Abstract
Multiple myeloma (MM) is a genetically complex hematological cancer that is characterized by proliferation of malignant plasma cells in the bone marrow. MM evolves from the clonal premalignant disorder monoclonal gammopathy of unknown significance (MGUS) by sequential genetic changes involving many different genes, resulting in dysregulated growth of multiple clones of plasma cells. The migration, survival, and proliferation of these clones require the direct and indirect interactions with the non-hematopoietic cells of the bone marrow. We develop a hybrid discrete-continuous model of MM development from the MGUS stage. The discrete aspect of the model is observed at the cellular level: cells are represented as individual objects which move, interact, divide, and die by apoptosis. Each of these actions is regulated by intracellular and extracellular processes as described by continuous models. The hybrid model consists of the following submodels that have been simplified from the much more complex state of evolving MM: cell motion due to chemotaxis, intracellular regulation of plasma cells, extracellular regulation in the bone marrow, and acquisition of mutations upon cell division. By extending a previous, simpler model in which the extracellular matrix was considered to be uniformly distributed, the new hybrid model provides a more accurate description in which cytokines are produced by the marrow microenvironment and consumed by the myeloma cells. The complex multiple genetic changes in MM cells and the numerous cell-cell and cytokine-mediated interactions between myeloma cells and their marrow microenviroment are simplified in the model such that four related but evolving MM clones can be studied as they compete for dominance in the setting of intraclonal heterogeneity.
1. Introduction
Multiple myeloma (MM) is initiated through the acquisition of genetic changes that transform the plasma cells from normal to malignant. These changes often result in the development of selective advantage, leading to the excessive proliferation of myeloma cells. The development of MM leads to several harmful clinical conditions including anemia, renal failure, recurrent infections, hypercalcemia, and osteoporosis with bone fractures. These pathological conditions can frequently result in the death of the patient.
The evolution of myeloma from the clonal premalignant plasma cell condition termed monoclonal gammopathy of unknown significance (MGUS) through the intermediate stage of smoldering myeloma to the malignant MM stage is mediated by multiple sequential genetic changes, including chromosomal translocations, hyperdiploidy, and mutations, which permit independent growth and spread of MM in the bone marrow [1,2]. A wide variety of genetic changes involving many different genes have been documented in MM cases. In addition to these sequential genetic changes, MM cells require specific interactions with the non-hematopoietic cells of the bone marrow including the stromal cells (BMSCs), osteoblasts, osteoclasts, and cells associated with vascular supply of the marrow [1,2]. These marrow microenvironment interactions with the MM cells include direct binding of cell surface adhesion proteins and their binding partners on the other cell types as well as diffusible, soluble molecules that are secreted by one cell type and are bound and internalized by another cell type. These soluble molecules are often chemokines and cytokines produced by other marrow cells, and their receptors are expressed on the MM cells. However, specific molecules produced by MM cells indirectly affect growth of the malignancy by inducing localized resorption of bone, death of hematopoietic cells, and expansion of vessels supplying blood to the MM. The combined effects of sequential genetic changes within the evolving myeloma cells and interactions of the MM cells with the marrow microenvironment affect the migration, proliferation, and survival of the MM cells.
There are three types of mathematical models of tumor growth and intra-clonal heterogeneity. The first type is the continuous model in which partial and ordinary differential equations are used to describe the density of premalignant and malignant cells [3,4,5]. Some continuous models study the effects of nutrients and inhibitors on tumor growth that may or may not be accompanied by necrosis [6,7], while others focus on the nonlinear three-dimensional spatial aspects of tumors [8]. The question of clonal competition within tumors has been studied by other continuous models [9,10], and the role of clonal heterogeneity in drug resistance has been previously explored [11]. While these continuous models sustain mathematical analysis and provide key insights in the development of tumors, they do not describe the cell-cell interactions and intracellular regulation of single cells. The second type of mathematical models describing tumor growth is the discrete model which usually represents cells as individual objects that interact with each other. These interactions determine the fate of each cell [12,13,14,15,16,17]. Cellular automatons provide a simple yet powerful tool to understand the mechanisms of tumor growth [18,19]. The discrete models usually describe cells in on-lattice or off-lattice grids. They focus on the interactions between cells and how they lead to the development of tumors, but discrete models do not link the different processes taking place at different scales. The third type of tumor growth model, which is known as a hybrid model, combines the features of both continuous and discrete models, thereby benefiting from the advantages of both of them [20,21,22]. In hybrid models, the concentrations of intracellular and extracellular proteins are described by partial and ordinary differential equations while the cells are introduced as individual objects. The heterogeneous nature of tumors has been studied by some of these models [23,24,25,26]. The models describing MM development usually study the impact of MM on different biological systems such as bone remodeling [27] and erythropoiesis [28].
In this work, we develop a hybrid model describing the development and intra-clonal heterogeneity in MM as described in Walker et al. [29]. In this model, malignant cells are represented by elastic spheres that can move, grow, interact, divide and die by apoptosis. Their fate is determined by intracellular and extracellular regulation networks. The growth of the cellular radii between cell divisions and the formation of two daughter cells with each division lead to the expansion of the tumor. To investigate the role of the bone marrow (BM) microenvironment in the growth of myeloma tumors, we specifically consider another population of cells, the bone marrow stromal cells (BMSCs). These cells secrete cytokines such as stromal cell-derived factor 1 (SDF-1), insulin-like growth factor 1 (IGF-1),and interleukin 6 (IL-6) which are necessary to the survival, homing and growth of malignant plasma cells. The concentration of each of these cytokines is modelled by a reaction–diffusion equation. The intracellular regulation of malignant cells is represented by a system of ordinary differential equations which depends on the concentration of extracellular cytokines. We characterize each cell by a specific genotype, and we consider that after its division, daughter cells will either inherit the same genotype or acquire a slightly different one due to random mutations. As the result, aggressive clones emerge during tumor progression in a parallel pattern. Although experiments show the possibility of clones emerging in both linear and branching evolutionary patterns, we restrict this study to the parallel evolutionary case. The MM cells consume cytokines, leading to competition between clones for the available resources. We study the dynamics of clonal competition and its role in the progression of MM.
2. Hybrid Model of MM Development
We developed a hybrid discrete-continuous model of MGUS progression to MM. The discrete aspect of the model is observed at the cellular level: cells are represented as individual objects which move, interact, divide, and die by apoptosis. Each of these actions is regulated by intracellular and extracellular processes described by continuous models. The hybrid model consists of the following submodels: cell motion due to chemotaxis, intracellular regulation of plasma cells, extracellular regulation in the bone marrow, and acquisition of mutations upon cell division. It is an extension of a previously developed simpler model [30]. While the previous model considers the cytokines in the extracellular matrix to be uniformly distributed, the present study provides a more accurate description by considering that these cytokines are produced by the BMSCs and consumed by the myeloma cells. Furthermore, a more detailed description of the intracellular pathways regulating the fate of plasma cells is provided in the present model.
The model is based on the direct effects of sequential genetic changes and marrow microenvironmental chemokine and cytokine activity that influence the chemotaxis, proliferation and survival of MM, but does not include the MM effects on bone resorption, hematopoietic cell loss, or development of specialized vasculature. The complex multiple genetic changes in MM cells and the numerous cell-cell and cytokine-mediated interactions between myeloma cells and their marrow microenviroment are simplified in the model so that four related but evolving clones develop in a process termed intra-clonal heterogeneity [31,32] (See Figure 1). Competition among these four MM clones is based on differences in cellular growth and survival rates and interactions with the marrow microenvironment. This competition results in predominance of the more fit clones and decline and ultimate extinction of the less fit ones. An early event in the MM model (Figure 1a) is a standard-risk genetic change, a t(11;14) translocation that involves the immunoglobulin heavy chain switch region on chromosome 14, inducing overexpression of the gene encoding cyclin D1, a regulator of cell cycle progression located on chromosome 11 [33]. This t(11;14) clone has deregulated cell cycle progression due to the translocation, and it undergoes two separate secondary genetic events: mutations involving the oncogenes N-RAS and K-RAS, which are common secondary events in the development of multiple myeloma [2]. The resultant clone with t(11;14)/mutant N-RAS has significantly enhanced proliferation compared to the parent t(11;14) clone, while the t(11;14)/ mutant K-RAS clone has much less of an increase in proliferation relative to the t(11;14) parental clone. Thus, the two descendant clones have differing degrees of increased RAS activity, resulting in a proliferative advantage for the t(11;14)/mutant N-RAS clone compared to both the t(11;14) parental clone and t(11;14)/ mutant K-RAS clone. A subsequent genetic event in the t(11;14)/K-RAS clone is, however, a mutation in the gene encoding interferon regulatory factor 4 (IRF4). Mutant IRF4 is a protein that can enhance survival and proliferation of MM cells. IRF4 mutation has been associated with mutations of N-RAS or K-RAS [32], and it allows the t(11;14)/K-RAS mutant/IRF4 mutant clone in the model to compete more successfully with the t(11;14)/N-RAS clone than either the parent t(11;14) clone or the t(11;14)/K-RAS mutant clone.
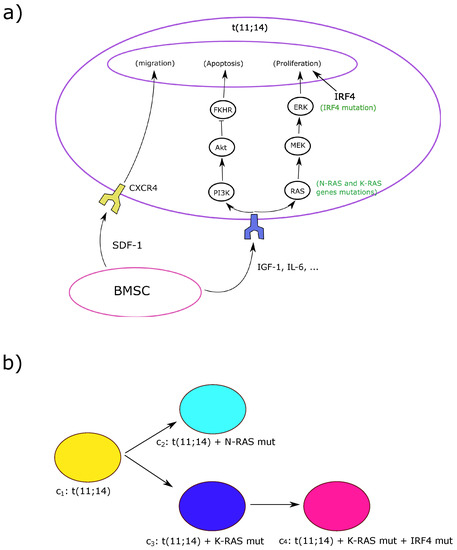
Figure 1.
(a) The intracellular regulation of myeloma cells as described in the model. Bone marrow stromal cells (BMSCs) secrete the cytokines insulin-like growth factor 1 (IGF-1), interleukin-6 (IL-6), and chemokine stromal cell-derived factor 1 (SDF-1) which are necessary to the survival, homing and proliferation of myeloma cells. Via their respective receptors, IGF-1 and IL-6 activate the RAS/ERK pathway which promotes the cell proliferation. They inhibit apoptosis through the phosphatidylinositol-3 kinase/protein kinase B/Forkhead in rabdomyosarcoma (Akt/FKHR) pathway. The cell migrates and homes to BMSCs through the SDF-1/CXCR4 axis. The IRF4 mutation, which has been associated with concomitant RAS mutations, promotes survival and proliferation. BMSCs, which are much larger cells than the myeloma cells, are shown in reduced in size in this figure as well as the receptors of both IGF-1 and IL-6; (b) The parallel evolution pattern of multiple myeloma clones resulting in intra-clonal heterogeneity. More aggressive clones result from a more aggressive N-RAS mutation in clone 2 or the acquisition of IRF4 mutation in addition to the less aggressive K-RAS in clone 4. Each clone is shown by its corresponding color in the model.
Among the many possible microenvironmental factors that may influence the growth of MM in the marrow, the simplified model includes the chemokine stromal cell-derived factor 1 (SDF-1). It is produced by multiple cell types but mainly by BMSCs. Other extracellular cytokines included in the model are interleukin 6 (IL-6) and insulin-like growth factor 1 (IGF-1), which are produced by other types of non-hematopoietic cells in the marrow as well as BMSCs. The marrow stromal cells are considered as sources of SDF-1, IL-6, and IGF-1 in the model (Figure 1a). SDF-1, through its receptor CXCR4, mediates the homing of circulating myeloma cells to the marrow and their migration within the marrow space [34,35]. IL-6 and IGF-1 induce multiple effects after binding to their respective specific surface receptors on the MM cells, but in the model we restrict our study to their effects on the RAS/Extracellular signal-regulated kinases (RAS/ERK) pathway that promotes proliferation and the phosphatidylinositol-3 kinase/protein kinase B/Forkhead in rabdomyosarcoma (Akt/FKHR) pathway that regulates apoptosis [2] (Figure 1a).
2.1. Cell Motion
We represent each cell as an elastic sphere. It contains two parts: an incompressible part and a compressible one. As the cell grows, its radius increases and hence it pushes the neighboring cells. The motion of each cell is determined by Newton’s second law. Let us denote the cell number by i. Then we have the following equation for the coordinate of the center of the cell:
where m is the mass of cell, μ is the friction coefficient, is the interaction force between the cells i and j, is the chemotactic force which depends on the concentration of SDF-1. We consider in the following form:
Here is the distance between the center of the cells i and j, is the sum of their radii, K is a positive parameter and represents the incompressible part of each cell. The numerical implementaton of Newton’s equation is presented in Appendix A.1. The chemotactic force represents the motion of the cell in response to SDF-1 stimulus. Let us denote the concentration of SDF-1 by S. Then the expression of is given by:
where κ is a positive constant and denotes the SDF-1 gradient.
2.2. Extracellular Regulation
We consider a square two-dimensional computational domain with the side equal to 1000 microns. The hybrid model contains two types of cells: myeloma cells and BMSCs. The former have a diameter of 10 microns while the latter are considered to have six-fold larger radii as observed in experimental data. Furthermore, they secrete cytokines that are necessary for the survival and proliferation of myeloma cells such as SDF-1, IL-6, and IGF-1. Let us denote the total concentration of the last two cytokines by I and the concentration of SDF-1 by S. Assuming that these cytokines have the same diffusion coefficient, production rate, consumption rate, and degradation rate, we describe their concentrations in the extracellular matrix by:
where D is the diffusion coefficient, W the production factor, λ the consumption rate by each myeloma cell, and σ the degradation rate. We set the Dirichlet boundary condition and at all boundaries. We have prescribed zero Dirichlet boundary condition to represent the local dynamics of tumor growth in a bone marrow site surrounded by adipose tissue. The finite difference method was used to implement the reaction-diffusion equations. The used numerical methods and implementation algorithm are presented in Appendix A.2.
2.3. Intracellular Regulation
Although myeloma cells are genetically complex [2], we will restrict the intracellular regulation to two pathways (Figure 1a). These pathways are the RAS/ERK pathway which is responsible for cell proliferation and the Akt/FKHR pathway which regulates its survival. Other pathways such as the janus kinase/signal transducer and activator of transcription (JAK/STAT) pathway are not considered in the model. We have chosen these two pathways in order to study the role of aquired RAS mutations in the progression of MM and how it is affected by the extracellular matrix. Let us denote the concentrations of ERK, Akt, and FKHR as e, a, and f, respectively. We describe these concentrations inside each cell as follows:
where and , and are positive constants. The coefficient depends on the genotype z, and this coefficient varies depending on the effect of RAS mutations. The cell will die by apoptosis if during its lifetime cycle. If the myeloma cell survives, then it will self-renew, if by the end of its life cycle. Otherwise, the cell will die by apoptosis. We consider that the proliferation threshold depends on the IRF4 gene expression. The values of both intracellular and extracellular regulation parameters are provided in Appendix B.
Myeloma Cells Division and Mutations
In the present study, we investigate the progression of MGUS into MM through RAS mutations [29]. We characterize each cell by an aggressiveness phenotype which represents the cumulative effects of acquired RAS mutations. We introduce an aggressiveness phenotype function resulting from the RAS genotype for each cell. After its division, the daughter myeloma cell will keep the genome of the mother cell with a probability of one-third. Otherwise, it will acquire a mutation and either increases or diminishes its aggressiveness by a positive value ϵ. We represent the genotype and resultant RAS activation that determine aggressiveness as due to describing the frequency and effects of RAS mutations in the resulting clones in Figure 2a. Furthermore, we assume that the ERK threshold for cell division is reduced upon the mutation of the IRF4 gene as shown in Figure 2b. We consider four clones of MM denoted by and, (Figure 1b). The clone is the initial clone which does not harbour any RAS mutations. The clone is more aggressive than clone . It consists of MM cells that acquired the N-RAS gene mutation. The clones and belong to a lineage that is independent from the clone . They emerge when the cells of acquire the K-RAS gene mutation for clone and an additional IRF4 gene mutation for clone . The latter clone is therefore more aggressive than the former. We show the relationships between these different clones in Figure 1b.
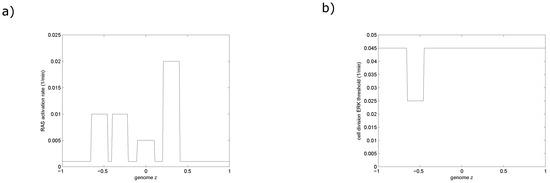
Figure 2.
(a) The activation rate of RAS protein as a function of the genotype function z; (b) The ERK threshold for division as a function of the genotype function z, it decreases due to the IRF4 mutation found in the clone .
3. Results
Initially, there are 47 myeloma cells in a bone marrow site containing a single BMSC, although the initial number of these cells is not essential for the results of the simulations. All of these malignant cells are considered to be tumor-initiating cells because they belong to the same initial clone. The motion of the malignant cells depends on their distance from the BMSC. Closer cells move faster than farther ones due to the increase in SDF-1 gradient near the BMSC. During their motion, myeloma cells consume the cytokines necessary for their survival such as IL-6 and IGF-1. These cytokines promote survival by activating the RAS/ERK pathway, and dowregulating the Akt/FKHR pathway. Depending upon its initial distance from the BMSC, the myeloma cell will either survive and get closer to the BMSC or it will die by apoptosis. The surviving myeloma cells will surround the BMSC and form the tumor niche. Initially, this niche consists of cells belonging to the same clone. After some time, other subclones will emerge in the course of tumor growth. We have shown the different steps of myeloma cells homing to a BMSC in Figure 3. The initial condition for the simulation is shown in Figure 3a. Due to the low number of myeloma cells in the beginning of the simulation, the BMSC secretes a high concentration of the chemokine SDF-1 which attracts the myeloma cells located close to it (Figure 3b). The surviving myeloma cells adhere to the BMSC and surround it while other cells divide and remain farther away (Figure 3c). During this process, cells whose aggressiveness phenotype belongs to the intermediate state between the clones and emerge (Figure 3d) and the global concentration of SDF-1 starts to reach stability due to its consumption by myeloma cells (Figure 4a).
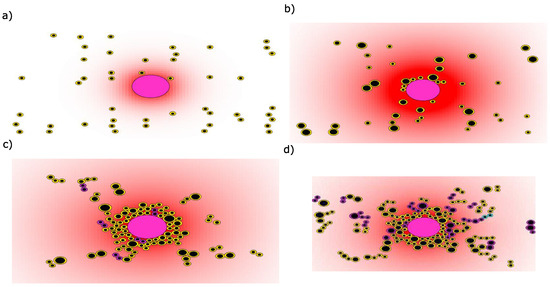
Figure 3.
The sequential steps of myeloma cells homing to a BMSC. Myeloma cells are represented by the cells with the smaller radii and the BMSC is the large cell in the middle. Each clone of multiple myeloma (MM) cells is denoted by a specific color. The red color gradient represents the summed concentrations of the cytokines SDF-1, IL-6, and IGF-1 shown with white (0) to red (1). (a) After their infiltration, the myeloma cells move towards the BMSC; (b) The surviving cells surround the BMSC; (c) The myeloma cells divide and form the tumor niche; (d) The tumor expands and more aggressive subclones start emerging.
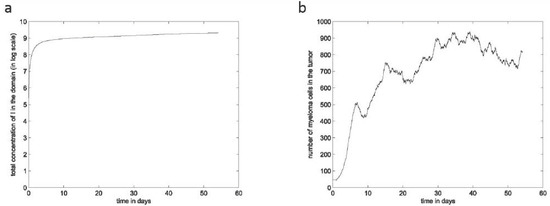
Figure 4.
(a) The total concentration of the IL-6 and IGF-1 cytokines over time in log scale; (b) The total number of malignant cells over time.
MGUS progresses to aggressive MM through the acquisition of random mutations by myeloma cells. In this respect, more aggressive subclones emerge in the course of MGUS progression due to the acquisition of RAS-associated mutations. The aggressive subclones need lower concentrations of IL-6 and IGF-1 to survive, resulting in the expansion and the persistence of the tumor (Figure 4b). Because of their rapid proliferation, they crowd out other clones and consume most of the available resources. Ultimately, these less aggressive subclones remain relatively stable in growth pattern. Still, a few of them manage to survive in proximity to the BMSC where there is a higher concentration of the IL-6 and IGF-1 cytokines, while the more aggressive subclones occupy the outer region of the tumor. The different steps of tumor development and intraclonal competition are shown in Figure 5. As the tumor progresses, the concentrations of IL-6 and IGF-1 become stable due to their consumption by the growing number of myeloma cells. As a result, the aggressive subclones expand to the detriment of the less aggressive ones. By the end of the simulation, we see that the tumor size is stable but the competition between clones is still in progress (Figure 5c vs. Figure 5d).
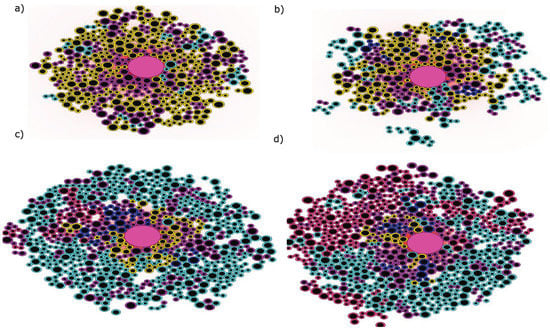

Figure 5.
Snapshots of a simulation showing the competition between clones. (a) The tumor consists mainly of the clone (shown in yellow) with few cells in the intermediate state (in purple) between clones and the emergence of the clone (in cyan) in the sides. The size of the tumor remains limited because clone cells need relatively high concentration of I to survive; (b) Compared to the clone , the clone cells expand and survive in areas with lower concentration of cytokines; (c) The clone cells surround the tumor and crowd out the cells of the clone leading to the reduction of their population. The clones (in blue) and (in magneta); (d) The subclone is as aggressive as the subclone due to the additional IRF4 mutation and it manages to coexist with it in the remote areas with fewer cytokines.
The emergence of more aggressive clones not only reduces the populations of the less aggressive ones but also increases the total number of malignant cells. These clones survive in areas farther from the BMSC, making the tumor expand. We show the total number of malignant cells over time in Figure 4b. The proportions of each clone in the total population show that in the end of the simulation, the clone only represents ≈5% of the total population. The subclone is predominant, with ≈60% of cells (Figure 6). Overall, the global population of malignant cells increases with the emergence of more resistant subclones. The emergence of the subclone leads to an increase in the number of malignant cells because the cells belonging to this subclone can survive far from the BMSCs. The subclone emerges few hours after but barely manage to survive because of limited resources. It gives rise to the subclone , which is as aggressive as the subclone due to the additional IRF4 mutation. As the tumor progresses, these two subclones and become predominant because they are better adapted to survive in sites with limited resources. By the end of the simulation, the number of malignant cells oscillates around a stable value as well as the total concentration of cytokines in the domain (Figure 4). We show the percent of the malignant cell population occupied by each clone over time in Figure 6.
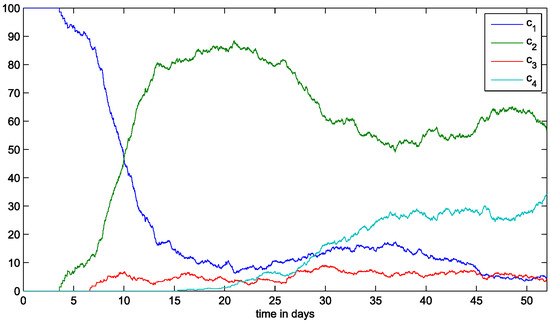
Figure 6.
The proportion of each clone in the total population over time.
4. Discussion
Bone marrow stromal cells play an important role in the pathogenesis of MM. After extravasation into the bone marrow, MGUS and myeloma cells migrate and home to the areas surrounding BMSCs. This homing process is mediated by the chemokine SDF-1. It promotes the chemotaxis and homing of myeloma cells upon the interaction with its receptor CXCR4 [36]. However, it is crucial for myeloma cells to be sufficiently close to BMSCs to be attracted by the secreted SDF-1. Otherwise, they will die if they are left without enough resources to survive and divide. The progression of MGUS to MM with t(11;14) translocation is marked by the emergence of new aggressive subclones which is crucial for the expansion of the tumor because it allows its adaptation to limited resources. As the tumor progresses, the aggressive subclones become predominant because lower concentrations of IL-6 and IGF-1 allow them to survive.
To study the development of MM tumors and its intraclonal heterogeneity, we developed a specific hybrid discrete-continuous model. The model was able to reproduce the experimental results presented in [29]. We have used it to investigate the central role of the BMSCs in myeloma cell homing and the progression of MGUS to MM. BMSCs participate in the homing of myeloma cells and provide them with the necessary cytokines for their survival and proliferation. Numerical simulation results suggest that the initial distance between the BMSC and infiltrating myeloma cells is of paramount importance for the survival of the latter. After their homing, new, more aggressive myeloma subclones emerge due to the acquisition of RAS mutations. Other gene mutations such as the one acquired by the IRF4 gene further increase the aggressiveness of some of these subclones. They compete with each other for the available cytokines, resulting in the predominance of the more fit subclones. To better quantify the heterogeneity of clones during MM progression, we show the kernel density plots at different stages of MGUS to MM progression in Figure 7. The x-axis shows the aggressiveness phenotype function scaled from 0 to 1. These results show the predominating subclones at the different stages of tumor progression. Our findings suggest that the total number of malignant cells oscillates around a stable value after a few weeks of MM development in agreement with the in vivo experiments conducted in [37]. However, we speculate that MM tumors can further expand due to other mechanisms such as stimulation of IL-6 and IGF-1 production by BMSC, the migration to other BMSCs [38], or the emegence of more resistant clones due to other mutations.
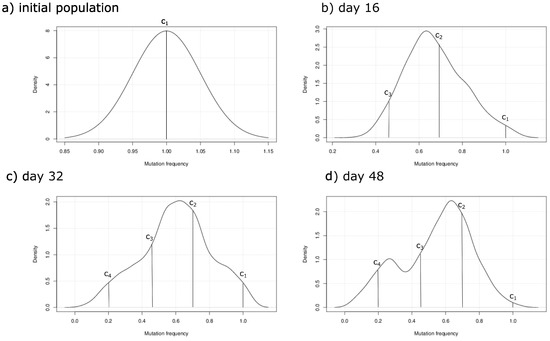
Figure 7.
Gaussian kernel density plots indicating the frequency of cells acquiring each specific mutation at the different stages of the simuation. (a) The initial population composed of cells belonging to the subclone ; (b) is the most predominant subclone after 16 hours of the beginning of the simulation; (c) The K-RAS subclones and emerge and start expanding; (d) The clones and represent the majority of the population because they are equally aggressive.
The multi-scale model follows a systems biology approach [39] by integrating different interfering biological processes in one model. To understand complex phenomena such as MGUS progression to MM, it is important to properly use available data in describing events at the single cell level where processes such as gene expression and mutations take place as well as the larger tissue level where each cell interacts with its environment. Systems biology approaches focus on the whole system rather than the sum of its components. Similarly, this study is more focused on the behavior of the global model of MM homing and tumor growth than the various individual processes which regulate it.
Although the model presented here was able to reproduce the experiments describing MM intra-clonal heterogeneity, it has several limitations. Firstly, the intracellular regulation network was limited to two pathways while in reality MM is more biochemically complex [2]. Furthermore, the number of considered mutations is also reduced compared to those found in MM cases. Interactions of the myeloma cells and BMSCs with other components of the bone marrow such as the osteoblasts and osteoclasts were not included in the present model. Other mechanisms affecting myeloma cell proliferation were introduced implicitly as a random perturbation of the cell cycle in the myeloma cells. The simplification of the myeloma cell transduction pathways and mutations included was due to limitations of computational power and the difficulty of studying complex models. We have limited the model to two pathways of intracellular regulation because we wanted to study the effect of cytokines IL-6 and IGF-1 on the fate of the myeloma cells and how that fate is affected by RAS mutations and the IRF4 mutation that is associated with mutant RAS genes. Also, we assumed that IL-6 and IGF-1 play the same role in the activation of the ERK/RAS and Akt/FKHR pathways. This hypothesis was made because it is difficult to distinguish the interfering actions of the different cytokines in the cell culture experiments. Finally, the study was restricted to a site of the bone marrow containing a single BMSC. Other configurations with more BMSCs and other marrow cell types can be considered as well. In future works, we will introduce the action of treatment and study the role of the intraclonal heterogeneity in MM drug resistance.
Acknowledgments
The last author was partially supported by the Russian Science Foundation (Grant No. 15-11-00029).
Author Contributions
All authors of the paper have contributed to the study design. A.B., M.J.K and V.V. conceived and developed the hybrid model. A.B., F.-E.B. and R.A. performed the numerical simulations. A.B., F.-E.B., R.A., M.J.K and V.V. wrote the paper and analyzed the results. All authors read and approved the final manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
Appendix A. Numerical Implementation
Appendix A.1. Equations for Cell Motion and Intracellular Regulation
We solve the following problem which describes the motion of each cell:
First, we write the late equation as a system of displacement and velocity :
We first determine the velocity using the first equation of the system, then we compute the displacement of the cell using the second equation. The forces term were introduced implicitly at each timestep. The equations for intracellular regulation are solved using the Euler explicit scheme.
Appendix A.2. Reaction-Diffusion Equations for Extracellular Cytokines Concentration
We consider a rectangular two-dimensional domain Ω and we denote its boundaries by Γ. We use the alternating direction implicit method to solve the following problem:
where D is the diffusion coefficient, W and σ are the production and degradation rates respectively, and represents the initial condition condition for I. We consider the two-dimensional grid where h and are the space and time steps respectively and we discretize it: and . We first rewrite the problem (P) in the following form:
We split the first equation of the problem () into two sub-steps as follows:
We solve the first equation for each fixed j to obtain . Next, we solve the second to obtain . Let us consider the first equation:
Rearranging the terms we obtain:
Therefore, we can write the first equation of the system (A2) in the form:
For each fixed , we solve numerically:
with the boundary conditions , . We solve the Equation (A3) using Thomas algorithm. For that, we write the left boundary condition as follows:
where and . Then, from the Equation (A3) for :
we obtain :
where we denote , . We continue for :
where , . We first obtain the coefficients , using the Equation (A3). Next, we find the solution for the sub-step by backward sweep using the Equation (A4). We apply the same procedure on the second equation of the system (A2) to compute the next step solution .
Appendix B. Parameter Values
Because of the simplifications made to the regulation network of myeloma cells, the parameters were fitted to reproduce the experimental results. The average cell cycle of the cancerous cells was considered to be stochastic. Both the extracellular and intracellular proteins concentrations are considered to be nondimensional.

Table A1.
Values of intracellular regulation parameters for myeloma cells. δ is an arbitrary length unit.
| Parameter | Value | Unit |
|---|---|---|
| Myeloma cell cycle length | 26 | h |
| Cell cycle variation | 13 | h |
| space variable and step | 1 | δ |
| time variable | 1 | min |
| time step | 0.01 | min |
| 0.001 | min | |
| 0.03 | min | |
| 0.002 | min | |
| 0.001 | min | |
| 0.01 | min | |
| 0.00083 | min |

Table A2.
Values of extracellular regulation parameters. δ is an arbitrary length unit.
| Parameter | Value | Unit |
|---|---|---|
| D | ·min | |
| W | 0.0003 | molecules··min |
| λ | 0.1 | NU |
| σ | min |
References
- Palumbo, A.; Anderson, K. Multiple myeloma. N. Engl. J. Med. 2011, 364, 1046–1060. [Google Scholar] [CrossRef] [PubMed]
- Morgan, G.J.; Walker, B.A.; Davies, F.E. The genetic architecture of multiple myeloma. Nat. Rev. Cancer 2012, 12, 335–348. [Google Scholar] [CrossRef] [PubMed]
- Adam, J.A. A simplified mathematical model of tumor growth. Math. Biosci. 1986, 81, 229–244. [Google Scholar] [CrossRef]
- Glass, L. Instability and mitotic patterns in tissue growth. J. Dyn. Syst. Meas. Control 1973, 95, 324–327. [Google Scholar] [CrossRef]
- McElwain, D.L.S.; Morris, L.E. Apoptosis as a volume loss mechanism in mathematical models of solid tumor growth. Math. Biosci. 1978, 39, 147–157. [Google Scholar] [CrossRef]
- Byrne, H.M.; Chaplain, M.A.J. Growth of non-necrotic tumours in the presence and absence of inhibitors. Math. Biosci. 1995, 130, 151–181. [Google Scholar] [CrossRef]
- Byrne, H.M.; Chaplain, M.A.J. Growth of Necrotic Tumours in the Presence and Absence of Inhibitors. Math. Biosci. 1996, 135, 187–216. [Google Scholar] [CrossRef]
- Wise, S.M.; Lowengrub, J.S.; Frieboes, H.B.; Cristini, V. Three-dimensional multispecies nonlinear tumor growth—I: Model and numerical method. J. Theor. Biol. 2008, 253, 524–543. [Google Scholar] [CrossRef] [PubMed]
- Stiehl, T.; Baran, N.; Ho, A.D.; Marciniak-Czohra, A. Clonal selection and therapy resistance in acute leukaemias: Mathematical modelling explains different proliferation patterns at diagnosis and relapse. J. R. Soc. Interface 2014, 11, 20140079. [Google Scholar] [CrossRef] [PubMed]
- Walenda, T.; Stiehl, T.; Braun, H.; Fröbel, J.; Ho, A.D.; Schroeder, T.; Goecke, T.W.; Rath, B.; Germing, U.; Marciniak-Czohra, A.; et al. Feedback Signals in Myelodysplastic Syndromes: Increased Self-Renewal of the Malignant Clone Suppresses Normal Hematopoiesis. PLoS Comp. Biol. 2014, 10, e1003599. [Google Scholar] [CrossRef] [PubMed]
- Panetta, J.C. A mathematical model of drug resistance: Heterogeneous tumors. Math. Biosci. 1998, 147, 42–61. [Google Scholar] [CrossRef]
- Basanta, D.; Haralambos, H.; Deutsch, A. Studying the emergence of invasiveness in tumours using game theory. Eur. Phys. J. B 2008, 63, 393–397. [Google Scholar] [CrossRef]
- Enderling, H.; Hlatky, L.; Hahnfeldt, P. Migration rules: Tumours are conglomerates of self-metastases. Br. J. Cancer 2009, 100, 1917–1925. [Google Scholar] [CrossRef] [PubMed]
- Piotrowska, M.J.; Angus, S.D. A quantitative cellular automaton model of in vitro multicellular spheroid tumour growth. J. Theor. Biol. 2009, 258, 165–178. [Google Scholar] [CrossRef] [PubMed]
- Drasdo, D.; Hoehme, S. A single-cell-based model of tumor growth in vitro: Monolayers and spheroids. Phys. Biol. 2005, 2, 133. [Google Scholar] [CrossRef] [PubMed]
- Shirinifard, A.; Gens, J.S.; Zaitlen, B.L.; Poplawski, N.J.; Swat, M.; Glazier, J.A. 3D Multi-Cell Simulation of Tumor Growth and Angiogenesis. PLoS ONE 2009, 4, e7190. [Google Scholar] [CrossRef] [PubMed]
- Swat, M.H.; Thomas, G.L.; Shirinifard, A.; Clandenon, S.G.; Glazier, J.A. Emergent Stratification in Solid Tumors Selects for Reduced Cohesion of Tumor Cells: A Multi-Cell, Virtual-Tissue Model of Tumor Evolution Using CompuCell3D. PLoS ONE 2015, 10, e0127972. [Google Scholar] [CrossRef] [PubMed]
- Hatzikirou, H.; Brusch, L.; Schaller, C.; Simon, M.; Deutsch, A. Prediction of traveling front behavior in a lattice-gas cellular automaton model for tumor invasion. Comput. Math. Appl. 2010, 59, 2326–2339. [Google Scholar] [CrossRef]
- Aubert, M.; Badoual, M.; Fereol, S.; Christov, C.; Grammaticos, B. A cellular automaton model for the migration of glioma cells. Phys. Biol. 2006, 3, 93–100. [Google Scholar] [CrossRef] [PubMed]
- Ramis-Conde, I.; Drasdo, D.; Chaplain, M.A.J.; Anderson, A.R.A. Modeling the influence of the E-cadherin-β-catenin pathway in cancer cell invasion: A multiscale approach. Biophys. J. 2008, 95, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Ramis-Conde, I.; Chaplain, M.A.J.; Anderson, A.R.A.; Drasdo, D. Multi-scale modelling of cancer cell intravasation: The role of cadherins in metastasis. Phys. Biol. 2009, 6, 016008. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Wang, Z.; Sagotsky, J.A.; Deisboeck, T.S. Multiscale agent-based cancer modeling. J. Math. Biol. 2009, 58, 545–559. [Google Scholar] [CrossRef] [PubMed]
- Anderson, A.R.A. A hybrid mathematical model of solid tumour invasion: the importance of cell adhesion. Math. Med. Biol. 2005, 22, 163–186. [Google Scholar] [CrossRef] [PubMed]
- Fang, J.S.; Gillies, R.D.; Gatenby, R.A. Adaptation to hypoxia and acidosis in carcinogenesis and tumor progression. Semin. Cancer Biol. 2008, 18, 330–337. [Google Scholar] [CrossRef] [PubMed]
- Vincent, T.L.; Gatenby, R.A. An evolutionary model for initiation, promotion, and progression in carcinogenesis. Int. J. Oncol. 2008, 32, 729–737. [Google Scholar] [PubMed]
- Chisholm, R.H.; Lorenzi, T.; Lorz, A.; Larsen, A.K.; de Almeida, L.N.; Escargueil, A.; Clairambault, J. Emergence of drug tolerance in cancer cell populations: An evolutionary outcome of selection, nongenetic instability, and stress-induced adaptation. Cancer Res. 2015, 75, 930–939. [Google Scholar] [CrossRef] [PubMed]
- Ayati, B.P.; Edwards, C.M.; Webb, G.F.; Wikswo, J.P. A mathematical model of bone remodeling dynamics for normal bone cell populations and myeloma bone disease. Biol. Direct 2010, 5, 28. [Google Scholar] [CrossRef] [PubMed]
- Bouchnita, A.; Eymard, N.; Moyo, T.K.; Koury, M.J.; Volpert, V. Bone marrow infiltration by multiple myeloma causes anemia by reversible disruption of erythropoiesis. Am. J. Hematol. 2016, 91, 371–378. [Google Scholar] [CrossRef] [PubMed]
- Walker, B.A.; Wardell, C.P.; Melchor, L.; Hulkki, S.; Potter, N.E.; Johnson, D.C.; Fenwick, K.; Kozarewa, I.; Gonzalez, D.; Lord, C.J.; et al. Intraclonal heterogeneity and distinct molecular mechanisms characterize the development of t(4;14) and t(11;14) myeloma. Blood 2012, 120, 1077–1086. [Google Scholar] [CrossRef] [PubMed]
- Bouchnita, A.; Belmaati, F.E.; Aboulaich, R.; Ellaia, R.; Volpert, V. Mathematical modelling of intra-clonal heterogeneity in multiple myeloma. In Proceedings of the CARI 2016, Hammamet, Tunisia, 11–14 October 2016.
- Brioli, A.; Melchor, L.; Cavo, M.; Morgan, G.J. The impact of intra-clonal heterogeneity on the treatment of multiple myeloma. Br. J. Haematol. 2014, 165, 441–454. [Google Scholar] [CrossRef] [PubMed]
- Melchor, L.; Brioli, A.; Wardell, C.P.; Murison, A.; Potter, N.E.; Kaiser, M.F.; Fryer, R.A.; Johnson, D.C.; Begum, D.B.; Wilson, S.H.; et al. Single-cell genetic analysis reveals the composition of initiating clones and phylogenetic patterns of branching and parallel evolution in myeloma. Leukemia 2014, 28, 1705–1715. [Google Scholar] [CrossRef] [PubMed]
- Chesi, M.; Bergsagel, P.L.; Brents, L.A.; Smith, C.M.; Gerhard, D.S.; Kuehl, W.M. Dysregulation of cyclin D1 by translocation into an IgH gamma switch region in two multiple myeloma cell lines. Blood 1996, 88, 674–681. [Google Scholar] [PubMed]
- Bouyssou, J.M.C.; Ghobrial, I.M.; Roccaro, A.M. Targeting SDF-1 in multiple myeloma tumor microenvironment. Cancer Lett. 2015, 380, 315–318. [Google Scholar] [CrossRef] [PubMed]
- Vanderkerken, K.; van Camp, B.; de Greef, C.; Broek, I.V.; Asosingh, K.; van Riet, I. Homing of the myeloma cell clone. Acta Oncol. 2000, 39, 771–776. [Google Scholar] [PubMed]
- Hideshima, T.; Mitsiades, C.; Tonon, G.; Richardson, P.G.; Anderson, K.C. Understanding multiple myeloma pathogenesis in the bone marrow to identify new therapeutic targets. Nat. Rev. Cancer 2007, 7, 585–598. [Google Scholar] [CrossRef] [PubMed]
- Rozemuller, H.; van der Spek, E.; Bogers-Boer, L.H.; Zwart, M.C.; Verweij, V.; Emmelot, M.; Groen, R.W.; Spaapen, R.; Bloem, A.C.; Lokhorst, H.M.; et al. A bioluminescence imaging based in vivo model for preclinical testing of novel cellular immunotherapy strategies to improve the graft-versus-myeloma effect. Haematologica 2008, 93, 1049–1057. [Google Scholar] [CrossRef] [PubMed]
- Manier, S.; Sacco, A.; Leleu, X.; Ghobrial, I.M.; Roccaro, A.M. Bone marrow microenvironment in multiple myeloma progression. BioMed Res. Int. 2012, 2012, 157496. [Google Scholar] [CrossRef] [PubMed]
- Kitano, H. Systems Biology: A brief overview. Science 2002, 295, 1662–1664. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).