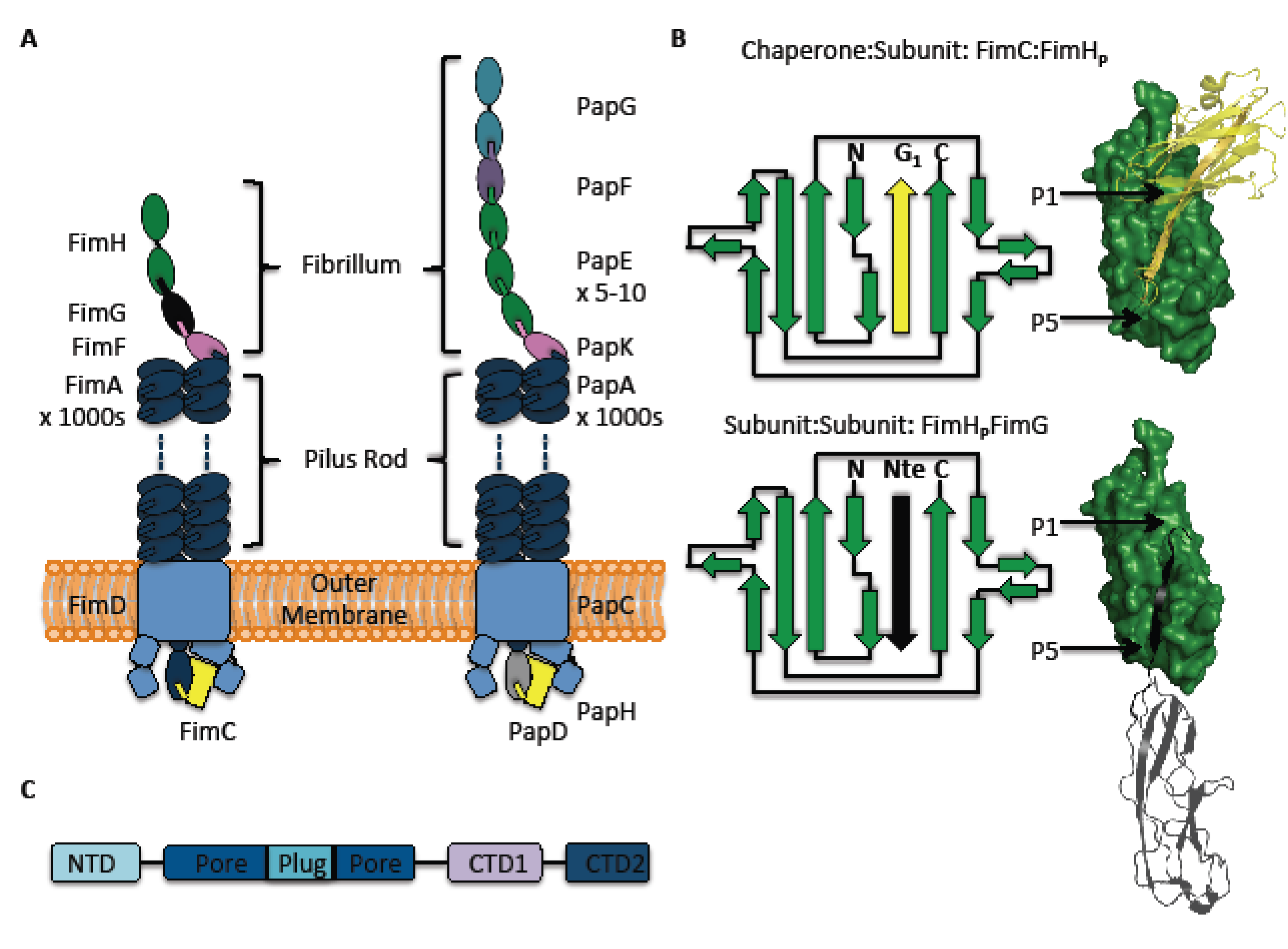
3. How the Subunits are Processed by the Usher
3.1. Mechanism of Subunit Polymerization at the Usher
3.2. Regulation of Ordering
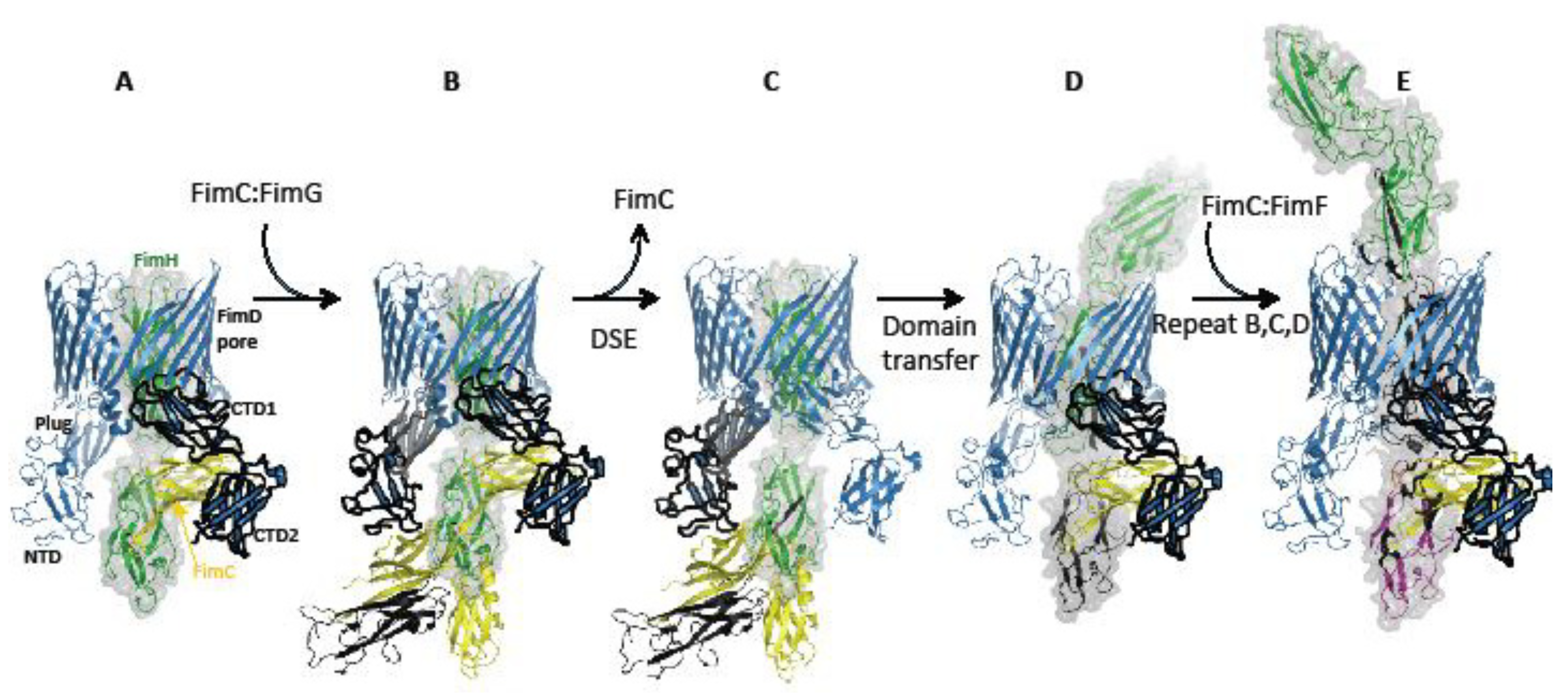
4. Passing of Subunits through the Usher
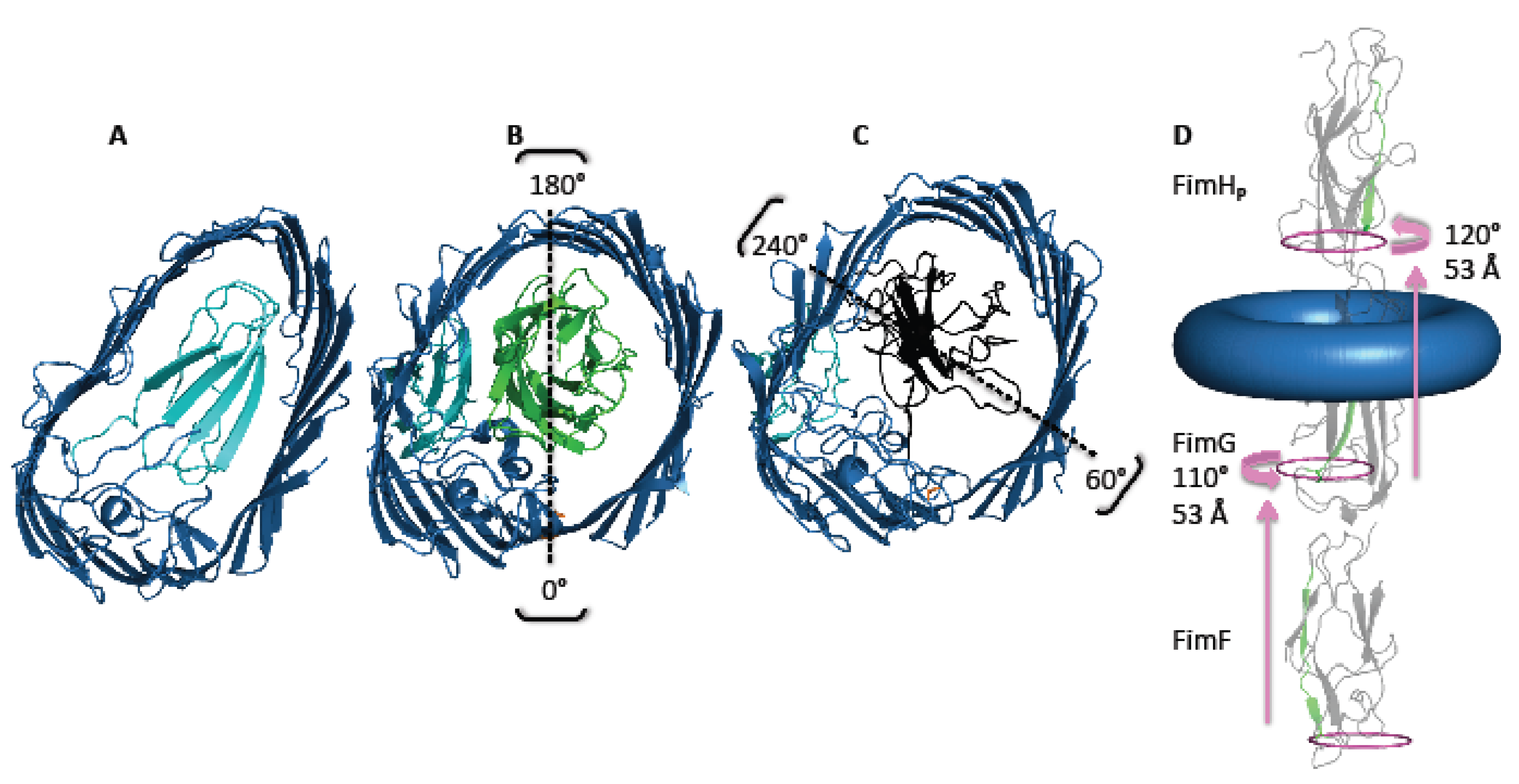
5. Emergence from the Pore and Adhesion
6. Preventing Adhesion

7. Concluding Remarks
Conflict of Interest
Acknowledgments
References and Notes
- Ulett, G.C.; Totsika, M.; Schaale, K.; Carey, A.J.; Sweet, M.J.; Schembri, M.A. Uropathogenic Escherichia coli virulence and innate immune responses during urinary tract infection. Curr. Opin. Microbiol. 2013, 16, 100–107. [Google Scholar]
- Hannan, T.J.; Totsika, M.; Mansfield, K.J.; Moore, K.H.; Schembri, M.A.; Hultgren, S.J. Host-pathogen checkpoints and population bottlenecks in persistent and intracellular uropathogenic Escherichia coli bladder infection. FEMS Microbiol. Rev. 2012, 36, 616–648. [Google Scholar] [CrossRef]
- Allen, W.J.; Phan, G.; Waksman, G. Pilus biogenesis at the outer membrane of Gram-negative bacterial pathogens. Curr. Opin. Struct. Biol. 2012, 22, 500–506. [Google Scholar] [CrossRef]
- Geibel, S.; Waksman, G. Crystallography and electron microscopy of chaperone/usher pilus systems. Adv. Exp. Med. Biol. 2011, 715, 159–174. [Google Scholar] [CrossRef]
- Thanassi, D.G.; Bliska, J.B.; Christie, P.J. Surface organelles assembled by secretion systems of Gram-negative bacteria: diversity in structure and function. FEMS Microbiol. Rev. 2012, 36, 1046–1082. [Google Scholar] [CrossRef]
- Dalbey, R.E.; Kuhn, A. Protein traffic in Gram-negative bacteria-how exported and secreted proteins find their way. FEMS Microbiol. Rev. 2012, 36, 1023–1045. [Google Scholar] [CrossRef]
- Silverman, P.M. Towards a structural biology of bacterial conjugation. Mol. Microbiol. 1997, 23, 423–429. [Google Scholar] [CrossRef]
- Clarke, M.; Maddera, L.; Harris, R.L.; Silverman, P.M. F-pili dynamics by live-cell imaging. PNAS 2008, 105, 17978–17981. [Google Scholar]
- Wallden, K.; Rivera-Calzada, A.; Waksman, G. Type IV secretion systems: versatility and diversity in function. Cell. Microbiol. 2010, 12, 1203–1212. [Google Scholar]
- Fronzes, R.; Christie, P.J.; Waksman, G. The structural biology of type IV secretion systems. Nature reviews. Microbiology 2009, 7, 703–714. [Google Scholar] [CrossRef]
- Deane, J.E.; Abrusci, P.; Johnson, S.; Lea, S.M. Timing is everything: the regulation of type III secretion. CMLS 2010, 67, 1065–1075. [Google Scholar] [CrossRef]
- Izoré, T.; Job, V.; Dessen, A. Biogenesis, regulation, and targeting of the type III secretion system. Structure 2011, 19, 603–612. [Google Scholar] [CrossRef]
- Burrows, L.L. Pseudomonas aeruginosa twitching motility: type IV pili in action. Annu. Rev. Microbiol. 2012, 66, 493–520. [Google Scholar] [CrossRef]
- Gibiansky, M.L.; Conrad, J.C.; Jin, F.; Gordon, V.D.; Motto, D.A.; Mathewson, M.A.; Stopka, W.G.; Zelasko, D.C.; Shrout, J.D.; Wong, G.C.L. Bacteria use type IV pili to walk upright and detach from surfaces. Science 2010, 330, 197. [Google Scholar] [CrossRef]
- Pelicic, V. Type IV pili: e pluribus unum? Mol. Microbiol. 2008, 68, 827–837. [Google Scholar] [CrossRef]
- Barnhart, M.M.; Chapman, M.R. Curli biogenesis and function. Annu. Rev. Microbiol. 2006, 60, 131–147. [Google Scholar] [CrossRef]
- Nuccio, S.-P.; Bäumler, A.J. Evolution of the chaperone/usher assembly pathway: Fimbrial classification goes Greek. MMBR 2007, 71, 551–575. [Google Scholar]
- Busch, A.; Waksman, G. Chaperone-usher pathways: diversity and pilus assembly mechanism. Philos. Trans. R. Soc. London Ser. B 2012, 367, 1112–1122. [Google Scholar] [CrossRef]
- Melican, K.; Sandoval, R.M.; Kader, A.; Josefsson, L.; Tanner, G.A.; Molitoris, B.A.; Richter-Dahlfors, A. Uropathogenic Escherichia coli P and Type 1 fimbriae act in synergy in a living host to facilitate renal colonization leading to nephron obstruction. PLoS Pathog. 2011, 7, e1001298. [Google Scholar] [CrossRef]
- Abraham, S.N.; Sun, D.; Dale, J.B.; Beachey, E.H. Conservation of the D-mannose-adhesion protein among type 1 fimbriated members of the family Enterobacteriaceae. Nature 1988, 336, 682–684. [Google Scholar] [CrossRef]
- Roberts, J.A.; Marklund, B.I.; Ilver, D.; Haslam, D.; Kaack, M.B.; Baskin, G.; Louis, M.; Möllby, R.; Winberg, J.; Normark, S. The Gal(alpha 1-4)Gal-specific tip adhesin of Escherichia coli P-fimbriae is needed for pyelonephritis to occur in the normal urinary tract. PNAS 1994, 91, 11889–11893. [Google Scholar] [CrossRef]
- Zhang, J.P.; Normark, S. Induction of gene expression in Escherichia coli after pilus-mediated adherence. Science 1996, 273, 1234–1236. [Google Scholar]
- Schilling, J.D.; Mulvey, M.A.; Vincent, C.D.; Lorenz, R.G.; Hultgren, S.J. Bacterial invasion augments epithelial cytokine responses to Escherichia coli through a lipopolysaccharide-dependent mechanism. J. Immunol. 2001, 166, 1148–1155. [Google Scholar]
- Henderson, N.S.; Thanassi, D.G. Purification of the outer membrane usher protein and periplasmic chaperone-subunit complexes from the P and type 1 pilus systems. Meth. Mol. Biol. 2013, 966, 37–52. [Google Scholar] [CrossRef]
- Phan, G.; Remaut, H.; Wang, T.; Allen, W.J.; Pirker, K.F.; Lebedev, A.; Henderson, N.S.; Geibel, S.; Volkan, E.; Yan, J.; Kunze, M.B.A.; Pinkner, J.S.; Ford, B.; Kay, C.W.M. Crystal structure of the FimD usher bound to its cognate FimC: FimH substrate. Nature 2011, 474, 49–53. [Google Scholar] [CrossRef]
- Kuehn, M.J.; Heuser, J.; Normark, S.; Hultgren, S.J. P pili in uropathogenic E. coli are composite fibres with distinct fibrillar adhesive tips. Nature 1992, 356, 252–255. [Google Scholar] [CrossRef]
- Bullitt, E.; Makowski, L. Structural polymorphism of bacterial adhesion pili. Nature 1995, 373, 164–167. [Google Scholar] [CrossRef]
- Hahn, E.; Wild, P.; Hermanns, U.; Sebbel, P.; Glockshuber, R.; Häner, M.; Taschner, N.; Burkhard, P.; Aebi, U.; Müller, S.A. Exploring the 3D molecular architecture of Escherichia coli type 1 pili. J. Mol. Biol. 2002, 323, 845–857. [Google Scholar] [CrossRef]
- Waksman, G.; Hultgren, S.J. Structural biology of the chaperone-usher pathway of pilus biogenesis. Nat. Rev. Microbiol. 2009, 7, 765–774. [Google Scholar] [CrossRef]
- Phan, G.; Waksman, G. Pilus biogenesis by the chaperone-usher pathway. Adv. Cell Mol. Microbiol 2013, in press. [Google Scholar]
- Mu, X.-Q.; Bullitt, E. Structure and assembly of P-pili: A protruding hinge region used for assembly of a bacterial adhesion filament. PNAS 2006, 103, 9861–9866. [Google Scholar] [CrossRef]
- Zav’yalov, V.; Zavialov, A.; Zav’yalova, G.; Korpela, T. Adhesive organelles of Gram-negative pathogens assembled with the classical chaperone/usher machinery: structure and function from a clinical standpoint. FEMS Microbiol. Rev. 2010, 34, 317–378. [Google Scholar] [CrossRef]
- Verger, D.; Miller, E.; Remaut, H.; Waksman, G.; Hultgren, S. Molecular mechanism of P pilus termination in uropathogenic Escherichia coli. EMBO Rep. 2006, 7, 1228–1232. [Google Scholar] [CrossRef]
- Rossolini, G.M.; Muscas, P.; Chiesurin, A.; Satta, G. Analysis of the Salmonella fim gene cluster: identification of a new gene (fimI) encoding a fimbrin-like protein and located downstream from the fimA gene. FEMS Microbiol. Lett. 1993, 114, 259–265. [Google Scholar] [CrossRef]
- Valenski, M.L.; Harris, S.L.; Spears, P.A.; Horton, J.R.; Orndorff, P.E. The Product of the fimI gene is necessary for Escherichia coli type 1 pilus biosynthesis. J. Bacteriol. 2003, 185, 5007–5011. [Google Scholar] [CrossRef]
- Stathopoulos, C.; Hendrixson, D.R.; Thanassi, D.G.; Hultgren, S.J.; St. Geme III, J.W.; Curtiss III, R. Secretion of virulence determinants by the general secretory pathway in Gram-negative pathogens: an evolving story. Microbes and Infection 2000, 2, 1061–1072. [Google Scholar] [CrossRef]
- Puorger, C.; Vetsch, M.; Wider, G.; Glockshuber, R. Structure, folding and stability of FimA, the main structural subunit of type 1 pili from uropathogenic Escherichia coli strain. J. Mol. Biol. 2011, 412, 520–535. [Google Scholar] [CrossRef]
- Vetsch, M.; Puorger, C.; Spirig, T.; Grauschopf, U.; Weber-Ban, E.U.; Glockshuber, R. Pilus chaperones represent a new type of protein-folding catalyst. Nature 2004, 431, 329–333. [Google Scholar] [CrossRef]
- Crespo, M.D.; Puorger, C.; Schärer, M.A.; Eidam, O.; Grütter, M.G.; Capitani, G.; Glockshuber, R. Quality control of disulfide bond formation in pilus subunits by the chaperone FimC. Nat. Chem. Biol. 2012, 8, 707–713. [Google Scholar] [CrossRef] [Green Version]
- Sauer, F.G.; Fütterer, K.; Pinkner, J.S.; Dodson, K.W.; Hultgren, S.J.; Waksman, G. Structural basis of chaperone function and pilus biogenesis. Science 1999, 285, 1058–1061. [Google Scholar] [CrossRef]
- Choudhury, D.; Thompson, A.; Stojanoff, V.; Langermann, S.; Pinkner, J.; Hultgren, S.J.; Knight, S.D. X-ray structure of the FimC-FimH chaperone-adhesin complex from uropathogenic Escherichia coli. Science 1999, 285, 1061–1066. [Google Scholar] [CrossRef]
- Ford, B.; Verger, D.; Dodson, K.; Volkan, E.; Kostakioti, M.; Elam, J.; Pinkner, J.; Waksman, G.; Hultgren, S. The structure of the PapD-PapGII pilin complex reveals an open and flexible P5 pocket. J. Bacteriol. 2012, 194, 6390–6397. [Google Scholar] [CrossRef]
- Remaut, H.; Rose, R.J.; Hannan, T.J.; Hultgren, S.J.; Radford, S.E.; Ashcroft, A.E.; Waksman, G. Donor-strand exchange in chaperone-assisted pilus assembly proceeds through a concerted beta strand displacement mechanism. Mol. Cell. 2006, 22, 831–842. [Google Scholar] [CrossRef]
- Zavialov, A.V.; Berglund, J.; Pudney, A.F.; Fooks, L.J.; Ibrahim, T.M.; MacIntyre, S.; Knight, S.D. Structure and biogenesis of the capsular F1 antigen from Yersinia pestis: Preserved folding energy drives fiber formation. Cell 2003, 113, 587–596. [Google Scholar] [CrossRef]
- Rose, R.J.; Welsh, T.S.; Waksman, G.; Ashcroft, A.E.; Radford, S.E.; Paci, E. Donor-strand exchange in chaperone-assisted pilus assembly revealed in atomic detail by molecular dynamics. J. Mol. Biol. 2008, 375, 908–919. [Google Scholar] [CrossRef]
- Puorger, C.; Eidam, O.; Capitani, G.; Erilov, D.; Grütter, M.G.; Glockshuber, R. Infinite kinetic stability against dissociation of supramolecular protein complexes through donor strand complementation. Structure 2008, 16, 631–642. [Google Scholar] [CrossRef]
- Leney, A.C.; Phan, G.; Allen, W.; Verger, D.; Waksman, G.; Radford, S.E.; Ashcroft, A.E. Second order rate constants of donor-strand exchange reveal individual amino acid residues important in determining the subunit specificity of pilus biogenesis. J. Am. Soc. Mass. Spectrom. 2011, 22, 1214–1223. [Google Scholar]
- Zavialov, A.V.; Tischenko, V.M.; Fooks, L.J.; Brandsdal, B.O.; Aqvist, J.; Zav’yalov, V.P.; Macintyre, S.; Knight, S.D. Resolving the energy paradox of chaperone/usher-mediated fibre assembly. Biochem. J. 2005, 389, 685–694. [Google Scholar] [CrossRef]
- Miller, E.; Garcia, T.; Hultgren, S.; Oberhauser, A.F. The mechanical properties of E. coli type 1 pili measured by atomic force microscopy techniques. Biophys. J. 2006, 91, 3848–3856. [Google Scholar] [CrossRef]
- Li, H.; Qian, L.; Chen, Z.; Thibault, D.; Liu, G.; Liu, T.; Thanassi, D.G. The outer membrane usher forms a twin-pore secretion complex. J. Mol. Biol. 2004, 344, 1397–1407. [Google Scholar] [CrossRef]
- Allen, W.J.; Phan, G.; Hultgren, S.J.; Waksman, G. Dissection of Pilus Tip Assembly by the FimD Usher Monomer. J. Mol. Biol. 2013, 425, 958–967. [Google Scholar] [CrossRef]
- Saulino, E.T.; Thanassi, D.G.; Pinkner, J.S.; Hultgren, S.J. Ramifications of kinetic partitioning on usher-mediated pilus biogenesis. EMBO J. 1998, 17, 2177–2185. [Google Scholar] [CrossRef]
- Nishiyama, M.; Vetsch, M.; Puorger, C.; Jelesarov, I.; Glockshuber, R. Identification and characterization of the chaperone-subunit complex-binding domain from the type 1 pilus assembly platform FimD. J. Mol. Biol. 2003, 330, 513–525. [Google Scholar] [CrossRef]
- Ng, T.W.; Akman, L.; Osisami, M.; Thanassi, D.G. The usher N terminus is the initial targeting site for chaperone-subunit complexes and participates in subsequent pilus biogenesis events. J. Bacteriol. 2004, 186, 5321–5331. [Google Scholar] [CrossRef]
- Volkan, E.; Ford, B.A.; Pinkner, J.S.; Dodson, K.W.; Henderson, N.S.; Thanassi, D.G.; Waksman, G.; Hultgren, S.J. Domain activities of PapC usher reveal the mechanism of action of an Escherichia coli molecular machine. PNAS 2012, 109, 9563–9568. [Google Scholar] [CrossRef]
- Di Yu, X.; Dubnovitsky, A.; Pudney, A.F.; Macintyre, S.; Knight, S.D.; Zavialov, A.V. Allosteric mechanism controls traffic in the chaperone/usher pathway. Structure 2012, 20, 1861–1871. [Google Scholar]
- Li, Q.; Ng, T.W.; Dodson, K.W.; So, S.S.K.; Bayle, K.-M.; Pinkner, J.S.; Scarlata, S.; Hultgren, S.J.; Thanassi, D.G. The differential affinity of the usher for chaperone-subunit complexes is required for assembly of complete pili. Mol. Microbiol. 2010, 76, 159–172. [Google Scholar] [CrossRef]
- Morrissey, B.; Leney, A.C.; Toste Rêgo, A.; Phan, G.; Allen, W.J.; Verger, D.; Waksman, G.; Ashcroft, A.E.; Radford, S.E. The role of chaperone-subunit usher domain interactions in the mechanism of bacterial pilus biogenesis revealed by ESI-MS. MCP 2012, 11. M111.015289. [Google Scholar]
- Geibel, S.; Procko, E.; Hultgren, S.J.; Baker, D.; Waksman, G. Structural and energetic basis of folded protein transport by the FimD usher. Nature 2013, 496, 243–246. [Google Scholar] [CrossRef]
- Lindberg, F.P.; Lund, B.; Normark, S. Genes of pyelonephritogenic E. coli required for digalactoside-specific agglutination of human cells. EMBO J. 1984, 3, 1167–1173. [Google Scholar]
- Lindberg, F.; Lund, B.; Johansson, L.; Normark, S. Localization of the receptor-binding protein adhesin at the tip of the bacterial pilus. Nature 1987, 328, 84–87. [Google Scholar] [CrossRef]
- Nishiyama, M.; Ishikawa, T.; Rechsteiner, H.; Glockshuber, R. Reconstitution of pilus assembly reveals a bacterial outer membrane catalyst. Science 2008, 320, 376–379. [Google Scholar] [CrossRef]
- Nishiyama, M.; Glockshuber, R. The outer membrane usher guarantees the formation of functional pili by selectively catalyzing donor-strand exchange between subunits that are adjacent in the mature pilus. J. Mol. Biol. 2010, 396, 1–8. [Google Scholar] [CrossRef]
- Vetsch, M.; Erilov, D.; Molière, N.; Nishiyama, M.; Ignatov, O.; Glockshuber, R. Mechanism of fibre assembly through the chaperone-usher pathway. EMBO R. 2006, 7, 734–738. [Google Scholar] [CrossRef]
- Rose, R.J.; Verger, D.; Daviter, T.; Remaut, H.; Paci, E.; Waksman, G.; Ashcroft, A.E.; Radford, S.E. Unraveling the molecular basis of subunit specificity in P pilus assembly by mass spectrometry. PNAS 2008, 105, 12873–12878. [Google Scholar] [CrossRef]
- Remaut, H.; Tang, C.; Henderson, N.S.; Pinkner, J.S.; Wang, T.; Hultgren, S.J.; Thanassi, D.G.; Waksman, G.; Li, H. Fiber formation across the bacterial outer membrane by the chaperone/usher pathway. Cell 2008, 133, 640–652. [Google Scholar] [CrossRef]
- Huang, Y.; Smith, B.S.; Chen, L.X.; Baxter, R.H.G.; Deisenhofer, J. Insights into pilus assembly and secretion from the structure and functional characterization of usher PapC. PNAS 2009, 106, 7403–7407. [Google Scholar]
- Mapingire, O.S.; Henderson, N.S.; Duret, G.; Thanassi, D.G.; Delcour, A.H. Modulating effects of the plug, helix, and N- and C-terminal domains on channel properties of the PapC usher. J. Biol. Chem. 2009, 284, 36324–36333. [Google Scholar]
- Anderson, K.L.; Billington, J.; Pettigrew, D.; Cota, E.; Simpson, P.; Roversi, P.; Chen, H.A.; Urvil, P.; Du Merle, L.; Barlow, P.N.; Medof, M.E.; Smith, R.A.G.; Nowicki, B.; Le Bouguénec, C.; Lea, S.M.; Matthews, S. An atomic resolution model for assembly, architecture, and function of the Dr adhesins. Mol. Cell 2004, 15, 647–657. [Google Scholar] [CrossRef]
- Pettigrew, D.; Anderson, K.L.; Billington, J.; Cota, E.; Simpson, P.; Urvil, P.; Rabuzin, F.; Roversi, P.; Nowicki, B.; Du Merle, L.; Le Bouguénec, C.; Matthews, S.; Lea, S.M. High resolution studies of the Afa/Dr adhesin DraE and its interaction with chloramphenicol. J. Biol. Chem. 2004, 279, 46851–46857. [Google Scholar] [CrossRef]
- Zavialov, A.; Zav’yalova, G.; Korpela, T.; Zav’yalov, V. FGL chaperone-assembled fimbrial polyadhesins: Anti-immune armament of Gram-negative bacterial pathogens. FEMS Microbiol. Rev. 2007, 31, 478–514. [Google Scholar] [CrossRef]
- Mulvey, M.A. Adhesion and entry of uropathogenic Escherichia coli. Cell. Microbiol. 2002, 4, 257–271. [Google Scholar] [CrossRef]
- Kurimura, Y.; Nishitani, C.; Ariki, S.; Saito, A.; Hasegawa, Y.; Takahashi, M.; Hashimoto, J.; Takahashi, S.; Tsukamoto, T.; Kuroki, Y. Surfactant protein D inhibits adherence of uropathogenic Escherichia coli to the bladder epithelial cells and the bacterium-induced cytotoxicity: A possible function in urinary tract. J. Biol. Chem. 2012, 287, 39578–88. [Google Scholar] [CrossRef]
- Lund, B.; Lindberg, F.; Marklund, B.I.; Normark, S. The PapG protein is the alpha-D-galactopyranosyl-(1-4)-beta-D-galactopyranose-binding adhesin of uropathogenic Escherichia coli. PNAS 1987, 84, 5898–5902. [Google Scholar] [CrossRef]
- Dodson, K.W.; Pinkner, J.S.; Rose, T.; Magnusson, G.; Hultgren, S.J.; Waksman, G. Structural basis of the interaction of the pyelonephritic E. coli adhesin to its human kidney receptor. Cell 2001, 105, 733–743. [Google Scholar] [CrossRef]
- Snyder, J.A.; Haugen, B.J.; Lockatell, C.V.; Maroncle, N.; Hagan, E.C.; Johnson, D.E.; Welch, R.A.; Mobley, H.L.T. Coordinate expression of fimbriae in uropathogenic Escherichia coli. Infection Immun. 2005, 73, 7588–7596. [Google Scholar] [CrossRef]
- Yakovenko, O.; Sharma, S.; Forero, M.; Tchesnokova, V.; Aprikian, P.; Kidd, B.; Mach, A.; Vogel, V.; Sokurenko, E.; Thomas, W.E. FimH forms catch bonds that are enhanced by mechanical force due to allosteric regulation. J. Biol. Chem. 2008, 283, 11596–11605. [Google Scholar] [CrossRef]
- Le Trong, I.; Aprikian, P.; Kidd, B.A.; Forero-Shelton, M.; Tchesnokova, V.; Rajagopal, P.; Rodriguez, V.; Interlandi, G.; Klevit, R.; Vogel, V.; Stenkamp, R.E.; Sokurenko, E.V.; Thomas, W.E. Structural basis for mechanical force regulation of the adhesin FimH via finger trap-like beta sheet twisting. Cell 2010, 141, 645–655. [Google Scholar] [CrossRef]
- Aprikian, P.; Interlandi, G.; Kidd, B.A.; Le Trong, I.; Tchesnokova, V.; Yakovenko, O.; Whitfield, M.J.; Bullitt, E.; Stenkamp, R.E.; Thomas, W.E.; Sokurenko, E. V The bacterial fimbrial tip acts as a mechanical force sensor. PLoS Biol. 2011, 9, e1000617. [Google Scholar] [CrossRef]
- Tchesnokova, V.; Aprikian, P.; Kisiela, D.; Gowey, S.; Korotkova, N.; Thomas, W.; Sokurenko, E. Type 1 fimbrial adhesin FimH elicits an immune response that enhances cell adhesion of Escherichia coli. Infection Immun. 2011, 79, 3895–3904. [Google Scholar] [CrossRef]
- Weissman, S.J.; Beskhlebnaya, V.; Chesnokova, V.; Chattopadhyay, S.; Stamm, W.E.; Hooton, T.M.; Sokurenko, E. V Differential stability and trade-off effects of pathoadaptive mutations in the Escherichia coli FimH adhesin. Infection Immun. 2007, 75, 3548–3555. [Google Scholar] [CrossRef]
- Stahlhut, S.G.; Tchesnokova, V.; Struve, C.; Weissman, S.J.; Chattopadhyay, S.; Yakovenko, O.; Aprikian, P.; Sokurenko, E.V.; Krogfelt, K.A. Comparative structure-function analysis of mannose-specific FimH adhesins from Klebsiella pneumoniae and Escherichia coli. J. Bacteriol. 2009, 191, 6592–6601. [Google Scholar] [CrossRef]
- Chen, S.L.; Hung, C.S.; Pinkner, J.S.; Walker, J.N.; Cusumano, C.K.; Li, Z.; Bouckaert, J.; Gordon, J.I.; Hultgren, S.J. Positive selection identifies an in vivo role for FimH during urinary tract infection in addition to mannose binding. PNAS 2009, 106, 22439–22444. [Google Scholar] [CrossRef]
- Martinez, J.J.; Mulvey, M.A.; Schilling, J.D.; Pinkner, J.S.; Hultgren, S.J. Type 1 pilus-mediated bacterial invasion of bladder epithelial cells. EMBO J. 2000, 19, 2803–2812. [Google Scholar] [CrossRef]
- Pratt, L.A.; Kolter, R. Genetic analysis of Escherichia coli biofilm formation: roles of flagella, motility, chemotaxis and type I pili. Mol. Microbiol. 1998, 30, 285–293. [Google Scholar] [CrossRef]
- Mulvey, M.A.; Lopez-Boado, Y.S.; Wilson, C.L.; Roth, R.; Parks, W.C.; Heuser, J.; Hultgren, S.J. Induction and evasion of host defenses by type 1-piliated uropathogenic Escherichia coli. Science 1998, 282, 1494–1497. [Google Scholar] [CrossRef]
- Aberg, V.; Almqvist, F. Pilicides-small molecules targeting bacterial virulence. Org. Biomol. Chem. 2007, 5, 1827–1834. [Google Scholar]
- Lo, A.W.H.; Moonens, K.; Remaut, H. Chemical attenuation of pilus function and assembly in Gram-negative bacteria. Curr. Opin. Microbiol. 2013, 16, 85–92. [Google Scholar]
- Durand, E.; Verger, D.; Rêgo, A.T.; Chandran, V.; Meng, G.; Fronzes, R.; Waksman, G. Structural biology of bacterial secretion systems in gram-negative pathogens-potential for new drug targets. Infect. Disorders Drug Targets 2009, 9, 518–547. [Google Scholar] [CrossRef]
- Pinkner, J.S.; Remaut, H.; Buelens, F.; Miller, E.; Aberg, V.; Pemberton, N.; Hedenström, M.; Larsson, A.; Seed, P.; Waksman, G.; Hultgren, S.J.; Almqvist, F. Rationally designed small compounds inhibit pilus biogenesis in uropathogenic bacteria. PNAS 2006, 103, 17897–17902. [Google Scholar] [CrossRef]
- Cusumano, C.K.; Hultgren, S.J. Bacterial adhesion-a source of alternate antibiotic targets. IDrugs 2009, 12, 699–705. [Google Scholar]
- Cusumano, C.K.; Pinkner, J.S.; Han, Z.; Greene, S.E.; Ford, B.A.; Crowley, J.R.; Henderson, J.P.; Janetka, J.W.; Hultgren, S.J. Treatment and prevention of urinary tract infection with orally active FimH inhibitors. Sci. Transl. Med. 2011, 3, 109ra115. [Google Scholar] [CrossRef]
- Jiang, X.; Abgottspon, D.; Kleeb, S.; Rabbani, S.; Scharenberg, M.; Wittwer, M.; Haug, M.; Schwardt, O.; Ernst, B. Antiadhesion therapy for urinary tract infections-a balanced PK/PD profile proved to be key for success. J. Med. Chem. 2012, 55, 4700–4713. [Google Scholar] [CrossRef]
- Larsson, A.; Ohlsson, J.; Dodson, K.W.; Hultgren, S.J.; Nilsson, U.; Kihlberg, J. Quantitative studies of the binding of the class II PapG adhesin from uropathogenic Escherichia coli to oligosaccharides. Bioorg. Med. Chem. 2003, 11, 2255–2261. [Google Scholar]
- Salminen, A.; Loimaranta, V.; Joosten, J.A.F.; Khan, A.S.; Hacker, J.; Pieters, R.J.; Finne, J. Inhibition of P-fimbriated Escherichia coli adhesion by multivalent galabiose derivatives studied by a live-bacteria application of surface plasmon resonance. J. Antimicrob. Chemother. 2007, 60, 495–501. [Google Scholar] [CrossRef]
- Watts, R.E.; Tan, C.K.; Ulett, G.C.; Carey, A.J.; Totsika, M.; Idris, A.; Paton, A.W.; Morona, R.; Paton, J.C.; Schembri, M.A. Escherichia coli 83972 expressing a P fimbriae oligosaccharide receptor mimic impairs adhesion of uropathogenic E.coli. J. Infectious Dis. 2012, 206, 1242–1249. [Google Scholar] [CrossRef]
- Klinth, J.E.; Castelain, M.; Uhlin, B.E.; Axner, O. The influence of pH on the specific adhesion of P piliated Escherichia coli. PLoS One 2012, 7, e38548. [Google Scholar]
- Klinth, J.E.; Pinkner, J.S.; Hultgren, S.J.; Almqvist, F.; Uhlin, B.E.; Axner, O. Impairment of the biomechanical compliance of P pili: A novel means of inhibiting uropathogenic bacterial infections? EBJ 2012, 41, 285–295. [Google Scholar] [CrossRef]
- Bouckaert, J.; Berglund, J.; Schembri, M.; De Genst, E.; Cools, L.; Wuhrer, M.; Hung, C.-S.; Pinkner, J.; Slättegård, R.; Zavialov, A.; Choudhury, D.; Langermann, S.; Hultgren, S.J.; Wyns, L.; Klemm, P.; Oscarson, S.; Knight, S.D.; De Greve, H. Receptor binding studies disclose a novel class of high-affinity inhibitors of the Escherichia coli FimH adhesin. Mol. Microbiol. 2005, 55, 441–455. [Google Scholar]
- Nishiyama, M.; Horst, R.; Eidam, O.; Herrmann, T.; Ignatov, O.; Vetsch, M.; Bettendorff, P.; Jelesarov, I.; Grütter, M.G.; Wüthrich, K.; Glockshuber, R.; Capitani, G. Structural basis of chaperone-subunit complex recognition by the type 1 pilus assembly platform FimD. EMBO J. 2005, 24, 2075–2086. [Google Scholar] [CrossRef]
- Chorell, E.; Pinkner, J.S.; Phan, G.; Edvinsson, S.; Buelens, F.; Remaut, H.; Waksman, G.; Hultgren, S.J.; Almqvist, F. Design and synthesis of C-2 substituted thiazolo and dihydrothiazolo ring-fused 2-pyridones: pilicides with increased antivirulence activity. J. Med. Chem. 2010, 53, 5690–5695. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lillington, J.; Waksman, G. Ordered and Ushered; the Assembly and Translocation of the Adhesive Type I and P Pili. Biology 2013, 2, 841-860. https://doi.org/10.3390/biology2030841
Lillington J, Waksman G. Ordered and Ushered; the Assembly and Translocation of the Adhesive Type I and P Pili. Biology. 2013; 2(3):841-860. https://doi.org/10.3390/biology2030841
Chicago/Turabian StyleLillington, James, and Gabriel Waksman. 2013. "Ordered and Ushered; the Assembly and Translocation of the Adhesive Type I and P Pili" Biology 2, no. 3: 841-860. https://doi.org/10.3390/biology2030841



